Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
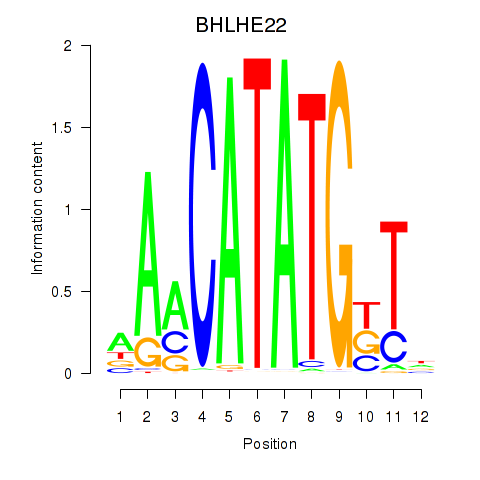
Results for BHLHE22_BHLHA15_BHLHE23
Z-value: 0.67
Transcription factors associated with BHLHE22_BHLHA15_BHLHE23
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
BHLHE22
|
ENSG00000180828.1 | BHLHE22 |
|
BHLHA15
|
ENSG00000180535.3 | BHLHA15 |
|
BHLHE23
|
ENSG00000125533.4 | BHLHE23 |
Activity profile of BHLHE22_BHLHA15_BHLHE23 motif
Sorted Z-values of BHLHE22_BHLHA15_BHLHE23 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of BHLHE22_BHLHA15_BHLHE23
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_111312622 | 2.92 |
ENST00000395634.3 |
NREP |
neuronal regeneration related protein |
| chr3_-_121379739 | 2.65 |
ENST00000428394.2 ENST00000314583.3 |
HCLS1 |
hematopoietic cell-specific Lyn substrate 1 |
| chr3_-_114477962 | 1.94 |
ENST00000471418.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr3_-_114477787 | 1.88 |
ENST00000464560.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr2_-_188378368 | 1.65 |
ENST00000392365.1 ENST00000435414.1 |
TFPI |
tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) |
| chr7_-_27205136 | 1.25 |
ENST00000396345.1 ENST00000343483.6 |
HOXA9 |
homeobox A9 |
| chr1_+_244214577 | 0.71 |
ENST00000358704.4 |
ZBTB18 |
zinc finger and BTB domain containing 18 |
| chr6_-_112115074 | 0.70 |
ENST00000368667.2 |
FYN |
FYN oncogene related to SRC, FGR, YES |
| chr11_+_65405556 | 0.63 |
ENST00000534313.1 ENST00000533361.1 ENST00000526137.1 |
SIPA1 |
signal-induced proliferation-associated 1 |
| chr18_-_52989525 | 0.60 |
ENST00000457482.3 |
TCF4 |
transcription factor 4 |
| chrX_+_128913906 | 0.59 |
ENST00000356892.3 |
SASH3 |
SAM and SH3 domain containing 3 |
| chr4_-_159094194 | 0.59 |
ENST00000592057.1 ENST00000585682.1 ENST00000393807.5 |
FAM198B |
family with sequence similarity 198, member B |
| chr7_+_116502605 | 0.54 |
ENST00000458284.2 ENST00000490693.1 |
CAPZA2 |
capping protein (actin filament) muscle Z-line, alpha 2 |
| chr15_-_55700522 | 0.48 |
ENST00000564092.1 ENST00000310958.6 |
CCPG1 |
cell cycle progression 1 |
| chr16_+_31119615 | 0.45 |
ENST00000394950.3 ENST00000287507.3 ENST00000219794.6 ENST00000561755.1 |
BCKDK |
branched chain ketoacid dehydrogenase kinase |
| chr4_-_41216619 | 0.40 |
ENST00000508676.1 ENST00000506352.1 ENST00000295974.8 |
APBB2 |
amyloid beta (A4) precursor protein-binding, family B, member 2 |
| chr15_-_55700216 | 0.37 |
ENST00000569205.1 |
CCPG1 |
cell cycle progression 1 |
| chr10_-_75415825 | 0.27 |
ENST00000394810.2 |
SYNPO2L |
synaptopodin 2-like |
| chr3_+_62304648 | 0.27 |
ENST00000462069.1 ENST00000232519.5 ENST00000465142.1 |
C3orf14 |
chromosome 3 open reading frame 14 |
| chr15_-_55700457 | 0.27 |
ENST00000442196.3 ENST00000563171.1 ENST00000425574.3 |
CCPG1 |
cell cycle progression 1 |
| chr7_+_116502527 | 0.26 |
ENST00000361183.3 |
CAPZA2 |
capping protein (actin filament) muscle Z-line, alpha 2 |
| chr15_-_63448973 | 0.26 |
ENST00000462430.1 |
RPS27L |
ribosomal protein S27-like |
| chr12_-_91573249 | 0.25 |
ENST00000550099.1 ENST00000546391.1 ENST00000551354.1 |
DCN |
decorin |
| chr15_+_63354769 | 0.25 |
ENST00000558910.1 |
TPM1 |
tropomyosin 1 (alpha) |
| chr12_-_91573316 | 0.24 |
ENST00000393155.1 |
DCN |
decorin |
| chr12_-_91573132 | 0.24 |
ENST00000550563.1 ENST00000546370.1 |
DCN |
decorin |
| chrX_+_107334895 | 0.24 |
ENST00000372232.3 ENST00000345734.3 ENST00000372254.3 |
ATG4A |
autophagy related 4A, cysteine peptidase |
| chr14_-_69864993 | 0.22 |
ENST00000555373.1 |
ERH |
enhancer of rudimentary homolog (Drosophila) |
| chr17_+_60758814 | 0.20 |
ENST00000579432.1 ENST00000446119.2 |
MRC2 |
mannose receptor, C type 2 |
| chr11_-_107582775 | 0.17 |
ENST00000305991.2 |
SLN |
sarcolipin |
| chr8_-_23315190 | 0.17 |
ENST00000356206.6 ENST00000358689.4 ENST00000417069.2 |
ENTPD4 |
ectonucleoside triphosphate diphosphohydrolase 4 |
| chr9_-_130635741 | 0.17 |
ENST00000223836.10 |
AK1 |
adenylate kinase 1 |
| chr9_+_2158443 | 0.15 |
ENST00000302401.3 ENST00000324954.5 ENST00000423555.1 ENST00000382186.1 |
SMARCA2 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr9_+_2158485 | 0.15 |
ENST00000417599.1 ENST00000382185.1 ENST00000382183.1 |
SMARCA2 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr3_+_113251143 | 0.14 |
ENST00000264852.4 ENST00000393830.3 |
SIDT1 |
SID1 transmembrane family, member 1 |
| chr12_-_108714412 | 0.14 |
ENST00000412676.1 ENST00000550573.1 |
CMKLR1 |
chemokine-like receptor 1 |
| chr2_-_163099885 | 0.13 |
ENST00000443424.1 |
FAP |
fibroblast activation protein, alpha |
| chr7_-_55930443 | 0.12 |
ENST00000388975.3 |
SEPT14 |
septin 14 |
| chr11_-_26743546 | 0.12 |
ENST00000280467.6 ENST00000396005.3 |
SLC5A12 |
solute carrier family 5 (sodium/monocarboxylate cotransporter), member 12 |
| chr4_+_2814011 | 0.12 |
ENST00000502260.1 ENST00000435136.2 |
SH3BP2 |
SH3-domain binding protein 2 |
| chr3_+_171758344 | 0.12 |
ENST00000336824.4 ENST00000423424.1 |
FNDC3B |
fibronectin type III domain containing 3B |
| chr7_-_38403077 | 0.11 |
ENST00000426402.2 |
TRGV2 |
T cell receptor gamma variable 2 |
| chr10_+_18629628 | 0.10 |
ENST00000377329.4 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr10_-_94257512 | 0.10 |
ENST00000371581.5 |
IDE |
insulin-degrading enzyme |
| chr4_+_2813946 | 0.10 |
ENST00000442312.2 |
SH3BP2 |
SH3-domain binding protein 2 |
| chr19_-_11450249 | 0.09 |
ENST00000222120.3 |
RAB3D |
RAB3D, member RAS oncogene family |
| chr12_-_102872317 | 0.09 |
ENST00000424202.2 |
IGF1 |
insulin-like growth factor 1 (somatomedin C) |
| chr11_-_44972476 | 0.08 |
ENST00000527685.1 ENST00000308212.5 |
TP53I11 |
tumor protein p53 inducible protein 11 |
| chr6_+_163148161 | 0.08 |
ENST00000337019.3 ENST00000366889.2 |
PACRG |
PARK2 co-regulated |
| chr4_+_144354644 | 0.08 |
ENST00000512843.1 |
GAB1 |
GRB2-associated binding protein 1 |
| chr14_+_69865401 | 0.08 |
ENST00000556605.1 ENST00000336643.5 ENST00000031146.4 |
SLC39A9 |
solute carrier family 39, member 9 |
| chrX_-_13835147 | 0.08 |
ENST00000493677.1 ENST00000355135.2 |
GPM6B |
glycoprotein M6B |
| chr2_+_219110149 | 0.08 |
ENST00000456575.1 |
ARPC2 |
actin related protein 2/3 complex, subunit 2, 34kDa |
| chr11_+_66008698 | 0.08 |
ENST00000531597.1 |
PACS1 |
phosphofurin acidic cluster sorting protein 1 |
| chr15_-_65503801 | 0.07 |
ENST00000261883.4 |
CILP |
cartilage intermediate layer protein, nucleotide pyrophosphohydrolase |
| chr1_+_104293028 | 0.06 |
ENST00000370079.3 |
AMY1C |
amylase, alpha 1C (salivary) |
| chr2_-_163100045 | 0.06 |
ENST00000188790.4 |
FAP |
fibroblast activation protein, alpha |
| chrX_+_107288239 | 0.05 |
ENST00000217957.5 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr19_+_57874835 | 0.05 |
ENST00000543226.1 ENST00000596755.1 ENST00000282282.3 ENST00000597658.1 |
TRAPPC2P1 ZNF547 AC003002.4 |
trafficking protein particle complex 2 pseudogene 1 zinc finger protein 547 Uncharacterized protein |
| chr6_+_73076432 | 0.04 |
ENST00000414192.2 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr12_+_58166370 | 0.04 |
ENST00000300209.8 |
METTL21B |
methyltransferase like 21B |
| chr8_-_67976509 | 0.03 |
ENST00000518747.1 |
COPS5 |
COP9 signalosome subunit 5 |
| chr7_+_107531580 | 0.03 |
ENST00000537148.1 ENST00000440410.1 ENST00000437604.2 |
DLD |
dihydrolipoamide dehydrogenase |
| chrX_+_107288197 | 0.03 |
ENST00000415430.3 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr10_-_74114714 | 0.03 |
ENST00000338820.3 ENST00000394903.2 ENST00000444643.2 |
DNAJB12 |
DnaJ (Hsp40) homolog, subfamily B, member 12 |
| chr5_-_171433819 | 0.02 |
ENST00000296933.6 |
FBXW11 |
F-box and WD repeat domain containing 11 |
| chrX_-_15402498 | 0.02 |
ENST00000297904.3 |
FIGF |
c-fos induced growth factor (vascular endothelial growth factor D) |
| chr2_+_210444748 | 0.02 |
ENST00000392194.1 |
MAP2 |
microtubule-associated protein 2 |
| chr1_-_44818599 | 0.02 |
ENST00000537474.1 |
ERI3 |
ERI1 exoribonuclease family member 3 |
| chr5_-_171433579 | 0.02 |
ENST00000265094.5 ENST00000393802.2 |
FBXW11 |
F-box and WD repeat domain containing 11 |
| chr11_-_44972418 | 0.02 |
ENST00000525680.1 ENST00000528290.1 ENST00000530035.1 |
TP53I11 |
tumor protein p53 inducible protein 11 |
| chr22_+_36113919 | 0.02 |
ENST00000249044.2 |
APOL5 |
apolipoprotein L, 5 |
| chr19_-_52227221 | 0.02 |
ENST00000222115.1 ENST00000540069.2 |
HAS1 |
hyaluronan synthase 1 |
| chr12_+_58166431 | 0.02 |
ENST00000333012.5 |
METTL21B |
methyltransferase like 21B |
| chrX_+_69509927 | 0.02 |
ENST00000374403.3 |
KIF4A |
kinesin family member 4A |
| chr1_+_186344883 | 0.01 |
ENST00000367470.3 |
C1orf27 |
chromosome 1 open reading frame 27 |
| chr16_-_4896205 | 0.01 |
ENST00000589389.1 |
GLYR1 |
glyoxylate reductase 1 homolog (Arabidopsis) |
| chr3_+_119298429 | 0.01 |
ENST00000478927.1 |
ADPRH |
ADP-ribosylarginine hydrolase |
| chr5_-_169725231 | 0.01 |
ENST00000046794.5 |
LCP2 |
lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa) |
| chr14_-_107283278 | 0.00 |
ENST00000390639.2 |
IGHV7-81 |
immunoglobulin heavy variable 7-81 (non-functional) |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 0.0 | 0.6 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.2 | GO:0045134 | uridine-diphosphatase activity(GO:0045134) |
| 0.0 | 0.1 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.0 | 3.8 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 2.9 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 0.3 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.0 | 1.7 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.2 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.0 | 0.1 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.1 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.0 | 0.1 | GO:0043559 | insulin binding(GO:0043559) |
| 0.0 | 0.1 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.0 | 0.7 | GO:0050840 | extracellular matrix binding(GO:0050840) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.7 | REACTOME FORMATION OF FIBRIN CLOT CLOTTING CASCADE | Genes involved in Formation of Fibrin Clot (Clotting Cascade) |
| 0.0 | 0.7 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.0 | 0.7 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.8 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.0 | 0.6 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.0 | 0.6 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.7 | PID SYNDECAN 4 PATHWAY | Syndecan-4-mediated signaling events |
| 0.0 | 3.3 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.0 | 0.7 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.1 | GO:2001106 | regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.2 | 1.7 | GO:0007598 | blood coagulation, extrinsic pathway(GO:0007598) |
| 0.2 | 2.6 | GO:0030854 | positive regulation of granulocyte differentiation(GO:0030854) |
| 0.1 | 0.6 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.1 | 1.3 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.1 | 0.2 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.1 | 3.8 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.1 | 0.7 | GO:1902951 | negative regulation of dendritic spine maintenance(GO:1902951) |
| 0.1 | 0.7 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.1 | 0.2 | GO:1901876 | regulation of calcium ion binding(GO:1901876) negative regulation of calcium ion binding(GO:1901877) |
| 0.0 | 0.2 | GO:0097325 | melanocyte proliferation(GO:0097325) |
| 0.0 | 0.1 | GO:1901143 | insulin catabolic process(GO:1901143) |
| 0.0 | 0.2 | GO:0003065 | positive regulation of heart rate by epinephrine(GO:0003065) |
| 0.0 | 0.8 | GO:0051016 | barbed-end actin filament capping(GO:0051016) |
| 0.0 | 0.2 | GO:0009191 | ribonucleoside diphosphate catabolic process(GO:0009191) |
| 0.0 | 0.6 | GO:0002726 | positive regulation of T cell cytokine production(GO:0002726) |
| 0.0 | 0.1 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.0 | 0.1 | GO:0018125 | peptidyl-cysteine methylation(GO:0018125) |
| 0.0 | 0.1 | GO:0001834 | trophectodermal cell proliferation(GO:0001834) regulation of trophectodermal cell proliferation(GO:1904073) positive regulation of trophectodermal cell proliferation(GO:1904075) |
| 0.0 | 0.1 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.0 | 0.5 | GO:0009083 | branched-chain amino acid catabolic process(GO:0009083) |
| 0.0 | 0.1 | GO:0035873 | lactate transport(GO:0015727) lactate transmembrane transport(GO:0035873) |
| 0.0 | 0.3 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) |
| 0.1 | 0.8 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.1 | 0.7 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.0 | 0.2 | GO:0031166 | integral component of vacuolar membrane(GO:0031166) |
| 0.0 | 0.1 | GO:0036195 | muscle cell projection(GO:0036194) muscle cell projection membrane(GO:0036195) |
| 0.0 | 0.2 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.0 | 0.2 | GO:0032059 | bleb(GO:0032059) |
| 0.0 | 0.1 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.0 | 0.3 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 0.7 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 0.2 | GO:0034709 | methylosome(GO:0034709) |
| 0.0 | 1.7 | GO:0031225 | anchored component of membrane(GO:0031225) |