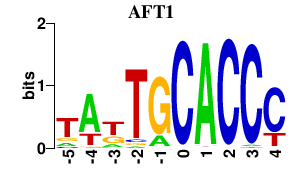
Results for AFT1
Z-value: 0.88
Transcription factors associated with AFT1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
AFT1
|
S000003039 | Transcription factor involved in iron utilization and homeostasis |
Activity-expression correlation:
Activity profile of AFT1 motif
Sorted Z-values of AFT1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| YOR383C | 13.45 |
FIT3
|
Mannoprotein that is incorporated into the cell wall via a glycosylphosphatidylinositol (GPI) anchor, involved in the retention of siderophore-iron in the cell wall |
|
| YOR382W | 13.17 |
FIT2
|
Mannoprotein that is incorporated into the cell wall via a glycosylphosphatidylinositol (GPI) anchor, involved in the retention of siderophore-iron in the cell wall |
|
| YDR536W | 7.21 |
STL1
|
Glycerol proton symporter of the plasma membrane, subject to glucose-induced inactivation, strongly but transiently induced when cells are subjected to osmotic shock |
|
| YLR136C | 5.99 |
TIS11
|
mRNA-binding protein expressed during iron starvation; binds to a sequence element in the 3'-untranslated regions of specific mRNAs to mediate their degradation; involved in iron homeostasis |
|
| YEL065W | 5.32 |
SIT1
|
Ferrioxamine B transporter, member of the ARN family of transporters that specifically recognize siderophore-iron chelates; transcription is induced during iron deprivation and diauxic shift; potentially phosphorylated by Cdc28p |
|
| YHL040C | 5.31 |
ARN1
|
Transporter, member of the ARN family of transporters that specifically recognize siderophore-iron chelates; responsible for uptake of iron bound to ferrirubin, ferrirhodin, and related siderophores |
|
| YLR205C | 5.24 |
HMX1
|
ER localized, heme-binding peroxidase involved in the degradation of heme; does not exhibit heme oxygenase activity despite similarity to heme oxygenases; expression regulated by AFT1 |
|
| YKL217W | 4.79 |
JEN1
|
Lactate transporter, required for uptake of lactate and pyruvate; phosphorylated; expression is derepressed by transcriptional activator Cat8p during respiratory growth, and repressed in the presence of glucose, fructose, and mannose |
|
| YAR053W | 4.36 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YER065C | 3.84 |
ICL1
|
Isocitrate lyase, catalyzes the formation of succinate and glyoxylate from isocitrate, a key reaction of the glyoxylate cycle; expression of ICL1 is induced by growth on ethanol and repressed by growth on glucose |
|
| YDL210W | 3.45 |
UGA4
|
Permease that serves as a gamma-aminobutyrate (GABA) transport protein involved in the utilization of GABA as a nitrogen source; catalyzes the transport of putrescine and delta-aminolevulinic acid (ALA); localized to the vacuolar membrane |
|
| YDR270W | 3.28 |
CCC2
|
Cu(+2)-transporting P-type ATPase, required for export of copper from the cytosol into an extracytosolic compartment; has similarity to human proteins involved in Menkes and Wilsons diseases |
|
| YLR327C | 3.23 |
TMA10
|
Protein of unknown function that associates with ribosomes |
|
| YDR269C | 3.19 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YPR151C | 3.17 |
SUE1
|
Mitochondrial protein required for degradation of unstable forms of cytochrome c |
|
| YAR060C | 3.04 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YIL057C | 2.82 |
Putative protein of unknown function; expression induced under carbon limitation and repressed under high glucose |
||
| YHR145C | 2.70 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YGR087C | 2.69 |
PDC6
|
Minor isoform of pyruvate decarboxylase, decarboxylates pyruvate to acetaldehyde, involved in amino acid catabolism; transcription is glucose- and ethanol-dependent, and is strongly induced during sulfur limitation |
|
| YFR053C | 2.65 |
HXK1
|
Hexokinase isoenzyme 1, a cytosolic protein that catalyzes phosphorylation of glucose during glucose metabolism; expression is highest during growth on non-glucose carbon sources; glucose-induced repression involves the hexokinase Hxk2p |
|
| YAL062W | 2.62 |
GDH3
|
NADP(+)-dependent glutamate dehydrogenase, synthesizes glutamate from ammonia and alpha-ketoglutarate; rate of alpha-ketoglutarate utilization differs from Gdh1p; expression regulated by nitrogen and carbon sources |
|
| YHR212C | 2.60 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YBL049W | 2.55 |
MOH1
|
Protein of unknown function, has homology to kinase Snf7p; not required for growth on nonfermentable carbon sources; essential for viability in stationary phase |
|
| YHR212W-A | 2.37 |
Putative protein of unknown function; identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching |
||
| YJR005C-A | 2.19 |
Putative protein of unknown function, originally identified as a syntenic homolog of an Ashbya gossypii gene |
||
| YER084W | 2.15 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YMR017W | 2.12 |
SPO20
|
Meiosis-specific subunit of the t-SNARE complex, required for prospore membrane formation during sporulation; similar to but not functionally redundant with Sec9p; SNAP-25 homolog |
|
| YGL146C | 2.11 |
Putative protein of unknown function, contains two putative transmembrane spans, shows no significant homology to any other known protein sequence, YGL146C is not an essential gene |
||
| YBR296C | 2.10 |
PHO89
|
Na+/Pi cotransporter, active in early growth phase; similar to phosphate transporters of Neurospora crassa; transcription regulated by inorganic phosphate concentrations and Pho4p |
|
| YER145C | 2.03 |
FTR1
|
High affinity iron permease involved in the transport of iron across the plasma membrane; forms complex with Fet3p; expression is regulated by iron |
|
| YCR021C | 2.00 |
HSP30
|
Hydrophobic plasma membrane localized, stress-responsive protein that negatively regulates the H(+)-ATPase Pma1p; induced by heat shock, ethanol treatment, weak organic acid, glucose limitation, and entry into stationary phase |
|
| YJL045W | 1.92 |
Minor succinate dehydrogenase isozyme; homologous to Sdh1p, the major isozyme reponsible for the oxidation of succinate and transfer of electrons to ubiquinone; induced during the diauxic shift in a Cat8p-dependent manner |
||
| YER014C-A | 1.90 |
BUD25
|
Protein involved in bud-site selection; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern |
|
| YNL237W | 1.86 |
YTP1
|
Probable type-III integral membrane protein of unknown function, has regions of similarity to mitochondrial electron transport proteins |
|
| YOR208W | 1.85 |
PTP2
|
Phosphotyrosine-specific protein phosphatase involved in the inactivation of mitogen-activated protein kinase (MAPK) during osmolarity sensing; dephosporylates Hog1p MAPK and regulates its localization; localized to the nucleus |
|
| YOR384W | 1.85 |
FRE5
|
Putative ferric reductase with similarity to Fre2p; expression induced by low iron levels; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YMR056C | 1.80 |
AAC1
|
Mitochondrial inner membrane ADP/ATP translocator, exchanges cytosolic ADP for mitochondrially synthesized ATP; phosphorylated; Aac1p is a minor isoform while Pet9p is the major ADP/ATP translocator |
|
| YDL067C | 1.79 |
COX9
|
Subunit VIIa of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain |
|
| YHL035C | 1.78 |
VMR1
|
Protein of unknown function that may interact with ribosomes, based on co-purification experiments; member of the ATP-binding cassette (ABC) family; potential Cdc28p substrate; detected in purified mitochondria in high-throughput studies |
|
| YBR292C | 1.78 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; YBR292C is not an essential gene |
||
| YEL060C | 1.76 |
PRB1
|
Vacuolar proteinase B (yscB), a serine protease of the subtilisin family; involved in protein degradation in the vacuole and required for full protein degradation during sporulation |
|
| YKL031W | 1.75 |
Dubious open reading frame, unlikely to encode a protein; not conserved in closely related Saccharomyces species |
||
| YEL070W | 1.75 |
DSF1
|
Deletion suppressor of mpt5 mutation |
|
| YFL051C | 1.70 |
Putative protein of unknown function; YFL051C is not an essential gene |
||
| YLL029W | 1.70 |
FRA1
|
Protein involved in negative regulation of transcription of iron regulon; forms an iron independent complex with Fra2p, Grx3p, and Grx4p; cytosolic; mutant fails to repress transcription of iron regulon and is defective in spore formation |
|
| YDR178W | 1.70 |
SDH4
|
Membrane anchor subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone |
|
| YOR381W | 1.69 |
FRE3
|
Ferric reductase, reduces siderophore-bound iron prior to uptake by transporters; expression induced by low iron levels |
|
| YGL072C | 1.69 |
Dubious open reading frame unlikely to encode a protein; partially overlaps the verified gene HSF1; null mutant displays increased resistance to antifungal agents gliotoxin, cycloheximide and H2O2 |
||
| YKR103W | 1.66 |
NFT1
|
Putative transporter of the multidrug resistance-associated protein (MRP) subfamily; adjacent ORFs YKR103W and YKR104W are merged in different strain backgrounds. |
|
| YKL038W | 1.63 |
RGT1
|
Glucose-responsive transcription factor that regulates expression of several glucose transporter (HXT) genes in response to glucose; binds to promoters and acts both as a transcriptional activator and repressor |
|
| YBL075C | 1.63 |
SSA3
|
ATPase involved in protein folding and the response to stress; plays a role in SRP-dependent cotranslational protein-membrane targeting and translocation; member of the heat shock protein 70 (HSP70) family; localized to the cytoplasm |
|
| YOL158C | 1.62 |
ENB1
|
Endosomal ferric enterobactin transporter, expressed under conditions of iron deprivation; member of the major facilitator superfamily; expression is regulated by Rcs1p and affected by chloroquine treatment |
|
| YBR047W | 1.60 |
FMP23
|
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YPL021W | 1.58 |
ECM23
|
Non-essential protein of unconfirmed function; affects pre-rRNA processing, may act as a negative regulator of the transcription of genes involved in pseudohyphal growth; homologous to Srd1p |
|
| YDR231C | 1.57 |
COX20
|
Mitochondrial inner membrane protein, required for proteolytic processing of Cox2p and its assembly into cytochrome c oxidase |
|
| YGL071W | 1.56 |
AFT1
|
Transcription factor involved in iron utilization and homeostasis; binds the consensus site PyPuCACCCPu and activates the expression of target genes in response to changes in iron availability |
|
| YLR326W | 1.54 |
Hypothetical protein |
||
| YHL047C | 1.52 |
ARN2
|
Transporter, member of the ARN family of transporters that specifically recognize siderophore-iron chelates; responsible for uptake of iron bound to the siderophore triacetylfusarinine C |
|
| YJR152W | 1.50 |
DAL5
|
Allantoin permease; ureidosuccinate permease; expression is constitutive but sensitive to nitrogen catabolite repression |
|
| YFL052W | 1.50 |
Putative zinc cluster protein that contains a DNA binding domain; null mutant sensitive to calcofluor white, low osmolarity and heat, suggesting a role for YFL052Wp in cell wall integrity |
||
| YHL046W-A | 1.49 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YLR047C | 1.48 |
FRE8
|
Protein with sequence similarity to iron/copper reductases, involved in iron homeostasis; deletion mutant has iron deficiency/accumulation growth defects; expression increased in the absence of copper-responsive transcription factor Mac1p |
|
| YFL063W | 1.43 |
Dubious open reading frame, based on available experimental and comparative sequence data |
||
| YDR441C | 1.41 |
APT2
|
Apparent pseudogene, not transcribed or translated under normal conditions; encodes a protein with similarity to adenine phosphoribosyltransferase, but artificially expressed protein exhibits no enzymatic activity |
|
| YOR186W | 1.40 |
Putative protein of unknown function; proper regulation of expression during heat stress is sphingolipid-dependent |
||
| YLR213C | 1.40 |
CRR1
|
Putative glycoside hydrolase of the spore wall envelope; required for normal spore wall assembly, possibly for cross-linking between the glucan and chitosan layers; expressed during sporulation |
|
| YPR007C | 1.40 |
REC8
|
Meiosis-specific component of sister chromatid cohesion complex; maintains cohesion between sister chromatids during meiosis I; maintains cohesion between centromeres of sister chromatids until meiosis II; homolog of S. pombe Rec8p |
|
| YJL163C | 1.39 |
Putative protein of unknown function |
||
| YOL084W | 1.37 |
PHM7
|
Protein of unknown function, expression is regulated by phosphate levels; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery and vacuole |
|
| YJL161W | 1.35 |
FMP33
|
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YKL102C | 1.34 |
Dubious open reading frame unlikely to encode a functional protein; deletion confers sensitivity to citric acid; predicted protein would include a thiol-disulfide oxidoreductase active site |
||
| YNL018C | 1.33 |
Putative protein of unknown function |
||
| YBR046C | 1.33 |
ZTA1
|
Zeta-crystallin homolog, found in the cytoplasm and nucleus; has similarity to E. coli quinone oxidoreductase and to human zeta-crystallin, which has quinone oxidoreductase activity |
|
| YKL103C | 1.33 |
LAP4
|
Vacuolar aminopeptidase, often used as a marker protein in studies of autophagy and cytosol to vacuole targeting (CVT) pathway |
|
| YGR065C | 1.28 |
VHT1
|
High-affinity plasma membrane H+-biotin (vitamin H) symporter; mutation results in fatty acid auxotrophy; 12 transmembrane domain containing major facilitator subfamily member; mRNA levels negatively regulated by iron deprivation and biotin |
|
| YLR004C | 1.28 |
THI73
|
Putative plasma membrane permease proposed to be involved in carboxylic acid uptake and repressed by thiamine; substrate of Dbf2p/Mob1p kinase; transcription is altered if mitochondrial dysfunction occurs |
|
| YLR334C | 1.26 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; overlaps a stand-alone long terminal repeat sequence whose presence indicates a retrotransposition event occurred here |
||
| YHR146W | 1.24 |
CRP1
|
Protein that binds to cruciform DNA structures |
|
| YDR034C | 1.21 |
LYS14
|
Transcriptional activator involved in regulation of genes of the lysine biosynthesis pathway; requires 2-aminoadipate semialdehyde as co-inducer |
|
| YLR174W | 1.21 |
IDP2
|
Cytosolic NADP-specific isocitrate dehydrogenase, catalyzes oxidation of isocitrate to alpha-ketoglutarate; levels are elevated during growth on non-fermentable carbon sources and reduced during growth on glucose |
|
| YOR020W-A | 1.18 |
Putative protein of unknown function, conserved in A. gossypii; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
||
| YFL011W | 1.17 |
HXT10
|
Putative hexose transporter, expressed at low levels and expression is repressed by glucose |
|
| YDR258C | 1.16 |
HSP78
|
Oligomeric mitochondrial matrix chaperone that cooperates with Ssc1p in mitochondrial thermotolerance after heat shock; prevents the aggregation of misfolded matrix proteins; component of the mitochondrial proteolysis system |
|
| YLR267W | 1.16 |
BOP2
|
Protein of unknown function, overproduction suppresses a pam1 slv3 double null mutation |
|
| YBR295W | 1.16 |
PCA1
|
Cadmium transporting P-type ATPase; may also have a role in copper and iron homeostasis; S288C and most other lab strains contain a G970R mutation which eliminates normal cadmium transport function |
|
| YOR139C | 1.15 |
Hypothetical protein |
||
| YLR142W | 1.14 |
PUT1
|
Proline oxidase, nuclear-encoded mitochondrial protein involved in utilization of proline as sole nitrogen source; PUT1 transcription is induced by Put3p in the presence of proline and the absence of a preferred nitrogen source |
|
| YDR342C | 1.13 |
HXT7
|
High-affinity glucose transporter of the major facilitator superfamily, nearly identical to Hxt6p, expressed at high basal levels relative to other HXTs, expression repressed by high glucose levels |
|
| YMR181C | 1.12 |
Protein of unknown function; mRNA transcribed as part of a bicistronic transcript with a predicted transcriptional repressor RGM1/YMR182C; mRNA is destroyed by nonsense-mediated decay (NMD); YMR181C is not an essential gene |
||
| YLR356W | 1.12 |
Putative protein of unknown function with similarity to SCM4; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YLR356W is not an essential gene |
||
| YGR043C | 1.12 |
NQM1
|
Protein of unknown function; transcription is repressed by Mot1p and induced by alpha-factor and during diauxic shift; null mutant non-quiescent cells exhibit reduced reproductive capacity |
|
| YHR211W | 1.10 |
FLO5
|
Lectin-like cell wall protein (flocculin) involved in flocculation, binds to mannose chains on the surface of other cells, confers floc-forming ability that is chymotrypsin resistant but heat labile; similar to Flo1p |
|
| YDR281C | 1.09 |
PHM6
|
Protein of unknown function, expression is regulated by phosphate levels |
|
| YFL064C | 1.09 |
Putative protein of unknown function |
||
| YPR169W-A | 1.08 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps two other dubious ORFs: YPR170C and YPR170W-B |
||
| YIL146C | 1.08 |
ECM37
|
Non-essential protein of unknown function |
|
| YDR379C-A | 1.08 |
Hypothetical protein identified by homology. See FEBS Letters [2000] 487:31-36. |
||
| YIL055C | 1.07 |
Putative protein of unknown function |
||
| YFL021W | 1.06 |
GAT1
|
Transcriptional activator of genes involved in nitrogen catabolite repression; contains a GATA-1-type zinc finger DNA-binding motif; activity and localization regulated by nitrogen limitation and Ure2p |
|
| YOR328W | 1.06 |
PDR10
|
ATP-binding cassette (ABC) transporter, multidrug transporter involved in the pleiotropic drug resistance network; regulated by Pdr1p and Pdr3p |
|
| YMR271C | 1.06 |
URA10
|
Minor orotate phosphoribosyltransferase (OPRTase) isozyme that catalyzes the fifth enzymatic step in the de novo biosynthesis of pyrimidines, converting orotate into orotidine-5'-phosphate; major OPRTase encoded by URA5 |
|
| YPR042C | 1.06 |
PUF2
|
Member of the PUF protein family, which is defined by the presence of Pumilio homology domains that confer RNA binding activity; preferentially binds mRNAs encoding membrane-associated proteins |
|
| YLR399C | 1.06 |
BDF1
|
Protein involved in transcription initiation at TATA-containing promoters; associates with the basal transcription factor TFIID; contains two bromodomains; corresponds to the C-terminal region of mammalian TAF1; redundant with Bdf2p |
|
| YKR011C | 1.04 |
Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus |
||
| YER101C | 1.04 |
AST2
|
Protein that may have a role in targeting of plasma membrane [H+]ATPase (Pma1p) to the plasma membrane, as suggested by analysis of genetic interactions |
|
| YOR084W | 1.02 |
LPX1
|
Oleic acid-inducible, peroxisomal matrix localized lipase; transcriptionally activated by Yrm1p along with genes involved in multidrug resistance; peroxisomal import is dependent on the PTS1 receptor, Pex5p and on self-interaction |
|
| YHR205W | 1.01 |
SCH9
|
Protein kinase involved in transcriptional activation of osmostress-responsive genes; regulates G1 progression, cAPK activity, nitrogen activation of the FGM pathway; involved in life span regulation; homologous to mammalian Akt/PKB |
|
| YNR073C | 1.01 |
Putative mannitol dehydrogenase |
||
| YGL089C | 1.00 |
MF(ALPHA)2
|
Mating pheromone alpha-factor, made by alpha cells; interacts with mating type a cells to induce cell cycle arrest and other responses leading to mating; also encoded by MF(ALPHA)1, which is more highly expressed than MF(ALPHA)2 |
|
| YHR095W | 1.00 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YDR406W | 1.00 |
PDR15
|
Plasma membrane ATP binding cassette (ABC) transporter, multidrug transporter and general stress response factor implicated in cellular detoxification; regulated by Pdr1p, Pdr3p and Pdr8p; promoter contains a PDR responsive element |
|
| YDL204W | 0.99 |
RTN2
|
Protein of unknown function; has similarity to mammalian reticulon proteins; member of the RTNLA (reticulon-like A) subfamily |
|
| YJL133C-A | 0.99 |
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
||
| YGR088W | 0.99 |
CTT1
|
Cytosolic catalase T, has a role in protection from oxidative damage by hydrogen peroxide |
|
| YOR140W | 0.99 |
SFL1
|
Transcriptional repressor and activator; involved in repression of flocculation-related genes, and activation of stress responsive genes; negatively regulated by cAMP-dependent protein kinase A subunit Tpk2p |
|
| YNL091W | 0.99 |
NST1
|
Protein of unknown function, mediates sensitivity to salt stress; interacts physically with the splicing factor Msl1p and also displays genetic interaction with MSL1 |
|
| YPL054W | 0.98 |
LEE1
|
Zinc-finger protein of unknown function |
|
| YKL202W | 0.98 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YPL057C | 0.98 |
SUR1
|
Probable catalytic subunit of a mannosylinositol phosphorylceramide (MIPC) synthase, forms a complex with probable regulatory subunit Csg2p; function in sphingolipid biosynthesis is overlapping with that of Csh1p |
|
| YCL001W-A | 0.98 |
Putative protein of unknown function; YCL001W-A is not an essential gene |
||
| YMR244W | 0.97 |
Putative protein of unknown function |
||
| YOR186C-A | 0.95 |
Identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching |
||
| YHR048W | 0.95 |
YHK8
|
Presumed antiporter of the DHA1 family of multidrug resistance transporters; contains 12 predicted transmembrane spans; expression of gene is up-regulated in cells exhibiting reduced susceptibility to azoles |
|
| YOR388C | 0.94 |
FDH1
|
NAD(+)-dependent formate dehydrogenase, may protect cells from exogenous formate |
|
| YBL073W | 0.94 |
Hypothetical protein; open reading frame overlaps 5' end of essential AAR2 gene encoding a component of the U5 snRNP required for splicing of U3 precursors |
||
| YPR065W | 0.94 |
ROX1
|
Heme-dependent repressor of hypoxic genes; contains an HMG domain that is responsible for DNA bending activity |
|
| YCR005C | 0.94 |
CIT2
|
Citrate synthase, catalyzes the condensation of acetyl coenzyme A and oxaloacetate to form citrate, peroxisomal isozyme involved in glyoxylate cycle; expression is controlled by Rtg1p and Rtg2p transcription factors |
|
| YJR155W | 0.93 |
AAD10
|
Putative aryl-alcohol dehydrogenase with similarity to P. chrysosporium aryl-alcohol dehydrogenase; mutational analysis has not yet revealed a physiological role |
|
| YBL100C | 0.93 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; almost completely overlaps the 5' end of ATP1 |
||
| YNL190W | 0.93 |
Cell wall protein of unknown function; proposed role as a hydrophilin induced by osmotic stress; contains a putative GPI-attachment site |
||
| YPL111W | 0.91 |
CAR1
|
Arginase, responsible for arginine degradation, expression responds to both induction by arginine and nitrogen catabolite repression; disruption enhances freeze tolerance |
|
| YJL149W | 0.91 |
DAS1
|
Putative SCF ubiquitin ligase F-box protein; interacts physically with both Cdc53p and Skp1 and genetically with CDC34; similar to putative F-box protein YDR131C; null mutant suppresses dst1delta sensitivity for 6-azauracil |
|
| YER175C | 0.90 |
TMT1
|
Trans-aconitate methyltransferase, cytosolic enzyme that catalyzes the methyl esterification of 3-isopropylmalate, an intermediate of the leucine biosynthetic pathway, and trans-aconitate, which inhibits the citric acid cycle |
|
| YLR425W | 0.89 |
TUS1
|
Guanine nucleotide exchange factor (GEF) that functions to modulate Rho1p activity as part of the cell integrity signaling pathway; multicopy suppressor of tor2 mutation and ypk1 ypk2 double mutation; potential Cdc28p substrate |
|
| YMR175W-A | 0.89 |
Putative protein of unknown function |
||
| YDR343C | 0.88 |
HXT6
|
High-affinity glucose transporter of the major facilitator superfamily, nearly identical to Hxt7p, expressed at high basal levels relative to other HXTs, repression of expression by high glucose requires SNF3 |
|
| YDR171W | 0.88 |
HSP42
|
Small heat shock protein (sHSP) with chaperone activity; forms barrel-shaped oligomers that suppress unfolded protein aggregation; involved in cytoskeleton reorganization after heat shock |
|
| YBL074C | 0.87 |
AAR2
|
Component of the U5 snRNP, required for splicing of U3 precursors; originally described as a splicing factor specifically required for splicing pre-mRNA of the MATa1 cistron |
|
| YEL030C-A | 0.86 |
Identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching |
||
| YHR217C | 0.86 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; located in the telomeric region TEL08R. |
||
| YML089C | 0.85 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data; expression induced by calcium shortage |
||
| YLR403W | 0.85 |
SFP1
|
Transcription factor that controls expression of many ribosome biogenesis genes in response to nutrients and stress, regulates G2/M transitions during mitotic cell cycle and DNA-damage response, involved in cell size modulation |
|
| YEL049W | 0.85 |
PAU2
|
Part of 23-member seripauperin multigene family encoded mainly in subtelomeric regions, active during alcoholic fermentation, regulated by anaerobiosis, negatively regulated by oxygen, repressed by heme |
|
| YDR442W | 0.84 |
Dubious open reading frame unlikely to encode a functional protein, based on available experimental and comparative sequence data |
||
| YLR402W | 0.83 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data |
||
| YIL013C | 0.83 |
PDR11
|
ATP-binding cassette (ABC) transporter, multidrug transporter involved in multiple drug resistance; mediates sterol uptake when sterol biosynthesis is compromisedregulated by Pdr1p; required for anaerobic growth |
|
| YNR069C | 0.83 |
BSC5
|
Protein of unknown function, ORF exhibits genomic organization compatible with a translational readthrough-dependent mode of expression |
|
| YIL144W | 0.82 |
TID3
|
Component of the evolutionarily conserved kinetochore-associated Ndc80 complex (Ndc80p-Nuf2p-Spc24p-Spc25p); conserved coiled-coil protein involved in chromosome segregation, spindle checkpoint activity, kinetochore assembly and clustering |
|
| YDL197C | 0.82 |
ASF2
|
Anti-silencing protein that causes derepression of silent loci when overexpressed |
|
| YLR122C | 0.82 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the dubious ORF YLR123C |
||
| YAL063C | 0.81 |
FLO9
|
Lectin-like protein with similarity to Flo1p, thought to be expressed and involved in flocculation |
|
| YBR071W | 0.80 |
Putative protein of unknown function; (GFP)-fusion and epitope-tagged proteins localize to the cytoplasm; mRNA expression may be regulated by the cell cycle and/or cell wall stress |
||
| YCR079W | 0.80 |
PTC6
|
Mitochondrial protein phosphatase of type 2C with similarity to mammalian PP1Ks; involved in mitophagy; null mutant is sensitive to rapamycin and has decreased phosphorylation of the Pda1 subunit of pyruvate dehydrogenase |
|
| YKL105C | 0.80 |
Putative protein of unknown function |
||
| YPL061W | 0.79 |
ALD6
|
Cytosolic aldehyde dehydrogenase, activated by Mg2+ and utilizes NADP+ as the preferred coenzyme; required for conversion of acetaldehyde to acetate; constitutively expressed; locates to the mitochondrial outer surface upon oxidative stress |
|
| YDR232W | 0.79 |
HEM1
|
5-aminolevulinate synthase, catalyzes the first step in the heme biosynthetic pathway; an N-terminal signal sequence is required for localization to the mitochondrial matrix; expression is regulated by Hap2p-Hap3p |
|
| YDL196W | 0.79 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; open reading frame is located in promoter region of essential gene SEC31 |
||
| YDR528W | 0.79 |
HLR1
|
Protein involved in regulation of cell wall composition and integrity and response to osmotic stress; overproduction suppresses a lysis sensitive PKC mutation; similar to Lre1p, which functions antagonistically to protein kinase A |
|
| YNL278W | 0.79 |
CAF120
|
Part of the evolutionarily-conserved CCR4-NOT transcriptional regulatory complex involved in controlling mRNA initiation, elongation, and degradation |
|
| YMR081C | 0.79 |
ISF1
|
Serine-rich, hydrophilic protein with similarity to Mbr1p; overexpression suppresses growth defects of hap2, hap3, and hap4 mutants; expression is under glucose control; cotranscribed with NAM7 in a cyp1 mutant |
|
| YDR452W | 0.78 |
PPN1
|
Vacuolar endopolyphosphatase with a role in phosphate metabolism; functions as a homodimer |
|
| YPR013C | 0.78 |
Putative zinc finger protein; YPR013C is not an essential gene |
||
| YKL046C | 0.77 |
DCW1
|
Putative mannosidase, GPI-anchored membrane protein required for cell wall biosynthesis in bud formation;homologous to Dfg5p |
|
| YMR141W-A | 0.77 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; overlaps the verified gene RPL13B/YMR142C |
||
| YMR057C | 0.76 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps verified ORF AAC1 |
||
| YDR077W | 0.76 |
SED1
|
Major stress-induced structural GPI-cell wall glycoprotein in stationary-phase cells, associates with translating ribosomes, possible role in mitochondrial genome maintenance; ORF contains two distinct variable minisatellites |
|
| YGL136C | 0.75 |
MRM2
|
Mitochondrial 2' O-ribose methyltransferase, required for methylation of U(2791) in 21S rRNA; MRM2 deletion confers thermosensitive respiration and loss of mitochondrial DNA; has similarity to Spb1p and Trm7p, and to E. coli FtsJ/RrmJ |
|
| YEL030W | 0.74 |
ECM10
|
Heat shock protein of the Hsp70 family, localized in mitochondrial nucleoids, plays a role in protein translocation, interacts with Mge1p in an ATP-dependent manner; overexpression induces extensive mitochondrial DNA aggregations |
|
| YBL060W | 0.74 |
YEL1
|
Guanine nucleotide exchange factor specific for Arf3p; localized to the bud neck and tip; required for localization of Arf3p to the bud neck and tip |
|
| YHR199C | 0.73 |
AIM46
|
Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss (petite formation) |
|
| YLR309C | 0.73 |
IMH1
|
Protein involved in vesicular transport, mediates transport between an endosomal compartment and the Golgi, contains a Golgi-localization (GRIP) domain that interacts with activated Arl1p-GTP to localize Imh1p to the Golgi |
|
| YLR123C | 0.73 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the dubious ORF YLR122C; contains characteristic aminoacyl-tRNA motif |
||
| YDR380W | 0.73 |
ARO10
|
Phenylpyruvate decarboxylase, catalyzes decarboxylation of phenylpyruvate to phenylacetaldehyde, which is the first specific step in the Ehrlich pathway |
|
| YKR067W | 0.72 |
GPT2
|
Glycerol-3-phosphate/dihydroxyacetone phosphate dual substrate-specific sn-1 acyltransferase located in lipid particles and the ER; involved in the stepwise acylation of glycerol-3-phosphate and dihydroxyacetone in lipid biosynthesis |
|
| YBR284W | 0.71 |
Putative protein of unknown function; YBR284W is not an essential gene; null mutant exhibits decreased resistance to rapamycin and wortmannin |
||
| YOR378W | 0.71 |
Putative protein of unknown function; belongs to the DHA2 family of drug:H+ antiporters; YOR378W is not an essential gene |
||
| YDR169C | 0.70 |
STB3
|
Protein that binds Sin3p in a two-hybrid assay |
|
| YHL005C | 0.70 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the verified ORF YHL004W |
||
| YBR203W | 0.70 |
COS111
|
Protein required for resistance to the antifungal drug ciclopirox olamine; not related to the subtelomerically-encoded COS family; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
|
| YBR132C | 0.70 |
AGP2
|
High affinity polyamine permease, preferentially uses spermidine over putrescine; expression is down-regulated by osmotic stress; plasma membrane carnitine transporter, also functions as a low-affinity amino acid permease |
|
| YCL025C | 0.69 |
AGP1
|
Low-affinity amino acid permease with broad substrate range, involved in uptake of asparagine, glutamine, and other amino acids; expression is regulated by the SPS plasma membrane amino acid sensor system (Ssy1p-Ptr3p-Ssy5p) |
|
| YDL113C | 0.69 |
ATG20
|
Sorting nexin family member required for the cytoplasm-to-vacuole targeting (Cvt) pathway and for endosomal sorting; has a Phox homology domain that binds phosphatidylinositol-3-phosphate; interacts with Snx4p; potential Cdc28p substrate |
|
| YLR187W | 0.69 |
SKG3
|
Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, bud, and bud neck; potential Cdc28p substrate; similar to Caf120p and Skg4p |
|
| YFL019C | 0.68 |
Dubious open reading frame unlikely to encode a protein; YFL019C is not an essential gene |
||
| YBL099W | 0.68 |
ATP1
|
Alpha subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated |
|
| YLL051C | 0.68 |
FRE6
|
Putative ferric reductase with similarity to Fre2p; expression induced by low iron levels |
|
| YPR040W | 0.67 |
TIP41
|
Protein that interacts physically and genetically with Tap42p, which regulates protein phosphatase 2A; component of the TOR (target of rapamycin) signaling pathway |
|
| YOL025W | 0.66 |
LAG2
|
Protein involved in determination of longevity; LAG2 gene is preferentially expressed in young cells; overexpression extends the mean and maximum life span of cells |
|
| YKR058W | 0.66 |
GLG1
|
Self-glucosylating initiator of glycogen synthesis, also glucosylates n-dodecyl-beta-D-maltoside; similar to mammalian glycogenin |
|
| YOL060C | 0.66 |
MAM3
|
Protein required for normal mitochondrial morphology, has similarity to hemolysins |
|
| YER004W | 0.65 |
FMP52
|
Protein of unknown function, localized to the mitochondrial outer membrane; induced by treatment with 8-methoxypsoralen and UVA irradiation |
|
| YDR540C | 0.65 |
IRC4
|
Putative protein of unknown function; null mutant displays increased levels of spontaneous Rad52p foci; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus |
|
| YHR097C | 0.65 |
Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus |
||
| YLL023C | 0.65 |
Protein of unknown function; highly conserved across species and orthologous to human TMEM33; may interact with ribosomes, based on co-purification experiments; GFP-fusion protein localizes to the endoplasmic reticulum |
||
| YKR095W | 0.64 |
MLP1
|
Myosin-like protein associated with the nuclear envelope, connects the nuclear pore complex with the nuclear interior; involved with Tel1p in telomere length control; involved with Pml1p and Pml39p in nuclear retention of unspliced mRNAs |
|
| YHL041W | 0.64 |
Dubious open reading frame unlikely to encode a protein, based on experimental and comparative sequence data |
||
| YDR443C | 0.63 |
SSN2
|
Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; required for stable association of Srb10p-Srb11p kinase; essential for transcriptional regulation |
|
| YOR020C | 0.63 |
HSP10
|
Mitochondrial matrix co-chaperonin that inhibits the ATPase activity of Hsp60p, a mitochondrial chaperonin; involved in protein folding and sorting in the mitochondria; 10 kD heat shock protein with similarity to E. coli groES |
|
| YPL223C | 0.63 |
GRE1
|
Hydrophilin of unknown function; stress induced (osmotic, ionic, oxidative, heat shock and heavy metals); regulated by the HOG pathway |
|
| YOL085W-A | 0.63 |
Dubious open reading frame unlikely to encode a protein, based on available experimental and comparative sequence data; partially overlaps the dubious ORF YOL085C |
Network of associatons between targets according to the STRING database.
First level regulatory network of AFT1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.8 | 38.7 | GO:0015891 | siderophore transport(GO:0015891) |
| 1.6 | 1.6 | GO:0015688 | iron chelate transport(GO:0015688) |
| 1.5 | 7.6 | GO:0015793 | glycerol transport(GO:0015793) |
| 1.2 | 4.8 | GO:0006848 | pyruvate transport(GO:0006848) |
| 1.0 | 4.0 | GO:0046323 | glucose import(GO:0046323) |
| 0.9 | 3.7 | GO:0000949 | aromatic amino acid family catabolic process to alcohol via Ehrlich pathway(GO:0000949) |
| 0.9 | 2.6 | GO:0006827 | high-affinity iron ion transmembrane transport(GO:0006827) |
| 0.7 | 6.3 | GO:0006097 | glyoxylate cycle(GO:0006097) |
| 0.7 | 2.1 | GO:0019676 | ammonia assimilation cycle(GO:0019676) |
| 0.7 | 2.0 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.7 | 2.0 | GO:0010039 | response to iron ion(GO:0010039) regulation of transcription from RNA polymerase II promoter in response to iron(GO:0034395) cellular response to iron ion(GO:0071281) |
| 0.6 | 0.6 | GO:0015755 | fructose transport(GO:0015755) |
| 0.6 | 1.7 | GO:0043270 | positive regulation of ion transport(GO:0043270) |
| 0.5 | 2.2 | GO:0015847 | putrescine transport(GO:0015847) |
| 0.5 | 2.1 | GO:0000200 | inactivation of MAPK activity involved in cell wall organization or biogenesis(GO:0000200) regulation of MAPK cascade involved in cell wall organization or biogenesis(GO:1903137) negative regulation of MAPK cascade involved in cell wall organization or biogenesis(GO:1903138) negative regulation of cell wall organization or biogenesis(GO:1903339) |
| 0.5 | 12.6 | GO:0006879 | cellular iron ion homeostasis(GO:0006879) |
| 0.4 | 2.7 | GO:0000501 | flocculation(GO:0000128) flocculation via cell wall protein-carbohydrate interaction(GO:0000501) |
| 0.4 | 1.2 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.4 | 1.9 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.4 | 1.1 | GO:0010133 | proline catabolic process(GO:0006562) proline catabolic process to glutamate(GO:0010133) |
| 0.4 | 1.1 | GO:1900460 | negative regulation of invasive growth in response to glucose limitation by negative regulation of transcription from RNA polymerase II promoter(GO:1900460) |
| 0.4 | 1.1 | GO:0044209 | AMP salvage(GO:0044209) |
| 0.3 | 1.0 | GO:0042744 | hydrogen peroxide catabolic process(GO:0042744) |
| 0.3 | 1.4 | GO:0015886 | heme transport(GO:0015886) |
| 0.3 | 1.0 | GO:0006688 | glycosphingolipid biosynthetic process(GO:0006688) |
| 0.3 | 0.9 | GO:0019627 | urea metabolic process(GO:0019627) |
| 0.3 | 3.3 | GO:0015893 | drug transport(GO:0015893) |
| 0.3 | 0.3 | GO:0042743 | hydrogen peroxide metabolic process(GO:0042743) |
| 0.3 | 0.8 | GO:0034310 | ethanol catabolic process(GO:0006068) acetate biosynthetic process(GO:0019413) primary alcohol catabolic process(GO:0034310) |
| 0.3 | 0.8 | GO:0015888 | thiamine transport(GO:0015888) |
| 0.3 | 1.3 | GO:0051180 | vitamin transport(GO:0051180) |
| 0.3 | 0.5 | GO:0032006 | regulation of TOR signaling(GO:0032006) |
| 0.2 | 0.7 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.2 | 1.0 | GO:0043954 | cellular component maintenance(GO:0043954) |
| 0.2 | 1.2 | GO:0001080 | nitrogen catabolite activation of transcription from RNA polymerase II promoter(GO:0001080) nitrogen catabolite activation of transcription(GO:0090294) |
| 0.2 | 0.5 | GO:0045895 | positive regulation of mating-type specific transcription, DNA-templated(GO:0045895) |
| 0.2 | 0.9 | GO:0000256 | allantoin metabolic process(GO:0000255) allantoin catabolic process(GO:0000256) cellular amide catabolic process(GO:0043605) |
| 0.2 | 1.3 | GO:0007571 | age-dependent response to oxidative stress(GO:0001306) age-dependent general metabolic decline involved in chronological cell aging(GO:0001323) age-dependent response to oxidative stress involved in chronological cell aging(GO:0001324) age-dependent general metabolic decline(GO:0007571) |
| 0.2 | 2.0 | GO:0006616 | SRP-dependent cotranslational protein targeting to membrane, translocation(GO:0006616) |
| 0.2 | 1.9 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.2 | 0.4 | GO:0035337 | fatty-acyl-CoA metabolic process(GO:0035337) |
| 0.2 | 1.1 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.2 | 1.0 | GO:0045764 | positive regulation of cellular amino acid metabolic process(GO:0045764) positive regulation of cellular amino acid biosynthetic process(GO:2000284) |
| 0.2 | 0.6 | GO:0009107 | lipoate metabolic process(GO:0009106) lipoate biosynthetic process(GO:0009107) |
| 0.2 | 0.6 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.2 | 1.8 | GO:0007039 | protein catabolic process in the vacuole(GO:0007039) |
| 0.2 | 0.5 | GO:0000042 | protein targeting to Golgi(GO:0000042) establishment of protein localization to Golgi(GO:0072600) |
| 0.2 | 0.7 | GO:0001666 | response to hypoxia(GO:0001666) |
| 0.2 | 1.1 | GO:0006207 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.2 | 0.9 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.2 | 2.3 | GO:0071459 | protein localization to chromosome, centromeric region(GO:0071459) |
| 0.2 | 1.2 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.2 | 0.2 | GO:0046487 | glyoxylate metabolic process(GO:0046487) |
| 0.2 | 0.7 | GO:0015804 | neutral amino acid transport(GO:0015804) |
| 0.2 | 0.7 | GO:0051101 | positive regulation of DNA binding(GO:0043388) regulation of DNA binding(GO:0051101) |
| 0.2 | 0.7 | GO:0016024 | CDP-diacylglycerol biosynthetic process(GO:0016024) CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.2 | 0.3 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.2 | 0.6 | GO:0072396 | cellular response to biotic stimulus(GO:0071216) response to cell cycle checkpoint signaling(GO:0072396) response to mitotic cell cycle checkpoint signaling(GO:0072414) response to spindle checkpoint signaling(GO:0072417) response to mitotic spindle checkpoint signaling(GO:0072476) response to mitotic cell cycle spindle assembly checkpoint signaling(GO:0072479) response to spindle assembly checkpoint signaling(GO:0072485) negative regulation of protein import into nucleus during spindle assembly checkpoint(GO:1901925) |
| 0.2 | 0.5 | GO:0070304 | positive regulation of stress-activated MAPK cascade(GO:0032874) positive regulation of stress-activated protein kinase signaling cascade(GO:0070304) |
| 0.2 | 0.5 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 0.2 | 0.6 | GO:0000431 | regulation of transcription from RNA polymerase II promoter by galactose(GO:0000431) positive regulation of transcription from RNA polymerase II promoter by galactose(GO:0000435) |
| 0.2 | 0.5 | GO:0032079 | positive regulation of deoxyribonuclease activity(GO:0032077) positive regulation of endodeoxyribonuclease activity(GO:0032079) |
| 0.2 | 0.9 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 0.2 | 0.5 | GO:0006798 | polyphosphate catabolic process(GO:0006798) |
| 0.1 | 0.4 | GO:0030656 | regulation of vitamin metabolic process(GO:0030656) positive regulation of vitamin metabolic process(GO:0046136) regulation of thiamine biosynthetic process(GO:0070623) positive regulation of thiamine biosynthetic process(GO:0090180) |
| 0.1 | 0.4 | GO:0001196 | regulation of mating-type specific transcription from RNA polymerase II promoter(GO:0001196) negative regulation of mating-type specific transcription from RNA polymerase II promoter(GO:0001198) negative regulation of mating-type specific transcription, DNA-templated(GO:0045894) |
| 0.1 | 1.8 | GO:0033617 | mitochondrial respiratory chain complex IV assembly(GO:0033617) |
| 0.1 | 2.1 | GO:0031321 | ascospore-type prospore assembly(GO:0031321) |
| 0.1 | 0.5 | GO:0015809 | arginine transport(GO:0015809) |
| 0.1 | 0.6 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.1 | 3.8 | GO:0022900 | electron transport chain(GO:0022900) |
| 0.1 | 0.4 | GO:0046688 | response to copper ion(GO:0046688) |
| 0.1 | 0.6 | GO:0009756 | cellular glucose homeostasis(GO:0001678) carbohydrate mediated signaling(GO:0009756) hexose mediated signaling(GO:0009757) sugar mediated signaling pathway(GO:0010182) glucose mediated signaling pathway(GO:0010255) carbohydrate homeostasis(GO:0033500) glucose homeostasis(GO:0042593) cellular response to monosaccharide stimulus(GO:0071326) cellular response to hexose stimulus(GO:0071331) cellular response to glucose stimulus(GO:0071333) |
| 0.1 | 1.4 | GO:0000436 | carbon catabolite activation of transcription from RNA polymerase II promoter(GO:0000436) |
| 0.1 | 1.4 | GO:0015718 | monocarboxylic acid transport(GO:0015718) |
| 0.1 | 0.1 | GO:0015691 | cadmium ion transport(GO:0015691) |
| 0.1 | 0.6 | GO:0030026 | cellular manganese ion homeostasis(GO:0030026) manganese ion homeostasis(GO:0055071) |
| 0.1 | 1.4 | GO:0042026 | protein refolding(GO:0042026) |
| 0.1 | 1.5 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.1 | 0.6 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.1 | 0.9 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.1 | 0.4 | GO:0033683 | nucleotide-excision repair, DNA incision(GO:0033683) |
| 0.1 | 0.1 | GO:1900544 | positive regulation of nucleotide metabolic process(GO:0045981) positive regulation of purine nucleotide metabolic process(GO:1900544) |
| 0.1 | 0.4 | GO:0070054 | mRNA splicing via endonucleolytic cleavage and ligation involved in unfolded protein response(GO:0030969) IRE1-mediated unfolded protein response(GO:0036498) mRNA splicing, via endonucleolytic cleavage and ligation(GO:0070054) |
| 0.1 | 0.2 | GO:0046033 | AMP biosynthetic process(GO:0006167) AMP metabolic process(GO:0046033) |
| 0.1 | 0.6 | GO:0016241 | positive regulation of macroautophagy(GO:0016239) regulation of macroautophagy(GO:0016241) |
| 0.1 | 0.4 | GO:0001308 | negative regulation of chromatin silencing involved in replicative cell aging(GO:0001308) |
| 0.1 | 0.4 | GO:0015976 | carbon utilization(GO:0015976) |
| 0.1 | 1.0 | GO:0006817 | phosphate ion transport(GO:0006817) |
| 0.1 | 0.2 | GO:2000058 | regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000058) |
| 0.1 | 0.2 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.1 | 0.6 | GO:0015802 | basic amino acid transport(GO:0015802) |
| 0.1 | 1.2 | GO:0019682 | pentose-phosphate shunt(GO:0006098) glyceraldehyde-3-phosphate metabolic process(GO:0019682) |
| 0.1 | 0.3 | GO:0001113 | transcriptional open complex formation at RNA polymerase II promoter(GO:0001113) |
| 0.1 | 0.3 | GO:0070874 | negative regulation of cellular carbohydrate metabolic process(GO:0010677) negative regulation of glycogen biosynthetic process(GO:0045719) negative regulation of carbohydrate metabolic process(GO:0045912) negative regulation of glycogen metabolic process(GO:0070874) |
| 0.1 | 0.7 | GO:0031167 | rRNA methylation(GO:0031167) |
| 0.1 | 0.3 | GO:0043200 | response to amino acid(GO:0043200) |
| 0.1 | 0.8 | GO:0006813 | potassium ion transport(GO:0006813) |
| 0.1 | 0.2 | GO:0090282 | positive regulation of transcription involved in G2/M transition of mitotic cell cycle(GO:0090282) |
| 0.1 | 0.4 | GO:0061187 | regulation of chromatin silencing at rDNA(GO:0061187) negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 0.1 | 1.7 | GO:0061726 | mitophagy(GO:0000422) mitochondrion disassembly(GO:0061726) |
| 0.1 | 0.4 | GO:0043937 | regulation of sporulation(GO:0043937) |
| 0.1 | 0.3 | GO:0009749 | response to hexose(GO:0009746) response to glucose(GO:0009749) response to monosaccharide(GO:0034284) |
| 0.1 | 0.3 | GO:0019184 | glutathione biosynthetic process(GO:0006750) nonribosomal peptide biosynthetic process(GO:0019184) |
| 0.1 | 0.1 | GO:0006739 | NADP metabolic process(GO:0006739) |
| 0.1 | 0.1 | GO:0006102 | isocitrate metabolic process(GO:0006102) |
| 0.1 | 1.3 | GO:0072329 | monocarboxylic acid catabolic process(GO:0072329) |
| 0.1 | 0.3 | GO:0000078 | obsolete cytokinesis after mitosis checkpoint(GO:0000078) |
| 0.1 | 0.2 | GO:0097201 | negative regulation of transcription from RNA polymerase II promoter in response to stress(GO:0097201) |
| 0.1 | 0.9 | GO:0010648 | negative regulation of signal transduction(GO:0009968) negative regulation of cell communication(GO:0010648) negative regulation of signaling(GO:0023057) |
| 0.1 | 0.1 | GO:0071469 | response to alkaline pH(GO:0010446) cellular response to alkaline pH(GO:0071469) |
| 0.1 | 0.3 | GO:0006075 | (1->3)-beta-D-glucan biosynthetic process(GO:0006075) |
| 0.1 | 0.2 | GO:0006546 | glycine catabolic process(GO:0006546) |
| 0.1 | 0.1 | GO:2001023 | cellular response to drug(GO:0035690) regulation of response to drug(GO:2001023) positive regulation of response to drug(GO:2001025) regulation of cellular response to drug(GO:2001038) positive regulation of cellular response to drug(GO:2001040) |
| 0.1 | 0.3 | GO:0000196 | MAPK cascade involved in cell wall organization or biogenesis(GO:0000196) |
| 0.1 | 0.3 | GO:0032443 | regulation of steroid metabolic process(GO:0019218) regulation of ergosterol biosynthetic process(GO:0032443) regulation of steroid biosynthetic process(GO:0050810) |
| 0.1 | 0.4 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.1 | 0.6 | GO:0019878 | lysine biosynthetic process via aminoadipic acid(GO:0019878) |
| 0.1 | 0.4 | GO:0000023 | maltose metabolic process(GO:0000023) |
| 0.1 | 1.7 | GO:0006312 | mitotic recombination(GO:0006312) |
| 0.1 | 0.4 | GO:0008277 | regulation of G-protein coupled receptor protein signaling pathway(GO:0008277) |
| 0.1 | 0.4 | GO:0070911 | global genome nucleotide-excision repair(GO:0070911) |
| 0.1 | 0.2 | GO:0000715 | nucleotide-excision repair, DNA damage recognition(GO:0000715) |
| 0.1 | 2.0 | GO:0070591 | ascospore wall assembly(GO:0030476) spore wall assembly(GO:0042244) spore wall biogenesis(GO:0070590) ascospore wall biogenesis(GO:0070591) fungal-type cell wall assembly(GO:0071940) |
| 0.1 | 0.2 | GO:0010688 | negative regulation of ribosomal protein gene transcription from RNA polymerase II promoter(GO:0010688) |
| 0.1 | 0.2 | GO:0042843 | D-xylose catabolic process(GO:0042843) |
| 0.1 | 1.4 | GO:0006200 | obsolete ATP catabolic process(GO:0006200) |
| 0.1 | 0.7 | GO:0018208 | protein peptidyl-prolyl isomerization(GO:0000413) peptidyl-proline modification(GO:0018208) |
| 0.1 | 0.2 | GO:0045991 | carbon catabolite activation of transcription(GO:0045991) |
| 0.1 | 0.4 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.1 | 0.4 | GO:0070987 | error-free translesion synthesis(GO:0070987) |
| 0.0 | 0.1 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.0 | 0.1 | GO:0036228 | protein targeting to nuclear inner membrane(GO:0036228) |
| 0.0 | 0.1 | GO:0099515 | actin filament-based movement(GO:0030048) actin filament-based transport(GO:0099515) |
| 0.0 | 0.1 | GO:0044819 | mitotic G1 DNA damage checkpoint(GO:0031571) G1 DNA damage checkpoint(GO:0044783) mitotic G1/S transition checkpoint(GO:0044819) |
| 0.0 | 0.2 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.0 | 0.3 | GO:0070096 | mitochondrial outer membrane translocase complex assembly(GO:0070096) |
| 0.0 | 0.1 | GO:0070932 | histone H3 deacetylation(GO:0070932) histone H4 deacetylation(GO:0070933) |
| 0.0 | 0.2 | GO:0006598 | polyamine catabolic process(GO:0006598) |
| 0.0 | 0.1 | GO:0005985 | sucrose metabolic process(GO:0005985) sucrose catabolic process(GO:0005987) |
| 0.0 | 0.3 | GO:0009231 | riboflavin metabolic process(GO:0006771) riboflavin biosynthetic process(GO:0009231) |
| 0.0 | 0.7 | GO:0030474 | spindle pole body duplication(GO:0030474) |
| 0.0 | 0.4 | GO:0031990 | NLS-bearing protein import into nucleus(GO:0006607) mRNA export from nucleus in response to heat stress(GO:0031990) |
| 0.0 | 0.3 | GO:0000266 | mitochondrial fission(GO:0000266) |
| 0.0 | 0.9 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.0 | 0.4 | GO:0007266 | Rho protein signal transduction(GO:0007266) |
| 0.0 | 0.2 | GO:0000272 | polysaccharide catabolic process(GO:0000272) |
| 0.0 | 0.6 | GO:0005978 | glycogen biosynthetic process(GO:0005978) |
| 0.0 | 0.1 | GO:0043619 | regulation of transcription from RNA polymerase II promoter in response to oxidative stress(GO:0043619) |
| 0.0 | 0.2 | GO:0031204 | posttranslational protein targeting to membrane, translocation(GO:0031204) |
| 0.0 | 0.5 | GO:0042724 | thiamine biosynthetic process(GO:0009228) thiamine-containing compound biosynthetic process(GO:0042724) |
| 0.0 | 0.7 | GO:0008645 | hexose transport(GO:0008645) monosaccharide transport(GO:0015749) |
| 0.0 | 0.1 | GO:0042148 | strand invasion(GO:0042148) |
| 0.0 | 0.2 | GO:0032048 | cardiolipin metabolic process(GO:0032048) |
| 0.0 | 0.1 | GO:2000219 | positive regulation of invasive growth in response to glucose limitation by positive regulation of transcription from RNA polymerase II promoter(GO:0036095) positive regulation of invasive growth in response to glucose limitation(GO:2000219) |
| 0.0 | 0.8 | GO:0008361 | regulation of cell size(GO:0008361) |
| 0.0 | 0.2 | GO:0070646 | protein modification by small protein removal(GO:0070646) |
| 0.0 | 0.3 | GO:0000320 | re-entry into mitotic cell cycle(GO:0000320) re-entry into mitotic cell cycle after pheromone arrest(GO:0000321) |
| 0.0 | 0.1 | GO:0015883 | FAD transport(GO:0015883) |
| 0.0 | 1.0 | GO:0000282 | cellular bud site selection(GO:0000282) mitotic cytokinesis, site selection(GO:1902408) |
| 0.0 | 0.2 | GO:0046461 | triglyceride catabolic process(GO:0019433) neutral lipid catabolic process(GO:0046461) acylglycerol catabolic process(GO:0046464) |
| 0.0 | 0.4 | GO:0001558 | regulation of cell growth(GO:0001558) |
| 0.0 | 0.1 | GO:0097271 | protein localization to bud neck(GO:0097271) |
| 0.0 | 0.3 | GO:0001100 | negative regulation of exit from mitosis(GO:0001100) |
| 0.0 | 1.2 | GO:0009060 | aerobic respiration(GO:0009060) |
| 0.0 | 0.1 | GO:0070676 | intralumenal vesicle formation(GO:0070676) |
| 0.0 | 0.2 | GO:0000753 | cell morphogenesis involved in conjugation with cellular fusion(GO:0000753) |
| 0.0 | 0.1 | GO:0006279 | premeiotic DNA replication(GO:0006279) |
| 0.0 | 0.2 | GO:0070058 | tRNA gene clustering(GO:0070058) |
| 0.0 | 0.1 | GO:0060195 | antisense RNA transcription(GO:0009300) regulation of antisense RNA transcription(GO:0060194) negative regulation of antisense RNA transcription(GO:0060195) |
| 0.0 | 0.1 | GO:0052547 | negative regulation of peptidase activity(GO:0010466) regulation of peptidase activity(GO:0052547) |
| 0.0 | 0.2 | GO:0046685 | response to arsenic-containing substance(GO:0046685) |
| 0.0 | 0.1 | GO:0006545 | glycine biosynthetic process(GO:0006545) |
| 0.0 | 0.1 | GO:0048211 | Golgi vesicle docking(GO:0048211) |
| 0.0 | 0.3 | GO:0006081 | cellular aldehyde metabolic process(GO:0006081) |
| 0.0 | 0.3 | GO:0045005 | DNA-dependent DNA replication maintenance of fidelity(GO:0045005) |
| 0.0 | 0.1 | GO:0035437 | protein retention in ER lumen(GO:0006621) maintenance of protein localization in endoplasmic reticulum(GO:0035437) |
| 0.0 | 0.3 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.0 | 0.1 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 0.1 | GO:0000114 | obsolete regulation of transcription involved in G1 phase of mitotic cell cycle(GO:0000114) |
| 0.0 | 0.2 | GO:0009651 | response to salt stress(GO:0009651) |
| 0.0 | 0.1 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.0 | 0.1 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 0.0 | 0.3 | GO:0007129 | synapsis(GO:0007129) |
| 0.0 | 0.4 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.0 | 0.1 | GO:0046834 | lipid phosphorylation(GO:0046834) phosphatidylinositol phosphorylation(GO:0046854) |
| 0.0 | 0.1 | GO:0000920 | cell separation after cytokinesis(GO:0000920) |
| 0.0 | 0.1 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.0 | 0.1 | GO:0016233 | telomere capping(GO:0016233) |
| 0.0 | 0.1 | GO:0000350 | generation of catalytic spliceosome for second transesterification step(GO:0000350) |
| 0.0 | 0.1 | GO:0006490 | oligosaccharide-lipid intermediate biosynthetic process(GO:0006490) |
| 0.0 | 0.1 | GO:0036003 | positive regulation of transcription from RNA polymerase II promoter in response to stress(GO:0036003) |
| 0.0 | 0.0 | GO:0017006 | protein-heme linkage(GO:0017003) protein-tetrapyrrole linkage(GO:0017006) cytochrome c-heme linkage(GO:0018063) |
| 0.0 | 0.2 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.0 | 0.6 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.0 | 0.3 | GO:0030466 | chromatin silencing at silent mating-type cassette(GO:0030466) |
| 0.0 | 0.1 | GO:0045041 | protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.0 | 0.2 | GO:0000209 | protein polyubiquitination(GO:0000209) |
| 0.0 | 0.7 | GO:0006457 | protein folding(GO:0006457) |
| 0.0 | 0.1 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.0 | 0.1 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 0.1 | GO:0043461 | mitochondrial proton-transporting ATP synthase complex assembly(GO:0033615) proton-transporting ATP synthase complex assembly(GO:0043461) |
| 0.0 | 0.1 | GO:0006077 | (1->6)-beta-D-glucan metabolic process(GO:0006077) |
| 0.0 | 0.0 | GO:0071850 | mitotic cell cycle arrest in response to pheromone(GO:0000751) mitotic cell cycle arrest(GO:0071850) |
| 0.0 | 0.0 | GO:0031331 | positive regulation of cellular catabolic process(GO:0031331) |
| 0.0 | 0.2 | GO:0048278 | vesicle docking(GO:0048278) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 5.2 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.5 | 25.8 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.4 | 1.9 | GO:0045273 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.4 | 3.3 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) |
| 0.3 | 3.7 | GO:0005619 | ascospore wall(GO:0005619) |
| 0.3 | 0.8 | GO:0000113 | nucleotide-excision repair factor 4 complex(GO:0000113) |
| 0.2 | 0.5 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.2 | 1.9 | GO:0045277 | mitochondrial respiratory chain complex IV(GO:0005751) respiratory chain complex IV(GO:0045277) |
| 0.2 | 1.0 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.2 | 5.4 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.2 | 2.5 | GO:0098797 | plasma membrane protein complex(GO:0098797) |
| 0.2 | 0.5 | GO:0031422 | RecQ helicase-Topo III complex(GO:0031422) |
| 0.2 | 12.9 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.2 | 0.8 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.1 | 0.4 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.1 | 0.4 | GO:0001400 | mating projection base(GO:0001400) |
| 0.1 | 0.2 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.1 | 0.5 | GO:0005960 | glycine cleavage complex(GO:0005960) |
| 0.1 | 0.3 | GO:0005784 | Sec61 translocon complex(GO:0005784) |
| 0.1 | 0.3 | GO:0045265 | mitochondrial proton-transporting ATP synthase, stator stalk(GO:0000274) proton-transporting ATP synthase, stator stalk(GO:0045265) |
| 0.1 | 0.3 | GO:0035649 | Nrd1 complex(GO:0035649) |
| 0.1 | 2.3 | GO:0005628 | prospore membrane(GO:0005628) intracellular immature spore(GO:0042763) ascospore-type prospore(GO:0042764) |
| 0.1 | 0.5 | GO:0070210 | Rpd3L-Expanded complex(GO:0070210) |
| 0.1 | 0.4 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 0.4 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.1 | 1.0 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.1 | 0.8 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.1 | 0.8 | GO:0000974 | Prp19 complex(GO:0000974) |
| 0.1 | 1.1 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.1 | 0.3 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 1.1 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.1 | 1.3 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.1 | 8.6 | GO:0000329 | fungal-type vacuole membrane(GO:0000329) lytic vacuole membrane(GO:0098852) |
| 0.1 | 1.5 | GO:0000784 | nuclear chromosome, telomeric region(GO:0000784) |
| 0.1 | 0.5 | GO:0000112 | nucleotide-excision repair factor 3 complex(GO:0000112) |
| 0.1 | 1.5 | GO:0005774 | vacuolar membrane(GO:0005774) |
| 0.1 | 0.4 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.1 | 0.1 | GO:0032126 | eisosome(GO:0032126) |
| 0.1 | 0.7 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.1 | 0.4 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.1 | 0.4 | GO:0034657 | GID complex(GO:0034657) |
| 0.1 | 0.1 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.1 | 0.8 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 0.2 | GO:0032545 | CURI complex(GO:0032545) |
| 0.0 | 0.6 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 0.2 | GO:0031205 | endoplasmic reticulum Sec complex(GO:0031205) Sec62/Sec63 complex(GO:0031207) |
| 0.0 | 2.0 | GO:0000780 | condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.0 | 0.1 | GO:0033551 | monopolin complex(GO:0033551) |
| 0.0 | 0.1 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 0.7 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.2 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 0.1 | GO:0044697 | HICS complex(GO:0044697) |
| 0.0 | 11.6 | GO:0005886 | plasma membrane(GO:0005886) |
| 0.0 | 0.1 | GO:0035267 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 0.1 | GO:0030062 | mitochondrial tricarboxylic acid cycle enzyme complex(GO:0030062) |
| 0.0 | 0.2 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.0 | 0.5 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.0 | 0.6 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 0.2 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 0.2 | GO:0030915 | Smc5-Smc6 complex(GO:0030915) |
| 0.0 | 0.2 | GO:0005940 | septin ring(GO:0005940) |
| 0.0 | 0.1 | GO:0043541 | UDP-N-acetylglucosamine transferase complex(GO:0043541) |
| 0.0 | 6.8 | GO:0005740 | mitochondrial envelope(GO:0005740) |
| 0.0 | 0.2 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.0 | 0.2 | GO:0016514 | SWI/SNF complex(GO:0016514) BAF-type complex(GO:0090544) |
| 0.0 | 0.5 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.1 | GO:0000837 | Doa10p ubiquitin ligase complex(GO:0000837) |
| 0.0 | 0.3 | GO:0016586 | RSC complex(GO:0016586) |
| 0.0 | 0.2 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 0.0 | GO:0070390 | nucleotide-excision repair factor 2 complex(GO:0000111) transcription export complex 2(GO:0070390) |
| 0.0 | 0.3 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.5 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 0.1 | GO:0005686 | U2 snRNP(GO:0005686) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.4 | 13.8 | GO:0042927 | siderophore transmembrane transporter activity(GO:0015343) iron chelate transmembrane transporter activity(GO:0015603) siderophore transporter activity(GO:0042927) |
| 3.1 | 15.4 | GO:0015295 | solute:proton symporter activity(GO:0015295) |
| 0.9 | 3.7 | GO:0004737 | pyruvate decarboxylase activity(GO:0004737) |
| 0.7 | 2.1 | GO:0015114 | phosphate ion transmembrane transporter activity(GO:0015114) |
| 0.7 | 6.2 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) oxidoreductase activity, oxidizing metal ions, NAD or NADP as acceptor(GO:0016723) |
| 0.7 | 2.7 | GO:0005537 | mannose binding(GO:0005537) |
| 0.6 | 2.3 | GO:0004396 | hexokinase activity(GO:0004396) |
| 0.6 | 1.7 | GO:0048038 | quinone binding(GO:0048038) |
| 0.5 | 3.7 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.5 | 2.1 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.5 | 1.5 | GO:0004450 | isocitrate dehydrogenase (NADP+) activity(GO:0004450) |
| 0.4 | 4.7 | GO:0015662 | ATPase activity, coupled to transmembrane movement of ions, phosphorylative mechanism(GO:0015662) |
| 0.4 | 1.5 | GO:0000772 | mating pheromone activity(GO:0000772) pheromone activity(GO:0005186) |
| 0.4 | 1.1 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.3 | 1.4 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) ATP:ADP antiporter activity(GO:0005471) ADP transmembrane transporter activity(GO:0015217) |
| 0.3 | 0.9 | GO:0036440 | citrate (Si)-synthase activity(GO:0004108) citrate synthase activity(GO:0036440) |
| 0.3 | 0.9 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.3 | 1.1 | GO:0008198 | ferrous iron binding(GO:0008198) |
| 0.3 | 1.6 | GO:0051183 | vitamin transporter activity(GO:0051183) |
| 0.3 | 1.1 | GO:0001094 | TFIID-class transcription factor binding(GO:0001094) |
| 0.2 | 4.1 | GO:0016705 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen(GO:0016705) |
| 0.2 | 1.1 | GO:0004559 | alpha-mannosidase activity(GO:0004559) mannosidase activity(GO:0015923) |
| 0.2 | 0.6 | GO:0005536 | glucose binding(GO:0005536) |
| 0.2 | 2.7 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.2 | 2.0 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.2 | 0.4 | GO:0015151 | alpha-glucoside transmembrane transporter activity(GO:0015151) glucoside transmembrane transporter activity(GO:0042947) |
| 0.2 | 1.0 | GO:0018456 | aryl-alcohol dehydrogenase (NAD+) activity(GO:0018456) |
| 0.2 | 1.9 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.2 | 0.8 | GO:0022821 | potassium:proton antiporter activity(GO:0015386) potassium ion antiporter activity(GO:0022821) |
| 0.2 | 1.1 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 0.2 | 0.9 | GO:0008171 | O-methyltransferase activity(GO:0008171) |
| 0.2 | 0.4 | GO:0015294 | solute:cation symporter activity(GO:0015294) |
| 0.2 | 1.8 | GO:0005216 | ion channel activity(GO:0005216) |
| 0.2 | 1.0 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) aldehyde dehydrogenase [NAD(P)+] activity(GO:0004030) |
| 0.2 | 0.6 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.1 | 0.9 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.1 | 2.2 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.1 | 1.4 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.1 | 0.3 | GO:0015293 | symporter activity(GO:0015293) |
| 0.1 | 0.6 | GO:0016655 | oxidoreductase activity, acting on NAD(P)H, quinone or similar compound as acceptor(GO:0016655) |
| 0.1 | 2.0 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.1 | 1.3 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.1 | 7.0 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.1 | 0.5 | GO:0016642 | glycine dehydrogenase (decarboxylating) activity(GO:0004375) oxidoreductase activity, acting on the CH-NH2 group of donors, disulfide as acceptor(GO:0016642) |
| 0.1 | 0.3 | GO:0004458 | D-lactate dehydrogenase (cytochrome) activity(GO:0004458) |
| 0.1 | 2.1 | GO:0001191 | transcriptional repressor activity, RNA polymerase II transcription factor binding(GO:0001191) |
| 0.1 | 0.3 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.1 | 0.4 | GO:0016885 | ligase activity, forming carbon-carbon bonds(GO:0016885) |
| 0.1 | 0.2 | GO:0005102 | receptor binding(GO:0005102) |
| 0.1 | 3.7 | GO:0016820 | hydrolase activity, acting on acid anhydrides, catalyzing transmembrane movement of substances(GO:0016820) ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.1 | 0.2 | GO:0022838 | substrate-specific channel activity(GO:0022838) |
| 0.1 | 0.3 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.1 | 2.0 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.1 | 0.4 | GO:0000156 | phosphorelay response regulator activity(GO:0000156) |
| 0.1 | 0.3 | GO:0001097 | TFIIH-class transcription factor binding(GO:0001097) |
| 0.1 | 0.3 | GO:0000400 | four-way junction DNA binding(GO:0000400) |
| 0.1 | 0.4 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.1 | 0.3 | GO:0003843 | 1,3-beta-D-glucan synthase activity(GO:0003843) |
| 0.1 | 0.4 | GO:0000099 | sulfur amino acid transmembrane transporter activity(GO:0000099) |
| 0.1 | 0.3 | GO:0003916 | DNA topoisomerase activity(GO:0003916) |
| 0.1 | 1.0 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.1 | 6.0 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 0.4 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.1 | 0.4 | GO:0000014 | single-stranded DNA endodeoxyribonuclease activity(GO:0000014) |
| 0.1 | 0.5 | GO:0016888 | endodeoxyribonuclease activity, producing 5'-phosphomonoesters(GO:0016888) |
| 0.1 | 0.7 | GO:0016645 | oxidoreductase activity, acting on the CH-NH group of donors(GO:0016645) |
| 0.1 | 0.4 | GO:0015205 | nucleobase transmembrane transporter activity(GO:0015205) |
| 0.1 | 0.8 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.1 | 0.1 | GO:0016615 | malate dehydrogenase activity(GO:0016615) |
| 0.1 | 1.3 | GO:0004221 | obsolete ubiquitin thiolesterase activity(GO:0004221) |
| 0.1 | 0.7 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.1 | 0.6 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.1 | 0.1 | GO:0004448 | isocitrate dehydrogenase activity(GO:0004448) |
| 0.1 | 0.2 | GO:0008177 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) |
| 0.1 | 0.8 | GO:0020037 | heme binding(GO:0020037) tetrapyrrole binding(GO:0046906) |
| 0.1 | 0.1 | GO:0016208 | AMP binding(GO:0016208) |
| 0.1 | 0.3 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) ATP-dependent 5'-3' DNA helicase activity(GO:0043141) |
| 0.1 | 0.3 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.1 | 0.4 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.1 | 0.2 | GO:0097027 | cyclin binding(GO:0030332) ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.1 | 1.5 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.1 | 0.5 | GO:0008121 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors(GO:0016679) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.1 | 0.2 | GO:0097372 | NAD-dependent histone deacetylase activity (H3-K18 specific)(GO:0097372) |
| 0.1 | 1.7 | GO:0001076 | transcription factor activity, RNA polymerase II transcription factor binding(GO:0001076) |
| 0.0 | 0.2 | GO:0019783 | ubiquitin-like protein-specific protease activity(GO:0019783) |
| 0.0 | 0.7 | GO:0016859 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) cis-trans isomerase activity(GO:0016859) |
| 0.0 | 0.5 | GO:0060089 | receptor activity(GO:0004872) transmembrane signaling receptor activity(GO:0004888) signaling receptor activity(GO:0038023) molecular transducer activity(GO:0060089) transmembrane receptor activity(GO:0099600) |
| 0.0 | 0.1 | GO:0003951 | NAD+ kinase activity(GO:0003951) NADH kinase activity(GO:0042736) |
| 0.0 | 0.2 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.6 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.0 | 0.6 | GO:0015578 | fructose transmembrane transporter activity(GO:0005353) mannose transmembrane transporter activity(GO:0015578) |
| 0.0 | 0.5 | GO:0000384 | first spliceosomal transesterification activity(GO:0000384) |
| 0.0 | 0.5 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 0.4 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 0.3 | GO:0008641 | small protein activating enzyme activity(GO:0008641) |
| 0.0 | 0.6 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.0 | 0.5 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.1 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.0 | 0.3 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.0 | 0.9 | GO:0043021 | ribonucleoprotein complex binding(GO:0043021) |
| 0.0 | 0.3 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.0 | 0.1 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.0 | 0.1 | GO:0015230 | FAD transmembrane transporter activity(GO:0015230) |
| 0.0 | 0.2 | GO:0017022 | myosin binding(GO:0017022) |
| 0.0 | 0.1 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 0.2 | GO:0050253 | triglyceride lipase activity(GO:0004806) retinyl-palmitate esterase activity(GO:0050253) |
| 0.0 | 0.2 | GO:0004713 | protein serine/threonine/tyrosine kinase activity(GO:0004712) protein tyrosine kinase activity(GO:0004713) |
| 0.0 | 0.1 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.0 | 0.1 | GO:0043813 | phosphatidylinositol-3,5-bisphosphate 5-phosphatase activity(GO:0043813) |
| 0.0 | 0.1 | GO:0004564 | beta-fructofuranosidase activity(GO:0004564) sucrose alpha-glucosidase activity(GO:0004575) |
| 0.0 | 0.5 | GO:0008081 | phosphoric diester hydrolase activity(GO:0008081) |
| 0.0 | 0.4 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.0 | 0.7 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.2 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.1 | GO:0070567 | cytidylyltransferase activity(GO:0070567) |
| 0.0 | 0.1 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.0 | 0.1 | GO:0070336 | flap-structured DNA binding(GO:0070336) |
| 0.0 | 0.1 | GO:0016289 | CoA hydrolase activity(GO:0016289) |
| 0.0 | 0.6 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 0.1 | GO:0008236 | serine-type peptidase activity(GO:0008236) |
| 0.0 | 0.5 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.0 | 0.2 | GO:0033558 | histone deacetylase activity(GO:0004407) protein deacetylase activity(GO:0033558) |
| 0.0 | 0.2 | GO:0097472 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) cyclin-dependent protein kinase activity(GO:0097472) |
| 0.0 | 0.4 | GO:0061733 | histone acetyltransferase activity(GO:0004402) peptide-lysine-N-acetyltransferase activity(GO:0061733) |
| 0.0 | 0.3 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 0.0 | GO:0008409 | 5'-3' exonuclease activity(GO:0008409) |
| 0.0 | 0.1 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 0.0 | 0.1 | GO:0000033 | alpha-1,3-mannosyltransferase activity(GO:0000033) |
| 0.0 | 0.0 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.2 | GO:0005507 | copper ion binding(GO:0005507) |
| 0.0 | 0.0 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.8 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.2 | 0.7 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.1 | 0.5 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.1 | 0.5 | PID P73PATHWAY | p73 transcription factor network |
| 0.1 | 0.2 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.1 | 0.1 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.1 | 0.4 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.1 | 0.2 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.1 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 0.3 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 34.5 | PID ERBB1 DOWNSTREAM PATHWAY | ErbB1 downstream signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.1 | 0.5 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.1 | 0.9 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 0.4 | REACTOME BIOLOGICAL OXIDATIONS | Genes involved in Biological oxidations |
| 0.1 | 0.3 | REACTOME APOPTOTIC EXECUTION PHASE | Genes involved in Apoptotic execution phase |
| 0.1 | 0.1 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.5 | REACTOME DNA REPAIR | Genes involved in DNA Repair |
| 0.0 | 0.1 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 0.1 | REACTOME PYRUVATE METABOLISM AND CITRIC ACID TCA CYCLE | Genes involved in Pyruvate metabolism and Citric Acid (TCA) cycle |
| 0.0 | 34.3 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 0.1 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.2 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.1 | REACTOME TRANSMEMBRANE TRANSPORT OF SMALL MOLECULES | Genes involved in Transmembrane transport of small molecules |