| 0.4 |
1.9 |
GO:0051611 |
negative regulation of neurotransmitter uptake(GO:0051581) negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of thrombin receptor signaling pathway(GO:0070494) negative regulation of thrombin receptor signaling pathway(GO:0070495) |
| 0.3 |
1.6 |
GO:0006789 |
bilirubin conjugation(GO:0006789) biphenyl metabolic process(GO:0018879) biphenyl catabolic process(GO:0070980) |
| 0.3 |
1.6 |
GO:0043312 |
neutrophil activation involved in immune response(GO:0002283) neutrophil degranulation(GO:0043312) |
| 0.3 |
0.8 |
GO:0010902 |
positive regulation of very-low-density lipoprotein particle remodeling(GO:0010902) positive regulation of phospholipid catabolic process(GO:0060697) |
| 0.3 |
1.0 |
GO:0006542 |
glutamine biosynthetic process(GO:0006542) |
| 0.2 |
0.7 |
GO:0033278 |
cell proliferation in midbrain(GO:0033278) |
| 0.2 |
0.6 |
GO:0003308 |
negative regulation of Wnt signaling pathway involved in heart development(GO:0003308) |
| 0.2 |
0.6 |
GO:0046010 |
positive regulation of circadian sleep/wake cycle, non-REM sleep(GO:0046010) |
| 0.2 |
0.7 |
GO:0060649 |
nipple development(GO:0060618) mammary gland bud elongation(GO:0060649) nipple morphogenesis(GO:0060658) nipple sheath formation(GO:0060659) |
| 0.2 |
0.6 |
GO:0051552 |
flavone metabolic process(GO:0051552) |
| 0.1 |
0.4 |
GO:0060689 |
cell differentiation involved in salivary gland development(GO:0060689) |
| 0.1 |
0.5 |
GO:0051503 |
purine ribonucleotide transport(GO:0015868) adenine nucleotide transport(GO:0051503) |
| 0.1 |
0.5 |
GO:0071400 |
cellular response to oleic acid(GO:0071400) |
| 0.1 |
0.4 |
GO:0002514 |
B cell tolerance induction(GO:0002514) regulation of B cell tolerance induction(GO:0002661) positive regulation of B cell tolerance induction(GO:0002663) |
| 0.1 |
0.5 |
GO:0001575 |
globoside metabolic process(GO:0001575) |
| 0.1 |
0.5 |
GO:0050653 |
UDP-N-acetylgalactosamine metabolic process(GO:0019276) chondroitin sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0050653) |
| 0.1 |
1.1 |
GO:0051918 |
negative regulation of fibrinolysis(GO:0051918) |
| 0.1 |
0.6 |
GO:0060242 |
contact inhibition(GO:0060242) |
| 0.1 |
3.1 |
GO:0031424 |
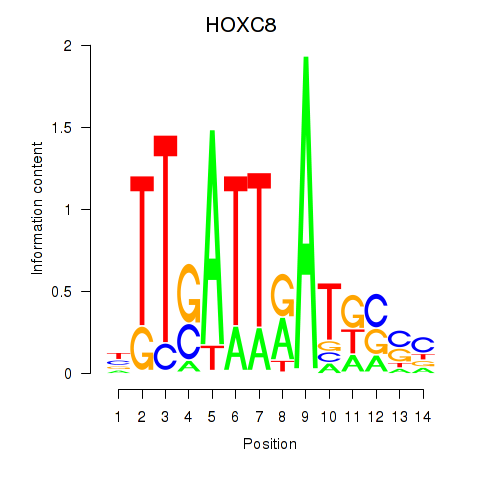
keratinization(GO:0031424) |
| 0.1 |
0.5 |
GO:0006927 |
obsolete transformed cell apoptotic process(GO:0006927) |
| 0.1 |
0.4 |
GO:0090161 |
Golgi ribbon formation(GO:0090161) |
| 0.1 |
0.4 |
GO:0008065 |
establishment of blood-nerve barrier(GO:0008065) |
| 0.1 |
2.5 |
GO:0031581 |
hemidesmosome assembly(GO:0031581) |
| 0.1 |
0.3 |
GO:0006530 |
asparagine catabolic process(GO:0006530) asparagine catabolic process via L-aspartate(GO:0033345) |
| 0.1 |
1.2 |
GO:0060391 |
positive regulation of SMAD protein import into nucleus(GO:0060391) |
| 0.1 |
1.1 |
GO:0030299 |
intestinal cholesterol absorption(GO:0030299) intestinal lipid absorption(GO:0098856) |
| 0.1 |
0.3 |
GO:0019371 |
cyclooxygenase pathway(GO:0019371) |
| 0.1 |
0.4 |
GO:0006883 |
cellular sodium ion homeostasis(GO:0006883) |
| 0.1 |
0.2 |
GO:0061302 |
smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 |
0.2 |
GO:0003350 |
pulmonary myocardium development(GO:0003350) |
| 0.1 |
0.3 |
GO:0045338 |
farnesyl diphosphate metabolic process(GO:0045338) |
| 0.1 |
0.2 |
GO:2000105 |
positive regulation of DNA-dependent DNA replication initiation(GO:0032298) positive regulation of DNA-dependent DNA replication(GO:2000105) |
| 0.1 |
0.3 |
GO:0009235 |
cobalamin metabolic process(GO:0009235) |
| 0.1 |
0.1 |
GO:0002675 |
positive regulation of acute inflammatory response(GO:0002675) |
| 0.1 |
0.3 |
GO:0042271 |
susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.1 |
0.2 |
GO:0032071 |
obsolete cleavage of lamin involved in execution phase of apoptosis(GO:0006922) regulation of endodeoxyribonuclease activity(GO:0032071) negative regulation of nuclease activity(GO:0032074) |
| 0.1 |
1.7 |
GO:0006693 |
prostaglandin metabolic process(GO:0006693) |
| 0.1 |
0.1 |
GO:0072201 |
negative regulation of mesenchymal cell proliferation(GO:0072201) |
| 0.1 |
0.6 |
GO:0045329 |
amino-acid betaine biosynthetic process(GO:0006578) carnitine biosynthetic process(GO:0045329) |
| 0.1 |
0.6 |
GO:0010739 |
positive regulation of protein kinase A signaling(GO:0010739) |
| 0.1 |
0.3 |
GO:0001778 |
plasma membrane repair(GO:0001778) |
| 0.1 |
0.5 |
GO:0006685 |
sphingomyelin catabolic process(GO:0006685) |
| 0.1 |
0.2 |
GO:0046102 |
nicotinamide riboside catabolic process(GO:0006738) inosine metabolic process(GO:0046102) nicotinamide riboside metabolic process(GO:0046495) pyridine nucleoside metabolic process(GO:0070637) pyridine nucleoside catabolic process(GO:0070638) |
| 0.1 |
0.2 |
GO:0019695 |
choline metabolic process(GO:0019695) |
| 0.1 |
0.2 |
GO:0060729 |
negative regulation of interleukin-23 production(GO:0032707) intestinal epithelial structure maintenance(GO:0060729) |
| 0.1 |
0.2 |
GO:0035461 |
vitamin transmembrane transport(GO:0035461) |
| 0.1 |
0.2 |
GO:0006122 |
mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.1 |
0.2 |
GO:0001692 |
histamine metabolic process(GO:0001692) |
| 0.0 |
0.1 |
GO:1901419 |
regulation of response to alcohol(GO:1901419) |
| 0.0 |
0.3 |
GO:0000050 |
urea cycle(GO:0000050) |
| 0.0 |
0.8 |
GO:0042730 |
fibrinolysis(GO:0042730) |
| 0.0 |
1.8 |
GO:0060338 |
regulation of type I interferon-mediated signaling pathway(GO:0060338) |
| 0.0 |
0.4 |
GO:0015886 |
heme transport(GO:0015886) |
| 0.0 |
1.3 |
GO:0006730 |
one-carbon metabolic process(GO:0006730) |
| 0.0 |
0.3 |
GO:0006264 |
mitochondrial DNA replication(GO:0006264) |
| 0.0 |
0.4 |
GO:0010658 |
striated muscle cell apoptotic process(GO:0010658) cardiac muscle cell apoptotic process(GO:0010659) regulation of striated muscle cell apoptotic process(GO:0010662) negative regulation of striated muscle cell apoptotic process(GO:0010664) regulation of cardiac muscle cell apoptotic process(GO:0010665) negative regulation of cardiac muscle cell apoptotic process(GO:0010667) |
| 0.0 |
0.9 |
GO:0045745 |
positive regulation of G-protein coupled receptor protein signaling pathway(GO:0045745) |
| 0.0 |
0.1 |
GO:0035025 |
positive regulation of Rho protein signal transduction(GO:0035025) |
| 0.0 |
0.8 |
GO:0007172 |
signal complex assembly(GO:0007172) |
| 0.0 |
0.1 |
GO:0030997 |
regulation of centriole-centriole cohesion(GO:0030997) |
| 0.0 |
0.4 |
GO:0051014 |
actin filament severing(GO:0051014) |
| 0.0 |
0.6 |
GO:0008210 |
estrogen metabolic process(GO:0008210) |
| 0.0 |
0.0 |
GO:0019627 |
urea metabolic process(GO:0019627) |
| 0.0 |
0.1 |
GO:0006726 |
eye pigment biosynthetic process(GO:0006726) eye pigment metabolic process(GO:0042441) pigment metabolic process involved in developmental pigmentation(GO:0043324) pigment metabolic process involved in pigmentation(GO:0043474) |
| 0.0 |
0.1 |
GO:0051066 |
dihydrobiopterin metabolic process(GO:0051066) |
| 0.0 |
0.1 |
GO:0072205 |
collecting duct development(GO:0072044) metanephric collecting duct development(GO:0072205) |
| 0.0 |
1.1 |
GO:0050909 |
sensory perception of taste(GO:0050909) |
| 0.0 |
0.5 |
GO:0035024 |
negative regulation of Rho protein signal transduction(GO:0035024) |
| 0.0 |
0.1 |
GO:0045750 |
obsolete positive regulation of S phase of mitotic cell cycle(GO:0045750) |
| 0.0 |
0.2 |
GO:0060219 |
camera-type eye photoreceptor cell differentiation(GO:0060219) |
| 0.0 |
0.3 |
GO:0060907 |
cGMP-mediated signaling(GO:0019934) regulation of phagocytosis, engulfment(GO:0060099) positive regulation of phagocytosis, engulfment(GO:0060100) positive regulation of macrophage cytokine production(GO:0060907) regulation of membrane invagination(GO:1905153) positive regulation of membrane invagination(GO:1905155) |
| 0.0 |
0.4 |
GO:0046697 |
decidualization(GO:0046697) |
| 0.0 |
0.3 |
GO:0050892 |
intestinal absorption(GO:0050892) |
| 0.0 |
0.2 |
GO:0046968 |
peptide antigen transport(GO:0046968) |
| 0.0 |
0.7 |
GO:0030574 |
collagen catabolic process(GO:0030574) |
| 0.0 |
0.1 |
GO:0046621 |
negative regulation of organ growth(GO:0046621) |
| 0.0 |
0.5 |
GO:0006071 |
glycerol metabolic process(GO:0006071) |
| 0.0 |
0.1 |
GO:0034720 |
histone H3-K4 demethylation(GO:0034720) |
| 0.0 |
0.1 |
GO:0043006 |
activation of phospholipase A2 activity by calcium-mediated signaling(GO:0043006) |
| 0.0 |
0.3 |
GO:0002347 |
response to tumor cell(GO:0002347) |
| 0.0 |
0.3 |
GO:0006787 |
porphyrin-containing compound catabolic process(GO:0006787) tetrapyrrole catabolic process(GO:0033015) heme catabolic process(GO:0042167) pigment catabolic process(GO:0046149) |
| 0.0 |
0.1 |
GO:0010727 |
negative regulation of hydrogen peroxide metabolic process(GO:0010727) |
| 0.0 |
0.1 |
GO:0018125 |
peptidyl-cysteine methylation(GO:0018125) |
| 0.0 |
0.1 |
GO:0002523 |
leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.0 |
0.4 |
GO:1900047 |
negative regulation of blood coagulation(GO:0030195) negative regulation of wound healing(GO:0061045) negative regulation of hemostasis(GO:1900047) |
| 0.0 |
0.1 |
GO:0050955 |
thermoception(GO:0050955) |
| 0.0 |
0.1 |
GO:0050747 |
positive regulation of lipoprotein metabolic process(GO:0050747) |
| 0.0 |
0.2 |
GO:0006654 |
positive regulation of cytokine-mediated signaling pathway(GO:0001961) phosphatidic acid biosynthetic process(GO:0006654) |
| 0.0 |
0.1 |
GO:0015712 |
hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.0 |
0.1 |
GO:0034442 |
plasma lipoprotein particle oxidation(GO:0034441) regulation of lipoprotein oxidation(GO:0034442) negative regulation of lipoprotein oxidation(GO:0034443) regulation of plasma lipoprotein particle oxidation(GO:0034444) negative regulation of plasma lipoprotein particle oxidation(GO:0034445) lipoprotein modification(GO:0042160) lipoprotein oxidation(GO:0042161) |
| 0.0 |
0.8 |
GO:0006699 |
bile acid biosynthetic process(GO:0006699) |
| 0.0 |
0.1 |
GO:0018874 |
benzoate metabolic process(GO:0018874) |
| 0.0 |
0.2 |
GO:0006013 |
mannose metabolic process(GO:0006013) |
| 0.0 |
0.9 |
GO:0030216 |
keratinocyte differentiation(GO:0030216) |
| 0.0 |
3.1 |
GO:0010951 |
negative regulation of endopeptidase activity(GO:0010951) |
| 0.0 |
0.1 |
GO:0042519 |
tyrosine phosphorylation of Stat4 protein(GO:0042504) regulation of tyrosine phosphorylation of Stat4 protein(GO:0042519) positive regulation of tyrosine phosphorylation of Stat4 protein(GO:0042520) |
| 0.0 |
0.2 |
GO:0016998 |
cell wall macromolecule catabolic process(GO:0016998) |
| 0.0 |
0.1 |
GO:0006422 |
aspartyl-tRNA aminoacylation(GO:0006422) |
| 0.0 |
0.1 |
GO:0072284 |
visceral serous pericardium development(GO:0061032) S-shaped body morphogenesis(GO:0072050) metanephric S-shaped body morphogenesis(GO:0072284) |
| 0.0 |
0.1 |
GO:1903960 |
negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) negative regulation of anion transmembrane transport(GO:1903960) negative regulation of fatty acid transport(GO:2000192) |
| 0.0 |
0.2 |
GO:0043415 |
positive regulation of skeletal muscle tissue regeneration(GO:0043415) regulation of skeletal muscle tissue regeneration(GO:0043416) |
| 0.0 |
0.3 |
GO:1904893 |
negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 |
0.2 |
GO:0006525 |
arginine metabolic process(GO:0006525) |
| 0.0 |
0.1 |
GO:0042986 |
positive regulation of amyloid precursor protein biosynthetic process(GO:0042986) |
| 0.0 |
0.3 |
GO:0050872 |
white fat cell differentiation(GO:0050872) |
| 0.0 |
0.3 |
GO:0035329 |
hippo signaling(GO:0035329) |
| 0.0 |
0.1 |
GO:0009822 |
alkaloid catabolic process(GO:0009822) |
| 0.0 |
0.2 |
GO:0006776 |
vitamin A metabolic process(GO:0006776) |
| 0.0 |
0.1 |
GO:0033522 |
histone H2A ubiquitination(GO:0033522) |
| 0.0 |
0.1 |
GO:0003215 |
cardiac right ventricle morphogenesis(GO:0003215) |
| 0.0 |
0.1 |
GO:0035120 |
post-embryonic appendage morphogenesis(GO:0035120) |
| 0.0 |
0.1 |
GO:0006642 |
triglyceride mobilization(GO:0006642) |
| 0.0 |
0.4 |
GO:0008206 |
bile acid metabolic process(GO:0008206) |
| 0.0 |
0.1 |
GO:0045162 |
clustering of voltage-gated sodium channels(GO:0045162) |
| 0.0 |
0.7 |
GO:0030837 |
negative regulation of actin filament polymerization(GO:0030837) |
| 0.0 |
0.3 |
GO:0097031 |
NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.0 |
0.1 |
GO:0030277 |
epithelial structure maintenance(GO:0010669) maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.0 |
0.1 |
GO:0070634 |
transepithelial ammonium transport(GO:0070634) |
| 0.0 |
0.1 |
GO:0007549 |
dosage compensation(GO:0007549) |
| 0.0 |
0.1 |
GO:0051289 |
protein homotetramerization(GO:0051289) |
| 0.0 |
0.1 |
GO:0010700 |
negative regulation of norepinephrine secretion(GO:0010700) |
| 0.0 |
0.1 |
GO:0001845 |
phagolysosome assembly(GO:0001845) post-embryonic organ morphogenesis(GO:0048563) phagosome maturation(GO:0090382) |
| 0.0 |
0.2 |
GO:0060216 |
definitive hemopoiesis(GO:0060216) |
| 0.0 |
0.1 |
GO:0043267 |
negative regulation of potassium ion transport(GO:0043267) |
| 0.0 |
0.2 |
GO:0008631 |
intrinsic apoptotic signaling pathway in response to oxidative stress(GO:0008631) cell death in response to oxidative stress(GO:0036473) |
| 0.0 |
0.1 |
GO:0045056 |
transcytosis(GO:0045056) |
| 0.0 |
0.1 |
GO:0007206 |
phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.0 |
0.0 |
GO:2000008 |
regulation of protein localization to cell surface(GO:2000008) |
| 0.0 |
0.4 |
GO:0046854 |
phosphatidylinositol phosphorylation(GO:0046854) |
| 0.0 |
0.1 |
GO:0048387 |
negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.0 |
0.2 |
GO:0006862 |
nucleotide transport(GO:0006862) |
| 0.0 |
0.0 |
GO:0034036 |
purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.0 |
0.1 |
GO:0071318 |
cellular response to ATP(GO:0071318) |
| 0.0 |
0.2 |
GO:0048261 |
negative regulation of receptor-mediated endocytosis(GO:0048261) |
| 0.0 |
0.1 |
GO:0090073 |
positive regulation of protein homodimerization activity(GO:0090073) |
| 0.0 |
0.1 |
GO:0048147 |
negative regulation of fibroblast proliferation(GO:0048147) |
| 0.0 |
0.1 |
GO:0042517 |
positive regulation of tyrosine phosphorylation of Stat3 protein(GO:0042517) |
| 0.0 |
0.0 |
GO:0019372 |
lipoxygenase pathway(GO:0019372) |
| 0.0 |
0.1 |
GO:0051294 |
establishment of mitotic spindle orientation(GO:0000132) establishment of spindle orientation(GO:0051294) |
| 0.0 |
0.2 |
GO:0018298 |
protein-chromophore linkage(GO:0018298) |
| 0.0 |
0.2 |
GO:0009629 |
response to gravity(GO:0009629) |
| 0.0 |
0.2 |
GO:0009812 |
flavonoid metabolic process(GO:0009812) |
| 0.0 |
0.1 |
GO:0002885 |
positive regulation of hypersensitivity(GO:0002885) |
| 0.0 |
0.0 |
GO:0032621 |
interleukin-18 production(GO:0032621) regulation of interleukin-18 production(GO:0032661) |
| 0.0 |
0.1 |
GO:0006553 |
lysine metabolic process(GO:0006553) lysine catabolic process(GO:0006554) |
| 0.0 |
0.0 |
GO:0006577 |
amino-acid betaine metabolic process(GO:0006577) carnitine metabolic process(GO:0009437) |
| 0.0 |
0.2 |
GO:0035036 |
binding of sperm to zona pellucida(GO:0007339) sperm-egg recognition(GO:0035036) |
| 0.0 |
0.1 |
GO:0017183 |
peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.0 |
0.1 |
GO:0045059 |
positive thymic T cell selection(GO:0045059) |
| 0.0 |
0.0 |
GO:0051083 |
'de novo' cotranslational protein folding(GO:0051083) |
| 0.0 |
0.1 |
GO:0007342 |
fusion of sperm to egg plasma membrane(GO:0007342) |
| 0.0 |
0.1 |
GO:0042769 |
DNA damage response, detection of DNA damage(GO:0042769) |
| 0.0 |
0.1 |
GO:0070682 |
proteasome regulatory particle assembly(GO:0070682) |
| 0.0 |
0.0 |
GO:0009103 |
lipopolysaccharide biosynthetic process(GO:0009103) |
| 0.0 |
0.1 |
GO:0015014 |
heparan sulfate proteoglycan biosynthetic process, polysaccharide chain biosynthetic process(GO:0015014) |
| 0.0 |
0.3 |
GO:0015804 |
neutral amino acid transport(GO:0015804) |
| 0.0 |
0.1 |
GO:0042940 |
D-amino acid transport(GO:0042940) D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.0 |
0.0 |
GO:0070265 |
necrotic cell death(GO:0070265) |
| 0.0 |
0.1 |
GO:0061339 |
establishment of apical/basal cell polarity(GO:0035089) establishment of monopolar cell polarity(GO:0061162) establishment or maintenance of monopolar cell polarity(GO:0061339) |
| 0.0 |
0.1 |
GO:0046668 |
regulation of retinal cell programmed cell death(GO:0046668) |
| 0.0 |
0.3 |
GO:0042554 |
superoxide anion generation(GO:0042554) |
| 0.0 |
0.1 |
GO:0010226 |
response to lithium ion(GO:0010226) |
| 0.0 |
0.1 |
GO:0055015 |
ventricular cardiac muscle cell development(GO:0055015) |
| 0.0 |
0.1 |
GO:0017121 |
phospholipid scrambling(GO:0017121) |
| 0.0 |
0.2 |
GO:0045104 |
intermediate filament cytoskeleton organization(GO:0045104) |
| 0.0 |
0.6 |
GO:0061025 |
membrane fusion(GO:0061025) |
| 0.0 |
0.0 |
GO:0060423 |
foregut regionalization(GO:0060423) lung field specification(GO:0060424) lung induction(GO:0060492) regulation of branching involved in lung morphogenesis(GO:0061046) positive regulation of branching involved in lung morphogenesis(GO:0061047) iris morphogenesis(GO:0061072) cornea development in camera-type eye(GO:0061303) |
| 0.0 |
0.1 |
GO:0048388 |
endosomal lumen acidification(GO:0048388) |
| 0.0 |
0.1 |
GO:0015871 |
choline transport(GO:0015871) |
| 0.0 |
0.2 |
GO:0006700 |
C21-steroid hormone biosynthetic process(GO:0006700) |
| 0.0 |
0.1 |
GO:0046519 |
sphingoid metabolic process(GO:0046519) |
| 0.0 |
0.1 |
GO:0051546 |
keratinocyte migration(GO:0051546) |
| 0.0 |
0.2 |
GO:0000038 |
very long-chain fatty acid metabolic process(GO:0000038) |