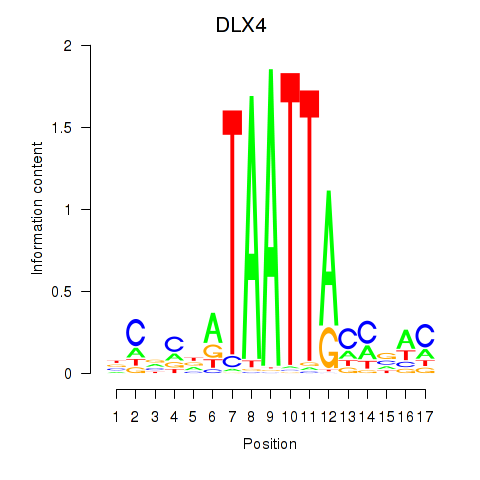
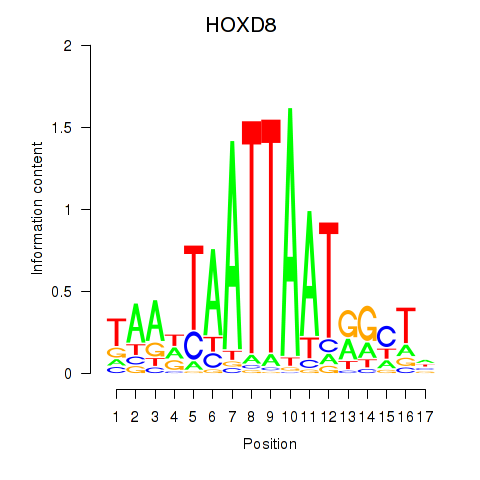
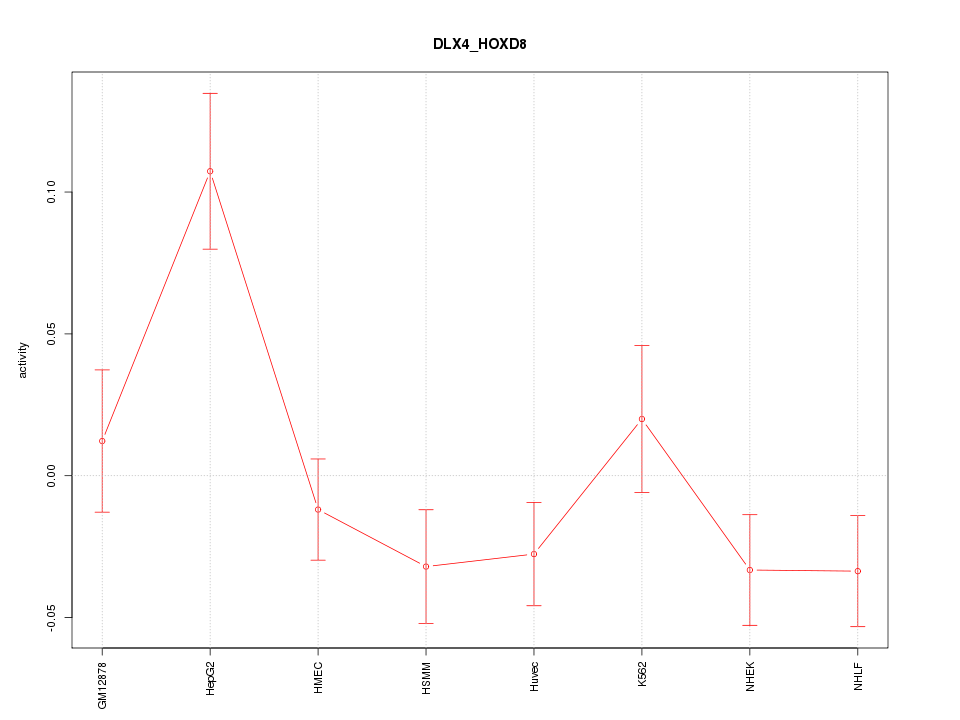
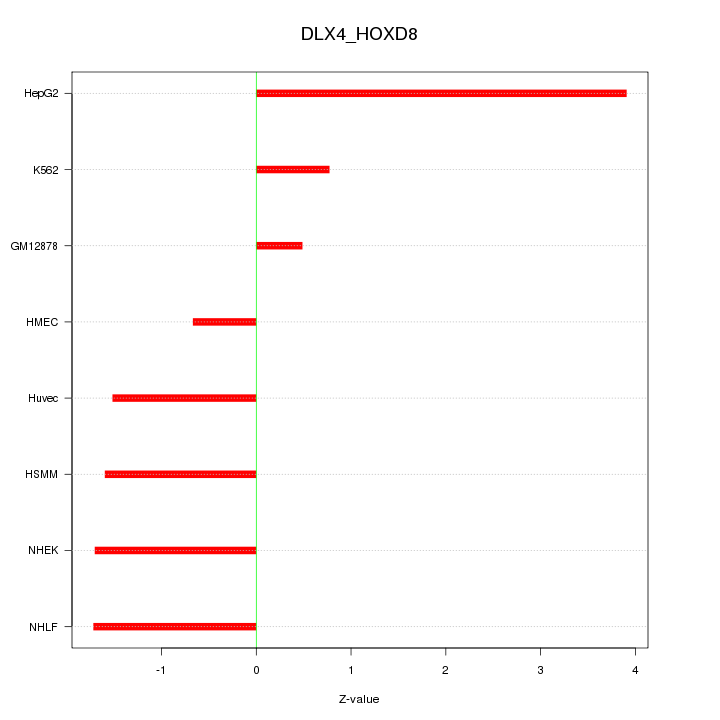
| 1.8 |
18.1 |
GO:0051918 |
negative regulation of fibrinolysis(GO:0051918) |
| 1.7 |
5.1 |
GO:0001907 |
cytolysis by symbiont of host cells(GO:0001897) killing by symbiont of host cells(GO:0001907) hemolysis by symbiont of host erythrocytes(GO:0019836) disruption by symbiont of host cell(GO:0044004) hemolysis in other organism(GO:0044179) positive regulation of circadian sleep/wake cycle, non-REM sleep(GO:0046010) cytolysis in other organism(GO:0051715) cytolysis in other organism involved in symbiotic interaction(GO:0051801) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 1.3 |
4.0 |
GO:0010902 |
positive regulation of very-low-density lipoprotein particle remodeling(GO:0010902) positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.9 |
13.1 |
GO:0030212 |
hyaluronan metabolic process(GO:0030212) |
| 0.7 |
2.7 |
GO:0010727 |
negative regulation of hydrogen peroxide metabolic process(GO:0010727) |
| 0.7 |
2.0 |
GO:1901419 |
regulation of response to alcohol(GO:1901419) |
| 0.6 |
1.8 |
GO:0010430 |
ethanol catabolic process(GO:0006068) fatty acid omega-oxidation(GO:0010430) |
| 0.6 |
5.1 |
GO:0034382 |
chylomicron remnant clearance(GO:0034382) triglyceride-rich lipoprotein particle clearance(GO:0071830) |
| 0.5 |
1.6 |
GO:0060318 |
definitive erythrocyte differentiation(GO:0060318) |
| 0.4 |
0.9 |
GO:0042905 |
9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.4 |
1.6 |
GO:0071400 |
cellular response to oleic acid(GO:0071400) |
| 0.4 |
1.2 |
GO:0042760 |
very long-chain fatty acid catabolic process(GO:0042760) |
| 0.4 |
5.5 |
GO:0030299 |
intestinal cholesterol absorption(GO:0030299) intestinal lipid absorption(GO:0098856) |
| 0.3 |
8.1 |
GO:0051923 |
sulfation(GO:0051923) |
| 0.3 |
1.0 |
GO:0002752 |
cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 0.3 |
0.9 |
GO:0009107 |
lipoate biosynthetic process(GO:0009107) |
| 0.3 |
2.7 |
GO:0042448 |
progesterone metabolic process(GO:0042448) |
| 0.3 |
1.7 |
GO:0006562 |
proline catabolic process(GO:0006562) |
| 0.2 |
0.7 |
GO:0090467 |
L-arginine import(GO:0043091) arginine import(GO:0090467) L-arginine transport(GO:1902023) |
| 0.2 |
0.7 |
GO:0006530 |
asparagine catabolic process(GO:0006530) asparagine catabolic process via L-aspartate(GO:0033345) |
| 0.2 |
1.2 |
GO:0035444 |
nickel cation transport(GO:0015675) vanadium ion transport(GO:0015676) ferrous iron transport(GO:0015684) lead ion transport(GO:0015692) nickel cation transmembrane transport(GO:0035444) ferrous iron import(GO:0070627) iron ion import(GO:0097286) |
| 0.2 |
0.7 |
GO:0048389 |
intermediate mesoderm development(GO:0048389) intermediate mesoderm morphogenesis(GO:0048390) intermediate mesoderm formation(GO:0048391) intermediate mesodermal cell differentiation(GO:0048392) bud dilation involved in lung branching(GO:0060503) branch elongation involved in ureteric bud branching(GO:0060681) regulation of transcription from RNA polymerase II promoter involved in kidney development(GO:0060994) BMP signaling pathway involved in ureter morphogenesis(GO:0061149) renal system segmentation(GO:0061150) BMP signaling pathway involved in renal system segmentation(GO:0061151) pulmonary artery endothelial tube morphogenesis(GO:0061155) pulmonary artery morphogenesis(GO:0061156) cell proliferation involved in mesonephros development(GO:0061209) regulation of transcription from RNA polymerase II promoter involved in mesonephros development(GO:0061216) negative regulation of mesonephros development(GO:0061218) pattern specification involved in mesonephros development(GO:0061227) renal system vasculature morphogenesis(GO:0061438) kidney vasculature morphogenesis(GO:0061439) BMP signaling pathway involved in nephric duct formation(GO:0071893) glomerular visceral epithelial cell development(GO:0072015) comma-shaped body morphogenesis(GO:0072049) regulation of branch elongation involved in ureteric bud branching(GO:0072095) negative regulation of branch elongation involved in ureteric bud branching(GO:0072096) negative regulation of branch elongation involved in ureteric bud branching by BMP signaling pathway(GO:0072097) anterior/posterior pattern specification involved in kidney development(GO:0072098) anterior/posterior pattern specification involved in ureteric bud development(GO:0072099) specification of ureteric bud anterior/posterior symmetry(GO:0072100) specification of ureteric bud anterior/posterior symmetry by BMP signaling pathway(GO:0072101) glomerulus morphogenesis(GO:0072102) glomerulus vasculature morphogenesis(GO:0072103) glomerular capillary formation(GO:0072104) mesenchymal cell proliferation involved in ureteric bud development(GO:0072138) nephric duct formation(GO:0072179) ureter urothelium development(GO:0072190) ureter epithelial cell differentiation(GO:0072192) ureter morphogenesis(GO:0072197) metanephric comma-shaped body morphogenesis(GO:0072278) glomerular epithelial cell development(GO:0072310) negative regulation of branching involved in ureteric bud morphogenesis(GO:0090191) regulation of metanephric S-shaped body morphogenesis(GO:2000004) negative regulation of metanephric S-shaped body morphogenesis(GO:2000005) regulation of metanephric comma-shaped body morphogenesis(GO:2000006) negative regulation of metanephric comma-shaped body morphogenesis(GO:2000007) |
| 0.2 |
1.6 |
GO:0006883 |
cellular sodium ion homeostasis(GO:0006883) |
| 0.2 |
1.1 |
GO:0007614 |
short-term memory(GO:0007614) |
| 0.2 |
1.0 |
GO:0030951 |
establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.2 |
3.1 |
GO:0018298 |
protein-chromophore linkage(GO:0018298) |
| 0.2 |
1.3 |
GO:0051026 |
chiasma assembly(GO:0051026) |
| 0.2 |
1.4 |
GO:0007351 |
tripartite regional subdivision(GO:0007351) anterior/posterior axis specification, embryo(GO:0008595) |
| 0.2 |
0.9 |
GO:0051852 |
disruption of cells of other organism involved in symbiotic interaction(GO:0051818) disruption by host of symbiont cells(GO:0051852) killing by host of symbiont cells(GO:0051873) killing of cells in other organism involved in symbiotic interaction(GO:0051883) |
| 0.2 |
0.5 |
GO:0045475 |
locomotor rhythm(GO:0045475) |
| 0.2 |
1.2 |
GO:0016584 |
nucleosome positioning(GO:0016584) |
| 0.2 |
0.5 |
GO:0007500 |
mesodermal cell fate determination(GO:0007500) |
| 0.2 |
1.9 |
GO:0042447 |
hormone catabolic process(GO:0042447) |
| 0.2 |
0.8 |
GO:0060164 |
regulation of timing of neuron differentiation(GO:0060164) |
| 0.2 |
0.6 |
GO:0006542 |
glutamine biosynthetic process(GO:0006542) |
| 0.1 |
1.6 |
GO:0006975 |
DNA damage induced protein phosphorylation(GO:0006975) |
| 0.1 |
0.6 |
GO:0000414 |
regulation of histone H3-K36 methylation(GO:0000414) |
| 0.1 |
0.4 |
GO:0046292 |
formaldehyde metabolic process(GO:0046292) |
| 0.1 |
0.7 |
GO:0015712 |
hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 |
1.3 |
GO:0030277 |
epithelial structure maintenance(GO:0010669) maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.1 |
0.7 |
GO:0050955 |
thermoception(GO:0050955) |
| 0.1 |
0.5 |
GO:0033522 |
histone H2A ubiquitination(GO:0033522) |
| 0.1 |
1.8 |
GO:0006491 |
N-glycan processing(GO:0006491) |
| 0.1 |
0.4 |
GO:0010212 |
response to ionizing radiation(GO:0010212) |
| 0.1 |
0.6 |
GO:0071168 |
protein localization to chromatin(GO:0071168) |
| 0.1 |
0.3 |
GO:0051083 |
'de novo' cotranslational protein folding(GO:0051083) |
| 0.1 |
1.6 |
GO:0051568 |
histone H3-K4 methylation(GO:0051568) |
| 0.1 |
1.1 |
GO:0031017 |
exocrine pancreas development(GO:0031017) |
| 0.1 |
1.1 |
GO:0007141 |
male meiosis I(GO:0007141) |
| 0.1 |
0.2 |
GO:0010896 |
regulation of triglyceride catabolic process(GO:0010896) positive regulation of triglyceride catabolic process(GO:0010898) |
| 0.1 |
0.6 |
GO:0000050 |
urea cycle(GO:0000050) |
| 0.1 |
0.3 |
GO:0051661 |
maintenance of centrosome location(GO:0051661) |
| 0.1 |
0.3 |
GO:0010107 |
potassium ion import(GO:0010107) |
| 0.1 |
0.7 |
GO:0048024 |
regulation of mRNA splicing, via spliceosome(GO:0048024) |
| 0.1 |
0.4 |
GO:0015917 |
aminophospholipid transport(GO:0015917) |
| 0.1 |
0.1 |
GO:0010700 |
negative regulation of norepinephrine secretion(GO:0010700) |
| 0.1 |
0.6 |
GO:0034501 |
protein localization to kinetochore(GO:0034501) |
| 0.1 |
2.7 |
GO:0051225 |
spindle assembly(GO:0051225) |
| 0.1 |
0.3 |
GO:0015959 |
diadenosine polyphosphate metabolic process(GO:0015959) |
| 0.1 |
0.2 |
GO:0070340 |
detection of bacterial lipoprotein(GO:0042494) detection of triacyl bacterial lipopeptide(GO:0042495) detection of bacterial lipopeptide(GO:0070340) response to triacyl bacterial lipopeptide(GO:0071725) cellular response to triacyl bacterial lipopeptide(GO:0071727) |
| 0.1 |
1.6 |
GO:0016601 |
Rac protein signal transduction(GO:0016601) |
| 0.1 |
0.5 |
GO:0042416 |
dopamine biosynthetic process(GO:0042416) |
| 0.1 |
0.6 |
GO:0006729 |
tetrahydrobiopterin biosynthetic process(GO:0006729) |
| 0.1 |
0.2 |
GO:0021836 |
chemorepulsion involved in postnatal olfactory bulb interneuron migration(GO:0021836) regulation of negative chemotaxis(GO:0050923) |
| 0.1 |
0.5 |
GO:0015811 |
sulfur amino acid transport(GO:0000101) L-cystine transport(GO:0015811) |
| 0.1 |
0.3 |
GO:0001832 |
blastocyst growth(GO:0001832) |
| 0.1 |
0.5 |
GO:0042797 |
5S class rRNA transcription from RNA polymerase III type 1 promoter(GO:0042791) tRNA transcription from RNA polymerase III promoter(GO:0042797) |
| 0.1 |
0.5 |
GO:0071480 |
contact inhibition(GO:0060242) cellular response to gamma radiation(GO:0071480) |
| 0.1 |
0.8 |
GO:0046784 |
viral mRNA export from host cell nucleus(GO:0046784) |
| 0.1 |
0.8 |
GO:0006570 |
tyrosine metabolic process(GO:0006570) |
| 0.1 |
0.8 |
GO:0006564 |
L-serine biosynthetic process(GO:0006564) |
| 0.1 |
2.3 |
GO:0006730 |
one-carbon metabolic process(GO:0006730) |
| 0.1 |
0.2 |
GO:0050917 |
sensory perception of umami taste(GO:0050917) |
| 0.1 |
1.2 |
GO:0006071 |
glycerol metabolic process(GO:0006071) |
| 0.1 |
0.9 |
GO:0046415 |
urate metabolic process(GO:0046415) |
| 0.1 |
0.4 |
GO:0046642 |
negative regulation of alpha-beta T cell proliferation(GO:0046642) |
| 0.1 |
0.9 |
GO:0090399 |
replicative senescence(GO:0090399) |
| 0.1 |
0.1 |
GO:0071460 |
cellular response to cell-matrix adhesion(GO:0071460) |
| 0.1 |
0.1 |
GO:0071502 |
cellular response to temperature stimulus(GO:0071502) |
| 0.1 |
0.1 |
GO:1904358 |
positive regulation of telomere maintenance via telomerase(GO:0032212) positive regulation of telomere maintenance via telomere lengthening(GO:1904358) |
| 0.1 |
0.6 |
GO:0006654 |
phosphatidic acid biosynthetic process(GO:0006654) |
| 0.1 |
0.2 |
GO:0006537 |
glutamate biosynthetic process(GO:0006537) |
| 0.1 |
0.3 |
GO:0034372 |
triglyceride-rich lipoprotein particle remodeling(GO:0034370) very-low-density lipoprotein particle remodeling(GO:0034372) |
| 0.1 |
0.3 |
GO:0070537 |
histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.1 |
0.3 |
GO:0071801 |
podosome assembly(GO:0071800) regulation of podosome assembly(GO:0071801) positive regulation of podosome assembly(GO:0071803) |
| 0.1 |
0.3 |
GO:0030035 |
microspike assembly(GO:0030035) |
| 0.1 |
0.5 |
GO:0051383 |
kinetochore assembly(GO:0051382) kinetochore organization(GO:0051383) |
| 0.1 |
1.1 |
GO:0050832 |
defense response to fungus(GO:0050832) |
| 0.1 |
0.3 |
GO:0008628 |
hormone-mediated apoptotic signaling pathway(GO:0008628) |
| 0.1 |
1.5 |
GO:0008211 |
glucocorticoid metabolic process(GO:0008211) |
| 0.1 |
0.6 |
GO:0033089 |
positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 0.1 |
0.2 |
GO:0006428 |
isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.1 |
0.2 |
GO:0050653 |
UDP-N-acetylgalactosamine metabolic process(GO:0019276) chondroitin sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0050653) |
| 0.1 |
1.5 |
GO:0051180 |
vitamin transport(GO:0051180) |
| 0.1 |
0.4 |
GO:0015886 |
heme transport(GO:0015886) |
| 0.1 |
0.4 |
GO:0033085 |
negative regulation of T cell differentiation in thymus(GO:0033085) negative regulation of thymocyte aggregation(GO:2000399) |
| 0.1 |
1.4 |
GO:0002504 |
antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002504) |
| 0.1 |
0.3 |
GO:0010957 |
negative regulation of calcidiol 1-monooxygenase activity(GO:0010956) negative regulation of vitamin D biosynthetic process(GO:0010957) negative regulation of vitamin metabolic process(GO:0046137) |
| 0.1 |
1.1 |
GO:0031000 |
response to caffeine(GO:0031000) |
| 0.1 |
0.3 |
GO:0007584 |
response to nutrient(GO:0007584) |
| 0.1 |
5.4 |
GO:0008360 |
regulation of cell shape(GO:0008360) |
| 0.1 |
0.8 |
GO:0030183 |
B cell differentiation(GO:0030183) |
| 0.1 |
0.4 |
GO:0042487 |
regulation of odontogenesis of dentin-containing tooth(GO:0042487) |
| 0.1 |
0.2 |
GO:0072205 |
collecting duct development(GO:0072044) metanephric collecting duct development(GO:0072205) |
| 0.1 |
0.4 |
GO:0050884 |
neuromuscular process controlling posture(GO:0050884) |
| 0.1 |
0.2 |
GO:0046477 |
glycosylceramide catabolic process(GO:0046477) |
| 0.1 |
0.4 |
GO:0045059 |
positive thymic T cell selection(GO:0045059) |
| 0.1 |
0.5 |
GO:0009249 |
protein lipoylation(GO:0009249) |
| 0.1 |
0.2 |
GO:0046621 |
negative regulation of organ growth(GO:0046621) |
| 0.1 |
1.0 |
GO:0005513 |
detection of calcium ion(GO:0005513) |
| 0.1 |
0.6 |
GO:0010923 |
negative regulation of phosphatase activity(GO:0010923) negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.1 |
0.5 |
GO:0045898 |
regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) |
| 0.1 |
1.0 |
GO:0000380 |
alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.1 |
1.5 |
GO:0006693 |
prostaglandin metabolic process(GO:0006693) |
| 0.0 |
0.2 |
GO:0042462 |
photoreceptor cell development(GO:0042461) eye photoreceptor cell development(GO:0042462) |
| 0.0 |
0.5 |
GO:0060046 |
regulation of acrosome reaction(GO:0060046) |
| 0.0 |
0.1 |
GO:0046984 |
regulation of hemoglobin biosynthetic process(GO:0046984) |
| 0.0 |
0.8 |
GO:0006359 |
regulation of transcription from RNA polymerase III promoter(GO:0006359) |
| 0.0 |
0.1 |
GO:0006655 |
phosphatidylglycerol biosynthetic process(GO:0006655) cardiolipin metabolic process(GO:0032048) cardiolipin biosynthetic process(GO:0032049) phosphatidylglycerol metabolic process(GO:0046471) |
| 0.0 |
0.1 |
GO:0009405 |
pathogenesis(GO:0009405) |
| 0.0 |
0.1 |
GO:0032074 |
negative regulation of nuclease activity(GO:0032074) |
| 0.0 |
0.4 |
GO:0006700 |
C21-steroid hormone biosynthetic process(GO:0006700) |
| 0.0 |
0.8 |
GO:0016578 |
histone deubiquitination(GO:0016578) |
| 0.0 |
1.1 |
GO:0015804 |
neutral amino acid transport(GO:0015804) |
| 0.0 |
2.6 |
GO:0006958 |
complement activation, classical pathway(GO:0006958) |
| 0.0 |
0.1 |
GO:0007028 |
cytoplasm organization(GO:0007028) ribonucleoprotein complex disassembly(GO:0032988) |
| 0.0 |
0.7 |
GO:0045652 |
regulation of megakaryocyte differentiation(GO:0045652) regulation of hematopoietic progenitor cell differentiation(GO:1901532) |
| 0.0 |
0.3 |
GO:0033504 |
floor plate development(GO:0033504) |
| 0.0 |
0.1 |
GO:0002331 |
immature B cell differentiation(GO:0002327) pre-B cell differentiation(GO:0002329) pre-B cell allelic exclusion(GO:0002331) |
| 0.0 |
0.2 |
GO:0050862 |
positive regulation of T cell receptor signaling pathway(GO:0050862) |
| 0.0 |
0.2 |
GO:0042519 |
tyrosine phosphorylation of Stat4 protein(GO:0042504) regulation of tyrosine phosphorylation of Stat4 protein(GO:0042519) positive regulation of tyrosine phosphorylation of Stat4 protein(GO:0042520) |
| 0.0 |
0.7 |
GO:0018214 |
peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.0 |
0.1 |
GO:0044240 |
multicellular organism lipid catabolic process(GO:0044240) |
| 0.0 |
0.2 |
GO:0051294 |
establishment of mitotic spindle orientation(GO:0000132) establishment of spindle orientation(GO:0051294) |
| 0.0 |
0.1 |
GO:0048752 |
semicircular canal morphogenesis(GO:0048752) |
| 0.0 |
0.2 |
GO:0048537 |
mucosal-associated lymphoid tissue development(GO:0048537) |
| 0.0 |
0.2 |
GO:0010890 |
positive regulation of sequestering of triglyceride(GO:0010890) |
| 0.0 |
1.7 |
GO:0060338 |
regulation of type I interferon-mediated signaling pathway(GO:0060338) |
| 0.0 |
0.9 |
GO:0071804 |
cellular potassium ion transport(GO:0071804) potassium ion transmembrane transport(GO:0071805) |
| 0.0 |
0.1 |
GO:0060748 |
tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
| 0.0 |
0.4 |
GO:0022027 |
interkinetic nuclear migration(GO:0022027) |
| 0.0 |
0.1 |
GO:0001692 |
histamine metabolic process(GO:0001692) |
| 0.0 |
0.1 |
GO:0051930 |
regulation of neurological system process(GO:0031644) regulation of sensory perception of pain(GO:0051930) regulation of sensory perception(GO:0051931) |
| 0.0 |
0.1 |
GO:0043985 |
histone H4-R3 methylation(GO:0043985) |
| 0.0 |
0.6 |
GO:0006400 |
tRNA modification(GO:0006400) |
| 0.0 |
0.1 |
GO:0010873 |
positive regulation of cholesterol esterification(GO:0010873) |
| 0.0 |
0.2 |
GO:0060023 |
soft palate development(GO:0060023) |
| 0.0 |
0.6 |
GO:0006906 |
vesicle fusion(GO:0006906) |
| 0.0 |
0.2 |
GO:0006422 |
aspartyl-tRNA aminoacylation(GO:0006422) |
| 0.0 |
0.2 |
GO:0046653 |
tetrahydrofolate metabolic process(GO:0046653) |
| 0.0 |
0.3 |
GO:0070584 |
mitochondrion morphogenesis(GO:0070584) |
| 0.0 |
0.4 |
GO:0031055 |
DNA replication-independent nucleosome assembly(GO:0006336) chromatin remodeling at centromere(GO:0031055) CENP-A containing nucleosome assembly(GO:0034080) centromere complex assembly(GO:0034508) DNA replication-independent nucleosome organization(GO:0034724) CENP-A containing chromatin organization(GO:0061641) |
| 0.0 |
0.2 |
GO:0010447 |
response to acidic pH(GO:0010447) |
| 0.0 |
0.1 |
GO:0008612 |
peptidyl-lysine modification to peptidyl-hypusine(GO:0008612) |
| 0.0 |
0.1 |
GO:0046080 |
dUTP metabolic process(GO:0046080) |
| 0.0 |
0.4 |
GO:0007217 |
tachykinin receptor signaling pathway(GO:0007217) |
| 0.0 |
0.6 |
GO:0000038 |
very long-chain fatty acid metabolic process(GO:0000038) |
| 0.0 |
2.6 |
GO:0051592 |
response to calcium ion(GO:0051592) |
| 0.0 |
0.7 |
GO:0019048 |
modulation by virus of host morphology or physiology(GO:0019048) |
| 0.0 |
0.5 |
GO:0014066 |
regulation of phosphatidylinositol 3-kinase signaling(GO:0014066) |
| 0.0 |
0.2 |
GO:0060512 |
prostate gland morphogenesis(GO:0060512) |
| 0.0 |
0.2 |
GO:0006189 |
'de novo' IMP biosynthetic process(GO:0006189) |
| 0.0 |
0.9 |
GO:0033209 |
tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.0 |
0.1 |
GO:0000114 |
obsolete regulation of transcription involved in G1 phase of mitotic cell cycle(GO:0000114) |
| 0.0 |
0.2 |
GO:0006610 |
ribosomal protein import into nucleus(GO:0006610) |
| 0.0 |
0.2 |
GO:0060009 |
Sertoli cell development(GO:0060009) |
| 0.0 |
1.0 |
GO:0042475 |
odontogenesis of dentin-containing tooth(GO:0042475) |
| 0.0 |
0.1 |
GO:0048934 |
peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.0 |
1.5 |
GO:0001541 |
ovarian follicle development(GO:0001541) |
| 0.0 |
0.1 |
GO:0010936 |
negative regulation of macrophage cytokine production(GO:0010936) |
| 0.0 |
0.1 |
GO:2000008 |
regulation of protein localization to cell surface(GO:2000008) |
| 0.0 |
0.1 |
GO:0006007 |
glucose catabolic process(GO:0006007) |
| 0.0 |
0.2 |
GO:0035414 |
negative regulation of catenin import into nucleus(GO:0035414) |
| 0.0 |
0.1 |
GO:0033326 |
cerebrospinal fluid secretion(GO:0033326) |
| 0.0 |
0.1 |
GO:0007352 |
zygotic specification of dorsal/ventral axis(GO:0007352) |
| 0.0 |
0.3 |
GO:0048013 |
ephrin receptor signaling pathway(GO:0048013) |
| 0.0 |
0.2 |
GO:0060999 |
positive regulation of dendritic spine development(GO:0060999) |
| 0.0 |
1.5 |
GO:0046324 |
regulation of glucose import(GO:0046324) |
| 0.0 |
0.1 |
GO:0090036 |
regulation of protein kinase C signaling(GO:0090036) positive regulation of protein kinase C signaling(GO:0090037) |
| 0.0 |
0.3 |
GO:0051319 |
mitotic G2 phase(GO:0000085) G2 phase(GO:0051319) |
| 0.0 |
0.1 |
GO:0005984 |
disaccharide metabolic process(GO:0005984) |
| 0.0 |
0.1 |
GO:0032747 |
positive regulation of interleukin-23 production(GO:0032747) |
| 0.0 |
0.9 |
GO:0043124 |
negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.0 |
0.2 |
GO:0031958 |
corticosteroid receptor signaling pathway(GO:0031958) glucocorticoid receptor signaling pathway(GO:0042921) |
| 0.0 |
0.1 |
GO:0010172 |
embryonic body morphogenesis(GO:0010172) |
| 0.0 |
0.3 |
GO:0032814 |
regulation of natural killer cell activation(GO:0032814) |
| 0.0 |
0.1 |
GO:0072367 |
regulation of low-density lipoprotein particle clearance(GO:0010988) positive regulation of low-density lipoprotein particle receptor biosynthetic process(GO:0045716) regulation of lipid transport by regulation of transcription from RNA polymerase II promoter(GO:0072367) regulation of lipid transport by positive regulation of transcription from RNA polymerase II promoter(GO:0072369) |
| 0.0 |
0.3 |
GO:0030889 |
negative regulation of B cell proliferation(GO:0030889) |
| 0.0 |
0.2 |
GO:0032148 |
activation of protein kinase B activity(GO:0032148) |
| 0.0 |
0.4 |
GO:0006386 |
transcription elongation from RNA polymerase III promoter(GO:0006385) termination of RNA polymerase III transcription(GO:0006386) |
| 0.0 |
0.1 |
GO:0034616 |
response to laminar fluid shear stress(GO:0034616) |
| 0.0 |
0.0 |
GO:0043486 |
histone exchange(GO:0043486) |
| 0.0 |
0.2 |
GO:0006020 |
inositol metabolic process(GO:0006020) |
| 0.0 |
0.7 |
GO:0030866 |
cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 |
0.1 |
GO:0019509 |
L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.0 |
0.2 |
GO:0030146 |
obsolete diuresis(GO:0030146) |
| 0.0 |
0.2 |
GO:0050830 |
defense response to Gram-positive bacterium(GO:0050830) |
| 0.0 |
0.4 |
GO:0016558 |
protein import into peroxisome matrix(GO:0016558) |
| 0.0 |
0.8 |
GO:0000387 |
spliceosomal snRNP assembly(GO:0000387) |
| 0.0 |
0.3 |
GO:0015939 |
pantothenate metabolic process(GO:0015939) |
| 0.0 |
0.1 |
GO:0009174 |
UMP biosynthetic process(GO:0006222) pyrimidine ribonucleoside monophosphate metabolic process(GO:0009173) pyrimidine ribonucleoside monophosphate biosynthetic process(GO:0009174) UMP metabolic process(GO:0046049) |
| 0.0 |
0.1 |
GO:0046952 |
ketone body catabolic process(GO:0046952) |
| 0.0 |
0.1 |
GO:0050858 |
negative regulation of antigen receptor-mediated signaling pathway(GO:0050858) negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.0 |
0.4 |
GO:0046854 |
phosphatidylinositol phosphorylation(GO:0046854) |
| 0.0 |
0.1 |
GO:0070071 |
proton-transporting two-sector ATPase complex assembly(GO:0070071) |
| 0.0 |
0.1 |
GO:1900087 |
traversing start control point of mitotic cell cycle(GO:0007089) positive regulation of G1/S transition of mitotic cell cycle(GO:1900087) positive regulation of cell cycle G1/S phase transition(GO:1902808) |
| 0.0 |
2.8 |
GO:0006814 |
sodium ion transport(GO:0006814) |
| 0.0 |
0.3 |
GO:0030321 |
positive regulation of smooth muscle cell migration(GO:0014911) transepithelial chloride transport(GO:0030321) |
| 0.0 |
0.0 |
GO:0000019 |
regulation of mitotic recombination(GO:0000019) |
| 0.0 |
0.1 |
GO:0002467 |
germinal center formation(GO:0002467) |
| 0.0 |
0.4 |
GO:1900047 |
negative regulation of blood coagulation(GO:0030195) negative regulation of wound healing(GO:0061045) negative regulation of hemostasis(GO:1900047) |
| 0.0 |
0.6 |
GO:0021675 |
nerve development(GO:0021675) |
| 0.0 |
1.3 |
GO:0031960 |
response to corticosteroid(GO:0031960) |
| 0.0 |
0.2 |
GO:0046051 |
GTP biosynthetic process(GO:0006183) UTP biosynthetic process(GO:0006228) CTP biosynthetic process(GO:0006241) pyrimidine ribonucleoside triphosphate metabolic process(GO:0009208) pyrimidine ribonucleoside triphosphate biosynthetic process(GO:0009209) CTP metabolic process(GO:0046036) UTP metabolic process(GO:0046051) pyrimidine ribonucleoside biosynthetic process(GO:0046132) |
| 0.0 |
0.1 |
GO:0060430 |
globus pallidus development(GO:0021759) forebrain dorsal/ventral pattern formation(GO:0021798) menarche(GO:0042696) lung saccule development(GO:0060430) Clara cell differentiation(GO:0060486) Type II pneumocyte differentiation(GO:0060510) |
| 0.0 |
0.1 |
GO:0060760 |
positive regulation of response to cytokine stimulus(GO:0060760) |
| 0.0 |
1.0 |
GO:0050821 |
protein stabilization(GO:0050821) |
| 0.0 |
0.4 |
GO:0006493 |
protein O-linked glycosylation(GO:0006493) |
| 0.0 |
0.1 |
GO:0045822 |
negative regulation of heart contraction(GO:0045822) |
| 0.0 |
0.4 |
GO:0016079 |
synaptic vesicle exocytosis(GO:0016079) |
| 0.0 |
0.1 |
GO:0070995 |
NADPH oxidation(GO:0070995) |
| 0.0 |
0.1 |
GO:0001831 |
trophectodermal cellular morphogenesis(GO:0001831) |
| 0.0 |
0.5 |
GO:0051925 |
obsolete regulation of calcium ion transport via voltage-gated calcium channel activity(GO:0051925) |
| 0.0 |
0.3 |
GO:0033108 |
mitochondrial respiratory chain complex assembly(GO:0033108) |
| 0.0 |
0.2 |
GO:0051090 |
regulation of sequence-specific DNA binding transcription factor activity(GO:0051090) |
| 0.0 |
0.8 |
GO:0032526 |
response to retinoic acid(GO:0032526) |
| 0.0 |
0.5 |
GO:0001707 |
mesoderm formation(GO:0001707) |
| 0.0 |
0.3 |
GO:0034332 |
adherens junction organization(GO:0034332) cell-cell junction organization(GO:0045216) |
| 0.0 |
1.0 |
GO:0061025 |
membrane fusion(GO:0061025) |
| 0.0 |
0.1 |
GO:0044070 |
regulation of anion transport(GO:0044070) |
| 0.0 |
0.2 |
GO:0071044 |
regulation of translational termination(GO:0006449) histone mRNA catabolic process(GO:0071044) |
| 0.0 |
0.4 |
GO:0006749 |
glutathione metabolic process(GO:0006749) |
| 0.0 |
0.2 |
GO:0032876 |
regulation of DNA endoreduplication(GO:0032875) negative regulation of DNA endoreduplication(GO:0032876) DNA endoreduplication(GO:0042023) |
| 0.0 |
0.1 |
GO:0014823 |
response to activity(GO:0014823) |
| 0.0 |
0.1 |
GO:0040015 |
cytoplasmic mRNA processing body assembly(GO:0033962) negative regulation of multicellular organism growth(GO:0040015) |
| 0.0 |
0.0 |
GO:0051541 |
regulation of oxidative phosphorylation(GO:0002082) detoxification of copper ion(GO:0010273) elastin metabolic process(GO:0051541) detoxification of inorganic compound(GO:0061687) stress response to metal ion(GO:0097501) stress response to copper ion(GO:1990169) |
| 0.0 |
0.1 |
GO:0010458 |
exit from mitosis(GO:0010458) |
| 0.0 |
0.7 |
GO:0001892 |
embryonic placenta development(GO:0001892) |
| 0.0 |
0.8 |
GO:0051781 |
positive regulation of cell division(GO:0051781) |
| 0.0 |
0.0 |
GO:0042727 |
flavin-containing compound biosynthetic process(GO:0042727) |
| 0.0 |
1.3 |
GO:0000087 |
mitotic M phase(GO:0000087) |
| 0.0 |
0.2 |
GO:0042558 |
pteridine-containing compound metabolic process(GO:0042558) |
| 0.0 |
0.1 |
GO:0006692 |
prostanoid metabolic process(GO:0006692) |
| 0.0 |
0.2 |
GO:0015695 |
organic cation transport(GO:0015695) |
| 0.0 |
0.1 |
GO:0045742 |
positive regulation of epidermal growth factor receptor signaling pathway(GO:0045742) positive regulation of ERBB signaling pathway(GO:1901186) |
| 0.0 |
0.7 |
GO:0008033 |
tRNA processing(GO:0008033) |
| 0.0 |
0.1 |
GO:0045039 |
protein import into mitochondrial inner membrane(GO:0045039) |
| 0.0 |
0.1 |
GO:0051497 |
negative regulation of stress fiber assembly(GO:0051497) |
| 0.0 |
0.2 |
GO:0050812 |
acetyl-CoA biosynthetic process from pyruvate(GO:0006086) regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) regulation of acyl-CoA biosynthetic process(GO:0050812) |
| 0.0 |
0.3 |
GO:0019674 |
NAD metabolic process(GO:0019674) |
| 0.0 |
0.2 |
GO:0009067 |
aspartate family amino acid biosynthetic process(GO:0009067) |
| 0.0 |
0.0 |
GO:0043438 |
acetoacetic acid metabolic process(GO:0043438) |
| 0.0 |
0.2 |
GO:0007158 |
neuron cell-cell adhesion(GO:0007158) |
| 0.0 |
0.2 |
GO:0040014 |
regulation of multicellular organism growth(GO:0040014) |
| 0.0 |
0.1 |
GO:0014909 |
smooth muscle cell migration(GO:0014909) |
| 0.0 |
0.3 |
GO:0051092 |
positive regulation of NF-kappaB transcription factor activity(GO:0051092) |
| 0.0 |
0.1 |
GO:0006506 |
GPI anchor biosynthetic process(GO:0006506) |
| 0.0 |
0.3 |
GO:0000186 |
activation of MAPKK activity(GO:0000186) |
| 0.0 |
0.1 |
GO:0047496 |
vesicle transport along microtubule(GO:0047496) vesicle cytoskeletal trafficking(GO:0099518) |
| 0.0 |
0.3 |
GO:0009083 |
branched-chain amino acid catabolic process(GO:0009083) |
| 0.0 |
0.3 |
GO:0000080 |
mitotic G1 phase(GO:0000080) |
| 0.0 |
0.3 |
GO:0030261 |
chromosome condensation(GO:0030261) |
| 0.0 |
0.0 |
GO:2000171 |
negative regulation of dendrite development(GO:2000171) |
| 0.0 |
0.1 |
GO:1901070 |
GMP biosynthetic process(GO:0006177) guanosine-containing compound biosynthetic process(GO:1901070) |
| 0.0 |
0.0 |
GO:0045349 |
interferon-alpha biosynthetic process(GO:0045349) regulation of interferon-alpha biosynthetic process(GO:0045354) positive regulation of interferon-alpha biosynthetic process(GO:0045356) |
| 0.0 |
0.0 |
GO:0006625 |
protein targeting to peroxisome(GO:0006625) protein localization to peroxisome(GO:0072662) establishment of protein localization to peroxisome(GO:0072663) |
| 0.0 |
0.2 |
GO:0008210 |
estrogen metabolic process(GO:0008210) |