Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
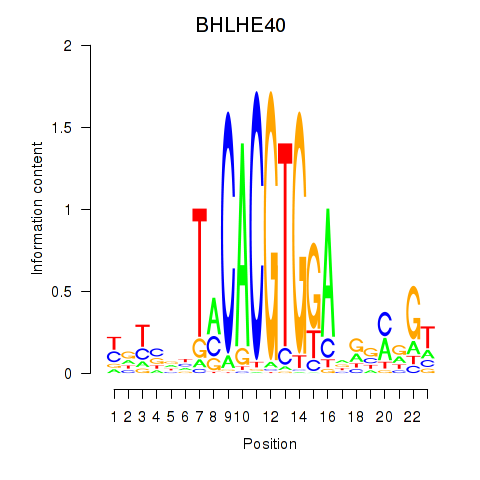
Results for BHLHE40
Z-value: 0.76
Transcription factors associated with BHLHE40
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
BHLHE40
|
ENSG00000134107.4 | BHLHE40 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| BHLHE40 | hg19_v2_chr3_+_5020801_5020952 | -0.28 | 2.9e-01 | Click! |
Activity profile of BHLHE40 motif
Sorted Z-values of BHLHE40 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of BHLHE40
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_35645618 | 1.81 |
ENST00000392218.2 ENST00000543307.1 ENST00000392219.2 ENST00000541435.2 ENST00000590686.1 ENST00000342879.3 ENST00000588699.1 |
FXYD5 |
FXYD domain containing ion transport regulator 5 |
| chr1_+_35247859 | 1.65 |
ENST00000373362.3 |
GJB3 |
gap junction protein, beta 3, 31kDa |
| chr19_-_55652290 | 1.65 |
ENST00000589745.1 |
TNNT1 |
troponin T type 1 (skeletal, slow) |
| chr19_+_35645817 | 1.56 |
ENST00000423817.3 |
FXYD5 |
FXYD domain containing ion transport regulator 5 |
| chr12_-_48298785 | 1.49 |
ENST00000550325.1 ENST00000546653.1 ENST00000549336.1 ENST00000535672.1 ENST00000229022.3 ENST00000548664.1 |
VDR |
vitamin D (1,25- dihydroxyvitamin D3) receptor |
| chr6_-_136871957 | 1.39 |
ENST00000354570.3 |
MAP7 |
microtubule-associated protein 7 |
| chr15_-_35088340 | 1.36 |
ENST00000290378.4 |
ACTC1 |
actin, alpha, cardiac muscle 1 |
| chr8_+_32405785 | 1.35 |
ENST00000287842.3 |
NRG1 |
neuregulin 1 |
| chr8_+_32405728 | 1.32 |
ENST00000523079.1 ENST00000338921.4 ENST00000356819.4 ENST00000287845.5 ENST00000341377.5 |
NRG1 |
neuregulin 1 |
| chr7_-_151574191 | 1.29 |
ENST00000287878.4 |
PRKAG2 |
protein kinase, AMP-activated, gamma 2 non-catalytic subunit |
| chr16_-_56701933 | 1.24 |
ENST00000568675.1 ENST00000569500.1 ENST00000444837.2 ENST00000379811.3 |
MT1G |
metallothionein 1G |
| chr4_+_47487285 | 1.11 |
ENST00000273859.3 ENST00000504445.1 |
ATP10D |
ATPase, class V, type 10D |
| chr13_-_38172863 | 1.01 |
ENST00000541481.1 ENST00000379743.4 ENST00000379742.4 ENST00000379749.4 ENST00000541179.1 ENST00000379747.4 |
POSTN |
periostin, osteoblast specific factor |
| chr1_-_43637915 | 0.90 |
ENST00000236051.2 |
EBNA1BP2 |
EBNA1 binding protein 2 |
| chr8_+_99129513 | 0.77 |
ENST00000522319.1 ENST00000401707.2 |
POP1 |
processing of precursor 1, ribonuclease P/MRP subunit (S. cerevisiae) |
| chr1_-_43638168 | 0.76 |
ENST00000431635.2 |
EBNA1BP2 |
EBNA1 binding protein 2 |
| chr21_-_45079341 | 0.70 |
ENST00000443485.1 ENST00000291560.2 |
HSF2BP |
heat shock transcription factor 2 binding protein |
| chr19_-_45953983 | 0.62 |
ENST00000592083.1 |
ERCC1 |
excision repair cross-complementing rodent repair deficiency, complementation group 1 (includes overlapping antisense sequence) |
| chr19_-_45909585 | 0.61 |
ENST00000593226.1 ENST00000418234.2 |
PPP1R13L |
protein phosphatase 1, regulatory subunit 13 like |
| chr8_+_55047763 | 0.54 |
ENST00000260102.4 ENST00000519831.1 |
MRPL15 |
mitochondrial ribosomal protein L15 |
| chr2_-_10587897 | 0.51 |
ENST00000405333.1 ENST00000443218.1 |
ODC1 |
ornithine decarboxylase 1 |
| chr2_+_201676908 | 0.46 |
ENST00000409226.1 ENST00000452790.2 |
BZW1 |
basic leucine zipper and W2 domains 1 |
| chr2_-_10588630 | 0.45 |
ENST00000234111.4 |
ODC1 |
ornithine decarboxylase 1 |
| chr9_+_130565147 | 0.44 |
ENST00000373247.2 ENST00000373245.1 ENST00000393706.2 ENST00000373228.1 |
FPGS |
folylpolyglutamate synthase |
| chr13_-_99630233 | 0.44 |
ENST00000376460.1 ENST00000442173.1 |
DOCK9 |
dedicator of cytokinesis 9 |
| chr17_-_13505219 | 0.43 |
ENST00000284110.1 |
HS3ST3A1 |
heparan sulfate (glucosamine) 3-O-sulfotransferase 3A1 |
| chr11_+_6625046 | 0.38 |
ENST00000396751.2 |
ILK |
integrin-linked kinase |
| chr19_+_13106383 | 0.38 |
ENST00000397661.2 |
NFIX |
nuclear factor I/X (CCAAT-binding transcription factor) |
| chr1_+_21835858 | 0.37 |
ENST00000539907.1 ENST00000540617.1 ENST00000374840.3 |
ALPL |
alkaline phosphatase, liver/bone/kidney |
| chr10_-_105677427 | 0.37 |
ENST00000369764.1 |
OBFC1 |
oligonucleotide/oligosaccharide-binding fold containing 1 |
| chr11_+_6624955 | 0.37 |
ENST00000299421.4 ENST00000537806.1 |
ILK |
integrin-linked kinase |
| chrX_+_66764375 | 0.36 |
ENST00000374690.3 |
AR |
androgen receptor |
| chr11_+_6624970 | 0.36 |
ENST00000420936.2 ENST00000528995.1 |
ILK |
integrin-linked kinase |
| chr5_+_36152091 | 0.34 |
ENST00000274254.5 |
SKP2 |
S-phase kinase-associated protein 2, E3 ubiquitin protein ligase |
| chr1_+_210001309 | 0.34 |
ENST00000491415.2 |
DIEXF |
digestive organ expansion factor homolog (zebrafish) |
| chr1_+_173793777 | 0.34 |
ENST00000239457.5 |
DARS2 |
aspartyl-tRNA synthetase 2, mitochondrial |
| chr5_+_36152163 | 0.33 |
ENST00000274255.6 |
SKP2 |
S-phase kinase-associated protein 2, E3 ubiquitin protein ligase |
| chr8_-_11710979 | 0.33 |
ENST00000415599.2 |
CTSB |
cathepsin B |
| chr15_+_68924327 | 0.33 |
ENST00000543950.1 |
CORO2B |
coronin, actin binding protein, 2B |
| chr11_-_65626753 | 0.32 |
ENST00000526975.1 ENST00000531413.1 |
CFL1 |
cofilin 1 (non-muscle) |
| chr1_+_203830703 | 0.32 |
ENST00000414487.2 |
SNRPE |
small nuclear ribonucleoprotein polypeptide E |
| chr7_-_73133959 | 0.32 |
ENST00000395155.3 ENST00000395154.3 ENST00000222812.3 ENST00000395156.3 |
STX1A |
syntaxin 1A (brain) |
| chr9_+_706842 | 0.31 |
ENST00000382293.3 |
KANK1 |
KN motif and ankyrin repeat domains 1 |
| chr21_-_27107344 | 0.30 |
ENST00000457143.2 |
ATP5J |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr21_-_27107283 | 0.29 |
ENST00000284971.3 ENST00000400099.1 |
ATP5J |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr8_-_99129338 | 0.29 |
ENST00000520507.1 |
HRSP12 |
heat-responsive protein 12 |
| chr5_+_36152179 | 0.29 |
ENST00000508514.1 ENST00000513151.1 ENST00000546211.1 |
SKP2 |
S-phase kinase-associated protein 2, E3 ubiquitin protein ligase |
| chr8_-_99129384 | 0.29 |
ENST00000521560.1 ENST00000254878.3 |
HRSP12 |
heat-responsive protein 12 |
| chrX_-_16887963 | 0.26 |
ENST00000380084.4 |
RBBP7 |
retinoblastoma binding protein 7 |
| chr17_-_74733404 | 0.26 |
ENST00000508921.3 ENST00000583836.1 ENST00000358156.6 ENST00000392485.2 ENST00000359995.5 |
SRSF2 |
serine/arginine-rich splicing factor 2 |
| chr10_-_105677886 | 0.25 |
ENST00000224950.3 |
OBFC1 |
oligonucleotide/oligosaccharide-binding fold containing 1 |
| chr21_-_30391636 | 0.25 |
ENST00000493196.1 |
RWDD2B |
RWD domain containing 2B |
| chr12_-_6677422 | 0.25 |
ENST00000382421.3 ENST00000545200.1 ENST00000399466.2 ENST00000536124.1 ENST00000540228.1 ENST00000542867.1 ENST00000545492.1 ENST00000322166.5 ENST00000545915.1 |
NOP2 |
NOP2 nucleolar protein |
| chr21_+_38071430 | 0.23 |
ENST00000290399.6 |
SIM2 |
single-minded family bHLH transcription factor 2 |
| chr3_+_133293278 | 0.22 |
ENST00000508481.1 ENST00000420115.2 ENST00000504867.1 ENST00000507408.1 ENST00000511392.1 ENST00000515421.1 |
CDV3 |
CDV3 homolog (mouse) |
| chrX_+_129040122 | 0.21 |
ENST00000394422.3 ENST00000371051.5 |
UTP14A |
UTP14, U3 small nucleolar ribonucleoprotein, homolog A (yeast) |
| chr1_-_112046289 | 0.21 |
ENST00000241356.4 |
ADORA3 |
adenosine A3 receptor |
| chr16_+_4897632 | 0.21 |
ENST00000262376.6 |
UBN1 |
ubinuclein 1 |
| chr17_+_74733744 | 0.20 |
ENST00000586689.1 ENST00000587661.1 ENST00000593181.1 ENST00000336509.4 ENST00000355954.3 |
MFSD11 |
major facilitator superfamily domain containing 11 |
| chr11_-_6624801 | 0.18 |
ENST00000534343.1 ENST00000254605.6 |
RRP8 |
ribosomal RNA processing 8, methyltransferase, homolog (yeast) |
| chrX_+_129040094 | 0.16 |
ENST00000425117.2 |
UTP14A |
UTP14, U3 small nucleolar ribonucleoprotein, homolog A (yeast) |
| chr1_+_118472343 | 0.16 |
ENST00000369441.3 ENST00000349139.5 |
WDR3 |
WD repeat domain 3 |
| chr22_-_37505449 | 0.16 |
ENST00000406725.1 |
TMPRSS6 |
transmembrane protease, serine 6 |
| chr17_+_2699697 | 0.16 |
ENST00000254695.8 ENST00000366401.4 ENST00000542807.1 |
RAP1GAP2 |
RAP1 GTPase activating protein 2 |
| chr10_+_49514698 | 0.15 |
ENST00000432379.1 ENST00000429041.1 ENST00000374189.1 |
MAPK8 |
mitogen-activated protein kinase 8 |
| chr8_+_109455845 | 0.15 |
ENST00000220853.3 |
EMC2 |
ER membrane protein complex subunit 2 |
| chrX_-_47518498 | 0.14 |
ENST00000335890.2 |
UXT |
ubiquitously-expressed, prefoldin-like chaperone |
| chrX_-_47518527 | 0.14 |
ENST00000333119.3 |
UXT |
ubiquitously-expressed, prefoldin-like chaperone |
| chr21_-_27107881 | 0.13 |
ENST00000400090.3 ENST00000400087.3 ENST00000400093.3 |
ATP5J |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr1_+_20465805 | 0.13 |
ENST00000375102.3 |
PLA2G2F |
phospholipase A2, group IIF |
| chr1_+_44445549 | 0.13 |
ENST00000356836.6 |
B4GALT2 |
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 2 |
| chr9_-_95640218 | 0.12 |
ENST00000395506.3 ENST00000375495.3 ENST00000332591.6 |
ZNF484 |
zinc finger protein 484 |
| chr1_-_47407111 | 0.12 |
ENST00000371904.4 |
CYP4A11 |
cytochrome P450, family 4, subfamily A, polypeptide 11 |
| chr6_+_5261225 | 0.12 |
ENST00000324331.6 |
FARS2 |
phenylalanyl-tRNA synthetase 2, mitochondrial |
| chr5_-_137090028 | 0.12 |
ENST00000314940.4 |
HNRNPA0 |
heterogeneous nuclear ribonucleoprotein A0 |
| chr16_+_4897912 | 0.12 |
ENST00000545171.1 |
UBN1 |
ubinuclein 1 |
| chr14_+_22694606 | 0.11 |
ENST00000390463.3 |
TRAV36DV7 |
T cell receptor alpha variable 36/delta variable 7 |
| chr12_-_113772835 | 0.11 |
ENST00000552014.1 ENST00000548186.1 ENST00000202831.3 ENST00000549181.1 |
SLC8B1 |
solute carrier family 8 (sodium/lithium/calcium exchanger), member B1 |
| chr14_+_93389425 | 0.11 |
ENST00000216492.5 ENST00000334654.4 |
CHGA |
chromogranin A (parathyroid secretory protein 1) |
| chr16_-_4897266 | 0.10 |
ENST00000591451.1 ENST00000436648.5 ENST00000381983.3 ENST00000588297.1 ENST00000321919.9 |
GLYR1 |
glyoxylate reductase 1 homolog (Arabidopsis) |
| chr17_-_7193711 | 0.10 |
ENST00000571464.1 |
YBX2 |
Y box binding protein 2 |
| chrX_+_43515467 | 0.10 |
ENST00000338702.3 ENST00000542639.1 |
MAOA |
monoamine oxidase A |
| chr12_-_54673871 | 0.09 |
ENST00000209875.4 |
CBX5 |
chromobox homolog 5 |
| chr4_-_159094194 | 0.09 |
ENST00000592057.1 ENST00000585682.1 ENST00000393807.5 |
FAM198B |
family with sequence similarity 198, member B |
| chr7_+_147830776 | 0.08 |
ENST00000538075.1 |
CNTNAP2 |
contactin associated protein-like 2 |
| chr17_+_36908984 | 0.08 |
ENST00000225426.4 ENST00000579088.1 |
PSMB3 |
proteasome (prosome, macropain) subunit, beta type, 3 |
| chr4_-_100009856 | 0.08 |
ENST00000296412.8 |
ADH5 |
alcohol dehydrogenase 5 (class III), chi polypeptide |
| chr19_+_41497178 | 0.07 |
ENST00000324071.4 |
CYP2B6 |
cytochrome P450, family 2, subfamily B, polypeptide 6 |
| chr1_+_12079517 | 0.07 |
ENST00000235332.4 ENST00000436478.2 |
MIIP |
migration and invasion inhibitory protein |
| chr1_+_207070775 | 0.07 |
ENST00000391929.3 ENST00000294984.2 ENST00000367093.3 |
IL24 |
interleukin 24 |
| chr1_-_86174065 | 0.07 |
ENST00000370574.3 ENST00000431532.2 |
ZNHIT6 |
zinc finger, HIT-type containing 6 |
| chr15_-_74988281 | 0.07 |
ENST00000566828.1 ENST00000563009.1 ENST00000568176.1 ENST00000566243.1 ENST00000566219.1 ENST00000426797.3 ENST00000566119.1 ENST00000315127.4 |
EDC3 |
enhancer of mRNA decapping 3 |
| chr22_-_30987837 | 0.06 |
ENST00000335214.6 |
PES1 |
pescadillo ribosomal biogenesis factor 1 |
| chr16_+_19729586 | 0.06 |
ENST00000564186.1 ENST00000541926.1 ENST00000433597.2 |
IQCK |
IQ motif containing K |
| chr5_-_114880533 | 0.06 |
ENST00000274457.3 |
FEM1C |
fem-1 homolog c (C. elegans) |
| chr17_-_53046058 | 0.06 |
ENST00000571584.1 ENST00000299335.3 |
COX11 |
cytochrome c oxidase assembly homolog 11 (yeast) |
| chr22_-_50523760 | 0.06 |
ENST00000395876.2 |
MLC1 |
megalencephalic leukoencephalopathy with subcortical cysts 1 |
| chr21_+_27107672 | 0.06 |
ENST00000400075.3 |
GABPA |
GA binding protein transcription factor, alpha subunit 60kDa |
| chr10_-_35379524 | 0.06 |
ENST00000374751.3 ENST00000374742.1 ENST00000602371.1 |
CUL2 |
cullin 2 |
| chr22_-_30987849 | 0.05 |
ENST00000402284.3 ENST00000354694.7 |
PES1 |
pescadillo ribosomal biogenesis factor 1 |
| chr12_+_66217911 | 0.05 |
ENST00000403681.2 |
HMGA2 |
high mobility group AT-hook 2 |
| chr7_+_141463897 | 0.05 |
ENST00000247879.2 |
TAS2R3 |
taste receptor, type 2, member 3 |
| chr8_-_48872686 | 0.05 |
ENST00000314191.2 ENST00000338368.3 |
PRKDC |
protein kinase, DNA-activated, catalytic polypeptide |
| chr7_-_107642348 | 0.05 |
ENST00000393561.1 |
LAMB1 |
laminin, beta 1 |
| chr17_+_72772621 | 0.04 |
ENST00000335464.5 ENST00000417024.2 ENST00000578764.1 ENST00000582773.1 ENST00000582330.1 |
TMEM104 |
transmembrane protein 104 |
| chr11_+_60609537 | 0.04 |
ENST00000227520.5 |
CCDC86 |
coiled-coil domain containing 86 |
| chr17_+_61851157 | 0.04 |
ENST00000578681.1 ENST00000583590.1 |
DDX42 |
DEAD (Asp-Glu-Ala-Asp) box helicase 42 |
| chr2_-_27545921 | 0.04 |
ENST00000402310.1 ENST00000405983.1 ENST00000403262.2 ENST00000428910.1 ENST00000402722.1 ENST00000399052.4 ENST00000380044.1 ENST00000405076.1 |
MPV17 |
MpV17 mitochondrial inner membrane protein |
| chr5_+_140588269 | 0.04 |
ENST00000541609.1 ENST00000239450.2 |
PCDHB12 |
protocadherin beta 12 |
| chr8_+_22857048 | 0.03 |
ENST00000251822.6 |
RHOBTB2 |
Rho-related BTB domain containing 2 |
| chr11_-_65626797 | 0.03 |
ENST00000525451.2 |
CFL1 |
cofilin 1 (non-muscle) |
| chr16_-_20362147 | 0.03 |
ENST00000396142.2 |
UMOD |
uromodulin |
| chr2_+_207630081 | 0.03 |
ENST00000236980.6 ENST00000418289.1 ENST00000402774.3 ENST00000403094.3 |
FASTKD2 |
FAST kinase domains 2 |
| chr1_-_113498943 | 0.03 |
ENST00000369626.3 |
SLC16A1 |
solute carrier family 16 (monocarboxylate transporter), member 1 |
| chr18_+_580367 | 0.03 |
ENST00000327228.3 |
CETN1 |
centrin, EF-hand protein, 1 |
| chr10_-_35379241 | 0.02 |
ENST00000374748.1 ENST00000374749.3 |
CUL2 |
cullin 2 |
| chr17_+_66624280 | 0.02 |
ENST00000585484.1 |
RP11-118B18.1 |
RP11-118B18.1 |
| chr2_+_74154032 | 0.02 |
ENST00000356837.6 |
DGUOK |
deoxyguanosine kinase |
| chr2_+_103236004 | 0.01 |
ENST00000233969.2 |
SLC9A2 |
solute carrier family 9, subfamily A (NHE2, cation proton antiporter 2), member 2 |
| chr6_+_13615554 | 0.01 |
ENST00000451315.2 |
NOL7 |
nucleolar protein 7, 27kDa |
| chr8_+_58890917 | 0.01 |
ENST00000522992.1 |
RP11-1112C15.1 |
RP11-1112C15.1 |
| chrX_+_23685653 | 0.01 |
ENST00000379331.3 |
PRDX4 |
peroxiredoxin 4 |
| chr2_+_74153953 | 0.00 |
ENST00000264093.4 ENST00000348222.1 |
DGUOK |
deoxyguanosine kinase |
| chr12_-_58165870 | 0.00 |
ENST00000257848.7 |
METTL1 |
methyltransferase like 1 |
| chr12_+_93861282 | 0.00 |
ENST00000552217.1 ENST00000393128.4 ENST00000547098.1 |
MRPL42 |
mitochondrial ribosomal protein L42 |
| chr1_+_119957554 | 0.00 |
ENST00000543831.1 ENST00000433745.1 ENST00000369416.3 |
HSD3B2 |
hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 2 |
| chr9_-_100881466 | 0.00 |
ENST00000341469.2 ENST00000342043.3 ENST00000375098.3 |
TRIM14 |
tripartite motif containing 14 |
| chr12_-_25102252 | 0.00 |
ENST00000261192.7 |
BCAT1 |
branched chain amino-acid transaminase 1, cytosolic |
Gene Ontology Analysis
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.7 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 1.5 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 1.3 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.0 | 1.0 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.1 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 0.4 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.5 | GO:0038181 | bile acid receptor activity(GO:0038181) |
| 0.3 | 2.7 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.1 | 1.3 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.1 | 1.6 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.1 | 1.6 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.1 | 0.6 | GO:1990599 | 3' overhang single-stranded DNA endodeoxyribonuclease activity(GO:1990599) |
| 0.1 | 0.2 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.1 | 0.4 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.1 | 3.4 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.1 | 0.4 | GO:0004882 | androgen receptor activity(GO:0004882) |
| 0.1 | 0.3 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.1 | 0.4 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.0 | 0.1 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.0 | 0.8 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 1.1 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.1 | GO:0018685 | alkane 1-monooxygenase activity(GO:0018685) |
| 0.0 | 0.1 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 0.0 | 0.3 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 0.0 | 0.1 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.0 | 0.1 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.0 | 0.2 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.0 | 0.3 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.0 | 0.4 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.0 | 0.1 | GO:0035501 | MH1 domain binding(GO:0035501) |
| 0.0 | 0.0 | GO:0004677 | DNA-dependent protein kinase activity(GO:0004677) |
| 0.0 | 1.3 | GO:0017022 | myosin binding(GO:0017022) |
| 0.0 | 0.1 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 0.1 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 3.4 | GO:0046586 | regulation of calcium-dependent cell-cell adhesion(GO:0046586) |
| 0.5 | 1.5 | GO:0060058 | apoptotic process involved in mammary gland involution(GO:0060057) positive regulation of apoptotic process involved in mammary gland involution(GO:0060058) positive regulation of apoptotic process involved in morphogenesis(GO:1902339) regulation of mammary gland involution(GO:1903519) positive regulation of mammary gland involution(GO:1903521) positive regulation of apoptotic process involved in development(GO:1904747) |
| 0.4 | 2.7 | GO:0021842 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.3 | 1.0 | GO:1990523 | bone regeneration(GO:1990523) |
| 0.2 | 1.0 | GO:0033387 | putrescine biosynthetic process from ornithine(GO:0033387) |
| 0.2 | 1.4 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 0.2 | 1.0 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 0.2 | 1.6 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 0.1 | 0.4 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.1 | 0.8 | GO:0016078 | tRNA catabolic process(GO:0016078) |
| 0.1 | 1.2 | GO:0042117 | monocyte activation(GO:0042117) |
| 0.1 | 0.4 | GO:0071529 | cementum mineralization(GO:0071529) |
| 0.1 | 0.4 | GO:0060599 | lateral sprouting involved in mammary gland duct morphogenesis(GO:0060599) |
| 0.1 | 0.3 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 0.1 | 0.3 | GO:2000393 | negative regulation of lamellipodium morphogenesis(GO:2000393) |
| 0.1 | 0.6 | GO:0000720 | pyrimidine dimer repair by nucleotide-excision repair(GO:0000720) positive regulation of t-circle formation(GO:1904431) |
| 0.1 | 0.3 | GO:0010701 | positive regulation of norepinephrine secretion(GO:0010701) |
| 0.1 | 0.4 | GO:2000771 | regulation of unidimensional cell growth(GO:0051510) negative regulation of unidimensional cell growth(GO:0051511) establishment of cell polarity regulating cell shape(GO:0071964) regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000769) positive regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000771) regulation of establishment of cell polarity regulating cell shape(GO:2000782) positive regulation of establishment of cell polarity regulating cell shape(GO:2000784) regulation of barbed-end actin filament capping(GO:2000812) positive regulation of barbed-end actin filament capping(GO:2000814) |
| 0.1 | 1.3 | GO:0006853 | carnitine shuttle(GO:0006853) |
| 0.1 | 0.3 | GO:0070370 | heat acclimation(GO:0010286) cellular heat acclimation(GO:0070370) |
| 0.0 | 0.1 | GO:1902595 | regulation of DNA replication origin binding(GO:1902595) |
| 0.0 | 1.1 | GO:0045663 | positive regulation of myoblast differentiation(GO:0045663) |
| 0.0 | 0.1 | GO:0003095 | pressure natriuresis(GO:0003095) |
| 0.0 | 1.1 | GO:0034204 | lipid translocation(GO:0034204) phospholipid translocation(GO:0045332) |
| 0.0 | 0.1 | GO:0061110 | dense core granule biogenesis(GO:0061110) positive regulation of relaxation of cardiac muscle(GO:1901899) regulation of dense core granule biogenesis(GO:2000705) |
| 0.0 | 0.2 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.0 | 0.1 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.0 | 0.1 | GO:1903300 | negative regulation of glucokinase activity(GO:0033132) negative regulation of hexokinase activity(GO:1903300) |
| 0.0 | 0.1 | GO:0071109 | superior temporal gyrus development(GO:0071109) |
| 0.0 | 0.1 | GO:0046294 | formaldehyde catabolic process(GO:0046294) |
| 0.0 | 0.2 | GO:0046015 | regulation of transcription by glucose(GO:0046015) |
| 0.0 | 0.1 | GO:0009386 | translational attenuation(GO:0009386) |
| 0.0 | 0.6 | GO:0003215 | cardiac right ventricle morphogenesis(GO:0003215) |
| 0.0 | 0.3 | GO:0033197 | response to vitamin E(GO:0033197) |
| 0.0 | 0.7 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.0 | 1.8 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.0 | 0.1 | GO:2000683 | mesodermal-endodermal cell signaling(GO:0003131) programmed DNA elimination(GO:0031049) chromosome breakage(GO:0031052) histone H2A-S139 phosphorylation(GO:0035978) regulation of cellular response to X-ray(GO:2000683) positive regulation of cellular response to X-ray(GO:2000685) |
| 0.0 | 0.2 | GO:0001973 | adenosine receptor signaling pathway(GO:0001973) |
| 0.0 | 1.6 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.9 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.0 | 0.1 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.0 | 0.3 | GO:0046697 | decidualization(GO:0046697) |
| 0.0 | 0.0 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 0.0 | 0.3 | GO:0034643 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) |
| 0.0 | 0.1 | GO:0048254 | snoRNA localization(GO:0048254) |
| 0.0 | 0.2 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.0 | GO:0046122 | purine deoxyribonucleoside metabolic process(GO:0046122) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 2.7 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.1 | 1.3 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 1.1 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 1.0 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.7 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.0 | 0.3 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.0 | 1.6 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.6 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.5 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.0 | 1.1 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 1.8 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.3 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.0 | 0.3 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 1.0 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.4 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | GO:0042643 | actomyosin, actin portion(GO:0042643) |
| 0.1 | 0.8 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.1 | 1.6 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 0.6 | GO:0070522 | nucleotide-excision repair factor 1 complex(GO:0000110) ERCC4-ERCC1 complex(GO:0070522) |
| 0.1 | 2.8 | GO:0030673 | axolemma(GO:0030673) |
| 0.1 | 1.6 | GO:0005861 | troponin complex(GO:0005861) |
| 0.1 | 1.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.1 | 0.3 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.1 | 0.4 | GO:0065010 | extracellular membrane-bounded organelle(GO:0065010) |
| 0.1 | 1.8 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.1 | 0.3 | GO:0035061 | interchromatin granule(GO:0035061) |
| 0.0 | 0.3 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.0 | 0.7 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 0.3 | GO:0034715 | U7 snRNP(GO:0005683) pICln-Sm protein complex(GO:0034715) |
| 0.0 | 0.2 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.0 | 0.2 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 0.7 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.0 | 1.1 | GO:0043034 | costamere(GO:0043034) |
| 0.0 | 0.1 | GO:0030891 | VCB complex(GO:0030891) |
| 0.0 | 1.0 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 0.1 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.0 | 0.1 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.0 | 0.0 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 0.0 | 0.5 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.6 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 0.8 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.3 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 0.3 | GO:0032587 | ruffle membrane(GO:0032587) |