Project
Epithelial-Mesenchymal Transition, human (Scheel, 2011)
Navigation
Downloads
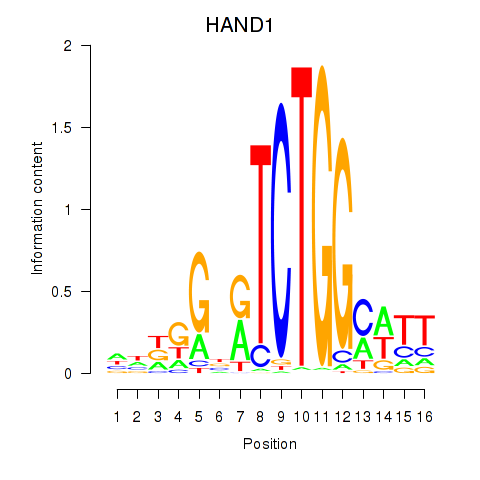
Results for HAND1
Z-value: 0.25
Transcription factors associated with HAND1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HAND1
|
ENSG00000113196.2 | HAND1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HAND1 | hg19_v2_chr5_-_153857819_153857824 | 0.02 | 9.5e-01 | Click! |
Activity profile of HAND1 motif
Sorted Z-values of HAND1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HAND1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_+_141103634 | 0.16 |
ENST00000507722.1 |
ZBTB38 |
zinc finger and BTB domain containing 38 |
| chr19_+_12862604 | 0.16 |
ENST00000553030.1 |
BEST2 |
bestrophin 2 |
| chr19_+_12862486 | 0.15 |
ENST00000549706.1 |
BEST2 |
bestrophin 2 |
| chr19_+_36359341 | 0.15 |
ENST00000221891.4 |
APLP1 |
amyloid beta (A4) precursor-like protein 1 |
| chr10_+_99344071 | 0.14 |
ENST00000370647.4 ENST00000370646.4 |
HOGA1 |
4-hydroxy-2-oxoglutarate aldolase 1 |
| chr3_+_148415571 | 0.13 |
ENST00000497524.1 ENST00000349243.3 ENST00000542281.1 ENST00000418473.2 ENST00000404754.2 |
AGTR1 |
angiotensin II receptor, type 1 |
| chr14_-_53331239 | 0.11 |
ENST00000553663.1 |
FERMT2 |
fermitin family member 2 |
| chr5_-_147286065 | 0.11 |
ENST00000318315.4 ENST00000515291.1 |
C5orf46 |
chromosome 5 open reading frame 46 |
| chr2_+_210444142 | 0.10 |
ENST00000360351.4 ENST00000361559.4 |
MAP2 |
microtubule-associated protein 2 |
| chr20_-_56285595 | 0.10 |
ENST00000395816.3 ENST00000347215.4 |
PMEPA1 |
prostate transmembrane protein, androgen induced 1 |
| chr20_-_56286479 | 0.10 |
ENST00000265626.4 |
PMEPA1 |
prostate transmembrane protein, androgen induced 1 |
| chr20_-_1165117 | 0.10 |
ENST00000381894.3 |
TMEM74B |
transmembrane protein 74B |
| chr17_-_64216748 | 0.09 |
ENST00000585162.1 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chr6_-_39282221 | 0.09 |
ENST00000453413.2 |
KCNK17 |
potassium channel, subfamily K, member 17 |
| chr6_-_39282329 | 0.08 |
ENST00000373231.4 |
KCNK17 |
potassium channel, subfamily K, member 17 |
| chr1_+_110036728 | 0.08 |
ENST00000369868.3 ENST00000430195.2 |
CYB561D1 |
cytochrome b561 family, member D1 |
| chrX_+_149531524 | 0.08 |
ENST00000370401.2 |
MAMLD1 |
mastermind-like domain containing 1 |
| chr6_+_6588902 | 0.08 |
ENST00000230568.4 |
LY86 |
lymphocyte antigen 86 |
| chrX_+_49832231 | 0.08 |
ENST00000376108.3 |
CLCN5 |
chloride channel, voltage-sensitive 5 |
| chr1_+_164528866 | 0.07 |
ENST00000420696.2 |
PBX1 |
pre-B-cell leukemia homeobox 1 |
| chr12_+_110338063 | 0.07 |
ENST00000405876.4 |
TCHP |
trichoplein, keratin filament binding |
| chr1_+_65775204 | 0.07 |
ENST00000371069.4 |
DNAJC6 |
DnaJ (Hsp40) homolog, subfamily C, member 6 |
| chrX_-_34675391 | 0.07 |
ENST00000275954.3 |
TMEM47 |
transmembrane protein 47 |
| chr6_+_124125286 | 0.07 |
ENST00000368416.1 ENST00000368417.1 ENST00000546092.1 |
NKAIN2 |
Na+/K+ transporting ATPase interacting 2 |
| chr2_+_228678550 | 0.07 |
ENST00000409189.3 ENST00000358813.4 |
CCL20 |
chemokine (C-C motif) ligand 20 |
| chr1_+_110036699 | 0.06 |
ENST00000496961.1 ENST00000533024.1 ENST00000310611.4 ENST00000527072.1 ENST00000420578.2 ENST00000528785.1 |
CYB561D1 |
cytochrome b561 family, member D1 |
| chr8_+_79428539 | 0.06 |
ENST00000352966.5 |
PKIA |
protein kinase (cAMP-dependent, catalytic) inhibitor alpha |
| chr7_-_142207004 | 0.06 |
ENST00000426318.2 |
TRBV10-2 |
T cell receptor beta variable 10-2 |
| chr1_-_32210275 | 0.06 |
ENST00000440175.2 |
BAI2 |
brain-specific angiogenesis inhibitor 2 |
| chr6_-_87804815 | 0.06 |
ENST00000369582.2 |
CGA |
glycoprotein hormones, alpha polypeptide |
| chr2_-_197791441 | 0.06 |
ENST00000409475.1 ENST00000354764.4 ENST00000374738.3 |
PGAP1 |
post-GPI attachment to proteins 1 |
| chr10_+_72972281 | 0.06 |
ENST00000335350.6 |
UNC5B |
unc-5 homolog B (C. elegans) |
| chr16_-_10652993 | 0.06 |
ENST00000536829.1 |
EMP2 |
epithelial membrane protein 2 |
| chr2_+_201997676 | 0.06 |
ENST00000462763.1 ENST00000479953.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr11_+_67171358 | 0.05 |
ENST00000526387.1 |
TBC1D10C |
TBC1 domain family, member 10C |
| chr11_+_67171391 | 0.05 |
ENST00000312390.5 |
TBC1D10C |
TBC1 domain family, member 10C |
| chr5_-_41870621 | 0.05 |
ENST00000196371.5 |
OXCT1 |
3-oxoacid CoA transferase 1 |
| chr11_+_67171548 | 0.05 |
ENST00000542590.1 |
TBC1D10C |
TBC1 domain family, member 10C |
| chr12_+_121131970 | 0.05 |
ENST00000535656.1 |
MLEC |
malectin |
| chr16_-_89556942 | 0.05 |
ENST00000301030.4 |
ANKRD11 |
ankyrin repeat domain 11 |
| chr7_-_81399411 | 0.05 |
ENST00000423064.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr11_-_2323290 | 0.05 |
ENST00000381153.3 |
C11orf21 |
chromosome 11 open reading frame 21 |
| chr19_-_10213335 | 0.05 |
ENST00000592641.1 ENST00000253109.4 |
ANGPTL6 |
angiopoietin-like 6 |
| chr3_-_114035026 | 0.05 |
ENST00000570269.1 |
RP11-553L6.5 |
RP11-553L6.5 |
| chr6_+_134210243 | 0.05 |
ENST00000367882.4 |
TCF21 |
transcription factor 21 |
| chr20_+_29956369 | 0.05 |
ENST00000253381.2 |
DEFB118 |
defensin, beta 118 |
| chr9_+_72658490 | 0.04 |
ENST00000377182.4 |
MAMDC2 |
MAM domain containing 2 |
| chr9_-_35828576 | 0.04 |
ENST00000377984.2 ENST00000423537.2 ENST00000472182.1 |
FAM221B |
family with sequence similarity 221, member B |
| chr7_-_116963334 | 0.04 |
ENST00000265441.3 |
WNT2 |
wingless-type MMTV integration site family member 2 |
| chr22_+_31518938 | 0.04 |
ENST00000412985.1 ENST00000331075.5 ENST00000412277.2 ENST00000420017.1 ENST00000400294.2 ENST00000405300.1 ENST00000404390.3 |
INPP5J |
inositol polyphosphate-5-phosphatase J |
| chr5_+_167913450 | 0.04 |
ENST00000231572.3 ENST00000538719.1 |
RARS |
arginyl-tRNA synthetase |
| chr9_-_25678856 | 0.04 |
ENST00000358022.3 |
TUSC1 |
tumor suppressor candidate 1 |
| chr11_+_101785727 | 0.04 |
ENST00000263468.8 |
KIAA1377 |
KIAA1377 |
| chr12_+_26111823 | 0.04 |
ENST00000381352.3 ENST00000535907.1 ENST00000405154.2 |
RASSF8 |
Ras association (RalGDS/AF-6) domain family (N-terminal) member 8 |
| chr1_-_11007927 | 0.04 |
ENST00000468348.1 |
C1orf127 |
chromosome 1 open reading frame 127 |
| chr4_+_41614720 | 0.04 |
ENST00000509277.1 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr1_-_214638146 | 0.04 |
ENST00000543945.1 |
PTPN14 |
protein tyrosine phosphatase, non-receptor type 14 |
| chr11_-_64527425 | 0.03 |
ENST00000377432.3 |
PYGM |
phosphorylase, glycogen, muscle |
| chr1_-_155904187 | 0.03 |
ENST00000368321.3 ENST00000368320.3 |
KIAA0907 |
KIAA0907 |
| chr2_+_231860830 | 0.03 |
ENST00000424440.1 ENST00000452881.1 ENST00000433428.2 ENST00000455816.1 ENST00000440792.1 ENST00000423134.1 |
SPATA3 |
spermatogenesis associated 3 |
| chr14_+_74083548 | 0.03 |
ENST00000381139.1 |
ACOT6 |
acyl-CoA thioesterase 6 |
| chr8_-_93029520 | 0.03 |
ENST00000521553.1 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr7_-_2883928 | 0.03 |
ENST00000275364.3 |
GNA12 |
guanine nucleotide binding protein (G protein) alpha 12 |
| chr12_-_57630873 | 0.03 |
ENST00000556732.1 |
NDUFA4L2 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 4-like 2 |
| chr3_-_157251383 | 0.03 |
ENST00000487753.1 ENST00000489602.1 ENST00000461299.1 ENST00000479987.1 |
VEPH1 |
ventricular zone expressed PH domain-containing 1 |
| chr1_+_27719148 | 0.03 |
ENST00000374024.3 |
GPR3 |
G protein-coupled receptor 3 |
| chr5_+_143584814 | 0.03 |
ENST00000507359.3 |
KCTD16 |
potassium channel tetramerization domain containing 16 |
| chr10_+_118350468 | 0.03 |
ENST00000358834.4 ENST00000528052.1 ENST00000442761.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr10_+_118350522 | 0.03 |
ENST00000530319.1 ENST00000527980.1 ENST00000471549.1 ENST00000534537.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr7_+_76139833 | 0.03 |
ENST00000257632.5 |
UPK3B |
uroplakin 3B |
| chr6_+_44191507 | 0.03 |
ENST00000371724.1 ENST00000371713.1 |
SLC29A1 |
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr14_-_106642049 | 0.03 |
ENST00000390605.2 |
IGHV1-18 |
immunoglobulin heavy variable 1-18 |
| chr2_+_201997492 | 0.02 |
ENST00000494258.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr8_-_93029865 | 0.02 |
ENST00000422361.2 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr12_-_7261772 | 0.02 |
ENST00000545280.1 ENST00000543933.1 ENST00000545337.1 ENST00000544702.1 ENST00000266542.4 |
C1RL |
complement component 1, r subcomponent-like |
| chr11_-_26743546 | 0.02 |
ENST00000280467.6 ENST00000396005.3 |
SLC5A12 |
solute carrier family 5 (sodium/monocarboxylate cotransporter), member 12 |
| chr19_-_55866061 | 0.02 |
ENST00000588572.2 ENST00000593184.1 ENST00000589467.1 |
COX6B2 |
cytochrome c oxidase subunit VIb polypeptide 2 (testis) |
| chr2_+_201997595 | 0.02 |
ENST00000470178.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr3_-_20227619 | 0.02 |
ENST00000425061.1 ENST00000443724.1 ENST00000421451.1 ENST00000452020.1 ENST00000417364.1 ENST00000306698.2 ENST00000419233.2 ENST00000263753.4 ENST00000383774.1 ENST00000437051.1 ENST00000412868.1 ENST00000429446.3 ENST00000442720.1 |
SGOL1 |
shugoshin-like 1 (S. pombe) |
| chr17_-_36358166 | 0.02 |
ENST00000537432.1 |
TBC1D3 |
TBC1 domain family, member 3 |
| chr17_+_45908974 | 0.02 |
ENST00000269025.4 |
LRRC46 |
leucine rich repeat containing 46 |
| chr2_-_128784846 | 0.02 |
ENST00000259235.3 ENST00000357702.5 ENST00000424298.1 |
SAP130 |
Sin3A-associated protein, 130kDa |
| chr17_+_53342311 | 0.02 |
ENST00000226067.5 |
HLF |
hepatic leukemia factor |
| chr10_+_82297658 | 0.02 |
ENST00000339284.2 |
SH2D4B |
SH2 domain containing 4B |
| chr10_-_118032697 | 0.02 |
ENST00000439649.3 |
GFRA1 |
GDNF family receptor alpha 1 |
| chr2_-_159237472 | 0.02 |
ENST00000409187.1 |
CCDC148 |
coiled-coil domain containing 148 |
| chrX_+_70364667 | 0.02 |
ENST00000536169.1 ENST00000395855.2 ENST00000374051.3 ENST00000358741.3 |
NLGN3 |
neuroligin 3 |
| chr11_+_2323349 | 0.02 |
ENST00000381121.3 |
TSPAN32 |
tetraspanin 32 |
| chr20_+_22034809 | 0.02 |
ENST00000449427.1 |
RP11-125P18.1 |
RP11-125P18.1 |
| chr14_-_74892805 | 0.02 |
ENST00000331628.3 ENST00000554953.1 |
SYNDIG1L |
synapse differentiation inducing 1-like |
| chr20_+_123010 | 0.02 |
ENST00000382398.3 |
DEFB126 |
defensin, beta 126 |
| chr1_+_202091980 | 0.02 |
ENST00000367282.5 |
GPR37L1 |
G protein-coupled receptor 37 like 1 |
| chr6_+_6588316 | 0.02 |
ENST00000379953.2 |
LY86 |
lymphocyte antigen 86 |
| chr7_+_140378955 | 0.02 |
ENST00000473512.1 |
ADCK2 |
aarF domain containing kinase 2 |
| chr19_+_47421933 | 0.02 |
ENST00000404338.3 |
ARHGAP35 |
Rho GTPase activating protein 35 |
| chr21_+_34602377 | 0.01 |
ENST00000342101.3 ENST00000413881.1 ENST00000443073.1 |
IFNAR2 |
interferon (alpha, beta and omega) receptor 2 |
| chr1_-_52344471 | 0.01 |
ENST00000352171.7 ENST00000354831.7 |
NRD1 |
nardilysin (N-arginine dibasic convertase) |
| chr6_-_53013620 | 0.01 |
ENST00000259803.7 |
GCM1 |
glial cells missing homolog 1 (Drosophila) |
| chr12_+_68042517 | 0.01 |
ENST00000393555.3 |
DYRK2 |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 2 |
| chr15_-_31393910 | 0.01 |
ENST00000397795.2 ENST00000256552.6 ENST00000559179.1 |
TRPM1 |
transient receptor potential cation channel, subfamily M, member 1 |
| chr1_+_215747118 | 0.01 |
ENST00000448333.1 |
KCTD3 |
potassium channel tetramerization domain containing 3 |
| chr12_+_68042495 | 0.01 |
ENST00000344096.3 |
DYRK2 |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 2 |
| chr4_-_40632605 | 0.01 |
ENST00000514014.1 |
RBM47 |
RNA binding motif protein 47 |
| chr16_+_6069664 | 0.01 |
ENST00000422070.4 |
RBFOX1 |
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr12_-_53730147 | 0.01 |
ENST00000536324.2 |
SP7 |
Sp7 transcription factor |
| chr16_+_70328680 | 0.01 |
ENST00000563206.1 ENST00000451014.3 ENST00000568625.1 |
DDX19B |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 19B |
| chr17_+_7836398 | 0.01 |
ENST00000565740.1 |
CNTROB |
centrobin, centrosomal BRCA2 interacting protein |
| chr17_+_6347761 | 0.01 |
ENST00000250056.8 ENST00000571373.1 ENST00000570337.2 ENST00000572595.2 ENST00000576056.1 |
FAM64A |
family with sequence similarity 64, member A |
| chr7_-_141957847 | 0.01 |
ENST00000552471.1 ENST00000547058.2 |
PRSS58 |
protease, serine, 58 |
| chr17_+_6347729 | 0.01 |
ENST00000572447.1 |
FAM64A |
family with sequence similarity 64, member A |
| chr2_-_89597542 | 0.01 |
ENST00000465170.1 |
IGKV1-37 |
immunoglobulin kappa variable 1-37 (non-functional) |
| chr12_+_53848505 | 0.01 |
ENST00000552819.1 ENST00000455667.3 |
PCBP2 |
poly(rC) binding protein 2 |
| chr22_-_24093267 | 0.01 |
ENST00000341976.3 |
ZNF70 |
zinc finger protein 70 |
| chr12_-_53729525 | 0.01 |
ENST00000303846.3 |
SP7 |
Sp7 transcription factor |
| chr17_-_43025005 | 0.01 |
ENST00000587309.1 ENST00000593135.1 ENST00000339151.4 |
KIF18B |
kinesin family member 18B |
| chr16_-_28608424 | 0.01 |
ENST00000335715.4 |
SULT1A2 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 2 |
| chr1_-_54405773 | 0.01 |
ENST00000371376.1 |
HSPB11 |
heat shock protein family B (small), member 11 |
| chr2_+_204732487 | 0.01 |
ENST00000302823.3 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
| chr19_-_37701386 | 0.01 |
ENST00000527838.1 ENST00000591492.1 ENST00000532828.2 |
ZNF585B |
zinc finger protein 585B |
| chr6_-_117923610 | 0.01 |
ENST00000535237.1 ENST00000052569.6 |
GOPC |
golgi-associated PDZ and coiled-coil motif containing |
| chrX_-_133930285 | 0.01 |
ENST00000486347.1 ENST00000343004.5 |
FAM122B |
family with sequence similarity 122B |
| chr4_-_25161996 | 0.01 |
ENST00000513285.1 ENST00000382103.2 |
SEPSECS |
Sep (O-phosphoserine) tRNA:Sec (selenocysteine) tRNA synthase |
| chr3_+_112051994 | 0.01 |
ENST00000473539.1 ENST00000315711.8 ENST00000383681.3 |
CD200 |
CD200 molecule |
| chr10_+_103113802 | 0.01 |
ENST00000370187.3 |
BTRC |
beta-transducin repeat containing E3 ubiquitin protein ligase |
| chr18_+_12407895 | 0.01 |
ENST00000590956.1 ENST00000336990.4 ENST00000440960.1 ENST00000588729.1 |
SLMO1 |
slowmo homolog 1 (Drosophila) |
| chr12_-_25801478 | 0.01 |
ENST00000540106.1 ENST00000445693.1 ENST00000545543.1 ENST00000542224.1 |
IFLTD1 |
intermediate filament tail domain containing 1 |
| chr3_-_15382875 | 0.01 |
ENST00000408919.3 |
SH3BP5 |
SH3-domain binding protein 5 (BTK-associated) |
| chr2_+_204732666 | 0.01 |
ENST00000295854.6 ENST00000472206.1 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
| chr16_+_56969284 | 0.01 |
ENST00000568358.1 |
HERPUD1 |
homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 |
| chr14_-_93673353 | 0.01 |
ENST00000556566.1 ENST00000306954.4 |
C14orf142 |
chromosome 14 open reading frame 142 |
| chr12_+_67663056 | 0.01 |
ENST00000545606.1 |
CAND1 |
cullin-associated and neddylation-dissociated 1 |
| chr11_+_118826999 | 0.01 |
ENST00000264031.2 |
UPK2 |
uroplakin 2 |
| chr2_+_90153696 | 0.01 |
ENST00000417279.2 |
IGKV3D-15 |
immunoglobulin kappa variable 3D-15 (gene/pseudogene) |
| chr20_+_3713314 | 0.00 |
ENST00000254963.2 ENST00000542646.1 ENST00000399701.1 |
HSPA12B |
heat shock 70kD protein 12B |
| chr21_+_34602200 | 0.00 |
ENST00000382264.3 ENST00000382241.3 ENST00000404220.3 ENST00000342136.4 |
IFNAR2 |
interferon (alpha, beta and omega) receptor 2 |
| chr12_-_68647281 | 0.00 |
ENST00000328087.4 ENST00000538666.1 |
IL22 |
interleukin 22 |
| chr3_+_123813509 | 0.00 |
ENST00000460856.1 ENST00000240874.3 |
KALRN |
kalirin, RhoGEF kinase |
| chr15_+_45028753 | 0.00 |
ENST00000338264.4 |
TRIM69 |
tripartite motif containing 69 |
| chr1_-_102462565 | 0.00 |
ENST00000370103.4 |
OLFM3 |
olfactomedin 3 |
| chr3_+_46919235 | 0.00 |
ENST00000449590.1 |
PTH1R |
parathyroid hormone 1 receptor |
| chr11_+_369804 | 0.00 |
ENST00000329962.6 |
B4GALNT4 |
beta-1,4-N-acetyl-galactosaminyl transferase 4 |
| chr2_-_208031943 | 0.00 |
ENST00000421199.1 ENST00000457962.1 |
KLF7 |
Kruppel-like factor 7 (ubiquitous) |
| chr7_+_96634850 | 0.00 |
ENST00000518156.2 |
DLX6 |
distal-less homeobox 6 |
| chr4_-_25162188 | 0.00 |
ENST00000302922.3 |
SEPSECS |
Sep (O-phosphoserine) tRNA:Sec (selenocysteine) tRNA synthase |
| chr10_+_103113840 | 0.00 |
ENST00000393441.4 ENST00000408038.2 |
BTRC |
beta-transducin repeat containing E3 ubiquitin protein ligase |
| chr7_-_81399329 | 0.00 |
ENST00000453411.1 ENST00000444829.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0009436 | glyoxylate catabolic process(GO:0009436) |
| 0.0 | 0.1 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.0 | 0.1 | GO:0071874 | cellular response to norepinephrine stimulus(GO:0071874) |
| 0.0 | 0.2 | GO:0051534 | negative regulation of NFAT protein import into nucleus(GO:0051534) |
| 0.0 | 0.1 | GO:0045360 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 0.0 | 0.2 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.0 | 0.0 | GO:0060435 | branchiomeric skeletal muscle development(GO:0014707) bronchiole development(GO:0060435) |
| 0.0 | 0.1 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.0 | 0.1 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.0 | 0.1 | GO:1903944 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.0 | 0.1 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.0 | 0.1 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 0.0 | 0.1 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.0 | 0.1 | GO:0008410 | CoA-transferase activity(GO:0008410) |
| 0.0 | 0.1 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |