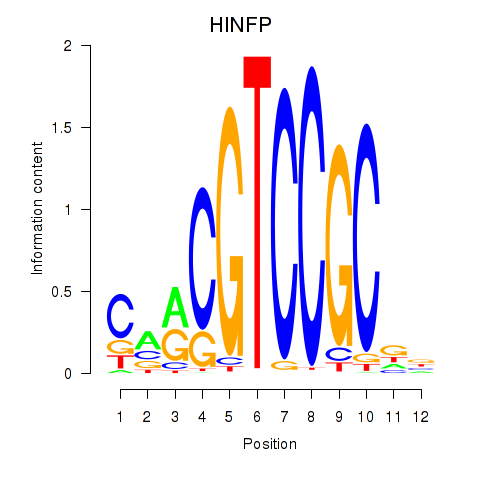
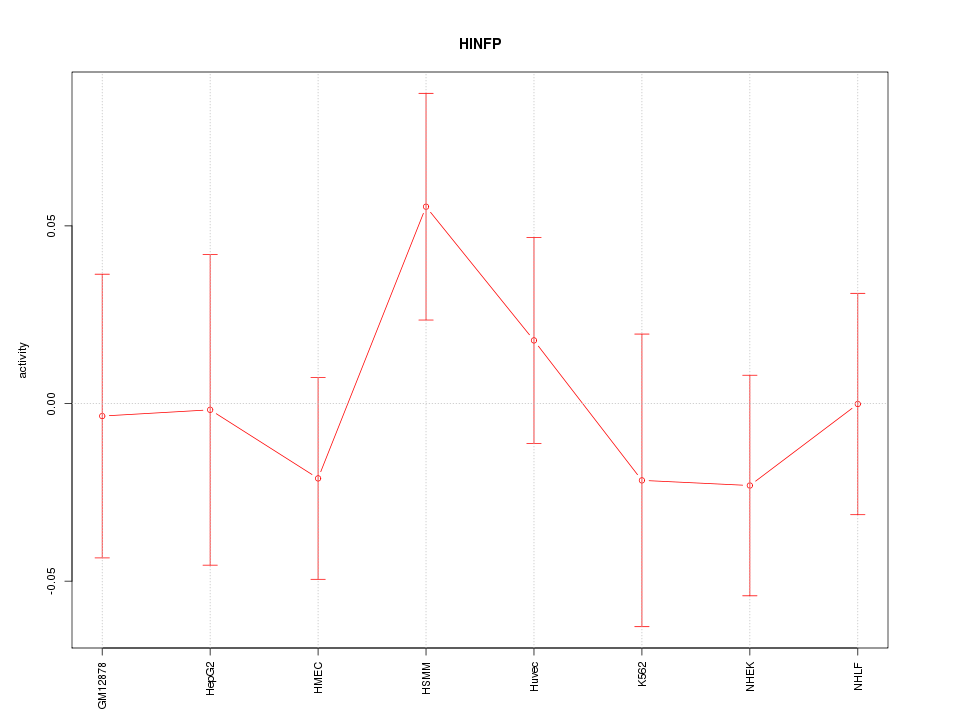
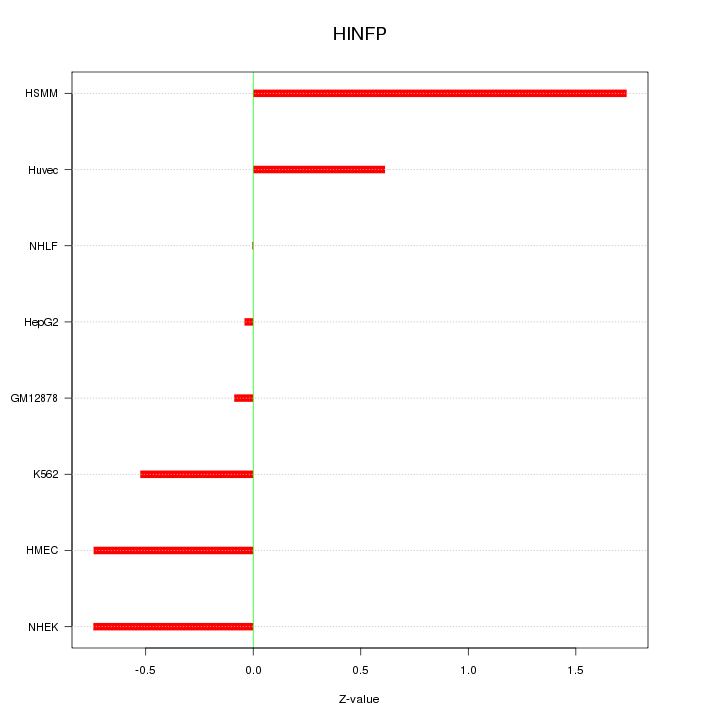
|
chr7_+_94023873
|
2.133
|
ENST00000297268.6
|
COL1A2
|
collagen, type I, alpha 2
|
|
chr5_-_136834982
|
1.042
|
ENST00000510689.1
ENST00000394945.1
|
SPOCK1
|
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 1
|
|
chr11_-_2162468
|
0.983
|
ENST00000434045.2
|
IGF2
|
insulin-like growth factor 2 (somatomedin A)
|
|
chr3_+_110790590
|
0.976
|
ENST00000485303.1
|
PVRL3
|
poliovirus receptor-related 3
|
|
chr11_-_2162162
|
0.958
|
ENST00000381389.1
|
IGF2
|
insulin-like growth factor 2 (somatomedin A)
|
|
chr10_-_62761188
|
0.925
|
ENST00000357917.4
|
RHOBTB1
|
Rho-related BTB domain containing 1
|
|
chr9_+_137533615
|
0.917
|
ENST00000371817.3
|
COL5A1
|
collagen, type V, alpha 1
|
|
chr3_-_38691119
|
0.859
|
ENST00000333535.4
ENST00000413689.1
ENST00000443581.1
ENST00000425664.1
ENST00000451551.2
|
SCN5A
|
sodium channel, voltage-gated, type V, alpha subunit
|
|
chr12_-_90102594
|
0.852
|
ENST00000428670.3
|
ATP2B1
|
ATPase, Ca++ transporting, plasma membrane 1
|
|
chr4_-_186733363
|
0.787
|
ENST00000393523.2
ENST00000393528.3
ENST00000449407.2
|
SORBS2
|
sorbin and SH3 domain containing 2
|
|
chr3_-_16555150
|
0.672
|
ENST00000334133.4
|
RFTN1
|
raftlin, lipid raft linker 1
|
|
chr17_-_46115122
|
0.625
|
ENST00000006101.4
|
COPZ2
|
coatomer protein complex, subunit zeta 2
|
|
chr6_-_139695757
|
0.566
|
ENST00000367651.2
|
CITED2
|
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2
|
|
chr16_+_6069586
|
0.476
|
ENST00000547372.1
|
RBFOX1
|
RNA binding protein, fox-1 homolog (C. elegans) 1
|
|
chr13_-_77460525
|
0.473
|
ENST00000377474.2
ENST00000317765.2
|
KCTD12
|
potassium channel tetramerization domain containing 12
|
|
chr3_+_49449636
|
0.454
|
ENST00000273590.3
|
TCTA
|
T-cell leukemia translocation altered
|
|
chr9_-_89562104
|
0.411
|
ENST00000298743.7
|
GAS1
|
growth arrest-specific 1
|
|
chr15_+_43803143
|
0.409
|
ENST00000382031.1
|
MAP1A
|
microtubule-associated protein 1A
|
|
chr13_-_36705425
|
0.391
|
ENST00000255448.4
ENST00000360631.3
ENST00000379892.4
|
DCLK1
|
doublecortin-like kinase 1
|
|
chr11_-_66336060
|
0.342
|
ENST00000310325.5
|
CTSF
|
cathepsin F
|
|
chr11_-_72353451
|
0.333
|
ENST00000376450.3
|
PDE2A
|
phosphodiesterase 2A, cGMP-stimulated
|
|
chr1_+_204797749
|
0.329
|
ENST00000367172.4
ENST00000367171.4
ENST00000367170.4
ENST00000338515.6
ENST00000339876.6
ENST00000338586.6
ENST00000539706.1
ENST00000360049.4
ENST00000367169.4
ENST00000446412.1
ENST00000403080.1
|
NFASC
|
neurofascin
|
|
chr5_+_72143988
|
0.286
|
ENST00000506351.2
|
TNPO1
|
transportin 1
|
|
chr1_-_94703118
|
0.285
|
ENST00000260526.6
ENST00000370217.3
|
ARHGAP29
|
Rho GTPase activating protein 29
|
|
chr17_-_1419878
|
0.282
|
ENST00000449479.1
ENST00000477910.1
ENST00000542125.1
ENST00000575172.1
|
INPP5K
|
inositol polyphosphate-5-phosphatase K
|
|
chr8_-_27462822
|
0.273
|
ENST00000522098.1
|
CLU
|
clusterin
|
|
chr4_+_123747834
|
0.266
|
ENST00000264498.3
|
FGF2
|
fibroblast growth factor 2 (basic)
|
|
chr3_-_49449521
|
0.262
|
ENST00000431929.1
ENST00000418115.1
|
RHOA
|
ras homolog family member A
|
|
chr13_+_25875785
|
0.259
|
ENST00000381747.3
|
NUPL1
|
nucleoporin like 1
|
|
chr13_-_41240717
|
0.258
|
ENST00000379561.5
|
FOXO1
|
forkhead box O1
|
|
chr17_-_1419914
|
0.251
|
ENST00000397335.3
ENST00000574561.1
|
INPP5K
|
inositol polyphosphate-5-phosphatase K
|
|
chr13_+_111806055
|
0.247
|
ENST00000218789.5
|
ARHGEF7
|
Rho guanine nucleotide exchange factor (GEF) 7
|
|
chr18_-_44336998
|
0.238
|
ENST00000315087.7
|
ST8SIA5
|
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 5
|
|
chr3_-_129612394
|
0.236
|
ENST00000505616.1
ENST00000426664.2
|
TMCC1
|
transmembrane and coiled-coil domain family 1
|
|
chr17_+_80186908
|
0.229
|
ENST00000582743.1
ENST00000578684.1
ENST00000577650.1
ENST00000582715.1
|
SLC16A3
|
solute carrier family 16 (monocarboxylate transporter), member 3
|
|
chr7_-_134143841
|
0.227
|
ENST00000285930.4
|
AKR1B1
|
aldo-keto reductase family 1, member B1 (aldose reductase)
|
|
chr3_-_49449350
|
0.225
|
ENST00000454011.2
ENST00000445425.1
ENST00000422781.1
|
RHOA
|
ras homolog family member A
|
|
chrX_-_119695279
|
0.221
|
ENST00000336592.6
|
CUL4B
|
cullin 4B
|
|
chr17_-_1420182
|
0.220
|
ENST00000421807.2
|
INPP5K
|
inositol polyphosphate-5-phosphatase K
|
|
chr8_+_42995548
|
0.218
|
ENST00000458501.2
ENST00000379644.4
|
HGSNAT
|
heparan-alpha-glucosaminide N-acetyltransferase
|
|
chr13_-_22178284
|
0.214
|
ENST00000468222.2
ENST00000382374.4
|
MICU2
|
mitochondrial calcium uptake 2
|
|
chr3_+_139654018
|
0.205
|
ENST00000458420.3
|
CLSTN2
|
calsyntenin 2
|
|
chr1_+_93913713
|
0.199
|
ENST00000604705.1
ENST00000370253.2
|
FNBP1L
|
formin binding protein 1-like
|
|
chrX_-_38186811
|
0.196
|
ENST00000318842.7
|
RPGR
|
retinitis pigmentosa GTPase regulator
|
|
chr17_+_46125707
|
0.191
|
ENST00000584137.1
ENST00000362042.3
ENST00000585291.1
ENST00000357480.5
|
NFE2L1
|
nuclear factor, erythroid 2-like 1
|
|
chr13_+_25875662
|
0.191
|
ENST00000381736.3
ENST00000463407.1
ENST00000381718.3
|
NUPL1
|
nucleoporin like 1
|
|
chr3_-_49377446
|
0.188
|
ENST00000351842.4
ENST00000416417.1
ENST00000415188.1
|
USP4
|
ubiquitin specific peptidase 4 (proto-oncogene)
|
|
chr12_+_102271129
|
0.188
|
ENST00000258534.8
|
DRAM1
|
DNA-damage regulated autophagy modulator 1
|
|
chr3_-_49377499
|
0.183
|
ENST00000265560.4
|
USP4
|
ubiquitin specific peptidase 4 (proto-oncogene)
|
|
chr3_-_156272924
|
0.177
|
ENST00000467789.1
ENST00000265044.2
|
SSR3
|
signal sequence receptor, gamma (translocon-associated protein gamma)
|
|
chr11_+_62104897
|
0.176
|
ENST00000415229.2
ENST00000535727.1
ENST00000301776.5
|
ASRGL1
|
asparaginase like 1
|
|
chr2_+_10442993
|
0.176
|
ENST00000423674.1
ENST00000307845.3
|
HPCAL1
|
hippocalcin-like 1
|
|
chr17_-_1420006
|
0.174
|
ENST00000320345.6
ENST00000406424.4
|
INPP5K
|
inositol polyphosphate-5-phosphatase K
|
|
chr6_+_42847348
|
0.172
|
ENST00000493763.1
ENST00000304734.5
|
RPL7L1
|
ribosomal protein L7-like 1
|
|
chr10_-_118502070
|
0.161
|
ENST00000369209.3
|
HSPA12A
|
heat shock 70kDa protein 12A
|
|
chr13_+_111806083
|
0.156
|
ENST00000375736.4
|
ARHGEF7
|
Rho guanine nucleotide exchange factor (GEF) 7
|
|
chr17_-_66453562
|
0.154
|
ENST00000262139.5
ENST00000546360.1
|
WIPI1
|
WD repeat domain, phosphoinositide interacting 1
|
|
chr11_-_6633799
|
0.153
|
ENST00000299424.4
|
TAF10
|
TAF10 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 30kDa
|
|
chr3_-_57583185
|
0.152
|
ENST00000463880.1
|
ARF4
|
ADP-ribosylation factor 4
|
|
chr4_+_123747979
|
0.152
|
ENST00000608478.1
|
FGF2
|
fibroblast growth factor 2 (basic)
|
|
chr2_+_204193149
|
0.150
|
ENST00000422511.2
|
ABI2
|
abl-interactor 2
|
|
chr12_+_21654714
|
0.148
|
ENST00000542038.1
ENST00000540141.1
ENST00000229314.5
|
GOLT1B
|
golgi transport 1B
|
|
chr5_+_14143728
|
0.146
|
ENST00000344204.4
ENST00000537187.1
|
TRIO
|
trio Rho guanine nucleotide exchange factor
|
|
chrX_+_40944871
|
0.146
|
ENST00000378308.2
ENST00000324545.8
|
USP9X
|
ubiquitin specific peptidase 9, X-linked
|
|
chrX_-_153602991
|
0.145
|
ENST00000369850.3
ENST00000422373.1
|
FLNA
|
filamin A, alpha
|
|
chr8_+_6565854
|
0.145
|
ENST00000285518.6
|
AGPAT5
|
1-acylglycerol-3-phosphate O-acyltransferase 5
|
|
chr17_-_42277203
|
0.145
|
ENST00000587097.1
|
ATXN7L3
|
ataxin 7-like 3
|
|
chr11_-_6440283
|
0.144
|
ENST00000299402.6
ENST00000609360.1
ENST00000389906.2
ENST00000532020.2
|
APBB1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65)
|
|
chrX_-_38186775
|
0.140
|
ENST00000339363.3
ENST00000309513.3
ENST00000338898.3
ENST00000342811.3
ENST00000378505.2
|
RPGR
|
retinitis pigmentosa GTPase regulator
|
|
chr17_+_46126135
|
0.136
|
ENST00000361665.3
ENST00000585062.1
|
NFE2L1
|
nuclear factor, erythroid 2-like 1
|
|
chr13_+_21714653
|
0.135
|
ENST00000382533.4
|
SAP18
|
Sin3A-associated protein, 18kDa
|
|
chr6_-_111804905
|
0.133
|
ENST00000358835.3
ENST00000435970.1
|
REV3L
|
REV3-like, polymerase (DNA directed), zeta, catalytic subunit
|
|
chr13_+_21714711
|
0.132
|
ENST00000607003.1
ENST00000492245.1
|
SAP18
|
Sin3A-associated protein, 18kDa
|
|
chr22_+_18593446
|
0.129
|
ENST00000316027.6
|
TUBA8
|
tubulin, alpha 8
|
|
chr2_+_120770645
|
0.128
|
ENST00000443902.2
|
EPB41L5
|
erythrocyte membrane protein band 4.1 like 5
|
|
chr2_+_204193129
|
0.126
|
ENST00000417864.1
|
ABI2
|
abl-interactor 2
|
|
chr2_-_175547571
|
0.122
|
ENST00000409415.3
ENST00000359761.3
ENST00000272746.5
|
WIPF1
|
WAS/WASL interacting protein family, member 1
|
|
chr4_+_166128735
|
0.119
|
ENST00000226725.6
|
KLHL2
|
kelch-like family member 2
|
|
chr2_+_120770581
|
0.117
|
ENST00000263713.5
|
EPB41L5
|
erythrocyte membrane protein band 4.1 like 5
|
|
chr11_-_6440624
|
0.116
|
ENST00000311051.3
|
APBB1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65)
|
|
chr20_+_42086525
|
0.115
|
ENST00000244020.3
|
SRSF6
|
serine/arginine-rich splicing factor 6
|
|
chr11_+_125462690
|
0.115
|
ENST00000392708.4
ENST00000529196.1
ENST00000531491.1
|
STT3A
|
STT3A, subunit of the oligosaccharyltransferase complex (catalytic)
|
|
chr8_+_26240414
|
0.110
|
ENST00000380629.2
|
BNIP3L
|
BCL2/adenovirus E1B 19kDa interacting protein 3-like
|
|
chr14_+_64971292
|
0.108
|
ENST00000358738.3
ENST00000394712.2
|
ZBTB1
|
zinc finger and BTB domain containing 1
|
|
chr11_-_62380199
|
0.103
|
ENST00000419857.1
ENST00000394773.2
|
EML3
|
echinoderm microtubule associated protein like 3
|
|
chr4_-_149363662
|
0.098
|
ENST00000355292.3
ENST00000358102.3
|
NR3C2
|
nuclear receptor subfamily 3, group C, member 2
|
|
chr3_-_57583052
|
0.093
|
ENST00000496292.1
ENST00000489843.1
|
ARF4
|
ADP-ribosylation factor 4
|
|
chr20_+_57267669
|
0.092
|
ENST00000356091.6
|
NPEPL1
|
aminopeptidase-like 1
|
|
chr19_-_14016877
|
0.089
|
ENST00000454313.1
ENST00000591586.1
ENST00000346736.2
|
C19orf57
|
chromosome 19 open reading frame 57
|
|
chr16_+_81069433
|
0.085
|
ENST00000299575.4
|
ATMIN
|
ATM interactor
|
|
chr10_-_135187193
|
0.082
|
ENST00000368547.3
|
ECHS1
|
enoyl CoA hydratase, short chain, 1, mitochondrial
|
|
chr15_-_43802769
|
0.080
|
ENST00000263801.3
|
TP53BP1
|
tumor protein p53 binding protein 1
|
|
chr3_-_57583130
|
0.074
|
ENST00000303436.6
|
ARF4
|
ADP-ribosylation factor 4
|
|
chr4_-_183838372
|
0.072
|
ENST00000503820.1
ENST00000503988.1
|
DCTD
|
dCMP deaminase
|
|
chr2_+_121010324
|
0.068
|
ENST00000272519.5
|
RALB
|
v-ral simian leukemia viral oncogene homolog B
|
|
chr14_-_103523745
|
0.064
|
ENST00000361246.2
|
CDC42BPB
|
CDC42 binding protein kinase beta (DMPK-like)
|
|
chr2_-_232791038
|
0.063
|
ENST00000295440.2
ENST00000409852.1
|
NPPC
|
natriuretic peptide C
|
|
chr13_+_21714913
|
0.062
|
ENST00000450573.1
ENST00000467636.1
|
SAP18
|
Sin3A-associated protein, 18kDa
|
|
chr4_-_183838596
|
0.059
|
ENST00000508994.1
ENST00000512766.1
|
DCTD
|
dCMP deaminase
|
|
chr2_+_121010370
|
0.059
|
ENST00000420510.1
|
RALB
|
v-ral simian leukemia viral oncogene homolog B
|
|
chr3_-_49131473
|
0.057
|
ENST00000430979.1
ENST00000357496.2
ENST00000437939.1
|
QRICH1
|
glutamine-rich 1
|
|
chr13_+_48877895
|
0.057
|
ENST00000267163.4
|
RB1
|
retinoblastoma 1
|
|
chr20_+_30946106
|
0.056
|
ENST00000375687.4
ENST00000542461.1
|
ASXL1
|
additional sex combs like 1 (Drosophila)
|
|
chr9_-_122131696
|
0.056
|
ENST00000373964.2
ENST00000265922.3
|
BRINP1
|
bone morphogenetic protein/retinoic acid inducible neural-specific 1
|
|
chr2_+_121010413
|
0.055
|
ENST00000404963.3
|
RALB
|
v-ral simian leukemia viral oncogene homolog B
|
|
chr8_+_32406179
|
0.052
|
ENST00000405005.3
|
NRG1
|
neuregulin 1
|
|
chr15_-_35047166
|
0.051
|
ENST00000290374.4
|
GJD2
|
gap junction protein, delta 2, 36kDa
|
|
chr10_+_94352956
|
0.050
|
ENST00000260731.3
|
KIF11
|
kinesin family member 11
|
|
chr5_+_142149932
|
0.049
|
ENST00000274498.4
|
ARHGAP26
|
Rho GTPase activating protein 26
|
|
chr2_-_33824382
|
0.048
|
ENST00000238823.8
|
FAM98A
|
family with sequence similarity 98, member A
|
|
chr1_+_29063119
|
0.045
|
ENST00000474884.1
ENST00000542507.1
|
YTHDF2
|
YTH domain family, member 2
|
|
chr12_+_113659234
|
0.045
|
ENST00000551096.1
ENST00000551099.1
ENST00000335509.6
ENST00000552897.1
ENST00000550785.1
ENST00000549279.1
|
TPCN1
|
two pore segment channel 1
|
|
chr19_-_55919087
|
0.043
|
ENST00000587845.1
ENST00000589978.1
ENST00000264552.9
|
UBE2S
|
ubiquitin-conjugating enzyme E2S
|
|
chr19_-_14117074
|
0.042
|
ENST00000588885.1
ENST00000254325.4
|
RFX1
|
regulatory factor X, 1 (influences HLA class II expression)
|
|
chr5_+_61602055
|
0.042
|
ENST00000381103.2
|
KIF2A
|
kinesin heavy chain member 2A
|
|
chr14_-_92572894
|
0.041
|
ENST00000532032.1
ENST00000506466.1
ENST00000555381.1
ENST00000557311.1
ENST00000554592.1
ENST00000554672.1
ENST00000553491.1
ENST00000556220.1
ENST00000502250.1
ENST00000503767.1
ENST00000393287.5
ENST00000340660.6
ENST00000545170.1
ENST00000429774.2
|
ATXN3
|
ataxin 3
|
|
chr17_-_7518145
|
0.041
|
ENST00000250113.7
ENST00000571597.1
|
FXR2
|
fragile X mental retardation, autosomal homolog 2
|
|
chr9_-_139094988
|
0.041
|
ENST00000371746.3
|
LHX3
|
LIM homeobox 3
|
|
chr17_-_73178599
|
0.040
|
ENST00000578238.1
|
SUMO2
|
small ubiquitin-like modifier 2
|
|
chr19_-_39881669
|
0.039
|
ENST00000221266.7
|
PAF1
|
Paf1, RNA polymerase II associated factor, homolog (S. cerevisiae)
|
|
chr5_+_142149955
|
0.039
|
ENST00000378004.3
|
ARHGAP26
|
Rho GTPase activating protein 26
|
|
chr1_+_29063439
|
0.038
|
ENST00000541996.1
ENST00000496288.1
|
YTHDF2
|
YTH domain family, member 2
|
|
chr5_+_113697983
|
0.038
|
ENST00000264773.3
|
KCNN2
|
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 2
|
|
chr22_-_43010928
|
0.037
|
ENST00000348657.2
ENST00000252115.5
|
POLDIP3
|
polymerase (DNA-directed), delta interacting protein 3
|
|
chr16_-_70557430
|
0.036
|
ENST00000393612.4
ENST00000564653.1
ENST00000323786.5
|
COG4
|
component of oligomeric golgi complex 4
|
|
chr7_-_99006443
|
0.036
|
ENST00000350498.3
|
PDAP1
|
PDGFA associated protein 1
|
|
chr5_-_132166579
|
0.035
|
ENST00000378679.3
|
SHROOM1
|
shroom family member 1
|
|
chr6_-_35464817
|
0.035
|
ENST00000338863.7
|
TEAD3
|
TEA domain family member 3
|
|
chr1_+_22351977
|
0.033
|
ENST00000420503.1
ENST00000416769.1
ENST00000404210.2
|
LINC00339
|
long intergenic non-protein coding RNA 339
|
|
chr7_-_26240357
|
0.032
|
ENST00000354667.4
ENST00000356674.7
|
HNRNPA2B1
|
heterogeneous nuclear ribonucleoprotein A2/B1
|
|
chr20_-_39946237
|
0.032
|
ENST00000441102.2
ENST00000559234.1
|
ZHX3
|
zinc fingers and homeoboxes 3
|
|
chr2_-_33824336
|
0.032
|
ENST00000431950.1
ENST00000403368.1
ENST00000441530.2
|
FAM98A
|
family with sequence similarity 98, member A
|
|
chr9_+_100174344
|
0.032
|
ENST00000422139.2
|
TDRD7
|
tudor domain containing 7
|
|
chr17_-_4544960
|
0.029
|
ENST00000293761.3
|
ALOX15
|
arachidonate 15-lipoxygenase
|
|
chr17_+_43971643
|
0.027
|
ENST00000344290.5
ENST00000262410.5
ENST00000351559.5
ENST00000340799.5
ENST00000535772.1
ENST00000347967.5
|
MAPT
|
microtubule-associated protein tau
|
|
chr6_+_42847649
|
0.027
|
ENST00000424341.2
ENST00000602561.1
|
RPL7L1
|
ribosomal protein L7-like 1
|
|
chr1_+_29063271
|
0.026
|
ENST00000373812.3
|
YTHDF2
|
YTH domain family, member 2
|
|
chr3_-_49761337
|
0.026
|
ENST00000308388.6
ENST00000480687.1
ENST00000308375.6
ENST00000535833.1
|
GMPPB
AMIGO3
|
GDP-mannose pyrophosphorylase B
adhesion molecule with Ig-like domain 3
|
|
chr9_+_77112244
|
0.025
|
ENST00000376896.3
|
RORB
|
RAR-related orphan receptor B
|
|
chr9_-_139922726
|
0.025
|
ENST00000265662.5
ENST00000371605.3
|
ABCA2
|
ATP-binding cassette, sub-family A (ABC1), member 2
|
|
chr10_+_99079008
|
0.023
|
ENST00000371021.3
|
FRAT1
|
frequently rearranged in advanced T-cell lymphomas
|
|
chr7_+_101459263
|
0.023
|
ENST00000292538.4
ENST00000393824.3
ENST00000547394.2
ENST00000360264.3
ENST00000425244.2
|
CUX1
|
cut-like homeobox 1
|
|
chr9_+_34458771
|
0.022
|
ENST00000437363.1
ENST00000242317.4
|
DNAI1
|
dynein, axonemal, intermediate chain 1
|
|
chr6_-_35464727
|
0.021
|
ENST00000402886.3
|
TEAD3
|
TEA domain family member 3
|
|
chr1_-_197115818
|
0.020
|
ENST00000367409.4
ENST00000294732.7
|
ASPM
|
asp (abnormal spindle) homolog, microcephaly associated (Drosophila)
|
|
chr14_-_99947168
|
0.019
|
ENST00000331768.5
|
SETD3
|
SET domain containing 3
|
|
chr11_-_73309228
|
0.019
|
ENST00000356467.4
ENST00000064778.4
|
FAM168A
|
family with sequence similarity 168, member A
|
|
chr5_+_176873446
|
0.018
|
ENST00000507881.1
|
PRR7
|
proline rich 7 (synaptic)
|
|
chr5_-_114598548
|
0.018
|
ENST00000379615.3
ENST00000419445.1
|
PGGT1B
|
protein geranylgeranyltransferase type I, beta subunit
|
|
chr8_-_100905363
|
0.018
|
ENST00000524245.1
|
COX6C
|
cytochrome c oxidase subunit VIc
|
|
chr1_-_40157345
|
0.017
|
ENST00000372844.3
|
HPCAL4
|
hippocalcin like 4
|
|
chrX_+_21392553
|
0.016
|
ENST00000279451.4
|
CNKSR2
|
connector enhancer of kinase suppressor of Ras 2
|
|
chr2_-_97304105
|
0.016
|
ENST00000599854.1
ENST00000441706.2
|
KANSL3
|
KAT8 regulatory NSL complex subunit 3
|
|
chr6_-_144329531
|
0.016
|
ENST00000429150.1
ENST00000392309.1
ENST00000416623.1
ENST00000392307.1
|
PLAGL1
|
pleiomorphic adenoma gene-like 1
|
|
chr8_-_30670053
|
0.016
|
ENST00000518564.1
|
PPP2CB
|
protein phosphatase 2, catalytic subunit, beta isozyme
|
|
chr4_-_2758015
|
0.015
|
ENST00000510267.1
ENST00000503235.1
ENST00000315423.7
|
TNIP2
|
TNFAIP3 interacting protein 2
|
|
chr19_-_39881777
|
0.014
|
ENST00000595564.1
ENST00000221265.3
|
PAF1
|
Paf1, RNA polymerase II associated factor, homolog (S. cerevisiae)
|
|
chr4_-_183838482
|
0.013
|
ENST00000357067.3
ENST00000510370.1
|
DCTD
|
dCMP deaminase
|
|
chr17_+_43972010
|
0.013
|
ENST00000334239.8
ENST00000446361.3
|
MAPT
|
microtubule-associated protein tau
|
|
chr16_+_56899114
|
0.013
|
ENST00000566786.1
ENST00000438926.2
ENST00000563236.1
ENST00000262502.5
|
SLC12A3
|
solute carrier family 12 (sodium/chloride transporter), member 3
|
|
chr2_+_64068074
|
0.012
|
ENST00000394417.2
ENST00000484142.1
ENST00000482668.1
ENST00000467648.2
|
UGP2
|
UDP-glucose pyrophosphorylase 2
|
|
chr19_-_45004556
|
0.011
|
ENST00000587047.1
ENST00000391956.4
ENST00000221327.4
ENST00000586637.1
ENST00000591064.1
ENST00000592529.1
|
ZNF180
|
zinc finger protein 180
|
|
chr3_-_48936272
|
0.011
|
ENST00000544097.1
ENST00000430379.1
ENST00000319017.4
|
SLC25A20
|
solute carrier family 25 (carnitine/acylcarnitine translocase), member 20
|
|
chr12_+_51985001
|
0.009
|
ENST00000354534.6
|
SCN8A
|
sodium channel, voltage gated, type VIII, alpha subunit
|
|
chr4_-_156298028
|
0.007
|
ENST00000433024.1
ENST00000379248.2
|
MAP9
|
microtubule-associated protein 9
|
|
chr1_+_6845384
|
0.007
|
ENST00000303635.7
|
CAMTA1
|
calmodulin binding transcription activator 1
|
|
chr4_-_183838747
|
0.006
|
ENST00000438320.2
|
DCTD
|
dCMP deaminase
|
|
chr11_+_28131821
|
0.005
|
ENST00000379199.2
ENST00000303459.6
|
METTL15
|
methyltransferase like 15
|
|
chr19_-_10530784
|
0.004
|
ENST00000593124.1
|
CDC37
|
cell division cycle 37
|
|
chr2_+_157292859
|
0.004
|
ENST00000438166.2
|
GPD2
|
glycerol-3-phosphate dehydrogenase 2 (mitochondrial)
|
|
chr2_+_42396574
|
0.003
|
ENST00000401738.3
|
EML4
|
echinoderm microtubule associated protein like 4
|
|
chr2_-_170219079
|
0.003
|
ENST00000263816.3
|
LRP2
|
low density lipoprotein receptor-related protein 2
|
|
chr12_+_57482665
|
0.002
|
ENST00000300131.3
|
NAB2
|
NGFI-A binding protein 2 (EGR1 binding protein 2)
|
|
chr11_-_62446527
|
0.001
|
ENST00000294119.2
ENST00000529640.1
ENST00000534176.1
ENST00000301935.5
|
UBXN1
|
UBX domain protein 1
|
|
chr12_+_70637494
|
0.001
|
ENST00000548159.1
ENST00000549750.1
ENST00000551043.1
|
CNOT2
|
CCR4-NOT transcription complex, subunit 2
|
|
chr6_-_33267101
|
0.001
|
ENST00000497454.1
|
RGL2
|
ral guanine nucleotide dissociation stimulator-like 2
|
|
chr7_+_128095900
|
0.000
|
ENST00000435296.2
|
HILPDA
|
hypoxia inducible lipid droplet-associated
|