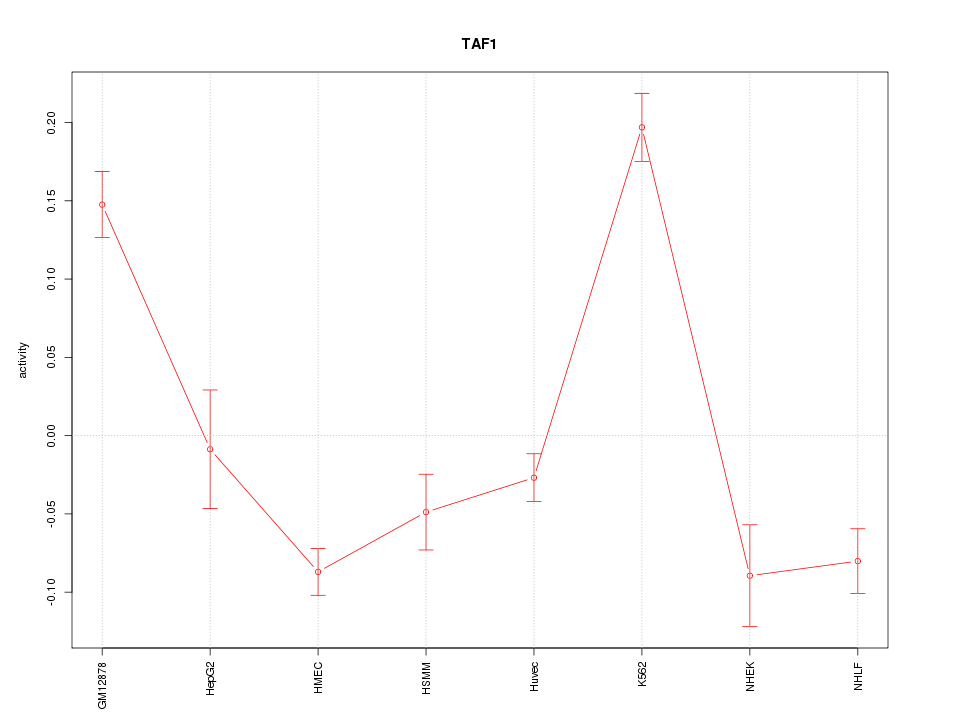
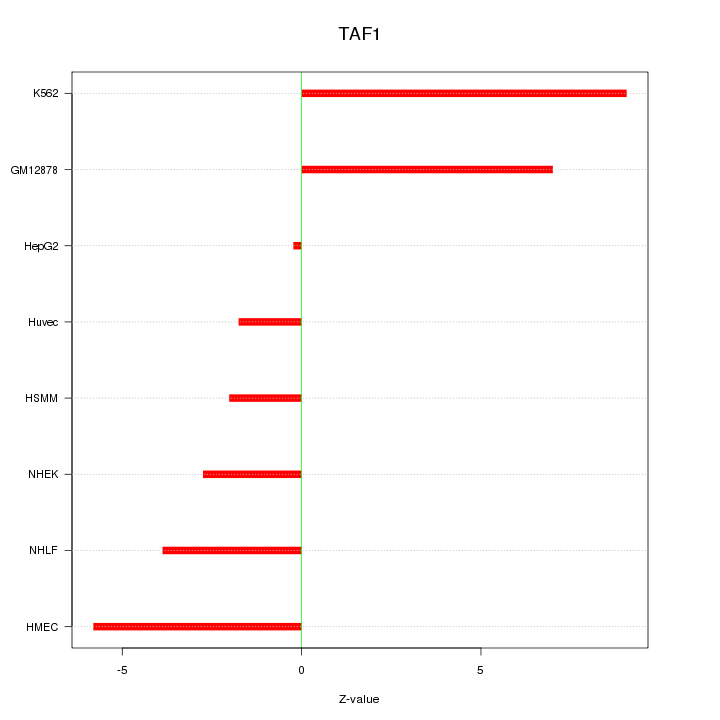
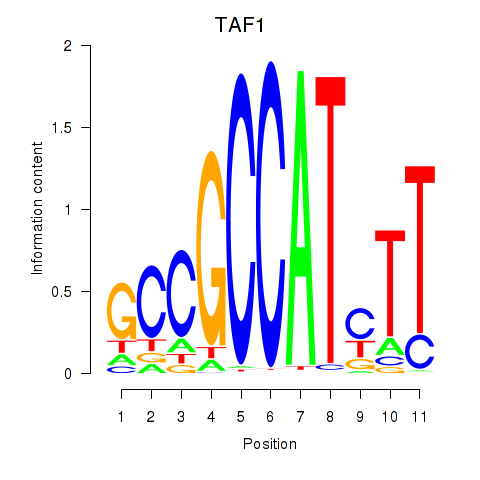
Motif ID: TAF1
Z-value: 4.933
Transcription factors associated with TAF1:
| Gene Symbol | Entrez ID | Gene Name |
|---|---|---|
| TAF1 | ENSG00000147133.11 | TAF1 |
Top targets:
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.9 | 11.7 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 1.8 | 5.5 | GO:0046015 | carbon catabolite regulation of transcription(GO:0045990) regulation of transcription by glucose(GO:0046015) positive regulation of transcription from RNA polymerase II promoter involved in cellular response to chemical stimulus(GO:1901522) |
| 1.6 | 4.7 | GO:0035461 | vitamin transmembrane transport(GO:0035461) |
| 1.4 | 4.3 | GO:0019858 | cytosine metabolic process(GO:0019858) |
| 1.3 | 6.3 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 1.1 | 7.6 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 1.1 | 5.4 | GO:0010508 | positive regulation of autophagy(GO:0010508) |
| 1.1 | 4.2 | GO:0071894 | histone H2B conserved C-terminal lysine ubiquitination(GO:0071894) |
| 1.0 | 4.1 | GO:0010520 | regulation of reciprocal meiotic recombination(GO:0010520) negative regulation of meiotic nuclear division(GO:0045835) regulation of meiosis I(GO:0060631) |
| 1.0 | 12.0 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
| 1.0 | 2.9 | GO:0097237 | cellular response to antibiotic(GO:0071236) cellular response to toxic substance(GO:0097237) |
| 0.9 | 4.7 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) |
| 0.9 | 1.8 | GO:0045648 | positive regulation of erythrocyte differentiation(GO:0045648) |
| 0.9 | 3.6 | GO:0031580 | membrane raft polarization(GO:0001766) membrane raft distribution(GO:0031580) |
| 0.9 | 3.6 | GO:0019348 | polyprenol metabolic process(GO:0016093) dolichol metabolic process(GO:0019348) |
| 0.9 | 2.6 | GO:0000059 | protein import into nucleus, docking(GO:0000059) |
| 0.9 | 2.6 | GO:0043461 | proton-transporting ATP synthase complex assembly(GO:0043461) proton-transporting ATP synthase complex biogenesis(GO:0070272) |
| 0.8 | 5.9 | GO:0071267 | amino acid salvage(GO:0043102) L-methionine biosynthetic process(GO:0071265) L-methionine salvage(GO:0071267) |
| 0.8 | 10.4 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |
| 0.8 | 3.9 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.7 | 16.9 | GO:0043486 | histone exchange(GO:0043486) |
| 0.7 | 2.9 | GO:0006122 | mitochondrial electron transport, ubiquinol to cytochrome c(GO:0006122) |
| 0.7 | 4.3 | GO:0015742 | alpha-ketoglutarate transport(GO:0015742) |
| 0.7 | 4.3 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.7 | 5.7 | GO:0033160 | positive regulation of protein import into nucleus, translocation(GO:0033160) |
| 0.7 | 2.1 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) DNA dealkylation(GO:0035510) |
| 0.7 | 2.1 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.7 | 2.1 | GO:0001845 | phagolysosome assembly(GO:0001845) phagosome maturation(GO:0090382) |
| 0.7 | 2.0 | GO:0070171 | negative regulation of tooth mineralization(GO:0070171) |
| 0.7 | 6.8 | GO:0051597 | response to methylmercury(GO:0051597) |
| 0.7 | 8.8 | GO:0032876 | regulation of DNA endoreduplication(GO:0032875) negative regulation of DNA endoreduplication(GO:0032876) DNA endoreduplication(GO:0042023) |
| 0.6 | 1.9 | GO:0060152 | peroxisome localization(GO:0060151) microtubule-based peroxisome localization(GO:0060152) |
| 0.6 | 1.2 | GO:0032070 | regulation of deoxyribonuclease activity(GO:0032070) |
| 0.6 | 3.0 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.6 | 1.8 | GO:0015959 | diadenosine polyphosphate metabolic process(GO:0015959) |
| 0.6 | 4.6 | GO:0043982 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.6 | 2.9 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.6 | 2.3 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) maturation of LSU-rRNA(GO:0000470) |
| 0.5 | 1.6 | GO:0031167 | rRNA methylation(GO:0031167) |
| 0.5 | 2.7 | GO:0051567 | histone H3-K9 methylation(GO:0051567) histone H3-K9 modification(GO:0061647) |
| 0.5 | 3.3 | GO:0061099 | negative regulation of protein tyrosine kinase activity(GO:0061099) |
| 0.5 | 3.2 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.5 | 1.6 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.5 | 3.6 | GO:0042797 | 5S class rRNA transcription from RNA polymerase III type 1 promoter(GO:0042791) tRNA transcription from RNA polymerase III promoter(GO:0042797) |
| 0.5 | 1.5 | GO:0006679 | glucosylceramide biosynthetic process(GO:0006679) |
| 0.5 | 1.5 | GO:0070734 | histone H3-K27 methylation(GO:0070734) |
| 0.5 | 9.8 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) extrinsic apoptotic signaling pathway(GO:0097191) |
| 0.5 | 19.5 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.5 | 1.5 | GO:0031033 | myosin filament organization(GO:0031033) |
| 0.5 | 2.4 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.5 | 2.4 | GO:0046501 | protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.5 | 2.9 | GO:0006422 | aspartyl-tRNA aminoacylation(GO:0006422) |
| 0.5 | 1.4 | GO:0061188 | regulation of chromatin silencing at rDNA(GO:0061187) negative regulation of chromatin silencing at rDNA(GO:0061188) |
| 0.5 | 1.9 | GO:0048024 | regulation of mRNA splicing, via spliceosome(GO:0048024) |
| 0.5 | 1.4 | GO:0019919 | peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) |
| 0.5 | 3.6 | GO:0051895 | negative regulation of focal adhesion assembly(GO:0051895) negative regulation of cell junction assembly(GO:1901889) negative regulation of adherens junction organization(GO:1903392) |
| 0.5 | 1.4 | GO:0035247 | peptidyl-arginine omega-N-methylation(GO:0035247) |
| 0.4 | 1.3 | GO:0034720 | histone H3-K4 demethylation(GO:0034720) |
| 0.4 | 1.3 | GO:0000354 | cis assembly of pre-catalytic spliceosome(GO:0000354) |
| 0.4 | 2.2 | GO:0032075 | positive regulation of nuclease activity(GO:0032075) |
| 0.4 | 1.3 | GO:0046599 | regulation of centriole replication(GO:0046599) |
| 0.4 | 1.7 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.4 | 1.7 | GO:0043490 | malate-aspartate shuttle(GO:0043490) |
| 0.4 | 2.1 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.4 | 1.2 | GO:0006015 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 0.4 | 4.5 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.4 | 2.4 | GO:0045722 | positive regulation of gluconeogenesis(GO:0045722) |
| 0.4 | 1.6 | GO:0009048 | dosage compensation by inactivation of X chromosome(GO:0009048) |
| 0.4 | 1.9 | GO:0072207 | metanephric tubule development(GO:0072170) metanephric epithelium development(GO:0072207) metanephric nephron tubule development(GO:0072234) metanephric nephron epithelium development(GO:0072243) |
| 0.4 | 1.1 | GO:0050983 | spermidine catabolic process(GO:0046203) deoxyhypusine biosynthetic process from spermidine(GO:0050983) |
| 0.4 | 1.9 | GO:0043922 | negative regulation by host of viral transcription(GO:0043922) |
| 0.4 | 1.5 | GO:0006999 | nuclear pore organization(GO:0006999) |
| 0.4 | 6.0 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.4 | 1.8 | GO:0010719 | negative regulation of epithelial to mesenchymal transition(GO:0010719) |
| 0.4 | 1.5 | GO:0008298 | intracellular mRNA localization(GO:0008298) |
| 0.4 | 8.6 | GO:0000291 | nuclear-transcribed mRNA catabolic process, exonucleolytic(GO:0000291) exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.4 | 1.8 | GO:0010447 | response to acidic pH(GO:0010447) |
| 0.4 | 0.7 | GO:0018343 | protein farnesylation(GO:0018343) |
| 0.3 | 2.8 | GO:0045003 | DNA recombinase assembly(GO:0000730) double-strand break repair via synthesis-dependent strand annealing(GO:0045003) |
| 0.3 | 1.4 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) embryonic cleavage(GO:0040016) |
| 0.3 | 1.0 | GO:0008612 | peptidyl-lysine modification to peptidyl-hypusine(GO:0008612) |
| 0.3 | 1.0 | GO:0006419 | alanyl-tRNA aminoacylation(GO:0006419) |
| 0.3 | 1.3 | GO:0046778 | modification by virus of host mRNA processing(GO:0046778) |
| 0.3 | 2.3 | GO:0033962 | cytoplasmic mRNA processing body assembly(GO:0033962) |
| 0.3 | 1.6 | GO:0045767 | obsolete regulation of anti-apoptosis(GO:0045767) |
| 0.3 | 1.0 | GO:0000114 | obsolete regulation of transcription involved in G1 phase of mitotic cell cycle(GO:0000114) |
| 0.3 | 2.6 | GO:0009303 | rRNA transcription(GO:0009303) |
| 0.3 | 1.3 | GO:0019046 | release from viral latency(GO:0019046) |
| 0.3 | 1.0 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.3 | 2.9 | GO:0022417 | protein maturation by protein folding(GO:0022417) |
| 0.3 | 2.8 | GO:0009148 | GTP biosynthetic process(GO:0006183) UTP biosynthetic process(GO:0006228) CTP biosynthetic process(GO:0006241) pyrimidine nucleoside triphosphate biosynthetic process(GO:0009148) pyrimidine ribonucleoside triphosphate metabolic process(GO:0009208) pyrimidine ribonucleoside triphosphate biosynthetic process(GO:0009209) CTP metabolic process(GO:0046036) UTP metabolic process(GO:0046051) |
| 0.3 | 1.9 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 0.3 | 3.4 | GO:0033327 | Leydig cell differentiation(GO:0033327) |
| 0.3 | 0.9 | GO:0072216 | response to heparin(GO:0071503) cellular response to heparin(GO:0071504) positive regulation of metanephros development(GO:0072216) positive regulation of cell proliferation involved in kidney development(GO:1901724) |
| 0.3 | 0.6 | GO:0014889 | muscle atrophy(GO:0014889) |
| 0.3 | 1.2 | GO:0043116 | negative regulation of vascular permeability(GO:0043116) |
| 0.3 | 1.8 | GO:0090231 | regulation of spindle checkpoint(GO:0090231) |
| 0.3 | 3.3 | GO:0031057 | negative regulation of histone modification(GO:0031057) negative regulation of histone acetylation(GO:0035067) negative regulation of protein acetylation(GO:1901984) negative regulation of peptidyl-lysine acetylation(GO:2000757) |
| 0.3 | 15.7 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.3 | 1.8 | GO:0007090 | obsolete regulation of S phase of mitotic cell cycle(GO:0007090) |
| 0.3 | 1.2 | GO:0033147 | negative regulation of intracellular estrogen receptor signaling pathway(GO:0033147) |
| 0.3 | 3.5 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.3 | 3.2 | GO:0042178 | xenobiotic catabolic process(GO:0042178) |
| 0.3 | 3.7 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.3 | 0.9 | GO:0001544 | initiation of primordial ovarian follicle growth(GO:0001544) |
| 0.3 | 1.7 | GO:0035964 | COPI-coated vesicle budding(GO:0035964) Golgi vesicle budding(GO:0048194) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 0.3 | 0.8 | GO:0072393 | microtubule anchoring at microtubule organizing center(GO:0072393) |
| 0.3 | 4.2 | GO:0000090 | mitotic anaphase(GO:0000090) |
| 0.3 | 1.7 | GO:0034501 | protein localization to kinetochore(GO:0034501) |
| 0.3 | 1.1 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.3 | 1.7 | GO:0043247 | protection from non-homologous end joining at telomere(GO:0031848) telomere maintenance in response to DNA damage(GO:0043247) |
| 0.3 | 1.7 | GO:0051294 | establishment of mitotic spindle orientation(GO:0000132) establishment of spindle orientation(GO:0051294) |
| 0.3 | 3.6 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.3 | 3.8 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.3 | 6.2 | GO:0000303 | response to superoxide(GO:0000303) |
| 0.3 | 1.9 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.3 | 1.1 | GO:0000447 | endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000447) maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000466) cleavage involved in rRNA processing(GO:0000469) endonucleolytic cleavage involved in rRNA processing(GO:0000478) endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000479) RNA 5'-end processing(GO:0000966) RNA phosphodiester bond hydrolysis, endonucleolytic(GO:0090502) |
| 0.3 | 6.4 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.3 | 1.6 | GO:0071699 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.3 | 3.9 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.3 | 15.7 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.3 | 7.6 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.3 | 2.1 | GO:0042362 | fat-soluble vitamin biosynthetic process(GO:0042362) |
| 0.3 | 4.6 | GO:0006386 | transcription elongation from RNA polymerase III promoter(GO:0006385) termination of RNA polymerase III transcription(GO:0006386) |
| 0.3 | 1.3 | GO:0060136 | embryonic process involved in female pregnancy(GO:0060136) |
| 0.3 | 7.1 | GO:0060334 | regulation of response to interferon-gamma(GO:0060330) regulation of interferon-gamma-mediated signaling pathway(GO:0060334) |
| 0.3 | 0.8 | GO:0006167 | AMP biosynthetic process(GO:0006167) |
| 0.3 | 1.3 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.2 | 12.2 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.2 | 2.2 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.2 | 2.4 | GO:0015074 | DNA integration(GO:0015074) |
| 0.2 | 0.9 | GO:0003057 | regulation of the force of heart contraction by chemical signal(GO:0003057) |
| 0.2 | 1.6 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.2 | 1.4 | GO:0043615 | astrocyte cell migration(GO:0043615) |
| 0.2 | 0.9 | GO:2000114 | regulation of establishment of cell polarity(GO:2000114) |
| 0.2 | 1.6 | GO:0016557 | peroxisome membrane biogenesis(GO:0016557) |
| 0.2 | 1.4 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.2 | 1.1 | GO:0097190 | apoptotic signaling pathway(GO:0097190) |
| 0.2 | 0.7 | GO:0032927 | positive regulation of activin receptor signaling pathway(GO:0032927) |
| 0.2 | 1.8 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.2 | 1.1 | GO:0071168 | protein localization to chromatin(GO:0071168) |
| 0.2 | 0.6 | GO:0003289 | septum primum development(GO:0003284) atrial septum primum morphogenesis(GO:0003289) |
| 0.2 | 0.2 | GO:0072367 | regulation of lipid transport by regulation of transcription from RNA polymerase II promoter(GO:0072367) |
| 0.2 | 0.6 | GO:0006434 | seryl-tRNA aminoacylation(GO:0006434) |
| 0.2 | 17.5 | GO:0006406 | mRNA export from nucleus(GO:0006406) mRNA-containing ribonucleoprotein complex export from nucleus(GO:0071427) |
| 0.2 | 1.0 | GO:0048550 | negative regulation of pinocytosis(GO:0048550) |
| 0.2 | 2.8 | GO:0006359 | regulation of transcription from RNA polymerase III promoter(GO:0006359) |
| 0.2 | 3.3 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.2 | 2.3 | GO:1901798 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) positive regulation of signal transduction by p53 class mediator(GO:1901798) |
| 0.2 | 0.6 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.2 | 0.6 | GO:0046958 | nonassociative learning(GO:0046958) |
| 0.2 | 0.6 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.2 | 3.1 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.2 | 1.9 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.2 | 1.5 | GO:0015886 | heme transport(GO:0015886) |
| 0.2 | 0.8 | GO:0032525 | somite rostral/caudal axis specification(GO:0032525) |
| 0.2 | 0.4 | GO:1903301 | regulation of glucokinase activity(GO:0033131) positive regulation of glucokinase activity(GO:0033133) regulation of hexokinase activity(GO:1903299) positive regulation of hexokinase activity(GO:1903301) |
| 0.2 | 8.7 | GO:0008033 | tRNA processing(GO:0008033) |
| 0.2 | 3.5 | GO:0018022 | peptidyl-lysine methylation(GO:0018022) |
| 0.2 | 4.6 | GO:0000060 | protein import into nucleus, translocation(GO:0000060) |
| 0.2 | 3.3 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.2 | 4.1 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.2 | 0.9 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.2 | 3.4 | GO:0006596 | polyamine biosynthetic process(GO:0006596) |
| 0.2 | 1.0 | GO:0007256 | activation of JNKK activity(GO:0007256) |
| 0.2 | 1.4 | GO:0061101 | neuroendocrine cell differentiation(GO:0061101) |
| 0.2 | 0.7 | GO:0045900 | negative regulation of translational elongation(GO:0045900) |
| 0.2 | 0.9 | GO:0060528 | secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development(GO:0060528) |
| 0.2 | 1.4 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.2 | 1.4 | GO:0009396 | folic acid-containing compound biosynthetic process(GO:0009396) |
| 0.2 | 12.2 | GO:0030521 | androgen receptor signaling pathway(GO:0030521) |
| 0.2 | 2.3 | GO:0042255 | ribosome assembly(GO:0042255) |
| 0.2 | 1.1 | GO:0048147 | negative regulation of fibroblast proliferation(GO:0048147) |
| 0.2 | 0.3 | GO:0051771 | negative regulation of nitric-oxide synthase biosynthetic process(GO:0051771) |
| 0.2 | 1.0 | GO:0043984 | histone H4-K16 acetylation(GO:0043984) |
| 0.2 | 0.6 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 0.2 | 1.9 | GO:0042274 | ribosomal small subunit biogenesis(GO:0042274) |
| 0.2 | 0.6 | GO:0045007 | depurination(GO:0045007) |
| 0.2 | 1.5 | GO:0009086 | methionine biosynthetic process(GO:0009086) |
| 0.1 | 4.3 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) positive regulation of proteasomal protein catabolic process(GO:1901800) |
| 0.1 | 0.6 | GO:0045176 | apical protein localization(GO:0045176) |
| 0.1 | 4.7 | GO:0033138 | positive regulation of peptidyl-serine phosphorylation(GO:0033138) |
| 0.1 | 1.7 | GO:0006378 | mRNA polyadenylation(GO:0006378) |
| 0.1 | 1.3 | GO:0046835 | carbohydrate phosphorylation(GO:0046835) |
| 0.1 | 3.8 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.1 | 3.9 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.1 | 2.1 | GO:0050812 | acetyl-CoA biosynthetic process from pyruvate(GO:0006086) regulation of acetyl-CoA biosynthetic process from pyruvate(GO:0010510) regulation of acyl-CoA biosynthetic process(GO:0050812) |
| 0.1 | 1.1 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 0.1 | 2.3 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.1 | 0.9 | GO:0060123 | regulation of growth hormone secretion(GO:0060123) positive regulation of growth hormone secretion(GO:0060124) |
| 0.1 | 1.8 | GO:0048169 | regulation of long-term neuronal synaptic plasticity(GO:0048169) |
| 0.1 | 2.6 | GO:0007628 | adult walking behavior(GO:0007628) walking behavior(GO:0090659) |
| 0.1 | 2.9 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.1 | 4.5 | GO:0022904 | respiratory electron transport chain(GO:0022904) |
| 0.1 | 6.8 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.1 | 0.4 | GO:0060279 | regulation of ovulation(GO:0060278) positive regulation of ovulation(GO:0060279) |
| 0.1 | 1.3 | GO:0006362 | transcription elongation from RNA polymerase I promoter(GO:0006362) |
| 0.1 | 3.4 | GO:0006413 | translational initiation(GO:0006413) |
| 0.1 | 1.1 | GO:0007020 | microtubule nucleation(GO:0007020) |
| 0.1 | 1.0 | GO:0051593 | response to folic acid(GO:0051593) |
| 0.1 | 21.3 | GO:0000398 | RNA splicing, via transesterification reactions with bulged adenosine as nucleophile(GO:0000377) mRNA splicing, via spliceosome(GO:0000398) |
| 0.1 | 0.4 | GO:0051881 | regulation of mitochondrial membrane potential(GO:0051881) |
| 0.1 | 0.5 | GO:0035405 | histone-threonine phosphorylation(GO:0035405) |
| 0.1 | 0.5 | GO:0000375 | RNA splicing, via transesterification reactions(GO:0000375) |
| 0.1 | 1.1 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 4.2 | GO:0006626 | protein targeting to mitochondrion(GO:0006626) |
| 0.1 | 0.3 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.1 | 12.6 | GO:0008380 | RNA splicing(GO:0008380) |
| 0.1 | 6.8 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.1 | 1.1 | GO:0007141 | male meiosis I(GO:0007141) |
| 0.1 | 2.4 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.1 | 1.0 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.1 | 0.2 | GO:0006655 | phosphatidylglycerol biosynthetic process(GO:0006655) cardiolipin metabolic process(GO:0032048) cardiolipin biosynthetic process(GO:0032049) phosphatidylglycerol metabolic process(GO:0046471) |
| 0.1 | 1.7 | GO:0045885 | obsolete positive regulation of survival gene product expression(GO:0045885) |
| 0.1 | 1.8 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.1 | 1.2 | GO:0002566 | somatic diversification of immune receptors via somatic mutation(GO:0002566) somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.1 | 0.4 | GO:0042483 | negative regulation of odontogenesis(GO:0042483) |
| 0.1 | 1.5 | GO:1902591 | vesicle coating(GO:0006901) vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) COPII-coated vesicle budding(GO:0090114) single-organism membrane budding(GO:1902591) |
| 0.1 | 0.3 | GO:0003351 | epithelial cilium movement(GO:0003351) epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) |
| 0.1 | 38.2 | GO:0006412 | translation(GO:0006412) |
| 0.1 | 0.9 | GO:0070189 | kynurenine metabolic process(GO:0070189) |
| 0.1 | 0.6 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.1 | 0.5 | GO:0006356 | regulation of transcription from RNA polymerase I promoter(GO:0006356) positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.1 | 0.6 | GO:0010390 | histone monoubiquitination(GO:0010390) |
| 0.1 | 0.3 | GO:0046132 | pyrimidine ribonucleoside biosynthetic process(GO:0046132) |
| 0.1 | 1.7 | GO:0090174 | organelle membrane fusion(GO:0090174) |
| 0.1 | 1.4 | GO:0070875 | positive regulation of glycogen biosynthetic process(GO:0045725) positive regulation of glycogen metabolic process(GO:0070875) |
| 0.1 | 5.0 | GO:0006289 | nucleotide-excision repair(GO:0006289) |
| 0.1 | 1.8 | GO:0008637 | apoptotic mitochondrial changes(GO:0008637) |
| 0.1 | 0.2 | GO:0098534 | centriole replication(GO:0007099) centriole assembly(GO:0098534) |
| 0.1 | 1.2 | GO:0048070 | regulation of developmental pigmentation(GO:0048070) |
| 0.1 | 7.1 | GO:0000279 | M phase(GO:0000279) |
| 0.1 | 5.0 | GO:0051084 | 'de novo' posttranslational protein folding(GO:0051084) |
| 0.1 | 0.9 | GO:0007062 | sister chromatid cohesion(GO:0007062) |
| 0.1 | 1.3 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 7.7 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.1 | 1.1 | GO:0070534 | protein K63-linked ubiquitination(GO:0070534) |
| 0.1 | 0.8 | GO:0046339 | diacylglycerol metabolic process(GO:0046339) |
| 0.1 | 0.7 | GO:0043545 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) molybdopterin cofactor biosynthetic process(GO:0032324) molybdopterin cofactor metabolic process(GO:0043545) prosthetic group metabolic process(GO:0051189) |
| 0.1 | 0.7 | GO:0007199 | G-protein coupled receptor signaling pathway coupled to cGMP nucleotide second messenger(GO:0007199) |
| 0.1 | 1.8 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.1 | 0.5 | GO:0032916 | transforming growth factor beta3 production(GO:0032907) regulation of transforming growth factor beta3 production(GO:0032910) positive regulation of transforming growth factor beta3 production(GO:0032916) |
| 0.1 | 3.8 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.1 | 1.2 | GO:0043523 | regulation of neuron apoptotic process(GO:0043523) regulation of neuron death(GO:1901214) |
| 0.1 | 1.7 | GO:0001569 | patterning of blood vessels(GO:0001569) |
| 0.1 | 4.3 | GO:0006354 | DNA-templated transcription, elongation(GO:0006354) |
| 0.1 | 1.5 | GO:0015012 | heparan sulfate proteoglycan biosynthetic process(GO:0015012) |
| 0.1 | 0.5 | GO:0000729 | DNA double-strand break processing(GO:0000729) |
| 0.1 | 2.5 | GO:0038179 | neurotrophin signaling pathway(GO:0038179) neurotrophin TRK receptor signaling pathway(GO:0048011) |
| 0.1 | 2.2 | GO:0072659 | protein localization to plasma membrane(GO:0072659) protein localization to cell periphery(GO:1990778) |
| 0.1 | 0.3 | GO:0030913 | paranodal junction assembly(GO:0030913) |
| 0.1 | 0.5 | GO:0043009 | embryo development ending in birth or egg hatching(GO:0009792) chordate embryonic development(GO:0043009) |
| 0.1 | 0.1 | GO:0003056 | regulation of vascular smooth muscle contraction(GO:0003056) |
| 0.1 | 0.1 | GO:0000212 | meiotic spindle organization(GO:0000212) |
| 0.1 | 0.5 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.1 | 1.3 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 0.4 | GO:0071569 | protein ufmylation(GO:0071569) |
| 0.1 | 2.5 | GO:0015991 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) ATP hydrolysis coupled proton transport(GO:0015991) ATP hydrolysis coupled transmembrane transport(GO:0090662) |
| 0.1 | 0.2 | GO:0060314 | regulation of ryanodine-sensitive calcium-release channel activity(GO:0060314) |
| 0.1 | 0.7 | GO:0001702 | gastrulation with mouth forming second(GO:0001702) |
| 0.1 | 3.8 | GO:0006986 | response to unfolded protein(GO:0006986) |
| 0.1 | 0.7 | GO:0033599 | regulation of mammary gland epithelial cell proliferation(GO:0033599) |
| 0.1 | 0.9 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.1 | 0.2 | GO:0001780 | neutrophil homeostasis(GO:0001780) |
| 0.1 | 0.7 | GO:0051353 | positive regulation of oxidoreductase activity(GO:0051353) |
| 0.1 | 4.1 | GO:0030833 | regulation of actin filament polymerization(GO:0030833) |
| 0.1 | 2.0 | GO:0006213 | pyrimidine nucleoside metabolic process(GO:0006213) |
| 0.1 | 4.0 | GO:0034339 | obsolete regulation of transcription from RNA polymerase II promoter by nuclear hormone receptor(GO:0034339) |
| 0.1 | 1.7 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.1 | 0.6 | GO:0016558 | protein import into peroxisome matrix(GO:0016558) |
| 0.1 | 0.7 | GO:0050858 | negative regulation of antigen receptor-mediated signaling pathway(GO:0050858) negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.1 | 1.0 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) negative regulation of cysteine-type endopeptidase activity(GO:2000117) |
| 0.1 | 0.2 | GO:0006669 | sphinganine-1-phosphate biosynthetic process(GO:0006669) |
| 0.1 | 0.4 | GO:0001556 | oocyte maturation(GO:0001556) |
| 0.1 | 0.4 | GO:0044060 | regulation of endocrine process(GO:0044060) |
| 0.1 | 0.8 | GO:0070206 | protein trimerization(GO:0070206) |
| 0.1 | 0.4 | GO:0006546 | glycine catabolic process(GO:0006546) |
| 0.1 | 0.4 | GO:0070498 | interleukin-1-mediated signaling pathway(GO:0070498) |
| 0.1 | 2.5 | GO:0060041 | retina development in camera-type eye(GO:0060041) |
| 0.1 | 1.0 | GO:0000045 | autophagosome assembly(GO:0000045) autophagosome organization(GO:1905037) |
| 0.1 | 0.2 | GO:0050000 | mitotic metaphase plate congression(GO:0007080) chromosome localization(GO:0050000) establishment of chromosome localization(GO:0051303) metaphase plate congression(GO:0051310) |
| 0.1 | 0.2 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.1 | 0.7 | GO:2000273 | positive regulation of receptor activity(GO:2000273) |
| 0.1 | 0.8 | GO:0051453 | regulation of intracellular pH(GO:0051453) |
| 0.1 | 0.4 | GO:0014741 | negative regulation of cardiac muscle hypertrophy(GO:0010614) negative regulation of muscle hypertrophy(GO:0014741) |
| 0.0 | 4.7 | GO:0006333 | chromatin assembly or disassembly(GO:0006333) |
| 0.0 | 0.3 | GO:0001885 | endothelial cell development(GO:0001885) |
| 0.0 | 1.5 | GO:0001508 | action potential(GO:0001508) |
| 0.0 | 2.6 | GO:0071772 | BMP signaling pathway(GO:0030509) response to BMP(GO:0071772) cellular response to BMP stimulus(GO:0071773) |
| 0.0 | 0.7 | GO:0001502 | cartilage condensation(GO:0001502) cell aggregation(GO:0098743) |
| 0.0 | 0.5 | GO:0000725 | double-strand break repair via homologous recombination(GO:0000724) recombinational repair(GO:0000725) |
| 0.0 | 1.2 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.1 | GO:0072334 | UDP-galactose transport(GO:0015785) UDP-galactose transmembrane transport(GO:0072334) pyrimidine-containing compound transmembrane transport(GO:0072531) pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) nucleotide transmembrane transport(GO:1901679) |
| 0.0 | 0.3 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.0 | 3.1 | GO:0006521 | regulation of cellular amino acid metabolic process(GO:0006521) |
| 0.0 | 0.5 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.0 | 1.1 | GO:1903320 | regulation of protein ubiquitination(GO:0031396) regulation of protein modification by small protein conjugation or removal(GO:1903320) |
| 0.0 | 2.4 | GO:0097193 | intrinsic apoptotic signaling pathway(GO:0097193) |
| 0.0 | 1.5 | GO:0007093 | mitotic cell cycle checkpoint(GO:0007093) |
| 0.0 | 0.2 | GO:0046579 | positive regulation of Ras protein signal transduction(GO:0046579) |
| 0.0 | 2.1 | GO:0051321 | meiotic cell cycle(GO:0051321) |
| 0.0 | 1.5 | GO:0070647 | protein modification by small protein conjugation or removal(GO:0070647) |
| 0.0 | 0.6 | GO:0042438 | melanin metabolic process(GO:0006582) melanin biosynthetic process(GO:0042438) |
| 0.0 | 0.5 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.0 | 0.4 | GO:0010821 | regulation of mitochondrion organization(GO:0010821) |
| 0.0 | 0.4 | GO:0006613 | cotranslational protein targeting to membrane(GO:0006613) |
| 0.0 | 2.1 | GO:0007219 | Notch signaling pathway(GO:0007219) |
| 0.0 | 1.6 | GO:0006826 | iron ion transport(GO:0006826) |
| 0.0 | 0.4 | GO:0001947 | heart looping(GO:0001947) embryonic heart tube morphogenesis(GO:0003143) determination of heart left/right asymmetry(GO:0061371) |
| 0.0 | 0.7 | GO:0060338 | regulation of type I interferon-mediated signaling pathway(GO:0060338) |
| 0.0 | 0.5 | GO:0001709 | cell fate determination(GO:0001709) |
| 0.0 | 2.4 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.0 | 0.8 | GO:0034332 | adherens junction organization(GO:0034332) |
| 0.0 | 0.1 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.0 | 0.1 | GO:0018377 | N-terminal protein myristoylation(GO:0006499) protein myristoylation(GO:0018377) |
| 0.0 | 0.3 | GO:0045599 | negative regulation of fat cell differentiation(GO:0045599) |
| 0.0 | 0.3 | GO:0006939 | smooth muscle contraction(GO:0006939) |
| 0.0 | 1.2 | GO:0043506 | regulation of JUN kinase activity(GO:0043506) positive regulation of JUN kinase activity(GO:0043507) |
| 0.0 | 2.3 | GO:0006325 | chromatin organization(GO:0006325) |
| 0.0 | 0.5 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.4 | GO:0006098 | pentose-phosphate shunt(GO:0006098) glyceraldehyde-3-phosphate metabolic process(GO:0019682) |
| 0.0 | 1.0 | GO:0007051 | spindle organization(GO:0007051) |
| 0.0 | 0.2 | GO:0040019 | positive regulation of embryonic development(GO:0040019) |
| 0.0 | 0.6 | GO:0006892 | post-Golgi vesicle-mediated transport(GO:0006892) |
| 0.0 | 0.6 | GO:0007528 | neuromuscular junction development(GO:0007528) |
| 0.0 | 3.1 | GO:0006511 | ubiquitin-dependent protein catabolic process(GO:0006511) |
| 0.0 | 0.2 | GO:0051297 | centrosome organization(GO:0051297) |
| 0.0 | 0.7 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.3 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.0 | 0.2 | GO:0007492 | endoderm development(GO:0007492) |
| 0.0 | 0.4 | GO:0035384 | thioester biosynthetic process(GO:0035384) acyl-CoA biosynthetic process(GO:0071616) |
| 0.0 | 0.8 | GO:0042254 | ribosome biogenesis(GO:0042254) |
| 0.0 | 0.5 | GO:1902593 | protein import into nucleus(GO:0006606) protein targeting to nucleus(GO:0044744) single-organism nuclear import(GO:1902593) |
| 0.0 | 0.8 | GO:0051591 | response to cAMP(GO:0051591) |
| 0.0 | 0.3 | GO:0050931 | melanocyte differentiation(GO:0030318) pigment cell differentiation(GO:0050931) |
| 0.0 | 0.8 | GO:0061025 | membrane fusion(GO:0061025) |
| 0.0 | 0.3 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 3.6 | GO:0043066 | negative regulation of apoptotic process(GO:0043066) |
| 0.0 | 0.6 | GO:0042552 | myelination(GO:0042552) |
| 0.0 | 1.2 | GO:0050852 | T cell receptor signaling pathway(GO:0050852) |
| 0.0 | 2.3 | GO:0007601 | visual perception(GO:0007601) |
| 0.0 | 0.5 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.0 | 0.2 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.0 | 0.9 | GO:0006813 | potassium ion transport(GO:0006813) |
| 0.0 | 2.3 | GO:0008284 | positive regulation of cell proliferation(GO:0008284) |
| 0.0 | 0.1 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.1 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.0 | 0.1 | GO:0019852 | L-ascorbic acid metabolic process(GO:0019852) |
| 0.0 | 0.5 | GO:0016358 | dendrite development(GO:0016358) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 4.1 | GO:0032302 | MutSbeta complex(GO:0032302) |
| 1.2 | 4.8 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 1.2 | 3.5 | GO:0005873 | plus-end kinesin complex(GO:0005873) |
| 1.2 | 4.7 | GO:0045257 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 1.1 | 8.0 | GO:0016589 | NURF complex(GO:0016589) |
| 1.1 | 3.3 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 1.0 | 12.6 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.9 | 7.2 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.9 | 7.7 | GO:0000796 | condensin complex(GO:0000796) |
| 0.8 | 4.8 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.7 | 2.9 | GO:0036125 | mitochondrial fatty acid beta-oxidation multienzyme complex(GO:0016507) fatty acid beta-oxidation multienzyme complex(GO:0036125) |
| 0.7 | 6.4 | GO:0005638 | lamin filament(GO:0005638) |
| 0.7 | 5.4 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.7 | 1.3 | GO:0031213 | RSF complex(GO:0031213) |
| 0.6 | 11.5 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.6 | 2.3 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.5 | 3.8 | GO:0000300 | obsolete peripheral to membrane of membrane fraction(GO:0000300) |
| 0.5 | 1.6 | GO:0031262 | Ndc80 complex(GO:0031262) |
| 0.5 | 3.6 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.5 | 4.4 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.5 | 3.4 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.5 | 7.6 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.4 | 3.5 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.4 | 2.6 | GO:0005750 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.4 | 3.4 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.4 | 4.6 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.4 | 3.3 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.4 | 2.0 | GO:0000444 | MIS12/MIND type complex(GO:0000444) |
| 0.4 | 3.1 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.4 | 7.5 | GO:0033276 | transcription factor TFTC complex(GO:0033276) |
| 0.4 | 2.6 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.4 | 3.7 | GO:0042555 | MCM complex(GO:0042555) |
| 0.4 | 1.5 | GO:0000800 | lateral element(GO:0000800) |
| 0.4 | 4.3 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.4 | 3.9 | GO:0010369 | chromocenter(GO:0010369) |
| 0.4 | 3.9 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.3 | 3.5 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.3 | 8.9 | GO:0005761 | organellar ribosome(GO:0000313) mitochondrial ribosome(GO:0005761) |
| 0.3 | 3.7 | GO:0016442 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.3 | 1.3 | GO:0071256 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.3 | 1.6 | GO:0001740 | Barr body(GO:0001740) |
| 0.3 | 17.3 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.3 | 0.9 | GO:0070557 | PCNA-p21 complex(GO:0070557) |
| 0.3 | 1.8 | GO:0044665 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.3 | 0.6 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.3 | 1.4 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.3 | 5.7 | GO:0070469 | respiratory chain(GO:0070469) |
| 0.3 | 1.1 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.3 | 5.3 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.3 | 3.3 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.2 | 1.7 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.2 | 1.0 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.2 | 1.7 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.2 | 1.7 | GO:0030870 | Mre11 complex(GO:0030870) |
| 0.2 | 1.6 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.2 | 2.3 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.2 | 4.7 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.2 | 1.5 | GO:0070820 | tertiary granule(GO:0070820) |
| 0.2 | 1.5 | GO:0002102 | podosome(GO:0002102) |
| 0.2 | 1.9 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.2 | 3.1 | GO:0030530 | obsolete heterogeneous nuclear ribonucleoprotein complex(GO:0030530) |
| 0.2 | 2.1 | GO:0000178 | exosome (RNase complex)(GO:0000178) |
| 0.2 | 2.9 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.2 | 1.9 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.2 | 1.1 | GO:0005827 | polar microtubule(GO:0005827) |
| 0.2 | 2.4 | GO:0016471 | vacuolar proton-transporting V-type ATPase complex(GO:0016471) |
| 0.2 | 6.5 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.2 | 0.9 | GO:0035097 | methyltransferase complex(GO:0034708) histone methyltransferase complex(GO:0035097) |
| 0.2 | 1.1 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.2 | 1.8 | GO:0070688 | MLL5-L complex(GO:0070688) |
| 0.2 | 2.7 | GO:0061200 | clathrin-sculpted gamma-aminobutyric acid transport vesicle(GO:0061200) clathrin-sculpted gamma-aminobutyric acid transport vesicle membrane(GO:0061202) |
| 0.2 | 2.6 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 0.2 | 2.2 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.2 | 1.3 | GO:0000109 | nucleotide-excision repair complex(GO:0000109) |
| 0.2 | 1.3 | GO:0042575 | DNA polymerase complex(GO:0042575) |
| 0.2 | 0.5 | GO:0031372 | UBC13-MMS2 complex(GO:0031372) |
| 0.2 | 21.8 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.2 | 1.0 | GO:0072487 | MSL complex(GO:0072487) |
| 0.2 | 6.6 | GO:0042645 | mitochondrial nucleoid(GO:0042645) |
| 0.2 | 2.1 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.2 | 3.5 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.2 | 0.5 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.2 | 2.2 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.2 | 2.1 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.2 | 18.1 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.2 | 9.0 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.1 | 4.6 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.1 | 3.9 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 2.6 | GO:0098800 | inner mitochondrial membrane protein complex(GO:0098800) |
| 0.1 | 5.6 | GO:0005814 | centriole(GO:0005814) |
| 0.1 | 1.4 | GO:0008021 | synaptic vesicle(GO:0008021) exocytic vesicle(GO:0070382) |
| 0.1 | 1.0 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.1 | 0.6 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.1 | 1.2 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 1.1 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.1 | 3.9 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 4.0 | GO:1990204 | oxidoreductase complex(GO:1990204) |
| 0.1 | 0.8 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.1 | 11.4 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.1 | 9.4 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.1 | 18.7 | GO:0005840 | ribosome(GO:0005840) |
| 0.1 | 3.0 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.1 | 1.0 | GO:0000780 | condensed nuclear chromosome, centromeric region(GO:0000780) |
| 0.1 | 0.6 | GO:0042599 | lamellar body(GO:0042599) |
| 0.1 | 2.9 | GO:0000792 | heterochromatin(GO:0000792) |
| 0.1 | 2.7 | GO:0090568 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) nuclear transcriptional repressor complex(GO:0090568) |
| 0.1 | 8.2 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.1 | 0.7 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.1 | 25.6 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.1 | 4.9 | GO:0000786 | nucleosome(GO:0000786) |
| 0.1 | 1.3 | GO:0030122 | AP-2 adaptor complex(GO:0030122) clathrin coat of endocytic vesicle(GO:0030128) |
| 0.1 | 1.1 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.1 | 6.5 | GO:0000785 | chromatin(GO:0000785) |
| 0.1 | 4.8 | GO:0005884 | actin filament(GO:0005884) |
| 0.1 | 3.5 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 0.4 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 0.6 | GO:0000445 | transcription export complex(GO:0000346) THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.1 | 1.1 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 1.5 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 0.6 | GO:0014069 | postsynaptic density(GO:0014069) postsynaptic specialization(GO:0099572) |
| 0.1 | 12.0 | GO:1990904 | intracellular ribonucleoprotein complex(GO:0030529) ribonucleoprotein complex(GO:1990904) |
| 0.1 | 1.2 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 0.3 | GO:0012507 | ER to Golgi transport vesicle membrane(GO:0012507) |
| 0.1 | 2.5 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.1 | 0.4 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.1 | 3.5 | GO:0000123 | histone acetyltransferase complex(GO:0000123) protein acetyltransferase complex(GO:0031248) acetyltransferase complex(GO:1902493) |
| 0.1 | 1.3 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.1 | 1.7 | GO:0044452 | nucleolar part(GO:0044452) |
| 0.1 | 0.4 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 0.1 | 15.7 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.1 | 0.2 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.1 | 5.1 | GO:0016605 | PML body(GO:0016605) |
| 0.1 | 0.4 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.1 | 0.8 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.1 | 81.0 | GO:0005730 | nucleolus(GO:0005730) |
| 0.1 | 2.7 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.1 | 0.5 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.1 | 2.7 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 0.2 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.1 | 0.8 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.1 | 0.2 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.1 | 4.3 | GO:0000502 | proteasome complex(GO:0000502) |
| 0.1 | 3.9 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.1 | 3.5 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.1 | 0.7 | GO:0098573 | intrinsic component of mitochondrial membrane(GO:0098573) |
| 0.1 | 0.5 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 19.7 | GO:0005625 | obsolete soluble fraction(GO:0005625) |
| 0.1 | 1.4 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.1 | 6.0 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.1 | 0.2 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.1 | 28.5 | GO:0005654 | nucleoplasm(GO:0005654) |
| 0.0 | 5.6 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
| 0.0 | 32.5 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 0.9 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 0.5 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 1.0 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 0.1 | GO:0031371 | ubiquitin conjugating enzyme complex(GO:0031371) |
| 0.0 | 0.6 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.0 | 4.5 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 0.4 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 0.3 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.0 | 0.2 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 0.9 | GO:0043197 | dendritic spine(GO:0043197) |
| 0.0 | 0.6 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 64.9 | GO:0005634 | nucleus(GO:0005634) |
| 0.0 | 0.9 | GO:0030427 | growth cone(GO:0030426) site of polarized growth(GO:0030427) |
| 0.0 | 2.2 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 0.4 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 1.2 | GO:0043025 | neuronal cell body(GO:0043025) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.9 | 11.7 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 1.6 | 4.8 | GO:0008523 | sodium-dependent multivitamin transmembrane transporter activity(GO:0008523) |
| 1.5 | 4.6 | GO:0004853 | uroporphyrinogen decarboxylase activity(GO:0004853) |
| 1.4 | 4.1 | GO:0000404 | heteroduplex DNA loop binding(GO:0000404) double-strand/single-strand DNA junction binding(GO:0000406) dinucleotide repeat insertion binding(GO:0032181) |
| 1.3 | 1.3 | GO:0050660 | flavin adenine dinucleotide binding(GO:0050660) |
| 1.2 | 3.5 | GO:0004809 | tRNA (guanine-N2-)-methyltransferase activity(GO:0004809) |
| 1.2 | 4.7 | GO:0000104 | succinate dehydrogenase activity(GO:0000104) |
| 1.0 | 3.9 | GO:0061731 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.9 | 2.7 | GO:0004609 | phosphatidylserine decarboxylase activity(GO:0004609) |
| 0.9 | 3.6 | GO:0016882 | cyclo-ligase activity(GO:0016882) |
| 0.9 | 2.6 | GO:0000033 | alpha-1,3-mannosyltransferase activity(GO:0000033) |
| 0.8 | 4.2 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.8 | 4.2 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.8 | 3.3 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.7 | 2.2 | GO:0019976 | interleukin-2 binding(GO:0019976) |
| 0.7 | 6.8 | GO:0061659 | ubiquitin-ubiquitin ligase activity(GO:0034450) ubiquitin protein ligase activity(GO:0061630) ubiquitin-like protein ligase activity(GO:0061659) |
| 0.6 | 0.6 | GO:0043175 | RNA polymerase core enzyme binding(GO:0043175) |
| 0.6 | 4.4 | GO:0030911 | TPR domain binding(GO:0030911) |
| 0.6 | 1.9 | GO:0045155 | electron transporter, transferring electrons from CoQH2-cytochrome c reductase complex and cytochrome c oxidase complex activity(GO:0045155) |
| 0.6 | 4.2 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.6 | 5.4 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.6 | 2.9 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.5 | 3.3 | GO:0004749 | ribose phosphate diphosphokinase activity(GO:0004749) |
| 0.5 | 0.5 | GO:0030611 | arsenate reductase activity(GO:0030611) |
| 0.5 | 1.6 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.5 | 7.6 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.5 | 3.0 | GO:0003960 | NADPH:quinone reductase activity(GO:0003960) |
| 0.5 | 3.4 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors(GO:0016679) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.5 | 3.4 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.5 | 2.9 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.5 | 1.0 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.5 | 3.3 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.5 | 3.7 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.5 | 1.4 | GO:0035243 | protein-arginine omega-N symmetric methyltransferase activity(GO:0035243) |
| 0.5 | 0.9 | GO:0032139 | dinucleotide insertion or deletion binding(GO:0032139) |
| 0.4 | 1.7 | GO:0003831 | beta-N-acetylglucosaminylglycopeptide beta-1,4-galactosyltransferase activity(GO:0003831) |
| 0.4 | 1.3 | GO:0033858 | N-acetylgalactosamine kinase activity(GO:0033858) |
| 0.4 | 1.3 | GO:0004476 | mannose-6-phosphate isomerase activity(GO:0004476) |
| 0.4 | 1.3 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 0.4 | 5.4 | GO:0008187 | poly-pyrimidine tract binding(GO:0008187) |
| 0.4 | 20.9 | GO:0036459 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.4 | 1.7 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.4 | 1.2 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.4 | 3.6 | GO:0016004 | phospholipase activator activity(GO:0016004) |
| 0.4 | 1.6 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.4 | 2.8 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.4 | 3.6 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.4 | 1.9 | GO:0004489 | methylenetetrahydrofolate reductase (NAD(P)H) activity(GO:0004489) |
| 0.4 | 1.9 | GO:0003720 | telomerase activity(GO:0003720) RNA-directed DNA polymerase activity(GO:0003964) |
| 0.4 | 1.1 | GO:0034038 | deoxyhypusine synthase activity(GO:0034038) |
| 0.4 | 10.5 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.4 | 1.5 | GO:0004738 | pyruvate dehydrogenase activity(GO:0004738) pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.4 | 3.2 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.4 | 1.8 | GO:0008486 | diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) |
| 0.3 | 4.5 | GO:0003709 | obsolete RNA polymerase III transcription factor activity(GO:0003709) |
| 0.3 | 1.4 | GO:0035242 | protein-arginine omega-N asymmetric methyltransferase activity(GO:0035242) |
| 0.3 | 1.3 | GO:0042808 | obsolete neuronal Cdc2-like kinase binding(GO:0042808) |
| 0.3 | 1.7 | GO:0031702 | type 1 angiotensin receptor binding(GO:0031702) |
| 0.3 | 1.0 | GO:0004813 | alanine-tRNA ligase activity(GO:0004813) |
| 0.3 | 1.3 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.3 | 4.9 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.3 | 1.0 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.3 | 1.9 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.3 | 16.4 | GO:0050136 | NADH dehydrogenase activity(GO:0003954) NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.3 | 5.9 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.3 | 14.8 | GO:0019212 | phosphatase inhibitor activity(GO:0019212) |
| 0.3 | 1.2 | GO:0004663 | Rab geranylgeranyltransferase activity(GO:0004663) |
| 0.3 | 0.9 | GO:0022897 | peptide:proton symporter activity(GO:0015333) proton-dependent peptide secondary active transmembrane transporter activity(GO:0022897) |
| 0.3 | 1.5 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.3 | 4.7 | GO:0005521 | lamin binding(GO:0005521) |
| 0.3 | 4.3 | GO:0016986 | obsolete transcription initiation factor activity(GO:0016986) |
| 0.3 | 5.1 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.3 | 1.1 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.3 | 5.4 | GO:0004004 | ATP-dependent RNA helicase activity(GO:0004004) |
| 0.3 | 2.8 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.3 | 0.8 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.3 | 1.6 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.3 | 20.5 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.3 | 6.5 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.3 | 0.8 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.3 | 3.1 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.3 | 2.0 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.3 | 1.0 | GO:0032810 | sterol response element binding(GO:0032810) |
| 0.2 | 1.5 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.2 | 3.4 | GO:0003857 | 3-hydroxyacyl-CoA dehydrogenase activity(GO:0003857) |
| 0.2 | 2.4 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.2 | 3.1 | GO:0070888 | E-box binding(GO:0070888) |
| 0.2 | 0.5 | GO:0003691 | double-stranded telomeric DNA binding(GO:0003691) |
| 0.2 | 0.7 | GO:0005047 | signal recognition particle binding(GO:0005047) |
| 0.2 | 9.7 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.2 | 1.1 | GO:0015140 | malate transmembrane transporter activity(GO:0015140) C4-dicarboxylate transmembrane transporter activity(GO:0015556) |
| 0.2 | 0.9 | GO:0070548 | 1-aminocyclopropane-1-carboxylate synthase activity(GO:0016847) L-glutamine aminotransferase activity(GO:0070548) |
| 0.2 | 0.7 | GO:0016362 | activin receptor activity, type II(GO:0016362) |
| 0.2 | 6.8 | GO:0016875 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.2 | 0.2 | GO:0046965 | retinoid X receptor binding(GO:0046965) |
| 0.2 | 2.3 | GO:0070513 | death domain binding(GO:0070513) |
| 0.2 | 1.4 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.2 | 0.6 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.2 | 1.2 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.2 | 0.8 | GO:0016531 | metallochaperone activity(GO:0016530) copper chaperone activity(GO:0016531) |
| 0.2 | 3.4 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.2 | 1.2 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.2 | 8.2 | GO:0002039 | p53 binding(GO:0002039) |
| 0.2 | 0.6 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.2 | 1.9 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.2 | 2.3 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.2 | 1.9 | GO:0003906 | DNA-(apurinic or apyrimidinic site) lyase activity(GO:0003906) |
| 0.2 | 1.4 | GO:0008143 | poly(A) binding(GO:0008143) poly-purine tract binding(GO:0070717) |
| 0.2 | 0.9 | GO:0004844 | uracil DNA N-glycosylase activity(GO:0004844) deaminated base DNA N-glycosylase activity(GO:0097506) |
| 0.2 | 1.5 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.2 | 8.6 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) RNA polymerase activity(GO:0034062) |
| 0.2 | 4.2 | GO:0004221 | obsolete ubiquitin thiolesterase activity(GO:0004221) |
| 0.2 | 0.7 | GO:0034190 | apolipoprotein receptor binding(GO:0034190) |
| 0.2 | 2.9 | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups(GO:0016780) |
| 0.2 | 0.5 | GO:0004532 | exoribonuclease activity(GO:0004532) |
| 0.2 | 10.0 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.2 | 6.8 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.2 | 3.2 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.2 | 2.9 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.2 | 0.6 | GO:0032408 | MutLbeta complex binding(GO:0032406) MutSbeta complex binding(GO:0032408) |
| 0.1 | 31.5 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 0.6 | GO:0033981 | D-dopachrome decarboxylase activity(GO:0033981) |
| 0.1 | 2.5 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.1 | 0.9 | GO:0008649 | rRNA methyltransferase activity(GO:0008649) |
| 0.1 | 4.1 | GO:0016278 | lysine N-methyltransferase activity(GO:0016278) protein-lysine N-methyltransferase activity(GO:0016279) |
| 0.1 | 1.2 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.1 | 1.6 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.1 | 3.7 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 2.2 | GO:0030528 | obsolete transcription regulator activity(GO:0030528) |
| 0.1 | 0.4 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 0.1 | 1.2 | GO:0008135 | translation factor activity, RNA binding(GO:0008135) |
| 0.1 | 2.2 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 0.4 | GO:0004119 | cGMP-inhibited cyclic-nucleotide phosphodiesterase activity(GO:0004119) |
| 0.1 | 1.8 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.1 | 1.1 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.1 | 0.5 | GO:0042608 | T cell receptor binding(GO:0042608) |
| 0.1 | 2.1 | GO:0043548 | phosphatidylinositol 3-kinase binding(GO:0043548) |
| 0.1 | 8.1 | GO:0042393 | histone binding(GO:0042393) |
| 0.1 | 0.7 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.1 | 0.6 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.1 | 2.3 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.1 | 8.6 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.1 | 1.2 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.1 | 2.0 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 3.1 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.1 | 1.4 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.1 | 0.8 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.1 | 1.5 | GO:0003711 | obsolete transcription elongation regulator activity(GO:0003711) |
| 0.1 | 2.9 | GO:0005310 | dicarboxylic acid transmembrane transporter activity(GO:0005310) |
| 0.1 | 5.0 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.1 | 1.4 | GO:0046625 | sphingolipid binding(GO:0046625) |
| 0.1 | 3.2 | GO:0019903 | protein phosphatase binding(GO:0019903) |
| 0.1 | 0.6 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.1 | 8.2 | GO:0031072 | heat shock protein binding(GO:0031072) |
| 0.1 | 0.8 | GO:0090599 | alpha-glucosidase activity(GO:0090599) |
| 0.1 | 8.0 | GO:0008026 | ATP-dependent helicase activity(GO:0008026) purine NTP-dependent helicase activity(GO:0070035) |
| 0.1 | 1.7 | GO:1905030 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.1 | 0.8 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.1 | 1.0 | GO:0016421 | CoA carboxylase activity(GO:0016421) |
| 0.1 | 0.5 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.1 | 0.6 | GO:0030275 | LRR domain binding(GO:0030275) |
| 0.1 | 8.6 | GO:0031625 | ubiquitin protein ligase binding(GO:0031625) ubiquitin-like protein ligase binding(GO:0044389) |
| 0.1 | 0.9 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.1 | 3.3 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.1 | 2.1 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.1 | 0.7 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.1 | 3.2 | GO:0008094 | DNA-dependent ATPase activity(GO:0008094) |
| 0.1 | 0.5 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) |
| 0.1 | 1.2 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.1 | 0.9 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.1 | 0.7 | GO:0005131 | growth hormone receptor binding(GO:0005131) |
| 0.1 | 1.0 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 1.8 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 0.4 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.1 | 0.7 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) histone deacetylase activity (H4-K16 specific)(GO:0034739) NAD-dependent histone deacetylase activity (H4-K16 specific)(GO:0046970) |
| 0.1 | 2.2 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.1 | 1.9 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.1 | 2.1 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.1 | 0.6 | GO:0015288 | porin activity(GO:0015288) |
| 0.1 | 0.3 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.1 | 1.0 | GO:0008320 | protein transmembrane transporter activity(GO:0008320) |
| 0.1 | 2.2 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.1 | 0.9 | GO:0019201 | nucleotide kinase activity(GO:0019201) |
| 0.1 | 1.6 | GO:0019003 | GDP binding(GO:0019003) |
| 0.1 | 6.9 | GO:0008168 | methyltransferase activity(GO:0008168) |
| 0.1 | 2.3 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 6.2 | GO:0015078 | hydrogen ion transmembrane transporter activity(GO:0015078) |
| 0.1 | 11.7 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.1 | 1.0 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.1 | 130.4 | GO:0003676 | nucleic acid binding(GO:0003676) |
| 0.1 | 1.1 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 0.3 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 0.1 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) |
| 0.0 | 0.8 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.0 | 0.4 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 2.8 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 0.1 | GO:0005459 | UDP-galactose transmembrane transporter activity(GO:0005459) |
| 0.0 | 1.7 | GO:0004177 | aminopeptidase activity(GO:0004177) |
| 0.0 | 0.5 | GO:0001619 | obsolete lysosphingolipid and lysophosphatidic acid receptor activity(GO:0001619) |
| 0.0 | 3.9 | GO:0008234 | cysteine-type peptidase activity(GO:0008234) |
| 0.0 | 2.4 | GO:0008565 | protein transporter activity(GO:0008565) |
| 0.0 | 1.8 | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.0 | 0.4 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.0 | 0.3 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.0 | 5.1 | GO:0042623 | ATPase activity, coupled(GO:0042623) |
| 0.0 | 0.5 | GO:0004437 | obsolete inositol or phosphatidylinositol phosphatase activity(GO:0004437) |
| 0.0 | 3.1 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 0.2 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.0 | 1.2 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 0.1 | GO:0004379 | glycylpeptide N-tetradecanoyltransferase activity(GO:0004379) myristoyltransferase activity(GO:0019107) |
| 0.0 | 1.6 | GO:0016874 | ligase activity(GO:0016874) |
| 0.0 | 0.4 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 1.8 | GO:0016651 | oxidoreductase activity, acting on NAD(P)H(GO:0016651) |
| 0.0 | 3.6 | GO:0005057 | receptor signaling protein activity(GO:0005057) |
| 0.0 | 0.3 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 1.6 | GO:0016563 | obsolete transcription activator activity(GO:0016563) |
| 0.0 | 0.2 | GO:0051766 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) inositol trisphosphate kinase activity(GO:0051766) |
| 0.0 | 4.2 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 0.8 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 0.1 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.0 | 0.1 | GO:0016972 | thiol oxidase activity(GO:0016972) |
| 0.0 | 0.4 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.0 | 0.2 | GO:0015114 | phosphate ion transmembrane transporter activity(GO:0015114) |
| 0.0 | 1.5 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 0.7 | GO:0016779 | nucleotidyltransferase activity(GO:0016779) |
| 0.0 | 1.3 | GO:0016782 | transferase activity, transferring sulfur-containing groups(GO:0016782) |
| 0.0 | 0.3 | GO:0009881 | photoreceptor activity(GO:0009881) |
| 0.0 | 0.6 | GO:0004518 | nuclease activity(GO:0004518) |
| 0.0 | 0.1 | GO:0003863 | alpha-ketoacid dehydrogenase activity(GO:0003826) 3-methyl-2-oxobutanoate dehydrogenase (2-methylpropanoyl-transferring) activity(GO:0003863) |
| 0.0 | 1.7 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 2.8 | GO:0004857 | enzyme inhibitor activity(GO:0004857) |
| 0.0 | 5.6 | GO:0005524 | ATP binding(GO:0005524) |
| 0.0 | 0.2 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.0 | 1.3 | GO:0019207 | kinase regulator activity(GO:0019207) |
| 0.0 | 1.4 | GO:0019787 | ubiquitin-like protein transferase activity(GO:0019787) |
| 0.0 | 0.2 | GO:0051018 | protein kinase A binding(GO:0051018) |
| 0.0 | 0.0 | GO:0016724 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.0 | 0.1 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 9.1 | ST_G_ALPHA_S_PATHWAY | G alpha s Pathway |
| 0.2 | 4.4 | SA_REG_CASCADE_OF_CYCLIN_EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 2.1 | SA_G1_AND_S_PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.1 | 2.6 | SA_FAS_SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.1 | 4.3 | ST_DIFFERENTIATION_PATHWAY_IN_PC12_CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.1 | 6.3 | ST_JNK_MAPK_PATHWAY | JNK MAPK Pathway |
| 0.1 | 0.8 | SIG_REGULATION_OF_THE_ACTIN_CYTOSKELETON_BY_RHO_GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 0.9 | ST_GAQ_PATHWAY | G alpha q Pathway |
| 0.1 | 0.7 | ST_IL_13_PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.0 | 3.4 | ST_T_CELL_SIGNAL_TRANSDUCTION | T Cell Signal Transduction |
| 0.0 | 1.9 | ST_GA13_PATHWAY | G alpha 13 Pathway |
| 0.0 | 0.6 | ST_INTERLEUKIN_4_PATHWAY | Interleukin 4 (IL-4) Pathway |
| 0.0 | 1.0 | SIG_CHEMOTAXIS | Genes related to chemotaxis |
| 0.0 | 0.4 | ST_ADRENERGIC | Adrenergic Pathway |