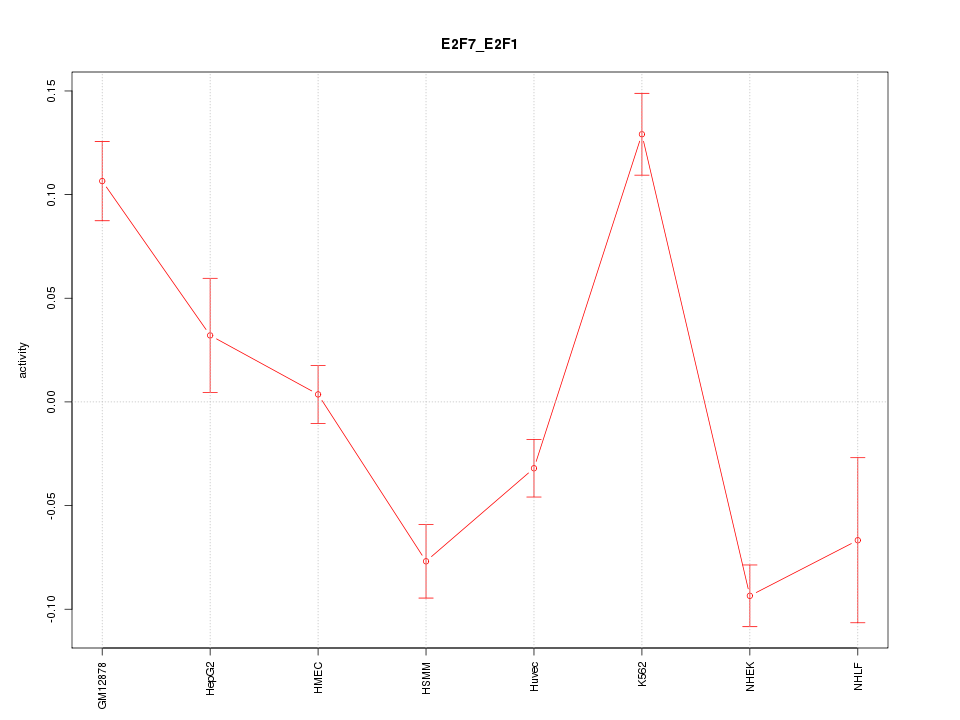
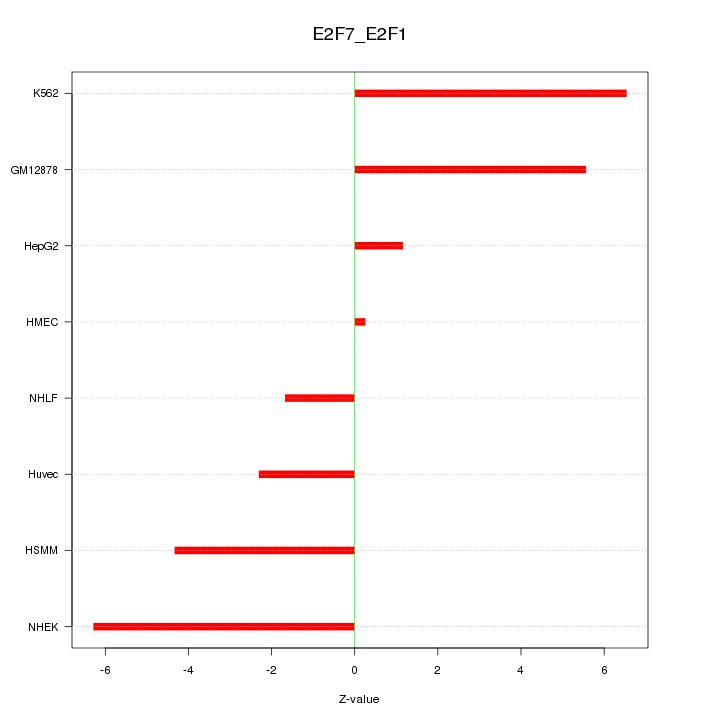
Motif ID: E2F7_E2F1
Z-value: 4.208
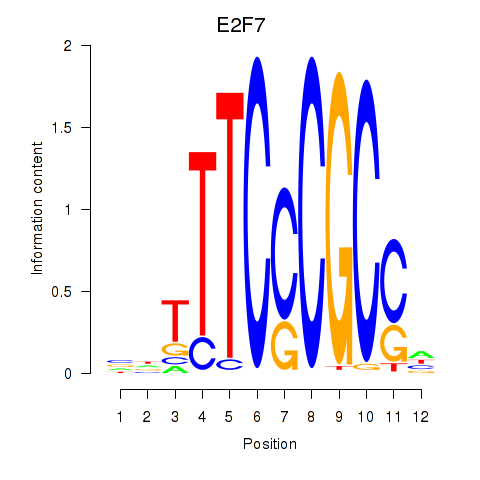
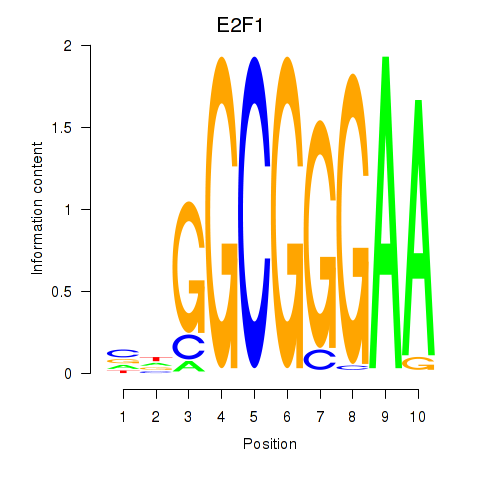
Transcription factors associated with E2F7_E2F1:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| E2F1 | ENSG00000101412.9 | E2F1 |
| E2F7 | ENSG00000165891.11 | E2F7 |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.4 | 13.5 | GO:0010520 | regulation of reciprocal meiotic recombination(GO:0010520) negative regulation of meiotic nuclear division(GO:0045835) regulation of meiosis I(GO:0060631) |
| 1.9 | 75.3 | GO:0006271 | DNA strand elongation involved in DNA replication(GO:0006271) |
| 1.7 | 6.8 | GO:0071313 | cellular response to caffeine(GO:0071313) |
| 1.7 | 13.4 | GO:0033261 | obsolete regulation of S phase(GO:0033261) |
| 1.5 | 4.6 | GO:0019087 | transformation of host cell by virus(GO:0019087) |
| 1.5 | 4.4 | GO:1904742 | regulation of single-stranded telomeric DNA binding(GO:0060380) positive regulation of single-stranded telomeric DNA binding(GO:0060381) regulation of telomeric DNA binding(GO:1904742) positive regulation of telomeric DNA binding(GO:1904744) |
| 1.5 | 4.4 | GO:0072216 | positive regulation of metanephros development(GO:0072216) positive regulation of cell proliferation involved in kidney development(GO:1901724) |
| 1.3 | 2.6 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 1.3 | 3.8 | GO:0009648 | photoperiodism(GO:0009648) |
| 1.2 | 10.5 | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 1.2 | 4.7 | GO:0072180 | mesonephric duct morphogenesis(GO:0072180) |
| 1.0 | 2.9 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 1.0 | 4.8 | GO:0032075 | positive regulation of nuclease activity(GO:0032075) |
| 1.0 | 5.7 | GO:0000710 | meiotic mismatch repair(GO:0000710) |
| 0.9 | 2.8 | GO:0042097 | interleukin-4 biosynthetic process(GO:0042097) positive regulation of interleukin-10 biosynthetic process(GO:0045082) regulation of interleukin-4 biosynthetic process(GO:0045402) positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 0.9 | 0.9 | GO:0009048 | dosage compensation by inactivation of X chromosome(GO:0009048) |
| 0.9 | 3.5 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 0.8 | 3.4 | GO:0060236 | regulation of mitotic spindle organization(GO:0060236) |
| 0.8 | 7.5 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 0.8 | 4.0 | GO:0050686 | negative regulation of mRNA processing(GO:0050686) negative regulation of mRNA metabolic process(GO:1903312) |
| 0.8 | 3.1 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.8 | 2.3 | GO:0035608 | protein deglutamylation(GO:0035608) |
| 0.7 | 5.9 | GO:0051383 | kinetochore assembly(GO:0051382) kinetochore organization(GO:0051383) |
| 0.7 | 10.3 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
| 0.7 | 2.2 | GO:0043366 | beta selection(GO:0043366) |
| 0.7 | 2.8 | GO:0061032 | visceral serous pericardium development(GO:0061032) |
| 0.7 | 2.1 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.6 | 1.9 | GO:0018916 | nitrobenzene metabolic process(GO:0018916) |
| 0.6 | 2.6 | GO:0006207 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) pyrimidine nucleobase biosynthetic process(GO:0019856) |
| 0.6 | 0.6 | GO:0006066 | alcohol metabolic process(GO:0006066) |
| 0.6 | 0.6 | GO:0009162 | pyrimidine nucleoside monophosphate metabolic process(GO:0009129) deoxyribonucleoside monophosphate metabolic process(GO:0009162) pyrimidine deoxyribonucleoside monophosphate metabolic process(GO:0009176) dUMP metabolic process(GO:0046078) |
| 0.6 | 3.1 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.6 | 1.8 | GO:0070734 | histone H3-K27 methylation(GO:0070734) |
| 0.6 | 12.7 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.5 | 1.6 | GO:0060061 | Spemann organizer formation(GO:0060061) |
| 0.5 | 2.2 | GO:0001575 | globoside metabolic process(GO:0001575) |
| 0.5 | 1.6 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) |
| 0.5 | 1.6 | GO:0015809 | arginine transport(GO:0015809) |
| 0.5 | 2.6 | GO:0046855 | phosphorylated carbohydrate dephosphorylation(GO:0046838) inositol phosphate dephosphorylation(GO:0046855) |
| 0.5 | 2.5 | GO:0046607 | positive regulation of centrosome cycle(GO:0046607) |
| 0.5 | 2.5 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.5 | 3.0 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.5 | 2.8 | GO:0007090 | obsolete regulation of S phase of mitotic cell cycle(GO:0007090) |
| 0.5 | 1.4 | GO:0051083 | 'de novo' cotranslational protein folding(GO:0051083) |
| 0.5 | 1.4 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.5 | 1.4 | GO:0097237 | cellular response to antibiotic(GO:0071236) cellular response to toxic substance(GO:0097237) |
| 0.4 | 1.3 | GO:0046080 | dUTP metabolic process(GO:0046080) |
| 0.4 | 1.3 | GO:0045875 | negative regulation of sister chromatid cohesion(GO:0045875) |
| 0.4 | 1.7 | GO:0071894 | histone H2B conserved C-terminal lysine ubiquitination(GO:0071894) |
| 0.4 | 1.3 | GO:0001830 | trophectodermal cell fate commitment(GO:0001830) |
| 0.4 | 1.7 | GO:0032898 | neurotrophin production(GO:0032898) |
| 0.4 | 1.3 | GO:0006344 | maintenance of chromatin silencing(GO:0006344) |
| 0.4 | 1.7 | GO:0042816 | pyridoxine metabolic process(GO:0008614) pyridoxine biosynthetic process(GO:0008615) vitamin B6 metabolic process(GO:0042816) vitamin B6 biosynthetic process(GO:0042819) |
| 0.4 | 1.2 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.4 | 1.2 | GO:0046732 | induction by symbiont of host defense response(GO:0044416) induction of host immune response by virus(GO:0046730) active induction of host immune response by virus(GO:0046732) modulation by virus of host immune response(GO:0075528) |
| 0.4 | 0.8 | GO:0035751 | lysosomal lumen acidification(GO:0007042) regulation of lysosomal lumen pH(GO:0035751) |
| 0.4 | 2.7 | GO:0033504 | floor plate development(GO:0033504) |
| 0.4 | 0.4 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.4 | 1.5 | GO:0071168 | protein localization to chromatin(GO:0071168) |
| 0.4 | 3.8 | GO:0001866 | NK T cell proliferation(GO:0001866) |
| 0.4 | 1.5 | GO:0033602 | negative regulation of gamma-aminobutyric acid secretion(GO:0014053) negative regulation of dopamine secretion(GO:0033602) |
| 0.4 | 1.1 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.4 | 1.1 | GO:0000354 | cis assembly of pre-catalytic spliceosome(GO:0000354) |
| 0.4 | 1.1 | GO:0006015 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 0.4 | 1.1 | GO:0046102 | inosine metabolic process(GO:0046102) |
| 0.4 | 1.1 | GO:0061117 | negative regulation of cardiac muscle tissue growth(GO:0055022) negative regulation of cardiac muscle tissue development(GO:0055026) negative regulation of cardiac muscle cell proliferation(GO:0060044) negative regulation of heart growth(GO:0061117) |
| 0.4 | 5.9 | GO:0007020 | microtubule nucleation(GO:0007020) |
| 0.4 | 1.1 | GO:0003289 | septum primum development(GO:0003284) atrial septum primum morphogenesis(GO:0003289) |
| 0.4 | 1.4 | GO:0071218 | cellular response to misfolded protein(GO:0071218) |
| 0.4 | 0.7 | GO:0070544 | histone H3-K36 demethylation(GO:0070544) |
| 0.3 | 3.1 | GO:1900087 | traversing start control point of mitotic cell cycle(GO:0007089) positive regulation of G1/S transition of mitotic cell cycle(GO:1900087) positive regulation of cell cycle G1/S phase transition(GO:1902808) |
| 0.3 | 8.4 | GO:0051225 | spindle assembly(GO:0051225) |
| 0.3 | 1.0 | GO:0002514 | B cell tolerance induction(GO:0002514) regulation of B cell tolerance induction(GO:0002661) positive regulation of B cell tolerance induction(GO:0002663) negative regulation of B cell mediated immunity(GO:0002713) negative regulation of immunoglobulin mediated immune response(GO:0002890) left/right pattern formation(GO:0060972) |
| 0.3 | 1.3 | GO:0060976 | positive regulation of vascular wound healing(GO:0035470) coronary vasculature development(GO:0060976) |
| 0.3 | 1.3 | GO:0046671 | regulation of nitrogen utilization(GO:0006808) positive regulation of neuron maturation(GO:0014042) nitrogen utilization(GO:0019740) cochlear nucleus development(GO:0021747) negative regulation of cellular pH reduction(GO:0032848) glial cell apoptotic process(GO:0034349) CD8-positive, alpha-beta T cell lineage commitment(GO:0043375) negative regulation of retinal cell programmed cell death(GO:0046671) positive regulation of cell maturation(GO:1903431) |
| 0.3 | 2.2 | GO:0031657 | regulation of cyclin-dependent protein serine/threonine kinase activity involved in G1/S transition of mitotic cell cycle(GO:0031657) positive regulation of cyclin-dependent protein serine/threonine kinase activity involved in G1/S transition of mitotic cell cycle(GO:0031659) |
| 0.3 | 0.6 | GO:0061188 | regulation of chromatin silencing at rDNA(GO:0061187) negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 0.3 | 0.6 | GO:0010834 | obsolete telomere maintenance via telomere shortening(GO:0010834) |
| 0.3 | 2.5 | GO:1901985 | positive regulation of histone acetylation(GO:0035066) positive regulation of protein acetylation(GO:1901985) positive regulation of peptidyl-lysine acetylation(GO:2000758) |
| 0.3 | 2.9 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.3 | 1.2 | GO:0034723 | DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.3 | 1.5 | GO:0007095 | mitotic G2 DNA damage checkpoint(GO:0007095) |
| 0.3 | 3.9 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.3 | 1.8 | GO:0006422 | aspartyl-tRNA aminoacylation(GO:0006422) |
| 0.3 | 1.2 | GO:0045199 | maintenance of epithelial cell apical/basal polarity(GO:0045199) |
| 0.3 | 1.2 | GO:0001705 | ectoderm formation(GO:0001705) |
| 0.3 | 0.9 | GO:0015959 | diadenosine polyphosphate metabolic process(GO:0015959) |
| 0.3 | 1.2 | GO:0048642 | negative regulation of skeletal muscle tissue development(GO:0048642) |
| 0.3 | 1.7 | GO:0061099 | negative regulation of protein tyrosine kinase activity(GO:0061099) |
| 0.3 | 2.6 | GO:0006546 | glycine catabolic process(GO:0006546) |
| 0.3 | 0.8 | GO:0032927 | positive regulation of activin receptor signaling pathway(GO:0032927) |
| 0.3 | 2.5 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.3 | 0.8 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.3 | 2.2 | GO:0051481 | negative regulation of cytosolic calcium ion concentration(GO:0051481) |
| 0.3 | 1.6 | GO:0034976 | response to endoplasmic reticulum stress(GO:0034976) |
| 0.3 | 0.5 | GO:0032201 | telomere maintenance via semi-conservative replication(GO:0032201) |
| 0.3 | 0.8 | GO:0060235 | lens induction in camera-type eye(GO:0060235) |
| 0.3 | 1.9 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.3 | 0.8 | GO:0007206 | phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.3 | 1.8 | GO:0060272 | embryonic skeletal joint morphogenesis(GO:0060272) embryonic skeletal joint development(GO:0072498) |
| 0.3 | 1.0 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.2 | 1.7 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.2 | 3.6 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.2 | 1.2 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.2 | 2.2 | GO:0043545 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) molybdopterin cofactor biosynthetic process(GO:0032324) molybdopterin cofactor metabolic process(GO:0043545) prosthetic group metabolic process(GO:0051189) |
| 0.2 | 3.3 | GO:0043097 | pyrimidine-containing compound salvage(GO:0008655) pyrimidine nucleoside salvage(GO:0043097) |
| 0.2 | 1.0 | GO:0021830 | interneuron migration from the subpallium to the cortex(GO:0021830) cerebral cortex GABAergic interneuron migration(GO:0021853) cerebral cortex GABAergic interneuron development(GO:0021894) interneuron migration(GO:1904936) |
| 0.2 | 1.2 | GO:0048304 | positive regulation of isotype switching to IgG isotypes(GO:0048304) |
| 0.2 | 3.8 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.2 | 13.6 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.2 | 2.1 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.2 | 0.7 | GO:0071280 | epithelial fluid transport(GO:0042045) cellular response to copper ion(GO:0071280) cellular response to mercury ion(GO:0071288) |
| 0.2 | 2.5 | GO:0031498 | nucleosome disassembly(GO:0006337) chromatin disassembly(GO:0031498) protein-DNA complex disassembly(GO:0032986) |
| 0.2 | 0.7 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.2 | 0.9 | GO:0015917 | aminophospholipid transport(GO:0015917) |
| 0.2 | 1.1 | GO:0045722 | positive regulation of gluconeogenesis(GO:0045722) |
| 0.2 | 7.3 | GO:0000725 | double-strand break repair via homologous recombination(GO:0000724) recombinational repair(GO:0000725) |
| 0.2 | 5.0 | GO:0034508 | centromere complex assembly(GO:0034508) |
| 0.2 | 0.9 | GO:0060430 | globus pallidus development(GO:0021759) menarche(GO:0042696) lung saccule development(GO:0060430) Clara cell differentiation(GO:0060486) Type II pneumocyte differentiation(GO:0060510) |
| 0.2 | 0.6 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.2 | 3.4 | GO:0045742 | positive regulation of epidermal growth factor receptor signaling pathway(GO:0045742) positive regulation of ERBB signaling pathway(GO:1901186) |
| 0.2 | 1.0 | GO:0001823 | ureteric bud development(GO:0001657) mesonephros development(GO:0001823) mesonephric epithelium development(GO:0072163) mesonephric tubule development(GO:0072164) |
| 0.2 | 0.8 | GO:2000104 | cell cycle DNA replication(GO:0044786) regulation of DNA-dependent DNA replication(GO:0090329) negative regulation of DNA-dependent DNA replication(GO:2000104) |
| 0.2 | 0.4 | GO:0039703 | single stranded viral RNA replication via double stranded DNA intermediate(GO:0039692) viral RNA genome replication(GO:0039694) RNA replication(GO:0039703) |
| 0.2 | 0.8 | GO:0006783 | heme biosynthetic process(GO:0006783) heme metabolic process(GO:0042168) pigment biosynthetic process(GO:0046148) |
| 0.2 | 1.8 | GO:0043507 | positive regulation of JUN kinase activity(GO:0043507) |
| 0.2 | 1.2 | GO:0031665 | negative regulation of lipopolysaccharide-mediated signaling pathway(GO:0031665) |
| 0.2 | 0.6 | GO:0034551 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.2 | 0.6 | GO:0051661 | maintenance of centrosome location(GO:0051661) |
| 0.2 | 1.4 | GO:0016075 | rRNA catabolic process(GO:0016075) |
| 0.2 | 1.7 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.2 | 0.7 | GO:0006540 | glutamate decarboxylation to succinate(GO:0006540) |
| 0.2 | 0.9 | GO:0032933 | SREBP signaling pathway(GO:0032933) cellular response to sterol depletion(GO:0071501) |
| 0.2 | 1.7 | GO:0048712 | negative regulation of astrocyte differentiation(GO:0048712) |
| 0.2 | 1.4 | GO:0009113 | purine nucleobase biosynthetic process(GO:0009113) |
| 0.2 | 0.7 | GO:0003057 | regulation of the force of heart contraction by chemical signal(GO:0003057) |
| 0.2 | 1.8 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.2 | 0.9 | GO:0060999 | positive regulation of dendritic spine development(GO:0060999) positive regulation of dendritic spine morphogenesis(GO:0061003) |
| 0.2 | 3.7 | GO:0007052 | mitotic spindle organization(GO:0007052) |
| 0.2 | 1.4 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.2 | 6.3 | GO:0043484 | regulation of RNA splicing(GO:0043484) |
| 0.2 | 0.9 | GO:0033326 | cerebrospinal fluid secretion(GO:0033326) |
| 0.2 | 0.5 | GO:0051026 | chiasma assembly(GO:0051026) |
| 0.2 | 1.4 | GO:0006072 | glycerol-3-phosphate metabolic process(GO:0006072) |
| 0.2 | 1.9 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.2 | 0.5 | GO:0009134 | nucleoside diphosphate catabolic process(GO:0009134) ribonucleoside diphosphate catabolic process(GO:0009191) |
| 0.2 | 0.7 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.2 | 0.2 | GO:1901659 | nucleoside biosynthetic process(GO:0009163) glycosyl compound biosynthetic process(GO:1901659) |
| 0.2 | 1.8 | GO:0060762 | regulation of branching involved in mammary gland duct morphogenesis(GO:0060762) |
| 0.2 | 1.6 | GO:0016246 | RNA interference(GO:0016246) |
| 0.2 | 0.5 | GO:0006106 | fumarate metabolic process(GO:0006106) |
| 0.2 | 3.7 | GO:0000303 | response to superoxide(GO:0000303) |
| 0.2 | 0.8 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.2 | 1.3 | GO:0048625 | myoblast fate commitment(GO:0048625) |
| 0.2 | 0.8 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.2 | 3.4 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.2 | 3.2 | GO:0000291 | nuclear-transcribed mRNA catabolic process, exonucleolytic(GO:0000291) exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.2 | 1.8 | GO:0055069 | cellular zinc ion homeostasis(GO:0006882) zinc ion homeostasis(GO:0055069) |
| 0.2 | 8.7 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.2 | 0.5 | GO:1901419 | regulation of response to alcohol(GO:1901419) |
| 0.2 | 0.3 | GO:0033147 | negative regulation of intracellular estrogen receptor signaling pathway(GO:0033147) |
| 0.2 | 1.1 | GO:0046548 | retinal rod cell development(GO:0046548) |
| 0.1 | 2.5 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.1 | 3.7 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.1 | 0.4 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.1 | 1.6 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.1 | 0.7 | GO:0014820 | tonic smooth muscle contraction(GO:0014820) artery smooth muscle contraction(GO:0014824) |
| 0.1 | 0.6 | GO:0071459 | protein localization to kinetochore(GO:0034501) protein localization to chromosome, centromeric region(GO:0071459) |
| 0.1 | 1.0 | GO:0032057 | negative regulation of translational initiation in response to stress(GO:0032057) |
| 0.1 | 1.2 | GO:0046498 | S-adenosylhomocysteine metabolic process(GO:0046498) |
| 0.1 | 2.6 | GO:0006596 | polyamine biosynthetic process(GO:0006596) |
| 0.1 | 0.8 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.1 | 0.4 | GO:0031167 | rRNA methylation(GO:0031167) |
| 0.1 | 0.8 | GO:0007352 | zygotic specification of dorsal/ventral axis(GO:0007352) |
| 0.1 | 0.7 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.1 | 1.9 | GO:0006787 | porphyrin-containing compound catabolic process(GO:0006787) tetrapyrrole catabolic process(GO:0033015) heme catabolic process(GO:0042167) pigment catabolic process(GO:0046149) |
| 0.1 | 0.8 | GO:0015871 | choline transport(GO:0015871) |
| 0.1 | 0.7 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.1 | 1.2 | GO:0070189 | kynurenine metabolic process(GO:0070189) |
| 0.1 | 2.7 | GO:0006349 | regulation of gene expression by genetic imprinting(GO:0006349) |
| 0.1 | 20.4 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.1 | 0.6 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.1 | 1.3 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 0.1 | 0.5 | GO:0090267 | positive regulation of spindle checkpoint(GO:0090232) regulation of mitotic cell cycle spindle assembly checkpoint(GO:0090266) positive regulation of mitotic cell cycle spindle assembly checkpoint(GO:0090267) positive regulation of cell cycle checkpoint(GO:1901978) regulation of mitotic spindle checkpoint(GO:1903504) |
| 0.1 | 0.5 | GO:0021819 | layer formation in cerebral cortex(GO:0021819) |
| 0.1 | 0.3 | GO:0007288 | sperm axoneme assembly(GO:0007288) |
| 0.1 | 0.7 | GO:0051973 | positive regulation of telomerase activity(GO:0051973) |
| 0.1 | 2.1 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.1 | 0.2 | GO:0072205 | collecting duct development(GO:0072044) metanephric collecting duct development(GO:0072205) |
| 0.1 | 0.5 | GO:0006006 | glucose metabolic process(GO:0006006) |
| 0.1 | 0.8 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 0.1 | 1.3 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.1 | 0.2 | GO:0010792 | DNA double-strand break processing involved in repair via single-strand annealing(GO:0010792) double-strand break repair via single-strand annealing(GO:0045002) |
| 0.1 | 0.5 | GO:0051319 | mitotic G2 phase(GO:0000085) G2 phase(GO:0051319) |
| 0.1 | 0.2 | GO:0048752 | semicircular canal morphogenesis(GO:0048752) |
| 0.1 | 0.6 | GO:0001510 | RNA methylation(GO:0001510) |
| 0.1 | 1.2 | GO:0006378 | mRNA polyadenylation(GO:0006378) |
| 0.1 | 0.7 | GO:0042362 | fat-soluble vitamin biosynthetic process(GO:0042362) |
| 0.1 | 3.7 | GO:0000288 | nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:0000288) |
| 0.1 | 0.6 | GO:0001906 | cell killing(GO:0001906) |
| 0.1 | 0.6 | GO:0006325 | chromatin organization(GO:0006325) |
| 0.1 | 0.7 | GO:0070779 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.1 | 0.2 | GO:0033002 | muscle cell proliferation(GO:0033002) |
| 0.1 | 0.5 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.1 | 0.5 | GO:0006839 | mitochondrial transport(GO:0006839) |
| 0.1 | 0.4 | GO:0000183 | chromatin silencing at rDNA(GO:0000183) |
| 0.1 | 0.9 | GO:0032355 | response to estradiol(GO:0032355) |
| 0.1 | 0.4 | GO:0043923 | positive regulation by host of viral transcription(GO:0043923) |
| 0.1 | 0.5 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 0.1 | 0.4 | GO:0001942 | hair follicle development(GO:0001942) molting cycle process(GO:0022404) hair cycle process(GO:0022405) molting cycle(GO:0042303) hair cycle(GO:0042633) skin epidermis development(GO:0098773) |
| 0.1 | 0.3 | GO:0006047 | UDP-N-acetylglucosamine metabolic process(GO:0006047) |
| 0.1 | 1.1 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.1 | 1.6 | GO:0043550 | regulation of lipid kinase activity(GO:0043550) |
| 0.1 | 0.3 | GO:0071344 | diphosphate metabolic process(GO:0071344) |
| 0.1 | 0.4 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 | 1.3 | GO:0008635 | activation of cysteine-type endopeptidase activity involved in apoptotic process by cytochrome c(GO:0008635) |
| 0.1 | 0.7 | GO:0043982 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.1 | 0.4 | GO:0007097 | nuclear migration(GO:0007097) |
| 0.1 | 1.8 | GO:0032011 | ARF protein signal transduction(GO:0032011) |
| 0.1 | 2.9 | GO:0006284 | base-excision repair(GO:0006284) |
| 0.1 | 0.4 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.1 | 1.0 | GO:0070059 | intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress(GO:0070059) |
| 0.1 | 0.5 | GO:0007035 | vacuolar acidification(GO:0007035) |
| 0.1 | 0.3 | GO:0071848 | regulation of fever generation by regulation of prostaglandin secretion(GO:0071810) positive regulation of fever generation by positive regulation of prostaglandin secretion(GO:0071812) TNFSF11-mediated signaling pathway(GO:0071847) positive regulation of ERK1 and ERK2 cascade via TNFSF11-mediated signaling(GO:0071848) regulation of fever generation by prostaglandin secretion(GO:0100009) |
| 0.1 | 2.2 | GO:0015949 | nucleobase-containing small molecule interconversion(GO:0015949) |
| 0.1 | 2.4 | GO:0030851 | granulocyte differentiation(GO:0030851) |
| 0.1 | 1.2 | GO:0014037 | Schwann cell differentiation(GO:0014037) |
| 0.1 | 0.4 | GO:0006107 | oxaloacetate metabolic process(GO:0006107) |
| 0.1 | 0.3 | GO:0000389 | mRNA 3'-splice site recognition(GO:0000389) |
| 0.1 | 0.7 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.1 | 0.2 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.1 | 0.9 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.1 | 0.7 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.1 | 0.2 | GO:0042339 | keratan sulfate metabolic process(GO:0042339) |
| 0.1 | 2.2 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.1 | 0.2 | GO:0070561 | vitamin D receptor signaling pathway(GO:0070561) |
| 0.1 | 0.5 | GO:0045070 | positive regulation of viral genome replication(GO:0045070) |
| 0.1 | 0.4 | GO:0048550 | negative regulation of pinocytosis(GO:0048550) |
| 0.1 | 0.2 | GO:0045603 | positive regulation of endothelial cell differentiation(GO:0045603) |
| 0.1 | 3.8 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.1 | 0.2 | GO:0021779 | oligodendrocyte cell fate commitment(GO:0021779) |
| 0.1 | 1.0 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.1 | 1.3 | GO:0032092 | positive regulation of protein binding(GO:0032092) |
| 0.1 | 2.0 | GO:0030308 | negative regulation of cell growth(GO:0030308) |
| 0.1 | 0.9 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.1 | 0.8 | GO:0007062 | sister chromatid cohesion(GO:0007062) |
| 0.1 | 1.4 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 0.1 | 0.1 | GO:0036260 | 7-methylguanosine mRNA capping(GO:0006370) 7-methylguanosine RNA capping(GO:0009452) RNA capping(GO:0036260) |
| 0.1 | 1.2 | GO:0008380 | RNA splicing(GO:0008380) |
| 0.1 | 0.3 | GO:0000956 | nuclear-transcribed mRNA catabolic process(GO:0000956) |
| 0.1 | 0.2 | GO:0032223 | negative regulation of synaptic transmission, cholinergic(GO:0032223) neurotransmitter receptor biosynthetic process(GO:0045212) |
| 0.1 | 0.4 | GO:0045657 | positive regulation of monocyte differentiation(GO:0045657) |
| 0.1 | 1.3 | GO:1903902 | positive regulation of viral process(GO:0048524) positive regulation of viral transcription(GO:0050434) positive regulation of viral life cycle(GO:1903902) |
| 0.1 | 0.5 | GO:0060412 | ventricular septum morphogenesis(GO:0060412) |
| 0.1 | 0.7 | GO:0032613 | interleukin-10 production(GO:0032613) |
| 0.1 | 0.9 | GO:0006972 | hyperosmotic response(GO:0006972) |
| 0.1 | 0.8 | GO:0044843 | G1/S transition of mitotic cell cycle(GO:0000082) cell cycle G1/S phase transition(GO:0044843) |
| 0.1 | 0.3 | GO:0006975 | DNA damage induced protein phosphorylation(GO:0006975) |
| 0.1 | 2.0 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.1 | 0.4 | GO:0009128 | AMP catabolic process(GO:0006196) nucleoside monophosphate catabolic process(GO:0009125) purine nucleoside monophosphate catabolic process(GO:0009128) ribonucleoside monophosphate catabolic process(GO:0009158) purine ribonucleoside monophosphate catabolic process(GO:0009169) |
| 0.1 | 1.8 | GO:0048709 | oligodendrocyte differentiation(GO:0048709) |
| 0.1 | 0.4 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.1 | 0.8 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.1 | 0.1 | GO:0045213 | neurotransmitter receptor metabolic process(GO:0045213) |
| 0.1 | 0.9 | GO:0050812 | acetyl-CoA biosynthetic process from pyruvate(GO:0006086) regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) regulation of acyl-CoA biosynthetic process(GO:0050812) |
| 0.1 | 4.0 | GO:0006959 | humoral immune response(GO:0006959) |
| 0.1 | 0.1 | GO:0035025 | positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.1 | 0.4 | GO:0007140 | male meiosis(GO:0007140) |
| 0.1 | 0.2 | GO:0031954 | positive regulation of protein autophosphorylation(GO:0031954) |
| 0.1 | 0.4 | GO:0032464 | regulation of protein homooligomerization(GO:0032462) positive regulation of protein homooligomerization(GO:0032464) |
| 0.1 | 0.6 | GO:0009404 | toxin metabolic process(GO:0009404) |
| 0.1 | 1.2 | GO:0043666 | regulation of phosphoprotein phosphatase activity(GO:0043666) |
| 0.1 | 1.4 | GO:0042274 | ribosomal small subunit biogenesis(GO:0042274) |
| 0.1 | 0.3 | GO:0006002 | fructose 6-phosphate metabolic process(GO:0006002) |
| 0.1 | 1.6 | GO:0048146 | positive regulation of fibroblast proliferation(GO:0048146) |
| 0.1 | 0.5 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.1 | 0.2 | GO:0000239 | pachytene(GO:0000239) |
| 0.1 | 0.4 | GO:0006477 | protein sulfation(GO:0006477) |
| 0.1 | 2.5 | GO:0000236 | mitotic prometaphase(GO:0000236) |
| 0.1 | 0.3 | GO:0015742 | alpha-ketoglutarate transport(GO:0015742) |
| 0.1 | 1.4 | GO:0006490 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) oligosaccharide-lipid intermediate biosynthetic process(GO:0006490) |
| 0.1 | 0.2 | GO:0045900 | negative regulation of translational elongation(GO:0045900) |
| 0.1 | 0.5 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) |
| 0.1 | 0.5 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.1 | 0.5 | GO:0006213 | pyrimidine nucleoside metabolic process(GO:0006213) |
| 0.1 | 0.8 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) extrinsic apoptotic signaling pathway(GO:0097191) |
| 0.1 | 0.2 | GO:0019919 | pathogenesis(GO:0009405) peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) |
| 0.1 | 0.3 | GO:0048015 | phosphatidylinositol-mediated signaling(GO:0048015) inositol lipid-mediated signaling(GO:0048017) |
| 0.1 | 0.4 | GO:0001701 | in utero embryonic development(GO:0001701) |
| 0.1 | 0.6 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.1 | 1.1 | GO:0030593 | neutrophil chemotaxis(GO:0030593) neutrophil migration(GO:1990266) |
| 0.1 | 0.6 | GO:0045351 | type I interferon biosynthetic process(GO:0045351) |
| 0.1 | 0.4 | GO:0001554 | luteolysis(GO:0001554) |
| 0.1 | 0.4 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.1 | 0.2 | GO:0046697 | decidualization(GO:0046697) |
| 0.0 | 0.1 | GO:0051053 | negative regulation of DNA metabolic process(GO:0051053) |
| 0.0 | 0.6 | GO:0009607 | response to biotic stimulus(GO:0009607) |
| 0.0 | 0.2 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.0 | 0.1 | GO:0034626 | fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.1 | GO:0010891 | negative regulation of sequestering of triglyceride(GO:0010891) |
| 0.0 | 0.8 | GO:0001836 | release of cytochrome c from mitochondria(GO:0001836) |
| 0.0 | 0.4 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 0.0 | 0.2 | GO:0019987 | obsolete negative regulation of anti-apoptosis(GO:0019987) |
| 0.0 | 1.1 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 0.7 | GO:0008543 | fibroblast growth factor receptor signaling pathway(GO:0008543) cellular response to fibroblast growth factor stimulus(GO:0044344) response to fibroblast growth factor(GO:0071774) |
| 0.0 | 1.1 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 6.9 | GO:0000375 | RNA splicing, via transesterification reactions(GO:0000375) |
| 0.0 | 0.2 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.0 | 2.6 | GO:0051781 | positive regulation of cell division(GO:0051781) |
| 0.0 | 1.6 | GO:0006289 | nucleotide-excision repair(GO:0006289) |
| 0.0 | 0.2 | GO:0030578 | nuclear body organization(GO:0030575) PML body organization(GO:0030578) |
| 0.0 | 0.3 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.0 | 0.2 | GO:0030238 | male sex determination(GO:0030238) |
| 0.0 | 0.1 | GO:0010259 | multicellular organism aging(GO:0010259) |
| 0.0 | 0.2 | GO:0050774 | negative regulation of dendrite morphogenesis(GO:0050774) |
| 0.0 | 0.9 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.0 | 5.4 | GO:0006397 | mRNA processing(GO:0006397) |
| 0.0 | 0.1 | GO:0061084 | regulation of protein refolding(GO:0061083) negative regulation of protein refolding(GO:0061084) negative regulation of protein folding(GO:1903333) |
| 0.0 | 0.5 | GO:0009060 | aerobic respiration(GO:0009060) |
| 0.0 | 0.0 | GO:0060295 | regulation of cilium movement(GO:0003352) regulation of cilium beat frequency(GO:0003356) cilium movement involved in cell motility(GO:0060294) regulation of cilium movement involved in cell motility(GO:0060295) regulation of cilium beat frequency involved in ciliary motility(GO:0060296) regulation of cilium-dependent cell motility(GO:1902019) |
| 0.0 | 0.1 | GO:0051168 | nuclear export(GO:0051168) |
| 0.0 | 0.6 | GO:0001502 | cartilage condensation(GO:0001502) cell aggregation(GO:0098743) |
| 0.0 | 0.2 | GO:0006405 | RNA export from nucleus(GO:0006405) |
| 0.0 | 0.3 | GO:0046827 | positive regulation of protein export from nucleus(GO:0046827) |
| 0.0 | 0.1 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.0 | 0.4 | GO:0042987 | amyloid precursor protein catabolic process(GO:0042987) |
| 0.0 | 0.2 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.0 | 0.6 | GO:0016601 | Rac protein signal transduction(GO:0016601) |
| 0.0 | 0.3 | GO:0022027 | interkinetic nuclear migration(GO:0022027) |
| 0.0 | 0.2 | GO:0021978 | telencephalon regionalization(GO:0021978) |
| 0.0 | 0.8 | GO:0006401 | RNA catabolic process(GO:0006401) |
| 0.0 | 0.2 | GO:0048286 | lung alveolus development(GO:0048286) |
| 0.0 | 0.5 | GO:0001954 | positive regulation of cell-matrix adhesion(GO:0001954) |
| 0.0 | 0.4 | GO:0006743 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) |
| 0.0 | 0.7 | GO:0019835 | cytolysis(GO:0019835) |
| 0.0 | 0.2 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.0 | 1.3 | GO:0008033 | tRNA processing(GO:0008033) |
| 0.0 | 0.2 | GO:0015853 | adenine transport(GO:0015853) |
| 0.0 | 0.2 | GO:0006547 | histidine metabolic process(GO:0006547) |
| 0.0 | 1.0 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 0.1 | GO:0097061 | dendritic spine morphogenesis(GO:0060997) regulation of dendritic spine development(GO:0060998) regulation of dendritic spine morphogenesis(GO:0061001) dendritic spine organization(GO:0097061) |
| 0.0 | 0.2 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 0.6 | GO:0046676 | negative regulation of insulin secretion(GO:0046676) |
| 0.0 | 0.2 | GO:0006541 | glutamine metabolic process(GO:0006541) |
| 0.0 | 0.2 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) |
| 0.0 | 0.8 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.0 | 0.4 | GO:0051384 | response to glucocorticoid(GO:0051384) |
| 0.0 | 2.8 | GO:0044819 | mitotic G1 DNA damage checkpoint(GO:0031571) G1 DNA damage checkpoint(GO:0044783) mitotic G1/S transition checkpoint(GO:0044819) |
| 0.0 | 0.2 | GO:0003032 | detection of oxygen(GO:0003032) |
| 0.0 | 0.7 | GO:0007257 | activation of JUN kinase activity(GO:0007257) |
| 0.0 | 0.2 | GO:0006305 | DNA alkylation(GO:0006305) DNA methylation(GO:0006306) |
| 0.0 | 1.5 | GO:0006413 | translational initiation(GO:0006413) |
| 0.0 | 0.1 | GO:0051958 | methotrexate transport(GO:0051958) |
| 0.0 | 0.2 | GO:0042976 | activation of Janus kinase activity(GO:0042976) |
| 0.0 | 0.2 | GO:0042493 | response to drug(GO:0042493) |
| 0.0 | 0.9 | GO:0000272 | polysaccharide catabolic process(GO:0000272) |
| 0.0 | 0.3 | GO:0031050 | dsRNA fragmentation(GO:0031050) production of miRNAs involved in gene silencing by miRNA(GO:0035196) production of small RNA involved in gene silencing by RNA(GO:0070918) |
| 0.0 | 0.3 | GO:0061025 | membrane fusion(GO:0061025) |
| 0.0 | 0.1 | GO:0051147 | regulation of muscle cell differentiation(GO:0051147) positive regulation of muscle cell differentiation(GO:0051149) |
| 0.0 | 0.3 | GO:0061098 | positive regulation of protein tyrosine kinase activity(GO:0061098) |
| 0.0 | 0.4 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.0 | 0.3 | GO:0045823 | positive regulation of heart contraction(GO:0045823) |
| 0.0 | 0.1 | GO:0032229 | negative regulation of synaptic transmission, GABAergic(GO:0032229) |
| 0.0 | 0.2 | GO:0010165 | response to X-ray(GO:0010165) |
| 0.0 | 0.1 | GO:0042517 | positive regulation of tyrosine phosphorylation of Stat3 protein(GO:0042517) |
| 0.0 | 0.1 | GO:0050666 | regulation of sulfur amino acid metabolic process(GO:0031335) regulation of homocysteine metabolic process(GO:0050666) |
| 0.0 | 0.6 | GO:0018208 | peptidyl-proline modification(GO:0018208) |
| 0.0 | 1.1 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.3 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.0 | 0.2 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0006919) activation of cysteine-type endopeptidase activity(GO:0097202) |
| 0.0 | 0.3 | GO:0006613 | cotranslational protein targeting to membrane(GO:0006613) |
| 0.0 | 0.4 | GO:0008633 | obsolete activation of pro-apoptotic gene products(GO:0008633) |
| 0.0 | 1.4 | GO:0034134 | toll-like receptor 1 signaling pathway(GO:0034130) toll-like receptor 2 signaling pathway(GO:0034134) |
| 0.0 | 0.5 | GO:0007595 | lactation(GO:0007595) |
| 0.0 | 0.3 | GO:0001662 | behavioral fear response(GO:0001662) |
| 0.0 | 0.5 | GO:0016458 | gene silencing(GO:0016458) |
| 0.0 | 0.4 | GO:0000279 | M phase(GO:0000279) |
| 0.0 | 0.7 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.7 | GO:0050772 | positive regulation of axonogenesis(GO:0050772) |
| 0.0 | 1.2 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.0 | 0.1 | GO:0009136 | ADP biosynthetic process(GO:0006172) nucleoside diphosphate biosynthetic process(GO:0009133) purine nucleoside diphosphate biosynthetic process(GO:0009136) purine ribonucleoside diphosphate biosynthetic process(GO:0009180) ribonucleoside diphosphate biosynthetic process(GO:0009188) |
| 0.0 | 1.1 | GO:0042593 | carbohydrate homeostasis(GO:0033500) glucose homeostasis(GO:0042593) |
| 0.0 | 3.4 | GO:0006813 | potassium ion transport(GO:0006813) |
| 0.0 | 0.5 | GO:0036075 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.0 | 0.4 | GO:0007215 | glutamate receptor signaling pathway(GO:0007215) |
| 0.0 | 0.3 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.3 | GO:0006662 | glycerol ether metabolic process(GO:0006662) ether metabolic process(GO:0018904) |
| 0.0 | 0.4 | GO:0051897 | positive regulation of protein kinase B signaling(GO:0051897) |
| 0.0 | 0.4 | GO:0034199 | activation of protein kinase A activity(GO:0034199) |
| 0.0 | 0.4 | GO:0007398 | ectoderm development(GO:0007398) |
| 0.0 | 1.5 | GO:0007605 | sensory perception of sound(GO:0007605) |
| 0.0 | 0.3 | GO:0015800 | acidic amino acid transport(GO:0015800) L-glutamate transport(GO:0015813) |
| 0.0 | 0.2 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.5 | GO:0006302 | double-strand break repair(GO:0006302) |
| 0.0 | 0.7 | GO:0006094 | gluconeogenesis(GO:0006094) |
| 0.0 | 0.2 | GO:0019674 | NAD metabolic process(GO:0019674) |
| 0.0 | 0.5 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.0 | 0.6 | GO:0043038 | tRNA aminoacylation for protein translation(GO:0006418) amino acid activation(GO:0043038) tRNA aminoacylation(GO:0043039) |
| 0.0 | 1.6 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.0 | 0.8 | GO:0019221 | cytokine-mediated signaling pathway(GO:0019221) |
| 0.0 | 0.0 | GO:0014887 | muscle hypertrophy in response to stress(GO:0003299) cardiac muscle adaptation(GO:0014887) cardiac muscle hypertrophy in response to stress(GO:0014898) |
| 0.0 | 0.2 | GO:0042771 | intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:0042771) |
| 0.0 | 0.2 | GO:0051693 | actin filament capping(GO:0051693) |
| 0.0 | 0.3 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.0 | 0.6 | GO:0006200 | obsolete ATP catabolic process(GO:0006200) |
| 0.0 | 0.4 | GO:0051925 | obsolete regulation of calcium ion transport via voltage-gated calcium channel activity(GO:0051925) |
| 0.0 | 0.4 | GO:0070830 | bicellular tight junction assembly(GO:0070830) |
| 0.0 | 0.1 | GO:0030520 | intracellular estrogen receptor signaling pathway(GO:0030520) |
| 0.0 | 0.9 | GO:0007608 | sensory perception of smell(GO:0007608) |
| 0.0 | 0.3 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 0.0 | GO:0003321 | positive regulation of blood pressure by epinephrine-norepinephrine(GO:0003321) |
| 0.0 | 0.2 | GO:0042417 | dopamine metabolic process(GO:0042417) |
| 0.0 | 0.3 | GO:0097193 | intrinsic apoptotic signaling pathway(GO:0097193) |
| 0.0 | 0.7 | GO:0034339 | obsolete regulation of transcription from RNA polymerase II promoter by nuclear hormone receptor(GO:0034339) |
| 0.0 | 1.5 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.1 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.0 | 0.1 | GO:0045060 | negative T cell selection(GO:0043383) negative thymic T cell selection(GO:0045060) |
| 0.0 | 0.1 | GO:0032097 | adult feeding behavior(GO:0008343) positive regulation of response to food(GO:0032097) positive regulation of appetite(GO:0032100) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.7 | 14.0 | GO:0032302 | MutSbeta complex(GO:0032302) |
| 2.8 | 33.5 | GO:0042555 | MCM complex(GO:0042555) |
| 2.5 | 5.0 | GO:0032301 | MutSalpha complex(GO:0032301) |
| 1.4 | 9.8 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 1.4 | 19.4 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 1.1 | 5.5 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 1.1 | 4.4 | GO:0031436 | BRCA1-BARD1 complex(GO:0031436) |
| 1.1 | 5.3 | GO:0000444 | MIS12/MIND type complex(GO:0000444) |
| 1.0 | 3.1 | GO:0031074 | nucleocytoplasmic shuttling complex(GO:0031074) |
| 1.0 | 9.0 | GO:0000796 | condensin complex(GO:0000796) |
| 1.0 | 2.9 | GO:0000806 | Y chromosome(GO:0000806) |
| 0.9 | 2.8 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.9 | 0.9 | GO:0001740 | Barr body(GO:0001740) |
| 0.9 | 7.2 | GO:0042575 | DNA polymerase complex(GO:0042575) |
| 0.8 | 7.6 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.7 | 5.9 | GO:0005638 | lamin filament(GO:0005638) |
| 0.6 | 2.5 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.6 | 6.6 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.6 | 4.6 | GO:0030894 | replisome(GO:0030894) nuclear replisome(GO:0043601) |
| 0.5 | 2.7 | GO:0030893 | meiotic cohesin complex(GO:0030893) |
| 0.4 | 4.4 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.4 | 1.2 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.4 | 2.2 | GO:0019815 | immunoglobulin complex(GO:0019814) B cell receptor complex(GO:0019815) |
| 0.4 | 1.1 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.3 | 6.6 | GO:0005657 | replication fork(GO:0005657) |
| 0.3 | 1.0 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.3 | 1.3 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.3 | 4.9 | GO:0000783 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) |
| 0.3 | 1.5 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.3 | 1.1 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.3 | 2.2 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.3 | 1.6 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.3 | 4.4 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.3 | 1.8 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.3 | 0.8 | GO:0031372 | UBC13-MMS2 complex(GO:0031372) |
| 0.2 | 0.5 | GO:0045239 | tricarboxylic acid cycle enzyme complex(GO:0045239) |
| 0.2 | 1.9 | GO:0005683 | U7 snRNP(GO:0005683) |
| 0.2 | 3.1 | GO:0030673 | axolemma(GO:0030673) |
| 0.2 | 2.6 | GO:0005678 | obsolete chromatin assembly complex(GO:0005678) |
| 0.2 | 1.9 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.2 | 1.2 | GO:0005688 | U6 snRNP(GO:0005688) |
| 0.2 | 2.3 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.2 | 0.6 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.2 | 9.8 | GO:0044815 | DNA packaging complex(GO:0044815) |
| 0.2 | 5.8 | GO:0030057 | desmosome(GO:0030057) |
| 0.2 | 1.2 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.2 | 0.6 | GO:0001652 | granular component(GO:0001652) |
| 0.2 | 2.1 | GO:0016442 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.2 | 1.1 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.2 | 2.0 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.2 | 1.0 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.2 | 1.9 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.2 | 2.5 | GO:0005746 | mitochondrial respiratory chain(GO:0005746) |
| 0.2 | 7.9 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.2 | 10.1 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.2 | 1.8 | GO:0032039 | integrator complex(GO:0032039) |
| 0.2 | 0.8 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.2 | 0.6 | GO:0031312 | extrinsic component of organelle membrane(GO:0031312) |
| 0.2 | 0.5 | GO:0033655 | host(GO:0018995) host cell cytoplasm(GO:0030430) host cell part(GO:0033643) host intracellular part(GO:0033646) host cell cytoplasm part(GO:0033655) intracellular region of host(GO:0043656) host cell(GO:0043657) other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) |
| 0.2 | 3.1 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.2 | 2.3 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.2 | 0.9 | GO:0042584 | chromaffin granule(GO:0042583) chromaffin granule membrane(GO:0042584) |
| 0.1 | 0.3 | GO:0060199 | clathrin-sculpted glutamate transport vesicle(GO:0060199) clathrin-sculpted glutamate transport vesicle membrane(GO:0060203) |
| 0.1 | 1.1 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.1 | 0.5 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 1.3 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.1 | 0.4 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.1 | 0.6 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.1 | 2.8 | GO:0030532 | small nuclear ribonucleoprotein complex(GO:0030532) |
| 0.1 | 1.0 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.1 | 1.6 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.1 | 0.8 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.1 | 1.6 | GO:0005838 | proteasome regulatory particle(GO:0005838) |
| 0.1 | 0.6 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.1 | 4.5 | GO:0030530 | obsolete heterogeneous nuclear ribonucleoprotein complex(GO:0030530) |
| 0.1 | 1.3 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.1 | 0.7 | GO:0000931 | gamma-tubulin large complex(GO:0000931) gamma-tubulin ring complex(GO:0008274) |
| 0.1 | 0.7 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.1 | 4.9 | GO:0005761 | organellar ribosome(GO:0000313) mitochondrial ribosome(GO:0005761) |
| 0.1 | 1.3 | GO:1903561 | extracellular organelle(GO:0043230) extracellular exosome(GO:0070062) extracellular vesicle(GO:1903561) |
| 0.1 | 0.8 | GO:0000300 | obsolete peripheral to membrane of membrane fraction(GO:0000300) |
| 0.1 | 15.7 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.1 | 3.5 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.1 | 1.1 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 1.2 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.1 | 5.6 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.1 | 0.3 | GO:0000153 | cytoplasmic ubiquitin ligase complex(GO:0000153) |
| 0.1 | 0.2 | GO:0000792 | heterochromatin(GO:0000792) |
| 0.1 | 2.8 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 9.1 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.1 | 2.4 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.1 | 0.6 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.1 | 0.8 | GO:0045211 | postsynaptic membrane(GO:0045211) postsynapse(GO:0098794) |
| 0.1 | 7.1 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 0.4 | GO:0071062 | alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 0.1 | 0.5 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 0.2 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.1 | 5.9 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.1 | 2.2 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.1 | 0.3 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 0.4 | GO:0005594 | collagen type IX trimer(GO:0005594) |
| 0.1 | 5.5 | GO:0000793 | condensed chromosome(GO:0000793) |
| 0.1 | 1.4 | GO:0000178 | exosome (RNase complex)(GO:0000178) |
| 0.1 | 2.2 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 4.3 | GO:0008287 | protein serine/threonine phosphatase complex(GO:0008287) phosphatase complex(GO:1903293) |
| 0.1 | 0.5 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.1 | 2.3 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.1 | 1.0 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.1 | 5.8 | GO:1990204 | oxidoreductase complex(GO:1990204) |
| 0.1 | 0.9 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.1 | 0.3 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.1 | 0.4 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.1 | 2.9 | GO:0019717 | obsolete synaptosome(GO:0019717) |
| 0.1 | 0.5 | GO:0033179 | proton-transporting V-type ATPase, V0 domain(GO:0033179) |
| 0.1 | 0.6 | GO:0005697 | telomerase holoenzyme complex(GO:0005697) |
| 0.1 | 0.8 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 1.8 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 0.8 | GO:0043034 | costamere(GO:0043034) |
| 0.1 | 0.5 | GO:0014704 | intercalated disc(GO:0014704) cell-cell contact zone(GO:0044291) |
| 0.1 | 0.5 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.1 | 19.0 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.1 | 1.3 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.1 | 0.3 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.1 | 5.2 | GO:0030141 | secretory granule(GO:0030141) |
| 0.1 | 0.7 | GO:0061200 | clathrin-sculpted gamma-aminobutyric acid transport vesicle(GO:0061200) clathrin-sculpted gamma-aminobutyric acid transport vesicle membrane(GO:0061202) |
| 0.1 | 0.1 | GO:0042597 | outer membrane-bounded periplasmic space(GO:0030288) periplasmic space(GO:0042597) |
| 0.0 | 1.3 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 0.6 | GO:0098878 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.0 | 0.6 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.0 | 3.7 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 0.4 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 0.8 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.0 | 0.3 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 0.8 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 2.6 | GO:0031228 | intrinsic component of Golgi membrane(GO:0031228) |
| 0.0 | 0.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 0.4 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.0 | 1.4 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.2 | GO:0002102 | podosome(GO:0002102) |
| 0.0 | 9.5 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 32.3 | GO:0005654 | nucleoplasm(GO:0005654) |
| 0.0 | 0.2 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) signal recognition particle(GO:0048500) |
| 0.0 | 0.6 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.2 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 30.2 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 0.2 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 1.4 | GO:0005626 | obsolete insoluble fraction(GO:0005626) |
| 0.0 | 0.7 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.3 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.0 | 1.6 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.0 | 0.8 | GO:0005605 | basal lamina(GO:0005605) |
| 0.0 | 0.6 | GO:1990904 | intracellular ribonucleoprotein complex(GO:0030529) ribonucleoprotein complex(GO:1990904) |
| 0.0 | 1.2 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.1 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.3 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.0 | 1.9 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 0.7 | GO:0043197 | dendritic spine(GO:0043197) |
| 0.0 | 0.3 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 1.2 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.0 | 2.3 | GO:0005840 | ribosome(GO:0005840) |
| 0.0 | 0.2 | GO:0046930 | pore complex(GO:0046930) |
| 0.0 | 0.6 | GO:0000922 | spindle pole(GO:0000922) |
| 0.0 | 1.6 | GO:0005694 | chromosome(GO:0005694) |
| 0.0 | 0.2 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.0 | 0.3 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.2 | GO:0098643 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.0 | 1.7 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 0.3 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 0.2 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.4 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.4 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 2.4 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 40.9 | GO:0005634 | nucleus(GO:0005634) |
| 0.0 | 0.4 | GO:0030496 | midbody(GO:0030496) |
| 0.0 | 0.2 | GO:0030175 | filopodium(GO:0030175) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.7 | 14.0 | GO:0000404 | heteroduplex DNA loop binding(GO:0000404) double-strand/single-strand DNA junction binding(GO:0000406) dinucleotide repeat insertion binding(GO:0032181) |
| 2.5 | 5.0 | GO:0032357 | single guanine insertion binding(GO:0032142) single thymine insertion binding(GO:0032143) oxidized DNA binding(GO:0032356) oxidized purine DNA binding(GO:0032357) |
| 1.9 | 7.5 | GO:0061731 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 1.8 | 9.2 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 1.8 | 5.5 | GO:0003896 | DNA primase activity(GO:0003896) |
| 1.6 | 9.7 | GO:0003910 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 1.3 | 3.9 | GO:0004146 | dihydrofolate reductase activity(GO:0004146) |
| 1.2 | 3.7 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 1.2 | 4.7 | GO:0060422 | peptidyl-dipeptidase inhibitor activity(GO:0060422) |
| 0.9 | 1.9 | GO:0070215 | obsolete MDM2 binding(GO:0070215) |
| 0.9 | 2.7 | GO:0004797 | thymidine kinase activity(GO:0004797) |
| 0.9 | 3.5 | GO:0016211 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.8 | 5.6 | GO:0008296 | 3'-5'-exodeoxyribonuclease activity(GO:0008296) |
| 0.8 | 6.2 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.7 | 4.4 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.6 | 1.8 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.6 | 23.0 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.5 | 3.8 | GO:0008107 | galactoside 2-alpha-L-fucosyltransferase activity(GO:0008107) alpha-(1,2)-fucosyltransferase activity(GO:0031127) |
| 0.5 | 2.2 | GO:0004727 | prenylated protein tyrosine phosphatase activity(GO:0004727) |
| 0.5 | 1.6 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.5 | 1.5 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.5 | 2.0 | GO:0004074 | biliverdin reductase activity(GO:0004074) |
| 0.5 | 2.0 | GO:0008934 | inositol monophosphate 1-phosphatase activity(GO:0008934) inositol monophosphate phosphatase activity(GO:0052834) |
| 0.5 | 1.4 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.5 | 1.8 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.4 | 1.3 | GO:0004170 | dUTP diphosphatase activity(GO:0004170) |
| 0.4 | 1.3 | GO:0045155 | electron transporter, transferring electrons from CoQH2-cytochrome c reductase complex and cytochrome c oxidase complex activity(GO:0045155) |
| 0.4 | 1.7 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.4 | 1.7 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.4 | 0.8 | GO:0016362 | activin receptor activity, type II(GO:0016362) |
| 0.4 | 1.2 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.4 | 2.7 | GO:0004439 | phosphatidylinositol-4,5-bisphosphate 5-phosphatase activity(GO:0004439) |
| 0.4 | 1.5 | GO:0046404 | ATP-dependent polydeoxyribonucleotide 5'-hydroxyl-kinase activity(GO:0046404) polydeoxyribonucleotide kinase activity(GO:0051733) ATP-dependent polynucleotide kinase activity(GO:0051734) |
| 0.4 | 2.6 | GO:0042978 | ornithine decarboxylase inhibitor activity(GO:0008073) ornithine decarboxylase activator activity(GO:0042978) |
| 0.4 | 1.1 | GO:0016534 | cyclin-dependent protein kinase 5 activator activity(GO:0016534) |
| 0.4 | 3.0 | GO:0015166 | polyol transmembrane transporter activity(GO:0015166) |
| 0.4 | 2.2 | GO:0004749 | ribose phosphate diphosphokinase activity(GO:0004749) |
| 0.4 | 5.9 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.4 | 9.1 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.4 | 5.4 | GO:0016565 | obsolete general transcriptional repressor activity(GO:0016565) |
| 0.4 | 18.9 | GO:0042054 | histone methyltransferase activity(GO:0042054) |
| 0.3 | 1.0 | GO:0030151 | molybdenum ion binding(GO:0030151) |
| 0.3 | 1.4 | GO:0004852 | uroporphyrinogen-III synthase activity(GO:0004852) |
| 0.3 | 1.4 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.3 | 1.0 | GO:0000033 | alpha-1,3-mannosyltransferase activity(GO:0000033) |
| 0.3 | 1.3 | GO:0002060 | purine nucleobase binding(GO:0002060) |
| 0.3 | 1.0 | GO:0016309 | 1-phosphatidylinositol-5-phosphate 4-kinase activity(GO:0016309) |
| 0.3 | 1.0 | GO:0003691 | double-stranded telomeric DNA binding(GO:0003691) |
| 0.3 | 0.6 | GO:0032139 | dinucleotide insertion or deletion binding(GO:0032139) |
| 0.3 | 2.2 | GO:0030020 | extracellular matrix structural constituent conferring tensile strength(GO:0030020) |
| 0.3 | 2.5 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.3 | 0.3 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.3 | 2.5 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.3 | 0.9 | GO:0015247 | aminophospholipid transporter activity(GO:0015247) |
| 0.3 | 1.8 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.3 | 1.2 | GO:0070548 | 1-aminocyclopropane-1-carboxylate synthase activity(GO:0016847) L-glutamine aminotransferase activity(GO:0070548) |
| 0.3 | 1.2 | GO:0052590 | sn-glycerol-3-phosphate:ubiquinone oxidoreductase activity(GO:0052590) sn-glycerol-3-phosphate:ubiquinone-8 oxidoreductase activity(GO:0052591) |
| 0.3 | 1.7 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.3 | 1.2 | GO:0043398 | HLH domain binding(GO:0043398) |
| 0.3 | 0.8 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.3 | 0.8 | GO:0016731 | ferredoxin-NADP+ reductase activity(GO:0004324) NADPH-adrenodoxin reductase activity(GO:0015039) oxidoreductase activity, acting on iron-sulfur proteins as donors(GO:0016730) oxidoreductase activity, acting on iron-sulfur proteins as donors, NAD or NADP as acceptor(GO:0016731) |
| 0.3 | 3.4 | GO:0016896 | exoribonuclease activity, producing 5'-phosphomonoesters(GO:0016896) |
| 0.3 | 2.6 | GO:0003701 | obsolete RNA polymerase I transcription factor activity(GO:0003701) |
| 0.3 | 2.5 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.3 | 2.0 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.2 | 1.0 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 0.2 | 0.7 | GO:0031755 | endothelial differentiation G-protein coupled receptor binding(GO:0031753) Edg-2 lysophosphatidic acid receptor binding(GO:0031755) |
| 0.2 | 1.2 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.2 | 3.3 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.2 | 1.4 | GO:0016564 | obsolete transcription repressor activity(GO:0016564) |
| 0.2 | 0.2 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 0.2 | 6.2 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.2 | 0.7 | GO:0016524 | latrotoxin receptor activity(GO:0016524) |
| 0.2 | 0.9 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.2 | 13.0 | GO:0003823 | antigen binding(GO:0003823) |
| 0.2 | 0.9 | GO:0004738 | pyruvate dehydrogenase activity(GO:0004738) pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.2 | 2.4 | GO:0061659 | ubiquitin-ubiquitin ligase activity(GO:0034450) ubiquitin protein ligase activity(GO:0061630) ubiquitin-like protein ligase activity(GO:0061659) |
| 0.2 | 0.6 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.2 | 1.9 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.2 | 3.9 | GO:0005542 | folic acid binding(GO:0005542) |
| 0.2 | 1.2 | GO:0002039 | p53 binding(GO:0002039) |
| 0.2 | 1.9 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.2 | 0.6 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.2 | 1.2 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.2 | 0.8 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.2 | 1.0 | GO:0008599 | protein phosphatase type 1 regulator activity(GO:0008599) |
| 0.2 | 2.3 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.2 | 4.6 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.2 | 1.5 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.2 | 2.4 | GO:0016918 | retinal binding(GO:0016918) |
| 0.2 | 2.8 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.2 | 0.7 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 0.2 | 1.1 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.2 | 0.4 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.2 | 0.5 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.2 | 1.0 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.2 | 4.8 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.2 | 9.0 | GO:0004180 | carboxypeptidase activity(GO:0004180) |
| 0.2 | 3.3 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.2 | 1.0 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.2 | 2.6 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.2 | 1.0 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.2 | 0.5 | GO:0004476 | mannose-6-phosphate isomerase activity(GO:0004476) |
| 0.2 | 0.5 | GO:0015217 | purine ribonucleotide transmembrane transporter activity(GO:0005346) ATP transmembrane transporter activity(GO:0005347) ATP:ADP antiporter activity(GO:0005471) ADP transmembrane transporter activity(GO:0015217) |
| 0.2 | 0.6 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.2 | 0.5 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 0.2 | 1.2 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.2 | 0.5 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.1 | 0.6 | GO:0000247 | C-8 sterol isomerase activity(GO:0000247) |
| 0.1 | 0.4 | GO:0042285 | UDP-xylosyltransferase activity(GO:0035252) xylosyltransferase activity(GO:0042285) |
| 0.1 | 2.5 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 2.4 | GO:0005521 | lamin binding(GO:0005521) |
| 0.1 | 1.8 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.1 | 0.7 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.1 | 6.0 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.1 | 4.8 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.1 | 3.0 | GO:0004716 | receptor signaling protein tyrosine kinase activity(GO:0004716) |
| 0.1 | 0.4 | GO:0004676 | 3-phosphoinositide-dependent protein kinase activity(GO:0004676) |
| 0.1 | 1.9 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.1 | 0.5 | GO:0032408 | MutLbeta complex binding(GO:0032406) MutSbeta complex binding(GO:0032408) |
| 0.1 | 0.4 | GO:0004613 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 0.1 | 1.6 | GO:0016634 | oxidoreductase activity, acting on the CH-CH group of donors, oxygen as acceptor(GO:0016634) |
| 0.1 | 0.7 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.1 | 0.8 | GO:0008409 | 5'-3' exonuclease activity(GO:0008409) |
| 0.1 | 0.3 | GO:0043008 | ATP-dependent protein binding(GO:0043008) |
| 0.1 | 0.7 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.1 | 1.0 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.1 | 0.6 | GO:0034061 | DNA polymerase activity(GO:0034061) |
| 0.1 | 0.1 | GO:0032564 | dATP binding(GO:0032564) |
| 0.1 | 3.9 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.1 | 0.9 | GO:0016151 | nickel cation binding(GO:0016151) |
| 0.1 | 0.6 | GO:0008967 | phosphoglycolate phosphatase activity(GO:0008967) |
| 0.1 | 13.7 | GO:0016566 | obsolete specific transcriptional repressor activity(GO:0016566) |
| 0.1 | 0.5 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.1 | 1.6 | GO:0050998 | nitric-oxide synthase binding(GO:0050998) |
| 0.1 | 1.8 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.1 | 3.7 | GO:0004527 | exonuclease activity(GO:0004527) |
| 0.1 | 0.7 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.1 | 3.8 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.1 | 0.3 | GO:0017130 | poly(C) RNA binding(GO:0017130) |
| 0.1 | 5.9 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.1 | 1.9 | GO:0004004 | ATP-dependent RNA helicase activity(GO:0004004) |
| 0.1 | 0.6 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.1 | 2.1 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 1.8 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.1 | 0.5 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 0.1 | 2.3 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.1 | 1.6 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.1 | 0.5 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) ubiquitin-like protein conjugating enzyme binding(GO:0044390) |
| 0.1 | 0.5 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 4.9 | GO:0042393 | histone binding(GO:0042393) |
| 0.1 | 0.7 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.1 | 0.6 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.1 | 0.4 | GO:0016404 | 15-hydroxyprostaglandin dehydrogenase (NAD+) activity(GO:0016404) |
| 0.1 | 0.3 | GO:0043199 | sulfate binding(GO:0043199) |
| 0.1 | 2.8 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.1 | 0.4 | GO:0004335 | galactokinase activity(GO:0004335) |
| 0.1 | 0.9 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.1 | 0.4 | GO:0032810 | sterol response element binding(GO:0032810) |
| 0.1 | 2.2 | GO:0030528 | obsolete transcription regulator activity(GO:0030528) |
| 0.1 | 1.5 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 1.0 | GO:0031369 | translation initiation factor binding(GO:0031369) |
| 0.1 | 1.3 | GO:0017153 | sodium:dicarboxylate symporter activity(GO:0017153) |
| 0.1 | 1.0 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.1 | 1.1 | GO:0019212 | phosphatase inhibitor activity(GO:0019212) |
| 0.1 | 0.6 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.1 | 0.7 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.1 | 0.4 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.1 | 0.3 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.1 | 13.0 | GO:0003702 | obsolete RNA polymerase II transcription factor activity(GO:0003702) |
| 0.1 | 0.4 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.1 | 0.5 | GO:0004047 | aminomethyltransferase activity(GO:0004047) |
| 0.1 | 2.5 | GO:0097472 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) cyclin-dependent protein kinase activity(GO:0097472) |
| 0.1 | 0.4 | GO:0001517 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) |
| 0.1 | 3.1 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.1 | 2.4 | GO:0016769 | transferase activity, transferring nitrogenous groups(GO:0016769) |
| 0.1 | 0.2 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 0.1 | 0.5 | GO:0034187 | obsolete apolipoprotein E binding(GO:0034187) |
| 0.1 | 0.8 | GO:0003995 | acyl-CoA dehydrogenase activity(GO:0003995) |
| 0.1 | 0.3 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.1 | 2.1 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 0.2 | GO:0005047 | signal recognition particle binding(GO:0005047) |
| 0.1 | 0.2 | GO:0004477 | methenyltetrahydrofolate cyclohydrolase activity(GO:0004477) |
| 0.1 | 2.1 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
| 0.1 | 1.2 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 3.4 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) RNA polymerase activity(GO:0034062) |
| 0.1 | 0.3 | GO:0070087 | chromo shadow domain binding(GO:0070087) |
| 0.1 | 0.7 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.1 | 4.0 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.1 | 0.5 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.1 | 0.7 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 0.1 | 5.2 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.1 | 0.5 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.1 | 0.6 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.1 | 0.7 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 1.2 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.1 | 0.3 | GO:0070061 | fructose binding(GO:0070061) |
| 0.1 | 0.3 | GO:0004427 | inorganic diphosphatase activity(GO:0004427) |
| 0.1 | 0.5 | GO:0017127 | cholesterol transporter activity(GO:0017127) |
| 0.1 | 1.0 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 0.7 | GO:0008187 | poly-pyrimidine tract binding(GO:0008187) |
| 0.1 | 0.9 | GO:0015605 | organophosphate ester transmembrane transporter activity(GO:0015605) |
| 0.1 | 0.3 | GO:0015140 | malate transmembrane transporter activity(GO:0015140) |
| 0.1 | 2.5 | GO:0050660 | flavin adenine dinucleotide binding(GO:0050660) |
| 0.1 | 0.9 | GO:0010181 | FMN binding(GO:0010181) |
| 0.1 | 1.1 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 0.6 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.1 | 0.4 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.1 | 0.4 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.1 | 2.0 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.1 | 1.0 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.1 | 0.7 | GO:0008641 | small protein activating enzyme activity(GO:0008641) |
| 0.1 | 0.4 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.1 | 0.4 | GO:0019104 | DNA N-glycosylase activity(GO:0019104) |
| 0.1 | 0.5 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.1 | 3.4 | GO:0004221 | obsolete ubiquitin thiolesterase activity(GO:0004221) |
| 0.1 | 0.4 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 0.3 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 0.1 | 1.8 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.1 | 1.1 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.1 | 0.4 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.1 | 0.4 | GO:0008321 | Ral guanyl-nucleotide exchange factor activity(GO:0008321) |
| 0.1 | 0.2 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 0.1 | 0.5 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.1 | 1.6 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 1.1 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 0.5 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.5 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 2.5 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 1.2 | GO:0001076 | transcription factor activity, RNA polymerase II transcription factor binding(GO:0001076) |
| 0.0 | 0.7 | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups(GO:0016780) |
| 0.0 | 0.2 | GO:0017163 | obsolete basal transcription repressor activity(GO:0017163) |
| 0.0 | 0.1 | GO:0033867 | Fas-activated serine/threonine kinase activity(GO:0033867) |
| 0.0 | 1.0 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 0.1 | GO:0032795 | heterotrimeric G-protein binding(GO:0032795) |
| 0.0 | 0.2 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.0 | 0.5 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.0 | 0.9 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.0 | 0.4 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.0 | 0.2 | GO:0003985 | acetyl-CoA C-acetyltransferase activity(GO:0003985) C-acetyltransferase activity(GO:0016453) |
| 0.0 | 0.4 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 0.4 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.0 | 1.9 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.2 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 0.2 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 0.5 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.0 | 2.1 | GO:0003704 | obsolete specific RNA polymerase II transcription factor activity(GO:0003704) |
| 0.0 | 0.4 | GO:0036459 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 1.9 | GO:0050661 | NADP binding(GO:0050661) |
| 0.0 | 0.3 | GO:0004835 | tubulin-tyrosine ligase activity(GO:0004835) |
| 0.0 | 0.9 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.0 | 0.3 | GO:0090599 | alpha-glucosidase activity(GO:0090599) |
| 0.0 | 0.7 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.2 | GO:0043548 | phosphatidylinositol 3-kinase binding(GO:0043548) |
| 0.0 | 0.1 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.5 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) lysophosphatidic acid acyltransferase activity(GO:0042171) lysophospholipid acyltransferase activity(GO:0071617) |
| 0.0 | 0.3 | GO:0005313 | L-glutamate transmembrane transporter activity(GO:0005313) acidic amino acid transmembrane transporter activity(GO:0015172) |
| 0.0 | 0.5 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.7 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.0 | 0.7 | GO:0008080 | N-acetyltransferase activity(GO:0008080) |
| 0.0 | 0.8 | GO:0000146 | microfilament motor activity(GO:0000146) |
| 0.0 | 1.7 | GO:0008026 | ATP-dependent helicase activity(GO:0008026) purine NTP-dependent helicase activity(GO:0070035) |
| 0.0 | 0.2 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) core promoter sequence-specific DNA binding(GO:0001046) core promoter binding(GO:0001047) |
| 0.0 | 0.2 | GO:0015288 | porin activity(GO:0015288) |
| 0.0 | 3.0 | GO:0001071 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
| 0.0 | 0.2 | GO:0070576 | vitamin D 24-hydroxylase activity(GO:0070576) |
| 0.0 | 0.9 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.1 | GO:0009922 | fatty acid elongase activity(GO:0009922) |
| 0.0 | 0.6 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 0.8 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.0 | 0.5 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.0 | 0.4 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.0 | 2.2 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 5.4 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.7 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 0.2 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.0 | 0.3 | GO:0046527 | glucosyltransferase activity(GO:0046527) |
| 0.0 | 0.0 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 0.5 | GO:0016646 | oxidoreductase activity, acting on the CH-NH group of donors, NAD or NADP as acceptor(GO:0016646) |
| 0.0 | 39.0 | GO:0003677 | DNA binding(GO:0003677) |
| 0.0 | 10.0 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.0 | 0.8 | GO:0017048 | Rho GTPase binding(GO:0017048) |
| 0.0 | 5.3 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.0 | 0.1 | GO:0004768 | stearoyl-CoA 9-desaturase activity(GO:0004768) acyl-CoA desaturase activity(GO:0016215) |
| 0.0 | 0.7 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.0 | 0.1 | GO:0003831 | beta-N-acetylglucosaminylglycopeptide beta-1,4-galactosyltransferase activity(GO:0003831) |
| 0.0 | 0.8 | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity(GO:0008757) |
| 0.0 | 0.1 | GO:0043560 | insulin receptor substrate binding(GO:0043560) |
| 0.0 | 0.4 | GO:0005048 | signal sequence binding(GO:0005048) |
| 0.0 | 0.1 | GO:0016802 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.5 | GO:0016814 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amidines(GO:0016814) |
| 0.0 | 0.3 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.3 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.0 | 0.4 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.0 | 0.7 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.0 | 0.1 | GO:0030371 | translation repressor activity(GO:0030371) |
| 0.0 | 0.2 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.0 | 0.5 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.5 | GO:0015405 | primary active transmembrane transporter activity(GO:0015399) P-P-bond-hydrolysis-driven transmembrane transporter activity(GO:0015405) ATPase activity, coupled to transmembrane movement of substances(GO:0042626) |
| 0.0 | 0.5 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 0.3 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.3 | GO:0016891 | endoribonuclease activity, producing 5'-phosphomonoesters(GO:0016891) |
| 0.0 | 0.1 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 0.3 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) |
| 0.0 | 0.2 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 0.2 | GO:0008320 | protein transmembrane transporter activity(GO:0008320) |
| 0.0 | 0.1 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.0 | 0.1 | GO:0042805 | actinin binding(GO:0042805) |
| 0.0 | 0.3 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.2 | GO:0004859 | phospholipase inhibitor activity(GO:0004859) |
| 0.0 | 0.0 | GO:0044620 | ACP phosphopantetheine attachment site binding involved in fatty acid biosynthetic process(GO:0000036) phosphopantetheine binding(GO:0031177) ACP phosphopantetheine attachment site binding(GO:0044620) prosthetic group binding(GO:0051192) |
| 0.0 | 0.2 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.0 | 0.1 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 0.5 | GO:0050136 | NADH dehydrogenase activity(GO:0003954) NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 6.8 | SA_G2_AND_M_PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.1 | 5.3 | ST_JNK_MAPK_PATHWAY | JNK MAPK Pathway |
| 0.1 | 6.2 | SIG_BCR_SIGNALING_PATHWAY | Members of the BCR signaling pathway |
| 0.1 | 1.6 | SA_CASPASE_CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 1.8 | ST_ERK1_ERK2_MAPK_PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 1.2 | ST_GAQ_PATHWAY | G alpha q Pathway |
| 0.0 | 0.7 | ST_P38_MAPK_PATHWAY | p38 MAPK Pathway |
| 0.0 | 1.1 | SA_MMP_CYTOKINE_CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.0 | 2.0 | ST_FAS_SIGNALING_PATHWAY | Fas Signaling Pathway |
| 0.0 | 0.4 | SA_PROGRAMMED_CELL_DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.0 | 1.5 | ST_T_CELL_SIGNAL_TRANSDUCTION | T Cell Signal Transduction |
| 0.0 | 0.2 | SA_PTEN_PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |