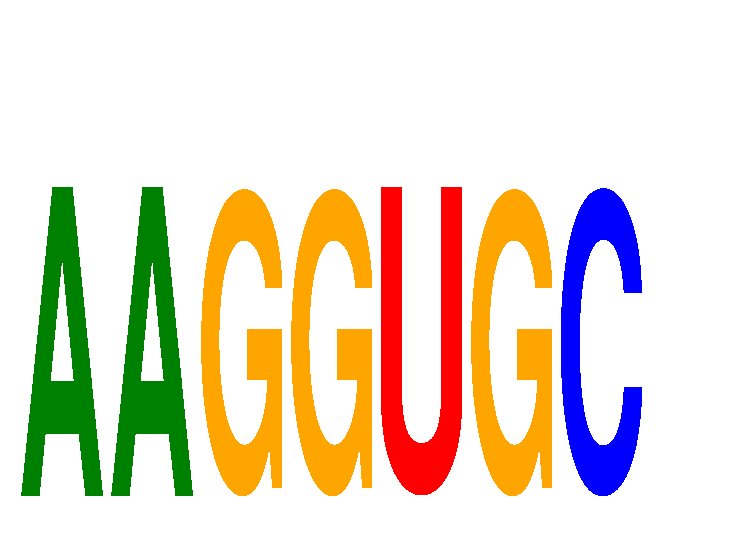Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
Results for AAGGUGC
Z-value: 0.86
miRNA associated with seed AAGGUGC
| Name | miRBASE accession |
|---|---|
|
hsa-miR-18a-5p
|
MIMAT0000072 |
|
hsa-miR-18b-5p
|
MIMAT0001412 |
|
hsa-miR-4735-3p
|
MIMAT0019861 |
Activity profile of AAGGUGC motif
Sorted Z-values of AAGGUGC motif
Network of associatons between targets according to the STRING database.
First level regulatory network of AAGGUGC
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_235860616 | 3.39 |
ENST00000392011.2 |
SH3BP4 |
SH3-domain binding protein 4 |
| chr3_-_123603137 | 3.19 |
ENST00000360304.3 ENST00000359169.1 ENST00000346322.5 ENST00000360772.3 |
MYLK |
myosin light chain kinase |
| chr1_-_20812690 | 3.15 |
ENST00000375078.3 |
CAMK2N1 |
calcium/calmodulin-dependent protein kinase II inhibitor 1 |
| chr15_+_39873268 | 2.43 |
ENST00000397591.2 ENST00000260356.5 |
THBS1 |
thrombospondin 1 |
| chr2_+_36582857 | 2.26 |
ENST00000280527.2 |
CRIM1 |
cysteine rich transmembrane BMP regulator 1 (chordin-like) |
| chr22_+_38093005 | 2.01 |
ENST00000406386.3 |
TRIOBP |
TRIO and F-actin binding protein |
| chr6_-_132272504 | 1.65 |
ENST00000367976.3 |
CTGF |
connective tissue growth factor |
| chr8_-_18871159 | 1.64 |
ENST00000327040.8 ENST00000440756.2 |
PSD3 |
pleckstrin and Sec7 domain containing 3 |
| chr9_-_74383799 | 1.57 |
ENST00000377044.4 |
TMEM2 |
transmembrane protein 2 |
| chr3_+_105085734 | 1.53 |
ENST00000306107.5 |
ALCAM |
activated leukocyte cell adhesion molecule |
| chr4_-_157892498 | 1.16 |
ENST00000502773.1 |
PDGFC |
platelet derived growth factor C |
| chr3_-_114790179 | 1.10 |
ENST00000462705.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr3_+_107241783 | 1.01 |
ENST00000415149.2 ENST00000402543.1 ENST00000325805.8 ENST00000427402.1 |
BBX |
bobby sox homolog (Drosophila) |
| chr3_+_187930719 | 0.97 |
ENST00000312675.4 |
LPP |
LIM domain containing preferred translocation partner in lipoma |
| chr1_-_120612240 | 0.84 |
ENST00000256646.2 |
NOTCH2 |
notch 2 |
| chr4_-_186877502 | 0.84 |
ENST00000431902.1 ENST00000284776.7 ENST00000415274.1 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr6_-_11232891 | 0.83 |
ENST00000379433.5 ENST00000379446.5 |
NEDD9 |
neural precursor cell expressed, developmentally down-regulated 9 |
| chr5_-_73937244 | 0.83 |
ENST00000302351.4 ENST00000510316.1 ENST00000508331.1 |
ENC1 |
ectodermal-neural cortex 1 (with BTB domain) |
| chr10_-_735553 | 0.77 |
ENST00000280886.6 ENST00000423550.1 |
DIP2C |
DIP2 disco-interacting protein 2 homolog C (Drosophila) |
| chr11_+_108093839 | 0.76 |
ENST00000452508.2 |
ATM |
ataxia telangiectasia mutated |
| chr15_-_56209306 | 0.76 |
ENST00000506154.1 ENST00000338963.2 ENST00000508342.1 |
NEDD4 |
neural precursor cell expressed, developmentally down-regulated 4, E3 ubiquitin protein ligase |
| chr20_+_34700333 | 0.74 |
ENST00000441639.1 |
EPB41L1 |
erythrocyte membrane protein band 4.1-like 1 |
| chr6_+_39760783 | 0.73 |
ENST00000398904.2 ENST00000538976.1 |
DAAM2 |
dishevelled associated activator of morphogenesis 2 |
| chr3_-_185542817 | 0.69 |
ENST00000382199.2 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr1_-_161993616 | 0.69 |
ENST00000294794.3 |
OLFML2B |
olfactomedin-like 2B |
| chr6_-_111136513 | 0.54 |
ENST00000368911.3 |
CDK19 |
cyclin-dependent kinase 19 |
| chr3_+_37903432 | 0.51 |
ENST00000443503.2 |
CTDSPL |
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase-like |
| chr11_-_34379546 | 0.51 |
ENST00000435224.2 |
ABTB2 |
ankyrin repeat and BTB (POZ) domain containing 2 |
| chr9_+_125703282 | 0.50 |
ENST00000373647.4 ENST00000402311.1 |
RABGAP1 |
RAB GTPase activating protein 1 |
| chr3_+_171758344 | 0.49 |
ENST00000336824.4 ENST00000423424.1 |
FNDC3B |
fibronectin type III domain containing 3B |
| chr1_-_95007193 | 0.43 |
ENST00000370207.4 ENST00000334047.7 |
F3 |
coagulation factor III (thromboplastin, tissue factor) |
| chr17_-_1395954 | 0.43 |
ENST00000359786.5 |
MYO1C |
myosin IC |
| chr12_-_44200052 | 0.39 |
ENST00000548315.1 ENST00000552521.1 ENST00000546662.1 ENST00000548403.1 ENST00000546506.1 |
TWF1 |
twinfilin actin-binding protein 1 |
| chr10_-_119806085 | 0.37 |
ENST00000355624.3 |
RAB11FIP2 |
RAB11 family interacting protein 2 (class I) |
| chr20_+_56884752 | 0.36 |
ENST00000244040.3 |
RAB22A |
RAB22A, member RAS oncogene family |
| chr17_+_48423453 | 0.36 |
ENST00000017003.2 ENST00000509778.1 ENST00000507602.1 |
XYLT2 |
xylosyltransferase II |
| chr17_+_40834580 | 0.35 |
ENST00000264638.4 |
CNTNAP1 |
contactin associated protein 1 |
| chr6_-_99873145 | 0.35 |
ENST00000369239.5 ENST00000438806.1 |
PNISR |
PNN-interacting serine/arginine-rich protein |
| chr21_-_36260980 | 0.31 |
ENST00000344691.4 ENST00000358356.5 |
RUNX1 |
runt-related transcription factor 1 |
| chr4_+_154125565 | 0.31 |
ENST00000338700.5 |
TRIM2 |
tripartite motif containing 2 |
| chr2_-_26101374 | 0.31 |
ENST00000435504.4 |
ASXL2 |
additional sex combs like 2 (Drosophila) |
| chr2_+_46926048 | 0.29 |
ENST00000306503.5 |
SOCS5 |
suppressor of cytokine signaling 5 |
| chr12_+_49212514 | 0.29 |
ENST00000301050.2 ENST00000548279.1 ENST00000547230.1 |
CACNB3 |
calcium channel, voltage-dependent, beta 3 subunit |
| chr2_+_28615669 | 0.27 |
ENST00000379619.1 ENST00000264716.4 |
FOSL2 |
FOS-like antigen 2 |
| chr7_-_111846435 | 0.26 |
ENST00000437633.1 ENST00000428084.1 |
DOCK4 |
dedicator of cytokinesis 4 |
| chr19_-_36523709 | 0.26 |
ENST00000592017.1 ENST00000360535.4 |
CLIP3 |
CAP-GLY domain containing linker protein 3 |
| chr5_-_59189545 | 0.24 |
ENST00000340635.6 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr11_-_72853091 | 0.24 |
ENST00000311172.7 ENST00000409314.1 |
FCHSD2 |
FCH and double SH3 domains 2 |
| chr4_+_183164574 | 0.23 |
ENST00000511685.1 |
TENM3 |
teneurin transmembrane protein 3 |
| chr16_+_69599861 | 0.23 |
ENST00000354436.2 |
NFAT5 |
nuclear factor of activated T-cells 5, tonicity-responsive |
| chr19_-_46000251 | 0.22 |
ENST00000590526.1 ENST00000344680.4 ENST00000245923.4 |
RTN2 |
reticulon 2 |
| chr10_-_81205373 | 0.22 |
ENST00000372336.3 |
ZCCHC24 |
zinc finger, CCHC domain containing 24 |
| chr16_-_74640986 | 0.19 |
ENST00000422840.2 ENST00000565260.1 ENST00000447066.2 ENST00000205061.5 |
GLG1 |
golgi glycoprotein 1 |
| chr7_-_121036337 | 0.18 |
ENST00000426156.1 ENST00000359943.3 ENST00000412653.1 |
FAM3C |
family with sequence similarity 3, member C |
| chr20_+_5107420 | 0.18 |
ENST00000460006.1 |
CDS2 |
CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 2 |
| chr11_-_79151695 | 0.16 |
ENST00000278550.7 |
TENM4 |
teneurin transmembrane protein 4 |
| chr1_+_84543734 | 0.16 |
ENST00000370689.2 |
PRKACB |
protein kinase, cAMP-dependent, catalytic, beta |
| chr16_+_30960375 | 0.15 |
ENST00000318663.4 ENST00000566237.1 ENST00000562699.1 |
ORAI3 |
ORAI calcium release-activated calcium modulator 3 |
| chr5_-_142783175 | 0.15 |
ENST00000231509.3 ENST00000394464.2 |
NR3C1 |
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr12_+_12764773 | 0.15 |
ENST00000228865.2 |
CREBL2 |
cAMP responsive element binding protein-like 2 |
| chr10_+_72575643 | 0.14 |
ENST00000373202.3 |
SGPL1 |
sphingosine-1-phosphate lyase 1 |
| chr10_+_102295616 | 0.14 |
ENST00000299163.6 |
HIF1AN |
hypoxia inducible factor 1, alpha subunit inhibitor |
| chr14_-_81687197 | 0.13 |
ENST00000553612.1 |
GTF2A1 |
general transcription factor IIA, 1, 19/37kDa |
| chr7_+_90225796 | 0.13 |
ENST00000380050.3 |
CDK14 |
cyclin-dependent kinase 14 |
| chr20_+_60697480 | 0.12 |
ENST00000370915.1 ENST00000253001.4 ENST00000400318.2 ENST00000279068.6 ENST00000279069.7 |
LSM14B |
LSM14B, SCD6 homolog B (S. cerevisiae) |
| chr10_+_111767720 | 0.10 |
ENST00000356080.4 ENST00000277900.8 |
ADD3 |
adducin 3 (gamma) |
| chr17_+_46985731 | 0.10 |
ENST00000360943.5 |
UBE2Z |
ubiquitin-conjugating enzyme E2Z |
| chr17_+_68165657 | 0.10 |
ENST00000243457.3 |
KCNJ2 |
potassium inwardly-rectifying channel, subfamily J, member 2 |
| chr11_+_94822968 | 0.10 |
ENST00000278505.4 |
ENDOD1 |
endonuclease domain containing 1 |
| chr17_-_62340581 | 0.09 |
ENST00000258991.3 ENST00000583738.1 ENST00000584379.1 |
TEX2 |
testis expressed 2 |
| chr20_+_30865429 | 0.08 |
ENST00000375712.3 |
KIF3B |
kinesin family member 3B |
| chr17_-_40306934 | 0.07 |
ENST00000592574.1 ENST00000550406.1 ENST00000547517.1 ENST00000393860.3 ENST00000346213.4 |
CTD-2132N18.3 RAB5C |
Uncharacterized protein RAB5C, member RAS oncogene family |
| chr16_-_17564738 | 0.06 |
ENST00000261381.6 |
XYLT1 |
xylosyltransferase I |
| chr5_+_31193847 | 0.06 |
ENST00000514738.1 ENST00000265071.2 |
CDH6 |
cadherin 6, type 2, K-cadherin (fetal kidney) |
| chr20_+_43104508 | 0.06 |
ENST00000262605.4 ENST00000372904.3 |
TTPAL |
tocopherol (alpha) transfer protein-like |
| chr20_+_43595115 | 0.06 |
ENST00000372806.3 ENST00000396731.4 ENST00000372801.1 ENST00000499879.2 |
STK4 |
serine/threonine kinase 4 |
| chr15_+_73344791 | 0.04 |
ENST00000261908.6 |
NEO1 |
neogenin 1 |
| chr9_+_91003271 | 0.04 |
ENST00000375859.3 ENST00000541629.1 |
SPIN1 |
spindlin 1 |
| chr19_+_3572925 | 0.04 |
ENST00000333651.6 ENST00000417382.1 ENST00000453933.1 ENST00000262949.7 |
HMG20B |
high mobility group 20B |
| chr3_-_169899504 | 0.04 |
ENST00000474275.1 ENST00000484931.1 ENST00000494943.1 ENST00000497658.1 ENST00000465896.1 ENST00000475729.1 ENST00000495893.2 ENST00000481639.1 ENST00000467570.1 ENST00000466189.1 |
PHC3 |
polyhomeotic homolog 3 (Drosophila) |
| chr1_-_204329013 | 0.01 |
ENST00000272203.3 ENST00000414478.1 |
PLEKHA6 |
pleckstrin homology domain containing, family A member 6 |
| chr18_-_45456930 | 0.01 |
ENST00000262160.6 ENST00000587269.1 |
SMAD2 |
SMAD family member 2 |
| chr2_+_24714729 | 0.01 |
ENST00000406961.1 ENST00000405141.1 |
NCOA1 |
nuclear receptor coactivator 1 |
| chr6_-_53409890 | 0.00 |
ENST00000229416.6 |
GCLC |
glutamate-cysteine ligase, catalytic subunit |
| chr1_+_78245303 | 0.00 |
ENST00000370791.3 ENST00000443751.2 |
FAM73A |
family with sequence similarity 73, member A |
Gene Ontology Analysis
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.2 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 2.4 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 1.6 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.7 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 0.7 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 0.8 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.7 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 2.3 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.4 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.1 | 0.4 | GO:0045160 | myosin I complex(GO:0045160) |
| 0.0 | 1.5 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.0 | 0.1 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.0 | 3.2 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 3.4 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.4 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.0 | 0.2 | GO:0014802 | terminal cisterna(GO:0014802) |
| 0.0 | 1.6 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.0 | 0.1 | GO:0005873 | plus-end kinesin complex(GO:0005873) |
| 0.0 | 0.8 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 0.3 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.4 | GO:0031233 | intrinsic component of external side of plasma membrane(GO:0031233) |
| 0.0 | 2.0 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.5 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 2.0 | GO:0030496 | midbody(GO:0030496) |
| 0.0 | 0.3 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 0.6 | GO:0055038 | recycling endosome membrane(GO:0055038) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.4 | GO:0002605 | negative regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002581) negative regulation of dendritic cell antigen processing and presentation(GO:0002605) |
| 0.5 | 1.6 | GO:0070318 | positive regulation of G0 to G1 transition(GO:0070318) |
| 0.5 | 2.4 | GO:0030047 | actin modification(GO:0030047) |
| 0.4 | 3.2 | GO:0060414 | aorta smooth muscle tissue morphogenesis(GO:0060414) |
| 0.3 | 0.8 | GO:1904884 | signal transduction involved in G2 DNA damage checkpoint(GO:0072425) signal transduction involved in mitotic G2 DNA damage checkpoint(GO:0072434) telomerase catalytic core complex assembly(GO:1904868) regulation of telomerase catalytic core complex assembly(GO:1904882) positive regulation of telomerase catalytic core complex assembly(GO:1904884) |
| 0.2 | 0.8 | GO:0044111 | development involved in symbiotic interaction(GO:0044111) |
| 0.1 | 0.4 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 0.1 | 0.8 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.1 | 3.4 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.1 | 1.6 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 0.8 | GO:0061314 | Notch signaling involved in heart development(GO:0061314) |
| 0.1 | 0.4 | GO:1900748 | positive regulation of vascular endothelial growth factor signaling pathway(GO:1900748) |
| 0.1 | 0.2 | GO:0032289 | central nervous system myelin formation(GO:0032289) |
| 0.1 | 1.6 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.1 | 0.3 | GO:0045906 | negative regulation of vasoconstriction(GO:0045906) |
| 0.1 | 0.3 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.1 | 1.5 | GO:0048846 | axon extension involved in axon guidance(GO:0048846) neuron projection extension involved in neuron projection guidance(GO:1902284) |
| 0.1 | 0.4 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.0 | 0.3 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.0 | 0.1 | GO:0042264 | peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.0 | 1.2 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 0.0 | 1.9 | GO:0048009 | insulin-like growth factor receptor signaling pathway(GO:0048009) |
| 0.0 | 0.4 | GO:0030202 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.0 | 0.2 | GO:0097338 | response to clozapine(GO:0097338) |
| 0.0 | 0.3 | GO:0035360 | positive regulation of peroxisome proliferator activated receptor signaling pathway(GO:0035360) |
| 0.0 | 0.8 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 0.2 | GO:0086024 | adrenergic receptor signaling pathway involved in positive regulation of heart rate(GO:0086024) |
| 0.0 | 0.2 | GO:0006655 | phosphatidylglycerol biosynthetic process(GO:0006655) |
| 0.0 | 0.1 | GO:0090076 | relaxation of skeletal muscle(GO:0090076) |
| 0.0 | 0.3 | GO:0061577 | calcium ion transmembrane transport via high voltage-gated calcium channel(GO:0061577) |
| 0.0 | 1.1 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.0 | 0.3 | GO:0030854 | positive regulation of granulocyte differentiation(GO:0030854) |
| 0.0 | 0.1 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 0.0 | 0.2 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.7 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 0.5 | GO:0097237 | cellular response to toxic substance(GO:0097237) |
| 0.0 | 0.1 | GO:0060800 | regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 0.0 | 0.1 | GO:0033327 | Leydig cell differentiation(GO:0033327) |
| 0.0 | 0.4 | GO:0003091 | renal water homeostasis(GO:0003091) |
| 0.0 | 0.1 | GO:0072383 | plus-end-directed vesicle transport along microtubule(GO:0072383) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.4 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.6 | 3.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.5 | 3.2 | GO:0010858 | calcium-dependent protein kinase regulator activity(GO:0010858) |
| 0.2 | 2.0 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.2 | 1.0 | GO:0038049 | transcription factor activity, ligand-activated RNA polymerase II transcription factor binding(GO:0038049) |
| 0.1 | 1.6 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.1 | 0.4 | GO:0030158 | protein xylosyltransferase activity(GO:0030158) |
| 0.1 | 3.4 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 0.8 | GO:0004677 | DNA-dependent protein kinase activity(GO:0004677) |
| 0.1 | 0.8 | GO:0050815 | phosphoserine binding(GO:0050815) |
| 0.1 | 3.9 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.1 | 0.3 | GO:0097001 | ceramide binding(GO:0097001) |
| 0.1 | 1.6 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 1.2 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.1 | 0.2 | GO:0004605 | phosphatidate cytidylyltransferase activity(GO:0004605) |
| 0.0 | 0.8 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.4 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.0 | 0.3 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.1 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.0 | 0.7 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 0.2 | GO:0031698 | beta-2 adrenergic receptor binding(GO:0031698) |
| 0.0 | 0.3 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 0.2 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 0.2 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.0 | 0.4 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 1.1 | GO:0017048 | Rho GTPase binding(GO:0017048) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.1 | 0.8 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.1 | 1.2 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 3.2 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 1.7 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.7 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.0 | 2.4 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 1.9 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 0.4 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 0.5 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |