Project
Mucociliary differentiation, bronchial epithelial cells, human (Ross 2007)
Navigation
Downloads
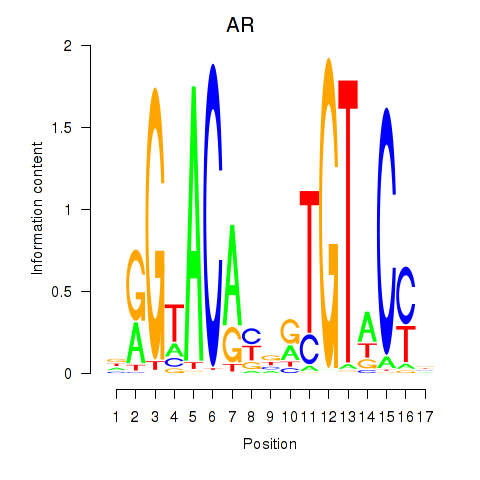
Results for AR_NR3C2
Z-value: 0.60
Transcription factors associated with AR_NR3C2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
AR
|
ENSG00000169083.11 | androgen receptor |
|
NR3C2
|
ENSG00000151623.10 | nuclear receptor subfamily 3 group C member 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR3C2 | hg19_v2_chr4_-_149365827_149365870 | -0.59 | 6.8e-04 | Click! |
| AR | hg19_v2_chrX_+_66764375_66764465 | -0.16 | 3.9e-01 | Click! |
Activity profile of AR_NR3C2 motif
Sorted Z-values of AR_NR3C2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_42916499 | 1.57 |
ENST00000601189.1
ENST00000599211.1 |
LIPE
|
lipase, hormone-sensitive |
| chr2_+_102953608 | 1.46 |
ENST00000311734.2
ENST00000409584.1 |
IL1RL1
|
interleukin 1 receptor-like 1 |
| chr4_+_75310851 | 1.26 |
ENST00000395748.3
ENST00000264487.2 |
AREG
|
amphiregulin |
| chr4_+_75311019 | 1.19 |
ENST00000502307.1
|
AREG
|
amphiregulin |
| chr4_+_75480629 | 1.17 |
ENST00000380846.3
|
AREGB
|
amphiregulin B |
| chr12_+_53443680 | 1.14 |
ENST00000314250.6
ENST00000451358.1 |
TENC1
|
tensin like C1 domain containing phosphatase (tensin 2) |
| chr12_+_53443963 | 1.13 |
ENST00000546602.1
ENST00000552570.1 ENST00000549700.1 |
TENC1
|
tensin like C1 domain containing phosphatase (tensin 2) |
| chr1_+_22979474 | 1.08 |
ENST00000509305.1
|
C1QB
|
complement component 1, q subcomponent, B chain |
| chr8_-_49833978 | 0.94 |
ENST00000020945.1
|
SNAI2
|
snail family zinc finger 2 |
| chr18_+_56530136 | 0.93 |
ENST00000591083.1
|
ZNF532
|
zinc finger protein 532 |
| chr1_+_22979676 | 0.91 |
ENST00000432749.2
ENST00000314933.6 |
C1QB
|
complement component 1, q subcomponent, B chain |
| chr10_+_696000 | 0.90 |
ENST00000381489.5
|
PRR26
|
proline rich 26 |
| chr16_+_56659687 | 0.88 |
ENST00000568293.1
ENST00000330439.6 |
MT1E
|
metallothionein 1E |
| chr16_+_56642041 | 0.86 |
ENST00000245185.5
|
MT2A
|
metallothionein 2A |
| chr8_-_49834299 | 0.83 |
ENST00000396822.1
|
SNAI2
|
snail family zinc finger 2 |
| chr11_+_18287721 | 0.78 |
ENST00000356524.4
|
SAA1
|
serum amyloid A1 |
| chr11_+_18287801 | 0.77 |
ENST00000532858.1
ENST00000405158.2 |
SAA1
|
serum amyloid A1 |
| chr16_+_56642489 | 0.74 |
ENST00000561491.1
|
MT2A
|
metallothionein 2A |
| chr7_-_94285402 | 0.74 |
ENST00000428696.2
ENST00000445866.2 |
SGCE
|
sarcoglycan, epsilon |
| chr7_-_94285511 | 0.65 |
ENST00000265735.7
|
SGCE
|
sarcoglycan, epsilon |
| chr1_-_24469602 | 0.60 |
ENST00000270800.1
|
IL22RA1
|
interleukin 22 receptor, alpha 1 |
| chr5_-_141392538 | 0.59 |
ENST00000503794.1
ENST00000510194.1 ENST00000504424.1 ENST00000513454.1 ENST00000458112.2 ENST00000542860.1 ENST00000503229.1 ENST00000500692.2 ENST00000311337.6 ENST00000504139.1 ENST00000505689.1 |
GNPDA1
|
glucosamine-6-phosphate deaminase 1 |
| chr8_-_23021533 | 0.53 |
ENST00000312584.3
|
TNFRSF10D
|
tumor necrosis factor receptor superfamily, member 10d, decoy with truncated death domain |
| chr16_+_56666563 | 0.50 |
ENST00000570233.1
|
MT1M
|
metallothionein 1M |
| chr2_-_231825668 | 0.50 |
ENST00000392039.2
|
GPR55
|
G protein-coupled receptor 55 |
| chr8_-_21669826 | 0.48 |
ENST00000517328.1
|
GFRA2
|
GDNF family receptor alpha 2 |
| chr3_-_98235962 | 0.46 |
ENST00000513873.1
|
CLDND1
|
claudin domain containing 1 |
| chr12_-_122241812 | 0.45 |
ENST00000538335.1
|
AC084018.1
|
AC084018.1 |
| chrX_+_17755563 | 0.45 |
ENST00000380045.3
ENST00000380041.3 ENST00000380043.3 ENST00000398080.1 |
SCML1
|
sex comb on midleg-like 1 (Drosophila) |
| chr14_+_105190514 | 0.43 |
ENST00000330877.2
|
ADSSL1
|
adenylosuccinate synthase like 1 |
| chr12_+_5153085 | 0.41 |
ENST00000252321.3
|
KCNA5
|
potassium voltage-gated channel, shaker-related subfamily, member 5 |
| chr18_-_48346298 | 0.38 |
ENST00000398439.3
|
MRO
|
maestro |
| chr19_+_35630628 | 0.38 |
ENST00000588715.1
ENST00000588607.1 |
FXYD1
|
FXYD domain containing ion transport regulator 1 |
| chr10_+_26932068 | 0.37 |
ENST00000544033.1
ENST00000454991.1 |
LINC00202-2
RP13-16H11.7
|
long intergenic non-protein coding RNA 202-2 RP13-16H11.7 |
| chr16_+_56716336 | 0.37 |
ENST00000394485.4
ENST00000562939.1 |
MT1X
|
metallothionein 1X |
| chr7_-_94285472 | 0.37 |
ENST00000437425.2
ENST00000447873.1 ENST00000415788.2 |
SGCE
|
sarcoglycan, epsilon |
| chr18_-_48346415 | 0.37 |
ENST00000431965.2
ENST00000436348.2 |
MRO
|
maestro |
| chrX_+_8432871 | 0.37 |
ENST00000381032.1
ENST00000453306.1 ENST00000444481.1 |
VCX3B
|
variable charge, X-linked 3B |
| chr19_+_35630926 | 0.36 |
ENST00000588081.1
ENST00000589121.1 |
FXYD1
|
FXYD domain containing ion transport regulator 1 |
| chr1_-_1149506 | 0.35 |
ENST00000379236.3
|
TNFRSF4
|
tumor necrosis factor receptor superfamily, member 4 |
| chr3_-_58563094 | 0.35 |
ENST00000464064.1
|
FAM107A
|
family with sequence similarity 107, member A |
| chr22_+_23046750 | 0.35 |
ENST00000390307.2
|
IGLV3-22
|
immunoglobulin lambda variable 3-22 (gene/pseudogene) |
| chr19_+_35630344 | 0.34 |
ENST00000455515.2
|
FXYD1
|
FXYD domain containing ion transport regulator 1 |
| chr16_+_32077386 | 0.33 |
ENST00000354689.6
|
IGHV3OR16-9
|
immunoglobulin heavy variable 3/OR16-9 (non-functional) |
| chr15_+_42694573 | 0.32 |
ENST00000397200.4
ENST00000569827.1 |
CAPN3
|
calpain 3, (p94) |
| chr1_+_1846519 | 0.32 |
ENST00000378604.3
|
CALML6
|
calmodulin-like 6 |
| chr3_-_108476231 | 0.31 |
ENST00000295755.6
|
RETNLB
|
resistin like beta |
| chr17_+_37824217 | 0.31 |
ENST00000394246.1
|
PNMT
|
phenylethanolamine N-methyltransferase |
| chr8_-_22014339 | 0.31 |
ENST00000306317.2
|
LGI3
|
leucine-rich repeat LGI family, member 3 |
| chr14_-_106845789 | 0.30 |
ENST00000390617.2
|
IGHV3-35
|
immunoglobulin heavy variable 3-35 (non-functional) |
| chrX_+_2609317 | 0.30 |
ENST00000381187.3
ENST00000381184.1 |
CD99
|
CD99 molecule |
| chr14_-_106668095 | 0.30 |
ENST00000390606.2
|
IGHV3-20
|
immunoglobulin heavy variable 3-20 |
| chr7_+_150147711 | 0.30 |
ENST00000307271.3
|
GIMAP8
|
GTPase, IMAP family member 8 |
| chr10_+_60759378 | 0.30 |
ENST00000432535.1
|
LINC00844
|
long intergenic non-protein coding RNA 844 |
| chr19_-_45657028 | 0.29 |
ENST00000429338.1
ENST00000589776.1 |
NKPD1
|
NTPase, KAP family P-loop domain containing 1 |
| chrX_+_2609356 | 0.28 |
ENST00000381180.3
ENST00000449611.1 |
CD99
|
CD99 molecule |
| chr6_-_111927062 | 0.28 |
ENST00000359831.4
|
TRAF3IP2
|
TRAF3 interacting protein 2 |
| chr2_+_242577097 | 0.28 |
ENST00000419606.1
ENST00000474739.2 ENST00000396411.3 ENST00000425239.1 ENST00000400771.3 ENST00000430617.2 |
ATG4B
|
autophagy related 4B, cysteine peptidase |
| chr8_+_38614807 | 0.28 |
ENST00000330691.6
ENST00000348567.4 |
TACC1
|
transforming, acidic coiled-coil containing protein 1 |
| chr16_+_56672571 | 0.26 |
ENST00000290705.8
|
MT1A
|
metallothionein 1A |
| chr14_-_106866934 | 0.26 |
ENST00000390618.2
|
IGHV3-38
|
immunoglobulin heavy variable 3-38 (non-functional) |
| chrX_+_2609207 | 0.25 |
ENST00000381192.3
|
CD99
|
CD99 molecule |
| chr12_+_66582919 | 0.23 |
ENST00000545837.1
ENST00000457197.2 |
IRAK3
|
interleukin-1 receptor-associated kinase 3 |
| chrY_-_16098393 | 0.23 |
ENST00000250825.4
|
VCY
|
variable charge, Y-linked |
| chr18_+_45778672 | 0.23 |
ENST00000600091.1
|
AC091150.1
|
HCG1818186; Uncharacterized protein |
| chr2_-_73298802 | 0.23 |
ENST00000411783.1
ENST00000410065.1 ENST00000442582.1 ENST00000272433.2 |
SFXN5
|
sideroflexin 5 |
| chrY_+_16168097 | 0.22 |
ENST00000250823.4
|
VCY1B
|
variable charge, Y-linked 1B |
| chr22_+_19939026 | 0.22 |
ENST00000406520.3
|
COMT
|
catechol-O-methyltransferase |
| chr19_+_56187987 | 0.21 |
ENST00000411543.2
|
EPN1
|
epsin 1 |
| chr18_-_48346130 | 0.21 |
ENST00000592966.1
|
MRO
|
maestro |
| chr16_+_67207838 | 0.21 |
ENST00000566871.1
ENST00000268605.7 |
NOL3
|
nucleolar protein 3 (apoptosis repressor with CARD domain) |
| chr9_-_106087316 | 0.20 |
ENST00000411575.1
|
RP11-341A22.2
|
RP11-341A22.2 |
| chr14_-_106552755 | 0.20 |
ENST00000390600.2
|
IGHV3-9
|
immunoglobulin heavy variable 3-9 |
| chr8_+_9046503 | 0.20 |
ENST00000512942.2
|
RP11-10A14.5
|
RP11-10A14.5 |
| chr17_-_16472483 | 0.20 |
ENST00000395824.1
ENST00000448349.2 ENST00000395825.3 |
ZNF287
|
zinc finger protein 287 |
| chr16_+_67207872 | 0.20 |
ENST00000563258.1
ENST00000568146.1 |
NOL3
|
nucleolar protein 3 (apoptosis repressor with CARD domain) |
| chr16_-_25269134 | 0.19 |
ENST00000328086.7
|
ZKSCAN2
|
zinc finger with KRAB and SCAN domains 2 |
| chr8_-_62602327 | 0.18 |
ENST00000445642.3
ENST00000517847.2 ENST00000389204.4 ENST00000517661.1 ENST00000517903.1 ENST00000522603.1 ENST00000522349.1 ENST00000522835.1 ENST00000541428.1 ENST00000518306.1 |
ASPH
|
aspartate beta-hydroxylase |
| chr19_-_3786253 | 0.18 |
ENST00000585778.1
|
MATK
|
megakaryocyte-associated tyrosine kinase |
| chr17_-_14140166 | 0.17 |
ENST00000420162.2
ENST00000431716.2 |
CDRT15
|
CMT1A duplicated region transcript 15 |
| chr16_+_25703274 | 0.17 |
ENST00000331351.5
|
HS3ST4
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 4 |
| chr19_-_3786354 | 0.16 |
ENST00000395040.2
ENST00000310132.6 |
MATK
|
megakaryocyte-associated tyrosine kinase |
| chr1_+_93811438 | 0.15 |
ENST00000370272.4
ENST00000370267.1 |
DR1
|
down-regulator of transcription 1, TBP-binding (negative cofactor 2) |
| chr7_+_94285637 | 0.15 |
ENST00000482108.1
ENST00000488574.1 |
PEG10
|
paternally expressed 10 |
| chr7_-_4877677 | 0.14 |
ENST00000538469.1
|
RADIL
|
Ras association and DIL domains |
| chr7_+_143701022 | 0.14 |
ENST00000408922.2
|
OR6B1
|
olfactory receptor, family 6, subfamily B, member 1 |
| chr14_+_22309368 | 0.14 |
ENST00000390433.1
|
TRAV12-1
|
T cell receptor alpha variable 12-1 |
| chr4_+_37892682 | 0.14 |
ENST00000508802.1
ENST00000261439.4 ENST00000402522.1 |
TBC1D1
|
TBC1 (tre-2/USP6, BUB2, cdc16) domain family, member 1 |
| chr14_-_45722605 | 0.14 |
ENST00000310806.4
|
MIS18BP1
|
MIS18 binding protein 1 |
| chr8_-_72756667 | 0.13 |
ENST00000325509.4
|
MSC
|
musculin |
| chrX_+_8433376 | 0.13 |
ENST00000440654.2
ENST00000381029.4 |
VCX3B
|
variable charge, X-linked 3B |
| chr19_-_8408139 | 0.12 |
ENST00000330915.3
ENST00000593649.1 ENST00000595639.1 |
KANK3
|
KN motif and ankyrin repeat domains 3 |
| chr4_-_69817481 | 0.12 |
ENST00000251566.4
|
UGT2A3
|
UDP glucuronosyltransferase 2 family, polypeptide A3 |
| chr11_+_236540 | 0.12 |
ENST00000532097.1
|
PSMD13
|
proteasome (prosome, macropain) 26S subunit, non-ATPase, 13 |
| chr13_-_26452679 | 0.12 |
ENST00000596729.1
|
AL138815.2
|
Uncharacterized protein |
| chrX_+_79270245 | 0.12 |
ENST00000373296.3
ENST00000442340.1 |
TBX22
|
T-box 22 |
| chr12_-_11002063 | 0.12 |
ENST00000544994.1
ENST00000228811.4 ENST00000540107.1 |
PRR4
|
proline rich 4 (lacrimal) |
| chr4_+_87928140 | 0.12 |
ENST00000307808.6
|
AFF1
|
AF4/FMR2 family, member 1 |
| chr1_-_209979465 | 0.11 |
ENST00000542854.1
|
IRF6
|
interferon regulatory factor 6 |
| chr3_-_24207039 | 0.11 |
ENST00000280696.5
|
THRB
|
thyroid hormone receptor, beta |
| chr3_-_190167571 | 0.10 |
ENST00000354905.2
|
TMEM207
|
transmembrane protein 207 |
| chr8_+_62200509 | 0.10 |
ENST00000519846.1
ENST00000518592.1 ENST00000325897.4 |
CLVS1
|
clavesin 1 |
| chr22_+_24030321 | 0.10 |
ENST00000401461.1
|
RGL4
|
ral guanine nucleotide dissociation stimulator-like 4 |
| chr4_-_186347099 | 0.10 |
ENST00000505357.1
ENST00000264689.6 |
UFSP2
|
UFM1-specific peptidase 2 |
| chr19_-_14117074 | 0.10 |
ENST00000588885.1
ENST00000254325.4 |
RFX1
|
regulatory factor X, 1 (influences HLA class II expression) |
| chr15_-_20193370 | 0.10 |
ENST00000558565.2
|
IGHV3OR15-7
|
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr17_+_38333263 | 0.10 |
ENST00000456989.2
ENST00000543876.1 ENST00000544503.1 ENST00000264644.6 ENST00000538884.1 |
RAPGEFL1
|
Rap guanine nucleotide exchange factor (GEF)-like 1 |
| chr8_-_145634731 | 0.09 |
ENST00000349769.3
|
CPSF1
|
cleavage and polyadenylation specific factor 1, 160kDa |
| chr9_+_131683174 | 0.09 |
ENST00000372592.3
ENST00000428610.1 |
PHYHD1
|
phytanoyl-CoA dioxygenase domain containing 1 |
| chr16_-_67978016 | 0.09 |
ENST00000264005.5
|
LCAT
|
lecithin-cholesterol acyltransferase |
| chr12_+_125549973 | 0.08 |
ENST00000536752.1
ENST00000261686.6 |
AACS
|
acetoacetyl-CoA synthetase |
| chr19_+_54704610 | 0.08 |
ENST00000302907.4
|
RPS9
|
ribosomal protein S9 |
| chr11_-_506739 | 0.08 |
ENST00000529306.1
ENST00000438658.2 ENST00000527485.1 ENST00000397615.2 ENST00000397614.1 |
RNH1
|
ribonuclease/angiogenin inhibitor 1 |
| chr12_+_1099675 | 0.07 |
ENST00000545318.2
|
ERC1
|
ELKS/RAB6-interacting/CAST family member 1 |
| chr1_-_145715565 | 0.07 |
ENST00000369288.2
ENST00000369290.1 ENST00000401557.3 |
CD160
|
CD160 molecule |
| chr18_+_7754957 | 0.07 |
ENST00000400053.4
|
PTPRM
|
protein tyrosine phosphatase, receptor type, M |
| chr17_-_61996136 | 0.07 |
ENST00000342364.4
|
GH1
|
growth hormone 1 |
| chr14_-_107211459 | 0.07 |
ENST00000390636.2
|
IGHV3-73
|
immunoglobulin heavy variable 3-73 |
| chr2_+_27435179 | 0.07 |
ENST00000606999.1
ENST00000405489.3 |
ATRAID
|
all-trans retinoic acid-induced differentiation factor |
| chr9_+_71650645 | 0.06 |
ENST00000396366.2
|
FXN
|
frataxin |
| chr11_+_114271251 | 0.06 |
ENST00000375490.5
|
RBM7
|
RNA binding motif protein 7 |
| chr11_+_48002076 | 0.06 |
ENST00000418331.2
ENST00000440289.2 |
PTPRJ
|
protein tyrosine phosphatase, receptor type, J |
| chr20_+_45947246 | 0.06 |
ENST00000599904.1
|
AL031666.2
|
HCG2018772; Uncharacterized protein; cDNA FLJ31609 fis, clone NT2RI2002852 |
| chr6_+_123317116 | 0.06 |
ENST00000275162.5
|
CLVS2
|
clavesin 2 |
| chr8_+_24298597 | 0.05 |
ENST00000380789.1
|
ADAM7
|
ADAM metallopeptidase domain 7 |
| chr4_+_8442496 | 0.05 |
ENST00000389737.4
ENST00000504134.1 |
TRMT44
|
tRNA methyltransferase 44 homolog (S. cerevisiae) |
| chr13_+_113777105 | 0.05 |
ENST00000409306.1
ENST00000375551.3 ENST00000375559.3 |
F10
|
coagulation factor X |
| chr12_+_125549925 | 0.05 |
ENST00000316519.6
|
AACS
|
acetoacetyl-CoA synthetase |
| chr2_+_27435734 | 0.04 |
ENST00000419744.1
|
ATRAID
|
all-trans retinoic acid-induced differentiation factor |
| chr4_-_8442438 | 0.03 |
ENST00000356406.5
ENST00000413009.2 |
ACOX3
|
acyl-CoA oxidase 3, pristanoyl |
| chrX_+_153029633 | 0.03 |
ENST00000538966.1
ENST00000361971.5 ENST00000538776.1 ENST00000538543.1 |
PLXNB3
|
plexin B3 |
| chr2_-_109605663 | 0.03 |
ENST00000409271.1
ENST00000258443.2 ENST00000376651.1 |
EDAR
|
ectodysplasin A receptor |
| chr14_-_23623577 | 0.02 |
ENST00000422941.2
ENST00000453702.1 |
SLC7A8
|
solute carrier family 7 (amino acid transporter light chain, L system), member 8 |
| chr8_+_24298531 | 0.02 |
ENST00000175238.6
|
ADAM7
|
ADAM metallopeptidase domain 7 |
| chr19_+_58570605 | 0.02 |
ENST00000359978.6
ENST00000401053.4 ENST00000439855.2 ENST00000313434.5 ENST00000511556.1 ENST00000506786.1 |
ZNF135
|
zinc finger protein 135 |
| chr11_+_114270752 | 0.01 |
ENST00000540163.1
|
RBM7
|
RNA binding motif protein 7 |
| chr11_-_506316 | 0.01 |
ENST00000532055.1
ENST00000531540.1 |
RNH1
|
ribonuclease/angiogenin inhibitor 1 |
| chr12_-_56615693 | 0.01 |
ENST00000394013.2
ENST00000345093.4 ENST00000551711.1 ENST00000552656.1 |
RNF41
|
ring finger protein 41 |
| chr16_+_32063311 | 0.00 |
ENST00000426099.1
|
AC142381.1
|
AC142381.1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of AR_NR3C2
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.8 | GO:0070563 | negative regulation of vitamin D receptor signaling pathway(GO:0070563) |
| 0.3 | 1.6 | GO:1990169 | detoxification of copper ion(GO:0010273) stress response to copper ion(GO:1990169) |
| 0.3 | 1.6 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.3 | 2.4 | GO:0042695 | thelarche(GO:0042695) mammary gland branching involved in thelarche(GO:0060744) |
| 0.2 | 1.5 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 0.2 | 1.1 | GO:0010731 | protein glutathionylation(GO:0010731) regulation of protein glutathionylation(GO:0010732) negative regulation of protein glutathionylation(GO:0010734) |
| 0.1 | 0.4 | GO:0014876 | response to injury involved in regulation of muscle adaptation(GO:0014876) |
| 0.1 | 0.6 | GO:0006041 | glucosamine metabolic process(GO:0006041) |
| 0.1 | 0.4 | GO:0044208 | 'de novo' AMP biosynthetic process(GO:0044208) |
| 0.1 | 0.3 | GO:0042418 | epinephrine metabolic process(GO:0042414) epinephrine biosynthetic process(GO:0042418) |
| 0.1 | 0.4 | GO:0036018 | cellular response to erythropoietin(GO:0036018) |
| 0.1 | 0.3 | GO:0014718 | positive regulation of satellite cell activation involved in skeletal muscle regeneration(GO:0014718) |
| 0.1 | 0.4 | GO:0060372 | regulation of atrial cardiac muscle cell membrane repolarization(GO:0060372) |
| 0.1 | 1.4 | GO:0071294 | cellular response to zinc ion(GO:0071294) |
| 0.0 | 0.2 | GO:0010936 | negative regulation of macrophage cytokine production(GO:0010936) |
| 0.0 | 0.2 | GO:0018197 | peptidyl-aspartic acid modification(GO:0018197) peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.0 | 0.2 | GO:0016036 | cellular response to phosphate starvation(GO:0016036) positive regulation of sulfur amino acid metabolic process(GO:0031337) positive regulation of homocysteine metabolic process(GO:0050668) |
| 0.0 | 0.1 | GO:0014707 | branchiomeric skeletal muscle development(GO:0014707) |
| 0.0 | 0.2 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.0 | 0.5 | GO:0038171 | cannabinoid signaling pathway(GO:0038171) |
| 0.0 | 0.2 | GO:0006842 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 0.0 | 0.5 | GO:0035860 | glial cell-derived neurotrophic factor receptor signaling pathway(GO:0035860) |
| 0.0 | 0.1 | GO:0008050 | female courtship behavior(GO:0008050) |
| 0.0 | 1.5 | GO:0050716 | positive regulation of interleukin-1 secretion(GO:0050716) |
| 0.0 | 3.5 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.0 | 0.1 | GO:0090107 | regulation of high-density lipoprotein particle assembly(GO:0090107) |
| 0.0 | 0.4 | GO:0051024 | positive regulation of immunoglobulin secretion(GO:0051024) |
| 0.0 | 0.1 | GO:0034201 | response to oleic acid(GO:0034201) |
| 0.0 | 0.1 | GO:0018282 | metal incorporation into metallo-sulfur cluster(GO:0018282) iron incorporation into metallo-sulfur cluster(GO:0018283) |
| 0.0 | 0.0 | GO:0010593 | negative regulation of lamellipodium assembly(GO:0010593) |
| 0.0 | 0.3 | GO:0048305 | immunoglobulin secretion(GO:0048305) |
| 0.0 | 0.1 | GO:0045903 | positive regulation of translational fidelity(GO:0045903) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.0 | GO:0005602 | complement component C1 complex(GO:0005602) |
| 0.2 | 1.8 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.1 | 1.5 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 1.1 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.0 | 1.3 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 2.4 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.2 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 2.0 | GO:0005901 | caveola(GO:0005901) |
| 0.0 | 0.1 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 0.0 | 0.1 | GO:0008541 | proteasome regulatory particle, lid subcomplex(GO:0008541) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.6 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.5 | 1.5 | GO:0002113 | interleukin-33 binding(GO:0002113) |
| 0.1 | 0.4 | GO:0004019 | adenylosuccinate synthase activity(GO:0004019) |
| 0.1 | 1.6 | GO:0046870 | cadmium ion binding(GO:0046870) |
| 0.1 | 0.3 | GO:0004603 | phenylethanolamine N-methyltransferase activity(GO:0004603) |
| 0.1 | 0.6 | GO:0042015 | interleukin-20 binding(GO:0042015) |
| 0.1 | 0.5 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.1 | 0.5 | GO:0045569 | TRAIL binding(GO:0045569) |
| 0.1 | 0.5 | GO:0016167 | glial cell-derived neurotrophic factor receptor activity(GO:0016167) |
| 0.1 | 0.4 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 0.1 | 0.2 | GO:0004597 | peptide-aspartate beta-dioxygenase activity(GO:0004597) |
| 0.0 | 0.6 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.0 | 2.4 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.4 | GO:0035877 | death effector domain binding(GO:0035877) |
| 0.0 | 0.2 | GO:0015137 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.0 | 1.5 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.0 | 0.2 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.0 | 1.3 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.1 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.0 | 0.3 | GO:0031432 | titin binding(GO:0031432) |
| 0.0 | 0.2 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.0 | 1.1 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 2.0 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 0.1 | GO:0047760 | butyrate-CoA ligase activity(GO:0047760) |
| 0.0 | 0.1 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.0 | 0.4 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.4 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 1.5 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 2.1 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 0.5 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.0 | 1.6 | PID AP1 PATHWAY | AP-1 transcription factor network |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.0 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.1 | 1.6 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 1.5 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.0 | 0.4 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.0 | 1.7 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 0.3 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |