Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
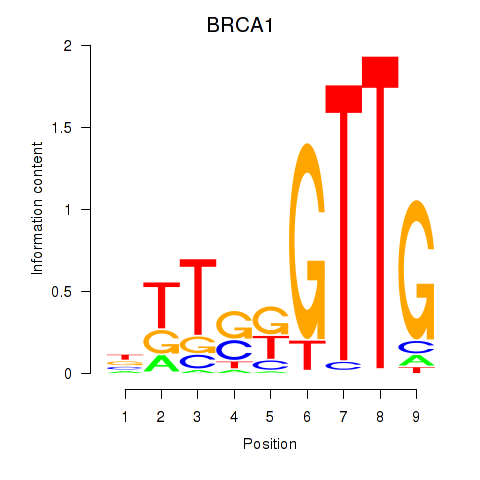
Results for BRCA1
Z-value: 0.68
Transcription factors associated with BRCA1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
BRCA1
|
ENSG00000012048.15 | BRCA1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| BRCA1 | hg19_v2_chr17_-_41277370_41277419 | 0.88 | 8.3e-06 | Click! |
Activity profile of BRCA1 motif
Sorted Z-values of BRCA1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of BRCA1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_69597810 | 1.47 |
ENST00000483798.2 |
DNAJC12 |
DnaJ (Hsp40) homolog, subfamily C, member 12 |
| chr17_+_74372662 | 1.43 |
ENST00000591651.1 ENST00000545180.1 |
SPHK1 |
sphingosine kinase 1 |
| chr15_-_58358607 | 1.31 |
ENST00000249750.4 |
ALDH1A2 |
aldehyde dehydrogenase 1 family, member A2 |
| chr14_+_103058948 | 1.03 |
ENST00000262241.6 |
RCOR1 |
REST corepressor 1 |
| chr7_-_23510086 | 0.96 |
ENST00000258729.3 |
IGF2BP3 |
insulin-like growth factor 2 mRNA binding protein 3 |
| chr6_+_16129308 | 0.87 |
ENST00000356840.3 ENST00000349606.4 |
MYLIP |
myosin regulatory light chain interacting protein |
| chr3_+_193853927 | 0.85 |
ENST00000232424.3 |
HES1 |
hes family bHLH transcription factor 1 |
| chr18_+_55888767 | 0.70 |
ENST00000431212.2 ENST00000586268.1 ENST00000587190.1 |
NEDD4L |
neural precursor cell expressed, developmentally down-regulated 4-like, E3 ubiquitin protein ligase |
| chr17_-_56296580 | 0.69 |
ENST00000313863.6 ENST00000546108.1 ENST00000337050.7 ENST00000393119.2 |
MKS1 |
Meckel syndrome, type 1 |
| chr4_-_140005443 | 0.56 |
ENST00000510408.1 ENST00000420916.2 ENST00000358635.3 |
ELF2 |
E74-like factor 2 (ets domain transcription factor) |
| chr6_+_33172407 | 0.52 |
ENST00000374662.3 |
HSD17B8 |
hydroxysteroid (17-beta) dehydrogenase 8 |
| chr1_+_10271674 | 0.50 |
ENST00000377086.1 |
KIF1B |
kinesin family member 1B |
| chr10_-_36813162 | 0.50 |
ENST00000440465.1 |
NAMPTL |
nicotinamide phosphoribosyltransferase-like |
| chr6_+_42981922 | 0.49 |
ENST00000326974.4 ENST00000244670.8 |
KLHDC3 |
kelch domain containing 3 |
| chr11_+_117947724 | 0.47 |
ENST00000534111.1 |
TMPRSS4 |
transmembrane protease, serine 4 |
| chr17_-_7082861 | 0.45 |
ENST00000269299.3 |
ASGR1 |
asialoglycoprotein receptor 1 |
| chr11_+_117947782 | 0.41 |
ENST00000522307.1 ENST00000523251.1 ENST00000437212.3 ENST00000522824.1 ENST00000522151.1 |
TMPRSS4 |
transmembrane protease, serine 4 |
| chr10_+_28822636 | 0.41 |
ENST00000442148.1 ENST00000448193.1 |
WAC |
WW domain containing adaptor with coiled-coil |
| chr3_+_185303962 | 0.41 |
ENST00000296257.5 |
SENP2 |
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr3_-_176914238 | 0.39 |
ENST00000430069.1 ENST00000428970.1 |
TBL1XR1 |
transducin (beta)-like 1 X-linked receptor 1 |
| chr7_+_73868439 | 0.36 |
ENST00000424337.2 |
GTF2IRD1 |
GTF2I repeat domain containing 1 |
| chr20_-_48530230 | 0.35 |
ENST00000422556.1 |
SPATA2 |
spermatogenesis associated 2 |
| chr20_+_52105495 | 0.33 |
ENST00000439873.2 |
AL354993.1 |
Cell growth-inhibiting protein 7; HCG1784586; Uncharacterized protein |
| chr8_+_22022800 | 0.33 |
ENST00000397814.3 |
BMP1 |
bone morphogenetic protein 1 |
| chr12_-_12491608 | 0.33 |
ENST00000545735.1 |
MANSC1 |
MANSC domain containing 1 |
| chr1_+_159141397 | 0.32 |
ENST00000368124.4 ENST00000368125.4 ENST00000416746.1 |
CADM3 |
cell adhesion molecule 3 |
| chr17_-_41738931 | 0.32 |
ENST00000329168.3 ENST00000549132.1 |
MEOX1 |
mesenchyme homeobox 1 |
| chr17_-_60142609 | 0.31 |
ENST00000397786.2 |
MED13 |
mediator complex subunit 13 |
| chr11_+_47291645 | 0.31 |
ENST00000395336.3 ENST00000402192.2 |
MADD |
MAP-kinase activating death domain |
| chr6_+_123100620 | 0.30 |
ENST00000368444.3 |
FABP7 |
fatty acid binding protein 7, brain |
| chr5_-_68665084 | 0.28 |
ENST00000509462.1 |
TAF9 |
TAF9 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 32kDa |
| chr8_-_81083341 | 0.28 |
ENST00000519303.2 |
TPD52 |
tumor protein D52 |
| chr11_-_6817128 | 0.28 |
ENST00000332601.3 |
OR6A2 |
olfactory receptor, family 6, subfamily A, member 2 |
| chrX_+_49644470 | 0.27 |
ENST00000508866.2 |
USP27X |
ubiquitin specific peptidase 27, X-linked |
| chr17_-_41739283 | 0.27 |
ENST00000393661.2 ENST00000318579.4 |
MEOX1 |
mesenchyme homeobox 1 |
| chr10_-_61900762 | 0.26 |
ENST00000355288.2 |
ANK3 |
ankyrin 3, node of Ranvier (ankyrin G) |
| chr19_-_48389651 | 0.26 |
ENST00000222002.3 |
SULT2A1 |
sulfotransferase family, cytosolic, 2A, dehydroepiandrosterone (DHEA)-preferring, member 1 |
| chr5_-_68665296 | 0.25 |
ENST00000512152.1 ENST00000503245.1 ENST00000512561.1 ENST00000380822.4 |
TAF9 |
TAF9 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 32kDa |
| chr13_-_31038370 | 0.25 |
ENST00000399489.1 ENST00000339872.4 |
HMGB1 |
high mobility group box 1 |
| chr19_+_46850320 | 0.25 |
ENST00000391919.1 |
PPP5C |
protein phosphatase 5, catalytic subunit |
| chr1_-_173020056 | 0.24 |
ENST00000239468.2 ENST00000404377.3 |
TNFSF18 |
tumor necrosis factor (ligand) superfamily, member 18 |
| chr2_-_152118352 | 0.23 |
ENST00000331426.5 |
RBM43 |
RNA binding motif protein 43 |
| chr19_+_46850251 | 0.22 |
ENST00000012443.4 |
PPP5C |
protein phosphatase 5, catalytic subunit |
| chr18_-_23671139 | 0.21 |
ENST00000579061.1 ENST00000542420.2 |
SS18 |
synovial sarcoma translocation, chromosome 18 |
| chr1_-_217250231 | 0.20 |
ENST00000493748.1 ENST00000463665.1 |
ESRRG |
estrogen-related receptor gamma |
| chr15_-_88799948 | 0.20 |
ENST00000394480.2 |
NTRK3 |
neurotrophic tyrosine kinase, receptor, type 3 |
| chr11_+_47291193 | 0.19 |
ENST00000428807.1 ENST00000402799.1 ENST00000406482.1 ENST00000349238.3 ENST00000311027.5 ENST00000407859.3 ENST00000395344.3 ENST00000444117.1 |
MADD |
MAP-kinase activating death domain |
| chr14_-_69261310 | 0.19 |
ENST00000336440.3 |
ZFP36L1 |
ZFP36 ring finger protein-like 1 |
| chr5_+_139493665 | 0.18 |
ENST00000331327.3 |
PURA |
purine-rich element binding protein A |
| chr1_-_227505289 | 0.18 |
ENST00000366765.3 |
CDC42BPA |
CDC42 binding protein kinase alpha (DMPK-like) |
| chr6_-_170862322 | 0.17 |
ENST00000262193.6 |
PSMB1 |
proteasome (prosome, macropain) subunit, beta type, 1 |
| chr12_-_67072714 | 0.17 |
ENST00000545666.1 ENST00000398016.3 ENST00000359742.4 ENST00000286445.7 ENST00000538211.1 |
GRIP1 |
glutamate receptor interacting protein 1 |
| chr5_-_68665469 | 0.16 |
ENST00000217893.5 |
TAF9 |
TAF9 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 32kDa |
| chr9_-_86593238 | 0.14 |
ENST00000351839.3 |
HNRNPK |
heterogeneous nuclear ribonucleoprotein K |
| chr1_-_160232312 | 0.13 |
ENST00000440682.1 |
DCAF8 |
DDB1 and CUL4 associated factor 8 |
| chr11_-_119234876 | 0.13 |
ENST00000525735.1 |
USP2 |
ubiquitin specific peptidase 2 |
| chr2_+_203879568 | 0.12 |
ENST00000449802.1 |
NBEAL1 |
neurobeachin-like 1 |
| chr4_+_619386 | 0.12 |
ENST00000496514.1 |
PDE6B |
phosphodiesterase 6B, cGMP-specific, rod, beta |
| chr14_-_60097297 | 0.11 |
ENST00000395090.1 |
RTN1 |
reticulon 1 |
| chr12_-_117319236 | 0.11 |
ENST00000257572.5 |
HRK |
harakiri, BCL2 interacting protein (contains only BH3 domain) |
| chr10_+_92631709 | 0.11 |
ENST00000413330.1 ENST00000277882.3 |
RPP30 |
ribonuclease P/MRP 30kDa subunit |
| chr14_+_32030582 | 0.11 |
ENST00000550649.1 ENST00000281081.7 |
NUBPL |
nucleotide binding protein-like |
| chr11_-_71823266 | 0.10 |
ENST00000538919.1 ENST00000539395.1 ENST00000542531.1 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr19_+_45394477 | 0.08 |
ENST00000252487.5 ENST00000405636.2 ENST00000592434.1 ENST00000426677.2 ENST00000589649.1 |
TOMM40 |
translocase of outer mitochondrial membrane 40 homolog (yeast) |
| chr6_-_25832254 | 0.08 |
ENST00000476801.1 ENST00000244527.4 ENST00000427328.1 |
SLC17A1 |
solute carrier family 17 (organic anion transporter), member 1 |
| chr8_-_16859690 | 0.08 |
ENST00000180166.5 |
FGF20 |
fibroblast growth factor 20 |
| chr19_-_40324767 | 0.06 |
ENST00000601972.1 ENST00000430012.2 ENST00000323039.5 ENST00000348817.3 |
DYRK1B |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1B |
| chr4_+_619347 | 0.05 |
ENST00000255622.6 |
PDE6B |
phosphodiesterase 6B, cGMP-specific, rod, beta |
| chr16_-_790982 | 0.04 |
ENST00000301694.5 ENST00000251588.2 |
NARFL |
nuclear prelamin A recognition factor-like |
| chr19_+_45971246 | 0.04 |
ENST00000585836.1 ENST00000417353.2 ENST00000353609.3 ENST00000591858.1 ENST00000443841.2 ENST00000590335.1 |
FOSB |
FBJ murine osteosarcoma viral oncogene homolog B |
| chr5_+_140186647 | 0.04 |
ENST00000512229.2 ENST00000356878.4 ENST00000530339.1 |
PCDHA4 |
protocadherin alpha 4 |
| chr12_-_49418407 | 0.03 |
ENST00000526209.1 |
KMT2D |
lysine (K)-specific methyltransferase 2D |
| chr1_-_160232197 | 0.03 |
ENST00000419626.1 ENST00000610139.1 ENST00000475733.1 ENST00000407642.2 ENST00000368073.3 ENST00000326837.2 |
DCAF8 |
DDB1 and CUL4 associated factor 8 |
| chr13_+_30002741 | 0.02 |
ENST00000380808.2 |
MTUS2 |
microtubule associated tumor suppressor candidate 2 |
| chr6_-_42981651 | 0.02 |
ENST00000244711.3 |
MEA1 |
male-enhanced antigen 1 |
| chr14_-_60097524 | 0.02 |
ENST00000342503.4 |
RTN1 |
reticulon 1 |
| chr17_+_36908984 | 0.01 |
ENST00000225426.4 ENST00000579088.1 |
PSMB3 |
proteasome (prosome, macropain) subunit, beta type, 3 |
| chr1_+_161736072 | 0.01 |
ENST00000367942.3 |
ATF6 |
activating transcription factor 6 |
| chrX_+_10124977 | 0.00 |
ENST00000380833.4 |
CLCN4 |
chloride channel, voltage-sensitive 4 |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.4 | GO:0008481 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.1 | 1.3 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.1 | 0.5 | GO:0070404 | NADH binding(GO:0070404) |
| 0.1 | 0.4 | GO:0070137 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.1 | 0.7 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.1 | 0.2 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.1 | 0.5 | GO:0004873 | asialoglycoprotein receptor activity(GO:0004873) |
| 0.0 | 0.7 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.0 | 0.2 | GO:0010858 | calcium-dependent protein kinase regulator activity(GO:0010858) |
| 0.0 | 0.3 | GO:0050294 | steroid sulfotransferase activity(GO:0050294) |
| 0.0 | 0.2 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.0 | 0.3 | GO:1990380 | Lys63-specific deubiquitinase activity(GO:0061578) Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.0 | 1.0 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 0.6 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.0 | 0.3 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.0 | 1.0 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.0 | 0.5 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.0 | 0.2 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) neurotrophin receptor activity(GO:0005030) |
| 0.0 | 0.5 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 0.9 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.4 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.0 | 0.2 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:0046521 | sphingoid catabolic process(GO:0046521) |
| 0.3 | 1.5 | GO:0061009 | common bile duct development(GO:0061009) |
| 0.2 | 0.9 | GO:0032803 | regulation of low-density lipoprotein particle receptor catabolic process(GO:0032803) |
| 0.2 | 0.5 | GO:1904647 | response to rotenone(GO:1904647) |
| 0.1 | 0.6 | GO:0001757 | somite specification(GO:0001757) sclerotome development(GO:0061056) |
| 0.1 | 1.3 | GO:0035799 | ureter maturation(GO:0035799) |
| 0.1 | 0.7 | GO:2001288 | positive regulation of caveolin-mediated endocytosis(GO:2001288) |
| 0.1 | 0.4 | GO:0071894 | histone H2B conserved C-terminal lysine ubiquitination(GO:0071894) |
| 0.1 | 1.0 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.1 | 0.5 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.1 | 0.5 | GO:2000324 | positive regulation of glucocorticoid receptor signaling pathway(GO:2000324) |
| 0.1 | 0.4 | GO:0014886 | transition between slow and fast fiber(GO:0014886) |
| 0.1 | 0.3 | GO:0010650 | positive regulation of cell communication by electrical coupling(GO:0010650) maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.1 | 0.2 | GO:0048687 | modulation by virus of host transcription(GO:0019056) positive regulation of sprouting of injured axon(GO:0048687) positive regulation of axon extension involved in regeneration(GO:0048691) modulation by symbiont of host transcription(GO:0052026) |
| 0.1 | 0.4 | GO:0060613 | fat pad development(GO:0060613) |
| 0.1 | 0.2 | GO:0048382 | mesendoderm development(GO:0048382) |
| 0.1 | 0.2 | GO:0002840 | plasmacytoid dendritic cell activation(GO:0002270) T cell mediated immune response to tumor cell(GO:0002424) regulation of T cell mediated immune response to tumor cell(GO:0002840) regulation of restriction endodeoxyribonuclease activity(GO:0032072) negative regulation of apoptotic cell clearance(GO:2000426) |
| 0.1 | 0.2 | GO:2000320 | negative regulation of T-helper 17 type immune response(GO:2000317) negative regulation of T-helper 17 cell differentiation(GO:2000320) |
| 0.1 | 0.7 | GO:0000492 | box C/D snoRNP assembly(GO:0000492) |
| 0.1 | 0.4 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.0 | 0.1 | GO:0072369 | regulation of lipid transport by positive regulation of transcription from RNA polymerase II promoter(GO:0072369) |
| 0.0 | 0.5 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.0 | 0.2 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.0 | 0.6 | GO:0050855 | regulation of B cell receptor signaling pathway(GO:0050855) |
| 0.0 | 0.1 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.0 | 0.3 | GO:0060134 | prepulse inhibition(GO:0060134) |
| 0.0 | 0.5 | GO:0035825 | reciprocal meiotic recombination(GO:0007131) reciprocal DNA recombination(GO:0035825) |
| 0.0 | 0.2 | GO:0034380 | high-density lipoprotein particle assembly(GO:0034380) |
| 0.0 | 0.3 | GO:0006068 | ethanol catabolic process(GO:0006068) steroid catabolic process(GO:0006706) |
| 0.0 | 0.3 | GO:0071108 | protein K63-linked deubiquitination(GO:0070536) protein K48-linked deubiquitination(GO:0071108) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 0.1 | 0.5 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.0 | 0.7 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 1.0 | GO:1990391 | DNA repair complex(GO:1990391) |
| 0.0 | 0.1 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 0.3 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 0.2 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 0.1 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.2 | GO:0019774 | proteasome core complex, beta-subunit complex(GO:0019774) |
| 0.0 | 0.7 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.0 | 0.5 | GO:1904115 | axon cytoplasm(GO:1904115) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.4 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.0 | 0.9 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 0.7 | PID MYC PATHWAY | C-MYC pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.4 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.9 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.0 | 0.4 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 0.2 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.2 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |