Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
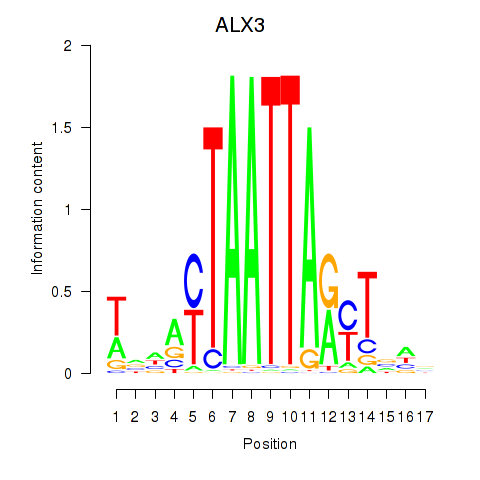
Results for ALX3
Z-value: 0.83
Transcription factors associated with ALX3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ALX3
|
ENSG00000156150.6 | ALX3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ALX3 | hg19_v2_chr1_-_110613276_110613322 | 0.14 | 6.1e-01 | Click! |
Activity profile of ALX3 motif
Sorted Z-values of ALX3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ALX3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_+_12059091 | 1.30 |
ENST00000562385.1 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr16_+_12059050 | 1.15 |
ENST00000396495.3 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr21_-_15918618 | 1.09 |
ENST00000400564.1 ENST00000400566.1 |
SAMSN1 |
SAM domain, SH3 domain and nuclear localization signals 1 |
| chr11_-_107729887 | 1.07 |
ENST00000525815.1 |
SLC35F2 |
solute carrier family 35, member F2 |
| chr3_-_141747950 | 0.98 |
ENST00000497579.1 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr7_+_138145076 | 0.84 |
ENST00000343526.4 |
TRIM24 |
tripartite motif containing 24 |
| chr7_-_87342564 | 0.83 |
ENST00000265724.3 ENST00000416177.1 |
ABCB1 |
ATP-binding cassette, sub-family B (MDR/TAP), member 1 |
| chr9_+_90112767 | 0.81 |
ENST00000408954.3 |
DAPK1 |
death-associated protein kinase 1 |
| chr15_-_20193370 | 0.79 |
ENST00000558565.2 |
IGHV3OR15-7 |
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr17_-_64225508 | 0.77 |
ENST00000205948.6 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chr9_+_90112590 | 0.73 |
ENST00000472284.1 |
DAPK1 |
death-associated protein kinase 1 |
| chr18_+_29171689 | 0.73 |
ENST00000237014.3 |
TTR |
transthyretin |
| chr3_-_141719195 | 0.71 |
ENST00000397991.4 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr5_-_94417339 | 0.69 |
ENST00000429576.2 ENST00000508509.1 ENST00000510732.1 |
MCTP1 |
multiple C2 domains, transmembrane 1 |
| chr14_-_106668095 | 0.68 |
ENST00000390606.2 |
IGHV3-20 |
immunoglobulin heavy variable 3-20 |
| chr9_+_90112741 | 0.67 |
ENST00000469640.2 |
DAPK1 |
death-associated protein kinase 1 |
| chr9_+_90112117 | 0.66 |
ENST00000358077.5 |
DAPK1 |
death-associated protein kinase 1 |
| chr1_-_24126023 | 0.64 |
ENST00000429356.1 |
GALE |
UDP-galactose-4-epimerase |
| chr16_+_15489603 | 0.61 |
ENST00000568766.1 ENST00000287594.7 |
RP11-1021N1.1 MPV17L |
Uncharacterized protein MPV17 mitochondrial membrane protein-like |
| chr18_-_67624160 | 0.61 |
ENST00000581982.1 ENST00000280200.4 |
CD226 |
CD226 molecule |
| chr4_+_80584903 | 0.60 |
ENST00000506460.1 |
RP11-452C8.1 |
RP11-452C8.1 |
| chr16_-_28634874 | 0.60 |
ENST00000395609.1 ENST00000350842.4 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr16_+_31885079 | 0.58 |
ENST00000300870.10 ENST00000394846.3 |
ZNF267 |
zinc finger protein 267 |
| chr11_+_7618413 | 0.54 |
ENST00000528883.1 |
PPFIBP2 |
PTPRF interacting protein, binding protein 2 (liprin beta 2) |
| chr1_+_62439037 | 0.50 |
ENST00000545929.1 |
INADL |
InaD-like (Drosophila) |
| chr11_-_62521614 | 0.50 |
ENST00000527994.1 ENST00000394807.3 |
ZBTB3 |
zinc finger and BTB domain containing 3 |
| chr8_-_102803163 | 0.50 |
ENST00000523645.1 ENST00000520346.1 ENST00000220931.6 ENST00000522448.1 ENST00000522951.1 ENST00000522252.1 ENST00000519098.1 |
NCALD |
neurocalcin delta |
| chr3_+_121774202 | 0.48 |
ENST00000469710.1 ENST00000493101.1 ENST00000330540.2 ENST00000264468.5 |
CD86 |
CD86 molecule |
| chr16_+_72088376 | 0.47 |
ENST00000570083.1 ENST00000355906.5 ENST00000398131.2 ENST00000569639.1 ENST00000564499.1 ENST00000357763.4 ENST00000562526.1 ENST00000565574.1 ENST00000568417.2 ENST00000356967.5 |
HP HPR |
haptoglobin haptoglobin-related protein |
| chr8_-_42623747 | 0.46 |
ENST00000534622.1 |
CHRNA6 |
cholinergic receptor, nicotinic, alpha 6 (neuronal) |
| chr18_+_29027696 | 0.46 |
ENST00000257189.4 |
DSG3 |
desmoglein 3 |
| chr4_+_113568207 | 0.45 |
ENST00000511529.1 |
LARP7 |
La ribonucleoprotein domain family, member 7 |
| chr14_-_107049312 | 0.44 |
ENST00000390627.2 |
IGHV3-53 |
immunoglobulin heavy variable 3-53 |
| chr1_-_150738261 | 0.43 |
ENST00000448301.2 ENST00000368985.3 |
CTSS |
cathepsin S |
| chr2_+_90077680 | 0.43 |
ENST00000390270.2 |
IGKV3D-20 |
immunoglobulin kappa variable 3D-20 |
| chr16_-_28608364 | 0.42 |
ENST00000533150.1 |
SULT1A2 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 2 |
| chr14_+_39736582 | 0.41 |
ENST00000556148.1 ENST00000348007.3 |
CTAGE5 |
CTAGE family, member 5 |
| chr4_-_25865159 | 0.41 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr19_-_36304201 | 0.40 |
ENST00000301175.3 |
PRODH2 |
proline dehydrogenase (oxidase) 2 |
| chr7_-_144435985 | 0.40 |
ENST00000549981.1 |
TPK1 |
thiamin pyrophosphokinase 1 |
| chr14_-_106552755 | 0.39 |
ENST00000390600.2 |
IGHV3-9 |
immunoglobulin heavy variable 3-9 |
| chr6_+_64345698 | 0.38 |
ENST00000506783.1 ENST00000481385.2 ENST00000515594.1 ENST00000494284.2 ENST00000262043.3 |
PHF3 |
PHD finger protein 3 |
| chrX_+_84258832 | 0.37 |
ENST00000373173.2 |
APOOL |
apolipoprotein O-like |
| chr12_-_22063787 | 0.37 |
ENST00000544039.1 |
ABCC9 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 9 |
| chr2_+_187371440 | 0.37 |
ENST00000445547.1 |
ZC3H15 |
zinc finger CCCH-type containing 15 |
| chrX_+_107288197 | 0.36 |
ENST00000415430.3 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr2_+_166095898 | 0.35 |
ENST00000424833.1 ENST00000375437.2 ENST00000357398.3 |
SCN2A |
sodium channel, voltage-gated, type II, alpha subunit |
| chr16_-_28621312 | 0.34 |
ENST00000314752.7 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chrX_-_71458802 | 0.34 |
ENST00000373657.1 ENST00000334463.3 |
ERCC6L |
excision repair cross-complementing rodent repair deficiency, complementation group 6-like |
| chr9_-_115095123 | 0.33 |
ENST00000458258.1 |
PTBP3 |
polypyrimidine tract binding protein 3 |
| chr2_+_102615416 | 0.33 |
ENST00000393414.2 |
IL1R2 |
interleukin 1 receptor, type II |
| chr5_-_98262240 | 0.33 |
ENST00000284049.3 |
CHD1 |
chromodomain helicase DNA binding protein 1 |
| chr15_+_58702742 | 0.32 |
ENST00000356113.6 ENST00000414170.3 |
LIPC |
lipase, hepatic |
| chr12_-_10282742 | 0.32 |
ENST00000298523.5 ENST00000396484.2 ENST00000310002.4 |
CLEC7A |
C-type lectin domain family 7, member A |
| chr16_-_28621298 | 0.32 |
ENST00000566189.1 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr1_+_12976450 | 0.32 |
ENST00000361079.2 |
PRAMEF7 |
PRAME family member 7 |
| chr3_-_151034734 | 0.31 |
ENST00000260843.4 |
GPR87 |
G protein-coupled receptor 87 |
| chr3_+_139063372 | 0.31 |
ENST00000478464.1 |
MRPS22 |
mitochondrial ribosomal protein S22 |
| chr3_-_164796269 | 0.30 |
ENST00000264382.3 |
SI |
sucrase-isomaltase (alpha-glucosidase) |
| chr1_-_92952433 | 0.30 |
ENST00000294702.5 |
GFI1 |
growth factor independent 1 transcription repressor |
| chrX_+_107288239 | 0.30 |
ENST00000217957.5 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr2_+_102953608 | 0.30 |
ENST00000311734.2 ENST00000409584.1 |
IL1RL1 |
interleukin 1 receptor-like 1 |
| chr10_+_35484793 | 0.30 |
ENST00000488741.1 ENST00000474931.1 ENST00000468236.1 ENST00000344351.5 ENST00000490511.1 |
CREM |
cAMP responsive element modulator |
| chr16_-_28608424 | 0.29 |
ENST00000335715.4 |
SULT1A2 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 2 |
| chrX_+_108779004 | 0.28 |
ENST00000218004.1 |
NXT2 |
nuclear transport factor 2-like export factor 2 |
| chr4_-_120243545 | 0.28 |
ENST00000274024.3 |
FABP2 |
fatty acid binding protein 2, intestinal |
| chr3_+_183853052 | 0.27 |
ENST00000273783.3 ENST00000432569.1 ENST00000444495.1 |
EIF2B5 |
eukaryotic translation initiation factor 2B, subunit 5 epsilon, 82kDa |
| chr11_-_71823715 | 0.27 |
ENST00000545944.1 ENST00000502597.2 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr10_-_105845674 | 0.27 |
ENST00000353479.5 ENST00000369733.3 |
COL17A1 |
collagen, type XVII, alpha 1 |
| chr16_-_55866997 | 0.27 |
ENST00000360526.3 ENST00000361503.4 |
CES1 |
carboxylesterase 1 |
| chr2_-_136678123 | 0.27 |
ENST00000422708.1 |
DARS |
aspartyl-tRNA synthetase |
| chr3_+_138340049 | 0.26 |
ENST00000464668.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr4_-_185275104 | 0.25 |
ENST00000317596.3 |
RP11-290F5.2 |
RP11-290F5.2 |
| chrX_-_77225135 | 0.25 |
ENST00000458128.1 |
PGAM4 |
phosphoglycerate mutase family member 4 |
| chr4_+_71859156 | 0.25 |
ENST00000286648.5 ENST00000504730.1 ENST00000504952.1 |
DCK |
deoxycytidine kinase |
| chr12_-_122985067 | 0.25 |
ENST00000540586.1 ENST00000543897.1 |
ZCCHC8 |
zinc finger, CCHC domain containing 8 |
| chr8_+_50824233 | 0.24 |
ENST00000522124.1 |
SNTG1 |
syntrophin, gamma 1 |
| chr2_+_68961934 | 0.24 |
ENST00000409202.3 |
ARHGAP25 |
Rho GTPase activating protein 25 |
| chr1_-_151431647 | 0.23 |
ENST00000368863.2 ENST00000409503.1 ENST00000491586.1 ENST00000533351.1 ENST00000540984.1 |
POGZ |
pogo transposable element with ZNF domain |
| chr7_-_44580861 | 0.23 |
ENST00000546276.1 ENST00000289547.4 ENST00000381160.3 ENST00000423141.1 |
NPC1L1 |
NPC1-like 1 |
| chr19_+_36139125 | 0.23 |
ENST00000246554.3 |
COX6B1 |
cytochrome c oxidase subunit VIb polypeptide 1 (ubiquitous) |
| chr8_-_91095099 | 0.23 |
ENST00000265431.3 |
CALB1 |
calbindin 1, 28kDa |
| chr10_-_28571015 | 0.23 |
ENST00000375719.3 ENST00000375732.1 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr11_-_71823796 | 0.23 |
ENST00000545680.1 ENST00000543587.1 ENST00000538393.1 ENST00000535234.1 ENST00000227618.4 ENST00000535503.1 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr16_-_28621353 | 0.22 |
ENST00000567512.1 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr2_-_89340242 | 0.22 |
ENST00000480492.1 |
IGKV1-12 |
immunoglobulin kappa variable 1-12 |
| chr7_+_150020329 | 0.22 |
ENST00000323078.7 |
LRRC61 |
leucine rich repeat containing 61 |
| chr14_-_54423529 | 0.22 |
ENST00000245451.4 ENST00000559087.1 |
BMP4 |
bone morphogenetic protein 4 |
| chr2_+_68961905 | 0.21 |
ENST00000295381.3 |
ARHGAP25 |
Rho GTPase activating protein 25 |
| chr1_-_190446759 | 0.21 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr11_-_128894053 | 0.21 |
ENST00000392657.3 |
ARHGAP32 |
Rho GTPase activating protein 32 |
| chr12_+_7014126 | 0.20 |
ENST00000415834.1 ENST00000436789.1 |
LRRC23 |
leucine rich repeat containing 23 |
| chr5_+_150639360 | 0.19 |
ENST00000523004.1 |
GM2A |
GM2 ganglioside activator |
| chr5_+_66300446 | 0.19 |
ENST00000261569.7 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr8_+_107738240 | 0.19 |
ENST00000449762.2 ENST00000297447.6 |
OXR1 |
oxidation resistance 1 |
| chr2_+_103089756 | 0.19 |
ENST00000295269.4 |
SLC9A4 |
solute carrier family 9, subfamily A (NHE4, cation proton antiporter 4), member 4 |
| chr6_+_34204642 | 0.19 |
ENST00000347617.6 ENST00000401473.3 ENST00000311487.5 ENST00000447654.1 ENST00000395004.3 |
HMGA1 |
high mobility group AT-hook 1 |
| chr1_+_186265399 | 0.18 |
ENST00000367486.3 ENST00000367484.3 ENST00000533951.1 ENST00000367482.4 ENST00000367483.4 ENST00000367485.4 ENST00000445192.2 |
PRG4 |
proteoglycan 4 |
| chr20_-_50722183 | 0.18 |
ENST00000371523.4 |
ZFP64 |
ZFP64 zinc finger protein |
| chr2_-_152118352 | 0.18 |
ENST00000331426.5 |
RBM43 |
RNA binding motif protein 43 |
| chr6_+_158733692 | 0.18 |
ENST00000367094.2 ENST00000367097.3 |
TULP4 |
tubby like protein 4 |
| chr7_+_150020363 | 0.17 |
ENST00000359623.4 ENST00000493307.1 |
LRRC61 |
leucine rich repeat containing 61 |
| chr6_+_26183958 | 0.17 |
ENST00000356530.3 |
HIST1H2BE |
histone cluster 1, H2be |
| chr6_+_26402517 | 0.17 |
ENST00000414912.2 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chr16_-_30122717 | 0.17 |
ENST00000566613.1 |
GDPD3 |
glycerophosphodiester phosphodiesterase domain containing 3 |
| chr14_+_32798547 | 0.17 |
ENST00000557354.1 ENST00000557102.1 ENST00000557272.1 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr12_-_112123524 | 0.16 |
ENST00000327551.6 |
BRAP |
BRCA1 associated protein |
| chr12_-_10978957 | 0.16 |
ENST00000240619.2 |
TAS2R10 |
taste receptor, type 2, member 10 |
| chr7_-_140482926 | 0.15 |
ENST00000496384.2 |
BRAF |
v-raf murine sarcoma viral oncogene homolog B |
| chrX_+_77154935 | 0.15 |
ENST00000481445.1 |
COX7B |
cytochrome c oxidase subunit VIIb |
| chr4_-_66536196 | 0.15 |
ENST00000511294.1 |
EPHA5 |
EPH receptor A5 |
| chr17_-_72772462 | 0.15 |
ENST00000582870.1 ENST00000581136.1 ENST00000357814.3 ENST00000579218.1 ENST00000583476.1 ENST00000580301.1 ENST00000583757.1 ENST00000582524.1 |
NAT9 |
N-acetyltransferase 9 (GCN5-related, putative) |
| chr12_+_122688090 | 0.15 |
ENST00000324189.4 ENST00000546192.1 |
B3GNT4 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 4 |
| chr12_-_10282836 | 0.15 |
ENST00000304084.8 ENST00000353231.5 ENST00000525605.1 |
CLEC7A |
C-type lectin domain family 7, member A |
| chr12_-_23737534 | 0.15 |
ENST00000396007.2 |
SOX5 |
SRY (sex determining region Y)-box 5 |
| chr9_+_117904097 | 0.14 |
ENST00000374016.1 |
DEC1 |
deleted in esophageal cancer 1 |
| chr17_-_3030875 | 0.14 |
ENST00000328890.2 |
OR1G1 |
olfactory receptor, family 1, subfamily G, member 1 |
| chr2_-_73520667 | 0.14 |
ENST00000545030.1 ENST00000436467.2 |
EGR4 |
early growth response 4 |
| chr11_-_71823266 | 0.14 |
ENST00000538919.1 ENST00000539395.1 ENST00000542531.1 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr17_-_39191107 | 0.13 |
ENST00000344363.5 |
KRTAP1-3 |
keratin associated protein 1-3 |
| chr1_+_115572415 | 0.13 |
ENST00000256592.1 |
TSHB |
thyroid stimulating hormone, beta |
| chr12_+_7014064 | 0.13 |
ENST00000443597.2 |
LRRC23 |
leucine rich repeat containing 23 |
| chr2_+_11682790 | 0.13 |
ENST00000389825.3 ENST00000381483.2 |
GREB1 |
growth regulation by estrogen in breast cancer 1 |
| chr5_+_140201183 | 0.13 |
ENST00000529619.1 ENST00000529859.1 ENST00000378126.3 |
PCDHA5 |
protocadherin alpha 5 |
| chr14_+_32798462 | 0.13 |
ENST00000280979.4 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr6_+_28317685 | 0.13 |
ENST00000252211.2 ENST00000341464.5 ENST00000377255.3 |
ZKSCAN3 |
zinc finger with KRAB and SCAN domains 3 |
| chrX_+_43515467 | 0.13 |
ENST00000338702.3 ENST00000542639.1 |
MAOA |
monoamine oxidase A |
| chr1_-_54411240 | 0.12 |
ENST00000371378.2 |
HSPB11 |
heat shock protein family B (small), member 11 |
| chr5_-_20575959 | 0.12 |
ENST00000507958.1 |
CDH18 |
cadherin 18, type 2 |
| chr4_+_88720698 | 0.12 |
ENST00000226284.5 |
IBSP |
integrin-binding sialoprotein |
| chr1_+_220267429 | 0.12 |
ENST00000366922.1 ENST00000302637.5 |
IARS2 |
isoleucyl-tRNA synthetase 2, mitochondrial |
| chr19_+_48949030 | 0.12 |
ENST00000253237.5 |
GRWD1 |
glutamate-rich WD repeat containing 1 |
| chr3_+_195447738 | 0.12 |
ENST00000447234.2 ENST00000320736.6 ENST00000436408.1 |
MUC20 |
mucin 20, cell surface associated |
| chr1_-_54411255 | 0.12 |
ENST00000371377.3 |
HSPB11 |
heat shock protein family B (small), member 11 |
| chr7_-_150020578 | 0.11 |
ENST00000478393.1 |
ACTR3C |
ARP3 actin-related protein 3 homolog C (yeast) |
| chr5_+_140557371 | 0.11 |
ENST00000239444.2 |
PCDHB8 |
protocadherin beta 8 |
| chr12_+_56435637 | 0.11 |
ENST00000356464.5 ENST00000552361.1 |
RPS26 |
ribosomal protein S26 |
| chr6_-_7313381 | 0.11 |
ENST00000489567.1 ENST00000479365.1 ENST00000462112.1 ENST00000397511.2 ENST00000534851.1 ENST00000474597.1 ENST00000244763.4 |
SSR1 |
signal sequence receptor, alpha |
| chr5_+_81575281 | 0.11 |
ENST00000380167.4 |
ATP6AP1L |
ATPase, H+ transporting, lysosomal accessory protein 1-like |
| chr2_-_3521518 | 0.11 |
ENST00000382093.5 |
ADI1 |
acireductone dioxygenase 1 |
| chr1_-_207226313 | 0.10 |
ENST00000367084.1 |
YOD1 |
YOD1 deubiquitinase |
| chr12_+_7013897 | 0.09 |
ENST00000007969.8 ENST00000323702.5 |
LRRC23 |
leucine rich repeat containing 23 |
| chr11_+_71900703 | 0.09 |
ENST00000393681.2 |
FOLR1 |
folate receptor 1 (adult) |
| chr2_+_90211643 | 0.09 |
ENST00000390277.2 |
IGKV3D-11 |
immunoglobulin kappa variable 3D-11 |
| chr4_-_66536057 | 0.09 |
ENST00000273854.3 |
EPHA5 |
EPH receptor A5 |
| chr6_+_154360357 | 0.09 |
ENST00000330432.7 ENST00000360422.4 |
OPRM1 |
opioid receptor, mu 1 |
| chr3_+_119298523 | 0.09 |
ENST00000357003.3 |
ADPRH |
ADP-ribosylarginine hydrolase |
| chr14_-_78083112 | 0.08 |
ENST00000216484.2 |
SPTLC2 |
serine palmitoyltransferase, long chain base subunit 2 |
| chr12_-_86650045 | 0.08 |
ENST00000604798.1 |
MGAT4C |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme C (putative) |
| chr1_+_28261492 | 0.08 |
ENST00000373894.3 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr2_-_89385283 | 0.08 |
ENST00000390252.2 |
IGKV3-15 |
immunoglobulin kappa variable 3-15 |
| chr9_+_105757590 | 0.08 |
ENST00000374798.3 ENST00000487798.1 |
CYLC2 |
cylicin, basic protein of sperm head cytoskeleton 2 |
| chr12_-_86650077 | 0.08 |
ENST00000552808.2 ENST00000547225.1 |
MGAT4C |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme C (putative) |
| chr3_+_45927994 | 0.08 |
ENST00000357632.2 ENST00000395963.2 |
CCR9 |
chemokine (C-C motif) receptor 9 |
| chr2_-_169887827 | 0.08 |
ENST00000263817.6 |
ABCB11 |
ATP-binding cassette, sub-family B (MDR/TAP), member 11 |
| chr1_+_28261533 | 0.08 |
ENST00000411604.1 ENST00000373888.4 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr6_+_160542821 | 0.08 |
ENST00000366963.4 |
SLC22A1 |
solute carrier family 22 (organic cation transporter), member 1 |
| chr15_+_75970672 | 0.08 |
ENST00000435356.1 |
AC105020.1 |
Uncharacterized protein; cDNA FLJ12988 fis, clone NT2RP3000080 |
| chr20_+_1099233 | 0.08 |
ENST00000246015.4 ENST00000335877.6 ENST00000438768.2 |
PSMF1 |
proteasome (prosome, macropain) inhibitor subunit 1 (PI31) |
| chr2_-_220264703 | 0.07 |
ENST00000519905.1 ENST00000523282.1 ENST00000434339.1 ENST00000457935.1 |
DNPEP |
aspartyl aminopeptidase |
| chr4_+_142142035 | 0.07 |
ENST00000262990.4 ENST00000512809.1 ENST00000503649.1 ENST00000512738.1 ENST00000421169.2 |
ZNF330 |
zinc finger protein 330 |
| chr4_+_5527117 | 0.07 |
ENST00000505296.1 |
C4orf6 |
chromosome 4 open reading frame 6 |
| chrX_+_153533275 | 0.07 |
ENST00000426989.1 ENST00000426203.1 ENST00000369912.2 |
TKTL1 |
transketolase-like 1 |
| chr2_+_103035102 | 0.07 |
ENST00000264260.2 |
IL18RAP |
interleukin 18 receptor accessory protein |
| chr17_-_2996290 | 0.06 |
ENST00000331459.1 |
OR1D2 |
olfactory receptor, family 1, subfamily D, member 2 |
| chr20_-_56265680 | 0.06 |
ENST00000414037.1 |
PMEPA1 |
prostate transmembrane protein, androgen induced 1 |
| chr6_+_161123270 | 0.06 |
ENST00000366924.2 ENST00000308192.9 ENST00000418964.1 |
PLG |
plasminogen |
| chr6_+_26402465 | 0.06 |
ENST00000476549.2 ENST00000289361.6 ENST00000450085.2 ENST00000425234.2 ENST00000427334.1 ENST00000506698.1 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chr6_+_160542870 | 0.06 |
ENST00000324965.4 ENST00000457470.2 |
SLC22A1 |
solute carrier family 22 (organic cation transporter), member 1 |
| chr9_+_111624577 | 0.06 |
ENST00000333999.3 |
ACTL7A |
actin-like 7A |
| chr4_+_110736659 | 0.05 |
ENST00000394631.3 ENST00000226796.6 |
GAR1 |
GAR1 ribonucleoprotein |
| chr2_+_234826016 | 0.05 |
ENST00000324695.4 ENST00000433712.2 |
TRPM8 |
transient receptor potential cation channel, subfamily M, member 8 |
| chr11_+_71900572 | 0.05 |
ENST00000312293.4 |
FOLR1 |
folate receptor 1 (adult) |
| chr19_-_14064114 | 0.05 |
ENST00000585607.1 ENST00000538517.2 ENST00000587458.1 ENST00000538371.2 |
PODNL1 |
podocan-like 1 |
| chr7_+_156433573 | 0.05 |
ENST00000311822.8 |
RNF32 |
ring finger protein 32 |
| chr11_-_13011081 | 0.04 |
ENST00000532541.1 ENST00000526388.1 ENST00000534477.1 ENST00000531402.1 ENST00000527945.1 ENST00000504230.2 |
LINC00958 |
long intergenic non-protein coding RNA 958 |
| chr14_-_67878917 | 0.04 |
ENST00000216446.4 |
PLEK2 |
pleckstrin 2 |
| chr19_+_42817527 | 0.04 |
ENST00000598766.1 |
TMEM145 |
transmembrane protein 145 |
| chr4_+_5526883 | 0.04 |
ENST00000195455.2 |
C4orf6 |
chromosome 4 open reading frame 6 |
| chr2_-_228497888 | 0.04 |
ENST00000264387.4 ENST00000409066.1 |
C2orf83 |
chromosome 2 open reading frame 83 |
| chr19_+_47421933 | 0.04 |
ENST00000404338.3 |
ARHGAP35 |
Rho GTPase activating protein 35 |
| chr11_-_89541743 | 0.03 |
ENST00000329758.1 |
TRIM49 |
tripartite motif containing 49 |
| chr12_+_104337515 | 0.03 |
ENST00000550595.1 |
HSP90B1 |
heat shock protein 90kDa beta (Grp94), member 1 |
| chr6_+_44355257 | 0.03 |
ENST00000371477.3 |
CDC5L |
cell division cycle 5-like |
| chr1_+_225600404 | 0.03 |
ENST00000366845.2 |
AC092811.1 |
AC092811.1 |
| chr9_-_5339873 | 0.03 |
ENST00000223862.1 ENST00000223858.4 |
RLN1 |
relaxin 1 |
| chr11_+_92702886 | 0.03 |
ENST00000257068.2 ENST00000528076.1 |
MTNR1B |
melatonin receptor 1B |
| chr9_-_99540328 | 0.03 |
ENST00000223428.4 ENST00000375231.1 ENST00000374641.3 |
ZNF510 |
zinc finger protein 510 |
| chr15_+_58724184 | 0.02 |
ENST00000433326.2 |
LIPC |
lipase, hepatic |
| chr10_-_75415825 | 0.02 |
ENST00000394810.2 |
SYNPO2L |
synaptopodin 2-like |
| chr20_-_33735070 | 0.02 |
ENST00000374491.3 ENST00000542871.1 ENST00000374492.3 |
EDEM2 |
ER degradation enhancer, mannosidase alpha-like 2 |
| chr3_+_138340067 | 0.02 |
ENST00000479848.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr1_+_28261621 | 0.02 |
ENST00000549094.1 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr1_-_24306798 | 0.02 |
ENST00000374452.5 ENST00000492112.2 ENST00000343255.5 ENST00000344989.6 |
SRSF10 |
serine/arginine-rich splicing factor 10 |
| chr10_+_18549645 | 0.01 |
ENST00000396576.2 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr17_+_40610862 | 0.01 |
ENST00000393829.2 ENST00000546249.1 ENST00000537728.1 ENST00000264649.6 ENST00000585525.1 ENST00000343619.4 ENST00000544137.1 ENST00000589727.1 ENST00000587824.1 |
ATP6V0A1 |
ATPase, H+ transporting, lysosomal V0 subunit a1 |
| chr2_+_90273679 | 0.01 |
ENST00000423080.2 |
IGKV3D-7 |
immunoglobulin kappa variable 3D-7 |
| chr3_-_126327398 | 0.01 |
ENST00000383572.2 |
TXNRD3NB |
thioredoxin reductase 3 neighbor |
| chr1_-_150669500 | 0.01 |
ENST00000271732.3 |
GOLPH3L |
golgi phosphoprotein 3-like |
| chr12_+_18414446 | 0.00 |
ENST00000433979.1 |
PIK3C2G |
phosphatidylinositol-4-phosphate 3-kinase, catalytic subunit type 2 gamma |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0002370 | natural killer cell cytokine production(GO:0002370) regulation of natural killer cell cytokine production(GO:0002727) positive regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060369) |
| 0.2 | 3.1 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.2 | 0.5 | GO:0042109 | negative regulation of tolerance induction(GO:0002644) lymphotoxin A production(GO:0032641) interleukin-4 biosynthetic process(GO:0042097) lymphotoxin A biosynthetic process(GO:0042109) regulation of interleukin-4 biosynthetic process(GO:0045402) positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 0.2 | 0.5 | GO:2000296 | negative regulation of hydrogen peroxide catabolic process(GO:2000296) |
| 0.2 | 0.6 | GO:0010730 | negative regulation of hydrogen peroxide biosynthetic process(GO:0010730) |
| 0.1 | 0.8 | GO:0034196 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 0.1 | 0.8 | GO:0070562 | regulation of vitamin D receptor signaling pathway(GO:0070562) |
| 0.1 | 0.8 | GO:1901529 | positive regulation of anion channel activity(GO:1901529) |
| 0.1 | 0.3 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.1 | 0.4 | GO:0042357 | thiamine diphosphate metabolic process(GO:0042357) |
| 0.1 | 0.3 | GO:0006422 | aspartyl-tRNA aminoacylation(GO:0006422) |
| 0.1 | 2.2 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.1 | 0.3 | GO:0050712 | negative regulation of interleukin-1 alpha production(GO:0032690) negative regulation of interleukin-1 alpha secretion(GO:0050712) |
| 0.1 | 0.7 | GO:0019722 | calcium-mediated signaling(GO:0019722) |
| 0.1 | 0.3 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 0.1 | 0.2 | GO:0055020 | apoptotic process involved in endocardial cushion morphogenesis(GO:0003277) intermediate mesoderm development(GO:0048389) intermediate mesoderm morphogenesis(GO:0048390) intermediate mesoderm formation(GO:0048391) intermediate mesodermal cell differentiation(GO:0048392) regulation of cardiac muscle fiber development(GO:0055018) positive regulation of cardiac muscle fiber development(GO:0055020) bud dilation involved in lung branching(GO:0060503) regulation of transcription from RNA polymerase II promoter involved in kidney development(GO:0060994) BMP signaling pathway involved in ureter morphogenesis(GO:0061149) renal system segmentation(GO:0061150) BMP signaling pathway involved in renal system segmentation(GO:0061151) pulmonary artery endothelial tube morphogenesis(GO:0061155) regulation of transcription from RNA polymerase II promoter involved in mesonephros development(GO:0061216) pattern specification involved in mesonephros development(GO:0061227) BMP signaling pathway involved in nephric duct formation(GO:0071893) negative regulation of branch elongation involved in ureteric bud branching(GO:0072096) negative regulation of branch elongation involved in ureteric bud branching by BMP signaling pathway(GO:0072097) anterior/posterior pattern specification involved in kidney development(GO:0072098) anterior/posterior pattern specification involved in ureteric bud development(GO:0072099) specification of ureteric bud anterior/posterior symmetry(GO:0072100) specification of ureteric bud anterior/posterior symmetry by BMP signaling pathway(GO:0072101) ureter epithelial cell differentiation(GO:0072192) negative regulation of mesenchymal cell proliferation involved in ureter development(GO:0072200) positive regulation of cell proliferation involved in outflow tract morphogenesis(GO:1901964) cardiac jelly development(GO:1905072) regulation of metanephric S-shaped body morphogenesis(GO:2000004) negative regulation of metanephric S-shaped body morphogenesis(GO:2000005) regulation of metanephric comma-shaped body morphogenesis(GO:2000006) negative regulation of metanephric comma-shaped body morphogenesis(GO:2000007) |
| 0.1 | 0.4 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.1 | 0.3 | GO:0043456 | regulation of pentose-phosphate shunt(GO:0043456) |
| 0.1 | 0.7 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 0.3 | GO:0038172 | interleukin-33-mediated signaling pathway(GO:0038172) |
| 0.0 | 0.1 | GO:0030309 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.0 | 0.6 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 0.0 | 0.1 | GO:0060974 | neural crest cell migration involved in heart formation(GO:0003147) cell migration involved in heart formation(GO:0060974) anterior neural tube closure(GO:0061713) |
| 0.0 | 0.2 | GO:0090402 | oncogene-induced cell senescence(GO:0090402) |
| 0.0 | 0.3 | GO:0044245 | polysaccharide digestion(GO:0044245) |
| 0.0 | 0.1 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.0 | 1.7 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.0 | 0.6 | GO:1903504 | regulation of mitotic cell cycle spindle assembly checkpoint(GO:0090266) regulation of mitotic spindle checkpoint(GO:1903504) |
| 0.0 | 0.4 | GO:1905097 | regulation of guanyl-nucleotide exchange factor activity(GO:1905097) |
| 0.0 | 1.1 | GO:0050869 | negative regulation of B cell activation(GO:0050869) |
| 0.0 | 0.3 | GO:0032057 | negative regulation of translational initiation in response to stress(GO:0032057) |
| 0.0 | 0.7 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.0 | 0.2 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.1 | GO:1990168 | protein K33-linked deubiquitination(GO:1990168) |
| 0.0 | 0.2 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.0 | 1.9 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 2.3 | GO:0002260 | lymphocyte homeostasis(GO:0002260) |
| 0.0 | 0.3 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.3 | GO:0008627 | intrinsic apoptotic signaling pathway in response to osmotic stress(GO:0008627) |
| 0.0 | 0.1 | GO:0051725 | protein de-ADP-ribosylation(GO:0051725) |
| 0.0 | 0.2 | GO:0072205 | metanephric part of ureteric bud development(GO:0035502) metanephric collecting duct development(GO:0072205) |
| 0.0 | 0.3 | GO:0051791 | medium-chain fatty acid metabolic process(GO:0051791) |
| 0.0 | 0.3 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) phosphatidylcholine catabolic process(GO:0034638) |
| 0.0 | 0.2 | GO:0030299 | intestinal cholesterol absorption(GO:0030299) intestinal lipid absorption(GO:0098856) |
| 0.0 | 0.1 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.0 | 0.2 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.0 | 0.1 | GO:0006772 | thiamine metabolic process(GO:0006772) |
| 0.0 | 0.1 | GO:1904504 | regulation of lipophagy(GO:1904502) positive regulation of lipophagy(GO:1904504) |
| 0.0 | 0.1 | GO:2000048 | interaction with other organism via secreted substance involved in symbiotic interaction(GO:0052047) negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 0.0 | 0.2 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 0.1 | GO:0048241 | epinephrine transport(GO:0048241) |
| 0.0 | 0.1 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.0 | 0.2 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.0 | 0.2 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.0 | 0.3 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.0 | 0.4 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.4 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.0 | 0.3 | GO:0043923 | positive regulation by host of viral transcription(GO:0043923) |
| 0.0 | 0.1 | GO:2000773 | negative regulation of cellular senescence(GO:2000773) |
| 0.0 | 0.1 | GO:0061358 | negative regulation of Wnt protein secretion(GO:0061358) |
| 0.0 | 1.6 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.0 | 0.2 | GO:0032793 | positive regulation of CREB transcription factor activity(GO:0032793) |
| 0.0 | 0.1 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.0 | 0.2 | GO:1990118 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 0.0 | GO:0015888 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 0.0 | 0.1 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) |
| 0.0 | 0.5 | GO:0014046 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.0 | 0.3 | GO:0043101 | purine-containing compound salvage(GO:0043101) |
| 0.0 | 0.0 | GO:1990637 | response to prolactin(GO:1990637) |
| 0.0 | 0.1 | GO:0050955 | thermoception(GO:0050955) |
| 0.0 | 0.2 | GO:0072643 | interferon-gamma secretion(GO:0072643) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.1 | 0.4 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.1 | 0.5 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.1 | 0.4 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.1 | 0.4 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 0.2 | GO:0005889 | hydrogen:potassium-exchanging ATPase complex(GO:0005889) |
| 0.0 | 0.8 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.0 | 0.3 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.0 | 0.3 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.0 | 0.5 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.0 | 1.5 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.1 | GO:0033011 | perinuclear theca(GO:0033011) cytoskeletal calyx(GO:0033150) |
| 0.0 | 0.2 | GO:0016013 | syntrophin complex(GO:0016013) |
| 0.0 | 0.1 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.0 | 0.6 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 0.3 | GO:0033270 | voltage-gated sodium channel complex(GO:0001518) paranode region of axon(GO:0033270) |
| 0.0 | 0.5 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 0.2 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.0 | 0.2 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.0 | 0.4 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.0 | 0.5 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 0.3 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.3 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.0 | 0.2 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.0 | 0.3 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.1 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.3 | GO:0097228 | sperm principal piece(GO:0097228) |
| 0.0 | 0.1 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.2 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) |
| 0.2 | 0.6 | GO:0003974 | UDP-N-acetylglucosamine 4-epimerase activity(GO:0003974) UDP-glucose 4-epimerase activity(GO:0003978) |
| 0.2 | 0.8 | GO:0090554 | phosphatidylcholine-translocating ATPase activity(GO:0090554) |
| 0.2 | 0.8 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.1 | 0.3 | GO:0004574 | oligo-1,6-glucosidase activity(GO:0004574) |
| 0.1 | 0.3 | GO:0002113 | interleukin-33 binding(GO:0002113) |
| 0.1 | 0.4 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 0.1 | 2.9 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 0.5 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.1 | 0.8 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.1 | 0.7 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 0.3 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 0.1 | 0.3 | GO:0004619 | bisphosphoglycerate mutase activity(GO:0004082) phosphoglycerate mutase activity(GO:0004619) 2,3-bisphosphoglycerate-dependent phosphoglycerate mutase activity(GO:0046538) |
| 0.1 | 0.3 | GO:0004771 | sterol esterase activity(GO:0004771) methylumbelliferyl-acetate deacetylase activity(GO:0047374) |
| 0.1 | 0.3 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.0 | 0.4 | GO:0016778 | diphosphotransferase activity(GO:0016778) |
| 0.0 | 0.1 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.0 | 0.1 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 0.0 | 0.2 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.0 | 0.1 | GO:0061714 | folic acid receptor activity(GO:0061714) |
| 0.0 | 0.1 | GO:0005277 | acetylcholine transmembrane transporter activity(GO:0005277) secondary active organic cation transmembrane transporter activity(GO:0008513) acetate ester transmembrane transporter activity(GO:1901375) |
| 0.0 | 0.1 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 0.0 | 0.4 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.0 | 0.1 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.0 | 1.5 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.2 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 0.2 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.2 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.1 | GO:0004802 | transketolase activity(GO:0004802) |
| 0.0 | 0.4 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.0 | 0.3 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.0 | 0.5 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.0 | 0.2 | GO:0015386 | sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.5 | GO:0008329 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.0 | 0.3 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.0 | 1.1 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.2 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.3 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 0.4 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.0 | 0.2 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.0 | 0.1 | GO:0038047 | beta-endorphin receptor activity(GO:0004979) morphine receptor activity(GO:0038047) |
| 0.0 | 0.1 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 0.5 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 0.3 | GO:1905030 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.0 | 0.2 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.2 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 0.7 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 0.4 | GO:0004129 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.0 | GO:0015234 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) |
| 0.0 | 0.2 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 0.5 | GO:0015026 | coreceptor activity(GO:0015026) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.9 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 2.2 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 1.0 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.0 | 0.4 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.0 | 0.3 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 0.5 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.5 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.5 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.0 | 0.8 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 0.2 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.0 | 0.5 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.3 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.5 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 0.4 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.3 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.9 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.3 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.9 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 1.9 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 0.5 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 0.7 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.2 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 0.8 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |