Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
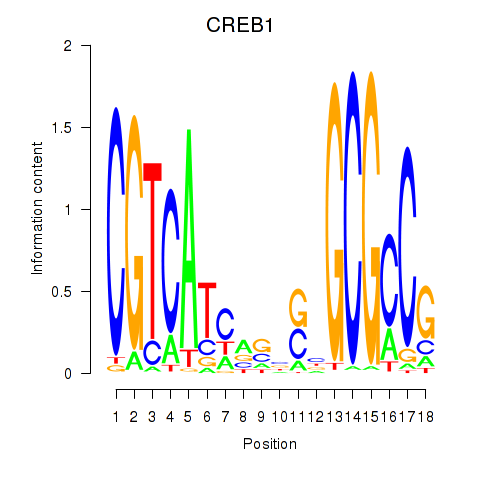
Results for CREB1
Z-value: 2.07
Transcription factors associated with CREB1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
CREB1
|
ENSG00000118260.10 | CREB1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| CREB1 | hg19_v2_chr2_+_208394455_208394471 | 0.27 | 3.2e-01 | Click! |
Activity profile of CREB1 motif
Sorted Z-values of CREB1 motif
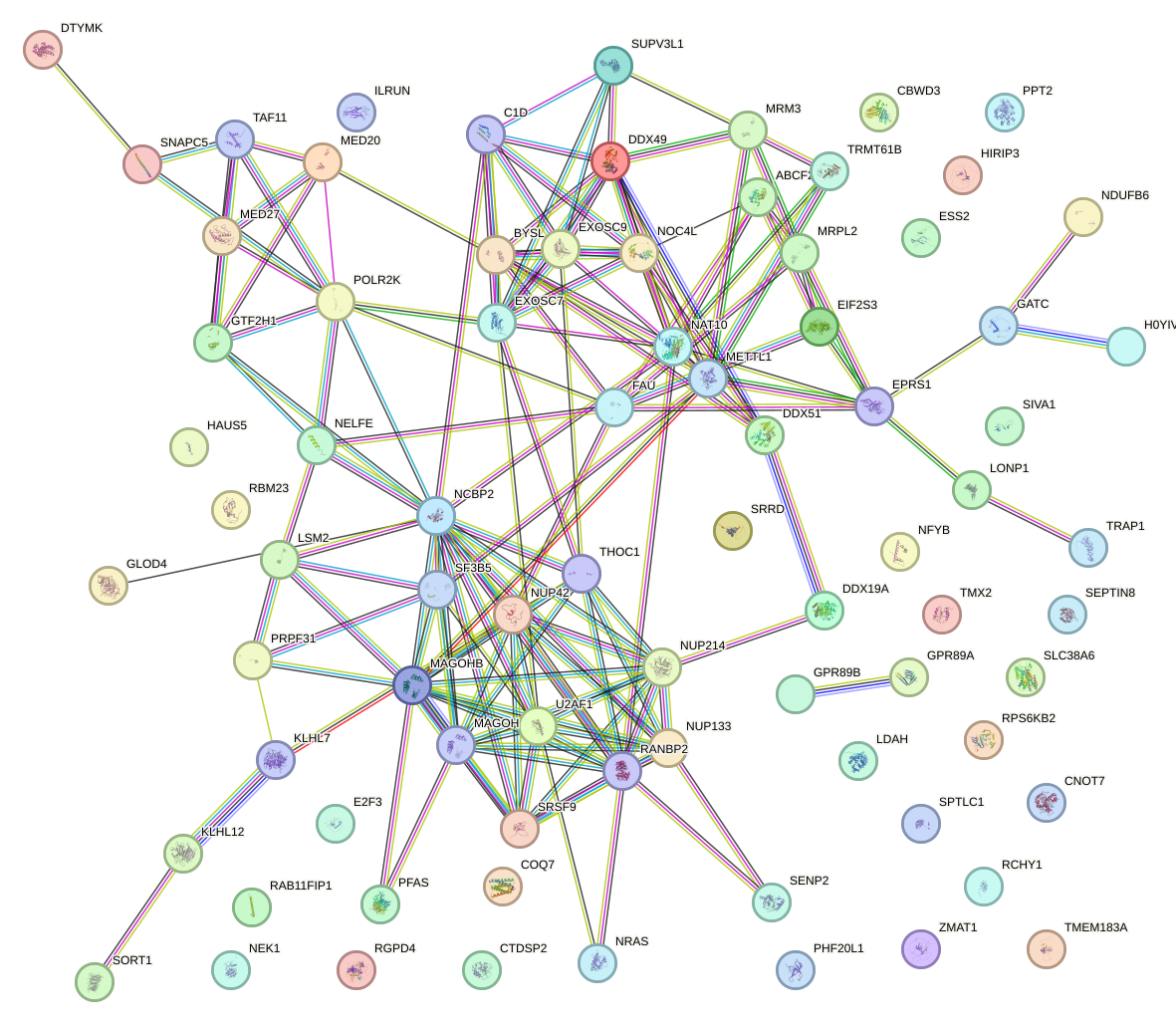
Network of associatons between targets according to the STRING database.
First level regulatory network of CREB1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr6_-_43027105 | 3.54 |
ENST00000230413.5 ENST00000487429.1 ENST00000489623.1 ENST00000468957.1 |
MRPL2 |
mitochondrial ribosomal protein L2 |
| chr12_+_132628963 | 3.36 |
ENST00000330579.1 |
NOC4L |
nucleolar complex associated 4 homolog (S. cerevisiae) |
| chr18_-_268019 | 2.68 |
ENST00000261600.6 |
THOC1 |
THO complex 1 |
| chr3_+_45017722 | 2.56 |
ENST00000265564.7 |
EXOSC7 |
exosome component 7 |
| chr1_-_53704157 | 2.43 |
ENST00000371466.4 ENST00000371470.3 |
MAGOH |
mago-nashi homolog, proliferation-associated (Drosophila) |
| chr16_-_3767506 | 2.42 |
ENST00000538171.1 |
TRAP1 |
TNF receptor-associated protein 1 |
| chr3_+_185303962 | 2.42 |
ENST00000296257.5 |
SENP2 |
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr6_-_31774714 | 2.36 |
ENST00000375661.5 |
LSM2 |
LSM2 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr3_+_185304059 | 2.29 |
ENST00000427465.2 |
SENP2 |
SUMO1/sentrin/SMT3 specific peptidase 2 |
| chr16_-_3767551 | 2.29 |
ENST00000246957.5 |
TRAP1 |
TNF receptor-associated protein 1 |
| chr11_-_62607036 | 2.26 |
ENST00000311713.7 ENST00000278856.4 |
WDR74 |
WD repeat domain 74 |
| chr9_+_134000948 | 2.22 |
ENST00000359428.5 ENST00000411637.2 ENST00000451030.1 |
NUP214 |
nucleoporin 214kDa |
| chr4_+_39460620 | 2.14 |
ENST00000340169.2 ENST00000261434.3 |
LIAS |
lipoic acid synthetase |
| chr4_+_39460659 | 2.13 |
ENST00000513731.1 |
LIAS |
lipoic acid synthetase |
| chr8_-_37756972 | 2.11 |
ENST00000330843.4 ENST00000522727.1 ENST00000287263.4 |
RAB11FIP1 |
RAB11 family interacting protein 1 (class I) |
| chr4_+_39460689 | 2.06 |
ENST00000381846.1 |
LIAS |
lipoic acid synthetase |
| chr6_-_34855773 | 2.02 |
ENST00000420584.2 ENST00000361288.4 |
TAF11 |
TAF11 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 28kDa |
| chr17_+_8152590 | 1.99 |
ENST00000584044.1 ENST00000314666.6 ENST00000545834.1 ENST00000581242.1 |
PFAS |
phosphoribosylformylglycinamidine synthase |
| chr7_+_23145347 | 1.94 |
ENST00000322231.7 |
KLHL7 |
kelch-like family member 7 |
| chr6_+_32121789 | 1.81 |
ENST00000437001.2 ENST00000375137.2 |
PPT2 |
palmitoyl-protein thioesterase 2 |
| chr1_+_12040238 | 1.74 |
ENST00000444836.1 ENST00000235329.5 |
MFN2 |
mitofusin 2 |
| chr6_+_32121908 | 1.61 |
ENST00000375143.2 ENST00000424499.1 |
PPT2 |
palmitoyl-protein thioesterase 2 |
| chr2_-_21022818 | 1.57 |
ENST00000440866.2 ENST00000541941.1 ENST00000402479.2 ENST00000435420.2 ENST00000432947.1 ENST00000403006.2 ENST00000419825.2 ENST00000381090.3 ENST00000237822.3 ENST00000412261.1 |
C2orf43 |
chromosome 2 open reading frame 43 |
| chr15_-_66790146 | 1.56 |
ENST00000316634.5 |
SNAPC5 |
small nuclear RNA activating complex, polypeptide 5, 19kDa |
| chr4_-_146019693 | 1.55 |
ENST00000514390.1 |
ANAPC10 |
anaphase promoting complex subunit 10 |
| chr7_+_23221613 | 1.49 |
ENST00000410002.3 ENST00000413919.1 |
NUPL2 |
nucleoporin like 2 |
| chr6_-_144416737 | 1.44 |
ENST00000367569.2 |
SF3B5 |
splicing factor 3b, subunit 5, 10kDa |
| chr22_+_20105012 | 1.44 |
ENST00000331821.3 ENST00000411892.1 |
RANBP1 |
RAN binding protein 1 |
| chr22_+_26879817 | 1.44 |
ENST00000215917.7 |
SRRD |
SRR1 domain containing |
| chr12_-_58240470 | 1.43 |
ENST00000548823.1 ENST00000398073.2 |
CTDSP2 |
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 2 |
| chr19_+_10362882 | 1.43 |
ENST00000393733.2 ENST00000588502.1 |
MRPL4 |
mitochondrial ribosomal protein L4 |
| chr11_+_34127142 | 1.43 |
ENST00000257829.3 ENST00000531159.2 |
NAT10 |
N-acetyltransferase 10 (GCN5-related) |
| chr7_+_23145366 | 1.39 |
ENST00000339077.5 ENST00000322275.5 ENST00000539124.1 ENST00000542558.1 |
KLHL7 |
kelch-like family member 7 |
| chr1_-_229644034 | 1.38 |
ENST00000366678.3 ENST00000261396.3 ENST00000537506.1 |
NUP133 |
nucleoporin 133kDa |
| chr19_+_10362577 | 1.37 |
ENST00000592514.1 ENST00000307422.5 ENST00000253099.6 ENST00000590150.1 ENST00000590669.1 |
MRPL4 |
mitochondrial ribosomal protein L4 |
| chr6_+_41888926 | 1.36 |
ENST00000230340.4 |
BYSL |
bystin-like |
| chr22_-_20104700 | 1.36 |
ENST00000439169.2 ENST00000445045.1 ENST00000404751.3 ENST00000252136.7 ENST00000403707.3 |
TRMT2A |
tRNA methyltransferase 2 homolog A (S. cerevisiae) |
| chr12_+_120884222 | 1.35 |
ENST00000551765.1 ENST00000229384.5 |
GATC |
glutamyl-tRNA(Gln) amidotransferase, subunit C |
| chr9_-_94877658 | 1.35 |
ENST00000262554.2 ENST00000337841.4 |
SPTLC1 |
serine palmitoyltransferase, long chain base subunit 1 |
| chr1_+_202976493 | 1.34 |
ENST00000367242.3 |
TMEM183A |
transmembrane protein 183A |
| chr7_-_150924121 | 1.34 |
ENST00000441774.1 ENST00000222388.2 ENST00000287844.2 |
ABCF2 |
ATP-binding cassette, sub-family F (GCN20), member 2 |
| chr19_+_19030478 | 1.33 |
ENST00000247003.4 |
DDX49 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 49 |
| chr14_+_61447927 | 1.32 |
ENST00000451406.1 |
SLC38A6 |
solute carrier family 38, member 6 |
| chr19_+_19030497 | 1.30 |
ENST00000438170.2 |
DDX49 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 49 |
| chrX_+_24711997 | 1.30 |
ENST00000379068.3 ENST00000379059.3 |
POLA1 |
polymerase (DNA directed), alpha 1, catalytic subunit |
| chr14_+_61447832 | 1.28 |
ENST00000354886.2 ENST00000267488.4 |
SLC38A6 |
solute carrier family 38, member 6 |
| chr19_-_5719860 | 1.27 |
ENST00000590729.1 |
LONP1 |
lon peptidase 1, mitochondrial |
| chr3_-_45017609 | 1.27 |
ENST00000342790.4 ENST00000424952.2 ENST00000296127.3 ENST00000455235.1 |
ZDHHC3 |
zinc finger, DHHC-type containing 3 |
| chr12_-_104531785 | 1.27 |
ENST00000551727.1 |
NFYB |
nuclear transcription factor Y, beta |
| chr6_+_37400974 | 1.24 |
ENST00000455891.1 ENST00000373451.4 |
CMTR1 |
cap methyltransferase 1 |
| chr1_-_220219775 | 1.24 |
ENST00000609181.1 |
EPRS |
glutamyl-prolyl-tRNA synthetase |
| chr6_+_20403997 | 1.22 |
ENST00000535432.1 |
E2F3 |
E2F transcription factor 3 |
| chr8_-_145743164 | 1.20 |
ENST00000428558.2 |
RECQL4 |
RecQ protein-like 4 |
| chr7_+_23221438 | 1.19 |
ENST00000258742.5 |
NUPL2 |
nucleoporin like 2 |
| chr10_+_70939983 | 1.19 |
ENST00000359655.4 ENST00000422378.1 |
SUPV3L1 |
suppressor of var1, 3-like 1 (S. cerevisiae) |
| chr19_+_54619125 | 1.17 |
ENST00000445811.1 ENST00000419967.1 ENST00000445124.1 ENST00000447810.1 |
PRPF31 |
pre-mRNA processing factor 31 |
| chr12_-_58165870 | 1.13 |
ENST00000257848.7 |
METTL1 |
methyltransferase like 1 |
| chr1_+_147400506 | 1.12 |
ENST00000314163.7 ENST00000468618.2 |
GPR89B |
G protein-coupled receptor 89B |
| chr6_-_34664612 | 1.12 |
ENST00000374023.3 ENST00000374026.3 |
C6orf106 |
chromosome 6 open reading frame 106 |
| chr2_-_68290106 | 1.12 |
ENST00000407324.1 ENST00000355848.3 ENST00000409302.1 ENST00000410067.3 |
C1D |
C1D nuclear receptor corepressor |
| chr7_+_102036798 | 1.11 |
ENST00000397912.3 ENST00000354783.4 |
PRKRIP1 |
PRKR interacting protein 1 (IL11 inducible) |
| chr4_-_146019287 | 1.10 |
ENST00000502847.1 ENST00000513054.1 |
ANAPC10 |
anaphase promoting complex subunit 10 |
| chr11_+_18343800 | 1.09 |
ENST00000453096.2 |
GTF2H1 |
general transcription factor IIH, polypeptide 1, 62kDa |
| chr12_-_104532062 | 1.08 |
ENST00000240055.3 |
NFYB |
nuclear transcription factor Y, beta |
| chr14_-_23388338 | 1.05 |
ENST00000555209.1 ENST00000554256.1 ENST00000557403.1 ENST00000557549.1 ENST00000555676.1 ENST00000557571.1 ENST00000557464.1 ENST00000554618.1 ENST00000556862.1 ENST00000555722.1 ENST00000346528.5 ENST00000542016.2 ENST00000399922.2 ENST00000557227.1 ENST00000359890.3 |
RBM23 |
RNA binding motif protein 23 |
| chr4_-_76439483 | 1.04 |
ENST00000380840.2 ENST00000513257.1 ENST00000507014.1 |
RCHY1 |
ring finger and CHY zinc finger domain containing 1, E3 ubiquitin protein ligase |
| chr1_-_202896310 | 1.04 |
ENST00000367261.3 |
KLHL12 |
kelch-like family member 12 |
| chr1_-_145827015 | 1.02 |
ENST00000534502.1 ENST00000313835.9 ENST00000454423.3 |
GPR89A |
G protein-coupled receptor 89A |
| chr4_+_122722466 | 1.02 |
ENST00000243498.5 ENST00000379663.3 ENST00000509800.1 |
EXOSC9 |
exosome component 9 |
| chr16_+_19079215 | 1.01 |
ENST00000544894.2 ENST00000561858.1 |
COQ7 |
coenzyme Q7 homolog, ubiquinone (yeast) |
| chr19_+_36103631 | 1.00 |
ENST00000203166.5 ENST00000379045.2 |
HAUS5 |
HAUS augmin-like complex, subunit 5 |
| chr2_-_242626127 | 1.00 |
ENST00000445261.1 |
DTYMK |
deoxythymidylate kinase (thymidylate kinase) |
| chr12_-_132628847 | 1.00 |
ENST00000397333.3 |
DDX51 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 51 |
| chr1_-_115259337 | 0.99 |
ENST00000369535.4 |
NRAS |
neuroblastoma RAS viral (v-ras) oncogene homolog |
| chr9_-_134955246 | 0.99 |
ENST00000357028.2 ENST00000474263.1 ENST00000292035.5 |
MED27 |
mediator complex subunit 27 |
| chr21_-_44299626 | 0.98 |
ENST00000330317.2 ENST00000398208.2 |
WDR4 |
WD repeat domain 4 |
| chrX_+_24072833 | 0.97 |
ENST00000253039.4 |
EIF2S3 |
eukaryotic translation initiation factor 2, subunit 3 gamma, 52kDa |
| chr5_-_132113036 | 0.96 |
ENST00000378706.1 |
SEPT8 |
septin 8 |
| chr8_-_17104356 | 0.96 |
ENST00000361272.4 ENST00000523917.1 |
CNOT7 |
CCR4-NOT transcription complex, subunit 7 |
| chr17_-_685559 | 0.96 |
ENST00000301329.6 |
GLOD4 |
glyoxalase domain containing 4 |
| chr7_+_44646218 | 0.93 |
ENST00000444676.1 ENST00000222673.5 |
OGDH |
oxoglutarate (alpha-ketoglutarate) dehydrogenase (lipoamide) |
| chr11_+_67195917 | 0.93 |
ENST00000524934.1 ENST00000539188.1 ENST00000312629.5 |
RPS6KB2 |
ribosomal protein S6 kinase, 70kDa, polypeptide 2 |
| chr1_-_220220000 | 0.93 |
ENST00000366923.3 |
EPRS |
glutamyl-prolyl-tRNA synthetase |
| chr16_+_70380732 | 0.92 |
ENST00000302243.7 |
DDX19A |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 19A |
| chr1_-_109940550 | 0.91 |
ENST00000256637.6 |
SORT1 |
sortilin 1 |
| chr11_+_57480046 | 0.90 |
ENST00000378312.4 ENST00000278422.4 |
TMX2 |
thioredoxin-related transmembrane protein 2 |
| chr11_-_64889649 | 0.90 |
ENST00000434372.2 |
FAU |
Finkel-Biskis-Reilly murine sarcoma virus (FBR-MuSV) ubiquitously expressed |
| chr5_-_132113083 | 0.90 |
ENST00000296873.7 |
SEPT8 |
septin 8 |
| chr4_-_146019335 | 0.89 |
ENST00000451299.2 ENST00000507656.1 ENST00000309439.5 |
ANAPC10 |
anaphase promoting complex subunit 10 |
| chr6_-_41888843 | 0.89 |
ENST00000434077.1 ENST00000409312.1 |
MED20 |
mediator complex subunit 20 |
| chr17_+_685513 | 0.89 |
ENST00000304478.4 |
RNMTL1 |
RNA methyltransferase like 1 |
| chr3_-_196669298 | 0.87 |
ENST00000411704.1 ENST00000452404.2 |
NCBP2 |
nuclear cap binding protein subunit 2, 20kDa |
| chr14_+_105219437 | 0.86 |
ENST00000329967.6 ENST00000347067.5 ENST00000553810.1 |
SIVA1 |
SIVA1, apoptosis-inducing factor |
| chr6_-_31926208 | 0.86 |
ENST00000454913.1 ENST00000436289.2 |
NELFE |
negative elongation factor complex member E |
| chr4_-_170533723 | 0.85 |
ENST00000510533.1 ENST00000439128.2 ENST00000511633.1 ENST00000512193.1 ENST00000507142.1 |
NEK1 |
NIMA-related kinase 1 |
| chr13_+_103249322 | 0.85 |
ENST00000376065.4 ENST00000376052.3 |
TPP2 |
tripeptidyl peptidase II |
| chr16_+_19078960 | 0.84 |
ENST00000568985.1 ENST00000566110.1 |
COQ7 |
coenzyme Q7 homolog, ubiquinone (yeast) |
| chr3_-_186288097 | 0.83 |
ENST00000446782.1 |
TBCCD1 |
TBCC domain containing 1 |
| chr2_+_109335929 | 0.80 |
ENST00000283195.6 |
RANBP2 |
RAN binding protein 2 |
| chr9_-_32573130 | 0.80 |
ENST00000350021.2 ENST00000379847.3 |
NDUFB6 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 6, 17kDa |
| chr8_+_101162812 | 0.78 |
ENST00000353107.3 ENST00000522439.1 |
POLR2K |
polymerase (RNA) II (DNA directed) polypeptide K, 7.0kDa |
| chr11_+_18344106 | 0.78 |
ENST00000534641.1 ENST00000525831.1 ENST00000265963.4 |
GTF2H1 |
general transcription factor IIH, polypeptide 1, 62kDa |
| chr11_-_62572901 | 0.78 |
ENST00000439713.2 ENST00000531131.1 ENST00000530875.1 ENST00000531709.2 ENST00000294172.2 |
NXF1 |
nuclear RNA export factor 1 |
| chr17_-_685493 | 0.77 |
ENST00000536578.1 ENST00000301328.5 ENST00000576419.1 |
GLOD4 |
glyoxalase domain containing 4 |
| chr2_-_29093132 | 0.77 |
ENST00000306108.5 |
TRMT61B |
tRNA methyltransferase 61 homolog B (S. cerevisiae) |
| chr8_+_133787586 | 0.77 |
ENST00000395379.1 ENST00000395386.2 ENST00000337920.4 |
PHF20L1 |
PHD finger protein 20-like 1 |
| chr16_-_30006922 | 0.74 |
ENST00000564026.1 |
HIRIP3 |
HIRA interacting protein 3 |
| chr21_-_44527613 | 0.72 |
ENST00000380276.2 ENST00000398137.1 ENST00000291552.4 |
U2AF1 |
U2 small nuclear RNA auxiliary factor 1 |
| chr15_-_66790047 | 0.72 |
ENST00000566658.1 ENST00000563480.2 ENST00000395589.2 ENST00000307979.7 |
SNAPC5 |
small nuclear RNA activating complex, polypeptide 5, 19kDa |
| chr9_+_70856899 | 0.72 |
ENST00000377342.5 ENST00000478048.1 |
CBWD3 |
COBW domain containing 3 |
| chr16_+_81348528 | 0.72 |
ENST00000568107.2 |
GAN |
gigaxonin |
| chr22_-_19132154 | 0.71 |
ENST00000252137.6 |
DGCR14 |
DiGeorge syndrome critical region gene 14 |
| chr1_+_213031570 | 0.70 |
ENST00000366971.4 |
FLVCR1 |
feline leukemia virus subgroup C cellular receptor 1 |
| chr12_+_69633317 | 0.70 |
ENST00000435070.2 |
CPSF6 |
cleavage and polyadenylation specific factor 6, 68kDa |
| chr12_+_69633407 | 0.70 |
ENST00000551516.1 |
CPSF6 |
cleavage and polyadenylation specific factor 6, 68kDa |
| chr3_+_141457030 | 0.69 |
ENST00000273480.3 |
RNF7 |
ring finger protein 7 |
| chr1_+_16085244 | 0.69 |
ENST00000400773.1 |
FBLIM1 |
filamin binding LIM protein 1 |
| chr5_-_132112921 | 0.68 |
ENST00000378721.4 ENST00000378701.1 |
SEPT8 |
septin 8 |
| chr18_+_268148 | 0.66 |
ENST00000581677.1 |
RP11-705O1.8 |
RP11-705O1.8 |
| chr2_+_101618965 | 0.66 |
ENST00000456292.1 ENST00000409711.1 ENST00000441435.1 |
RPL31 |
ribosomal protein L31 |
| chr11_-_66112555 | 0.66 |
ENST00000425825.2 ENST00000359957.3 |
BRMS1 |
breast cancer metastasis suppressor 1 |
| chr6_+_41755389 | 0.65 |
ENST00000398884.3 ENST00000398881.3 |
TOMM6 |
translocase of outer mitochondrial membrane 6 homolog (yeast) |
| chr22_+_41865109 | 0.64 |
ENST00000216254.4 ENST00000396512.3 |
ACO2 |
aconitase 2, mitochondrial |
| chr5_-_132112907 | 0.64 |
ENST00000458488.2 |
SEPT8 |
septin 8 |
| chr16_+_2022036 | 0.64 |
ENST00000568546.1 |
TBL3 |
transducin (beta)-like 3 |
| chr5_-_74807418 | 0.63 |
ENST00000405807.4 ENST00000261415.7 |
COL4A3BP |
collagen, type IV, alpha 3 (Goodpasture antigen) binding protein |
| chrX_-_153285395 | 0.63 |
ENST00000369980.3 |
IRAK1 |
interleukin-1 receptor-associated kinase 1 |
| chr9_-_33001520 | 0.63 |
ENST00000463596.1 ENST00000379819.1 ENST00000397172.3 ENST00000379812.5 ENST00000477119.1 ENST00000379813.3 ENST00000379825.2 ENST00000309615.3 ENST00000476858.1 ENST00000473221.1 |
APTX |
aprataxin |
| chr9_-_69262509 | 0.62 |
ENST00000377449.1 ENST00000382399.4 ENST00000377439.1 ENST00000377441.1 ENST00000377457.5 |
CBWD6 |
COBW domain containing 6 |
| chr19_+_58341656 | 0.62 |
ENST00000442832.4 ENST00000594901.1 |
ZNF587B |
zinc finger protein 587B |
| chr15_-_34394008 | 0.62 |
ENST00000527822.1 ENST00000528949.1 |
EMC7 |
ER membrane protein complex subunit 7 |
| chr9_-_179018 | 0.62 |
ENST00000431099.2 ENST00000382447.4 ENST00000382389.1 ENST00000377447.3 ENST00000314367.10 ENST00000356521.4 ENST00000382393.1 ENST00000377400.4 |
CBWD1 |
COBW domain containing 1 |
| chr12_-_120884175 | 0.61 |
ENST00000546954.1 |
TRIAP1 |
TP53 regulated inhibitor of apoptosis 1 |
| chr15_-_34394119 | 0.61 |
ENST00000256545.4 |
EMC7 |
ER membrane protein complex subunit 7 |
| chr12_-_124118296 | 0.60 |
ENST00000424014.2 ENST00000537073.1 |
EIF2B1 |
eukaryotic translation initiation factor 2B, subunit 1 alpha, 26kDa |
| chr4_-_74124502 | 0.59 |
ENST00000358602.4 ENST00000330838.6 ENST00000561029.1 |
ANKRD17 |
ankyrin repeat domain 17 |
| chrX_+_135579238 | 0.58 |
ENST00000535601.1 ENST00000448450.1 ENST00000425695.1 |
HTATSF1 |
HIV-1 Tat specific factor 1 |
| chr11_-_64889529 | 0.58 |
ENST00000531743.1 ENST00000527548.1 ENST00000526555.1 ENST00000279259.3 |
FAU |
Finkel-Biskis-Reilly murine sarcoma virus (FBR-MuSV) ubiquitously expressed |
| chr6_-_41888814 | 0.57 |
ENST00000409060.1 ENST00000265350.4 |
MED20 |
mediator complex subunit 20 |
| chr6_-_90062543 | 0.57 |
ENST00000435041.2 |
UBE2J1 |
ubiquitin-conjugating enzyme E2, J1 |
| chr2_+_135676381 | 0.56 |
ENST00000537343.1 ENST00000295238.6 ENST00000264157.5 |
CCNT2 |
cyclin T2 |
| chr16_-_20753114 | 0.55 |
ENST00000396083.2 |
THUMPD1 |
THUMP domain containing 1 |
| chr21_+_45079409 | 0.55 |
ENST00000340648.4 |
RRP1B |
ribosomal RNA processing 1B |
| chr3_+_141457105 | 0.54 |
ENST00000480908.1 ENST00000393000.3 |
RNF7 |
ring finger protein 7 |
| chr12_-_46384334 | 0.53 |
ENST00000369367.3 ENST00000266589.6 ENST00000395453.2 ENST00000395454.2 |
SCAF11 |
SR-related CTD-associated factor 11 |
| chr15_-_42840961 | 0.53 |
ENST00000563454.1 ENST00000397130.3 ENST00000570160.1 ENST00000323443.2 |
LRRC57 |
leucine rich repeat containing 57 |
| chr1_-_213031418 | 0.52 |
ENST00000356684.3 ENST00000426161.1 ENST00000424044.1 |
FLVCR1-AS1 |
FLVCR1 antisense RNA 1 (head to head) |
| chr16_+_29827285 | 0.52 |
ENST00000320330.6 |
PAGR1 |
PAXIP1 associated glutamate-rich protein 1 |
| chr14_+_23776024 | 0.52 |
ENST00000553781.1 ENST00000556100.1 ENST00000557236.1 ENST00000557579.1 |
BCL2L2-PABPN1 BCL2L2 |
BCL2L2-PABPN1 readthrough BCL2-like 2 |
| chr16_+_29827832 | 0.52 |
ENST00000609618.1 |
PAGR1 |
PAXIP1-associated glutamate-rich protein 1 |
| chr19_-_46234119 | 0.52 |
ENST00000317683.3 |
FBXO46 |
F-box protein 46 |
| chrX_-_47518527 | 0.51 |
ENST00000333119.3 |
UXT |
ubiquitously-expressed, prefoldin-like chaperone |
| chr7_-_19748640 | 0.50 |
ENST00000222567.5 |
TWISTNB |
TWIST neighbor |
| chr2_-_130939115 | 0.49 |
ENST00000441135.1 ENST00000339679.7 ENST00000426662.2 ENST00000443958.2 ENST00000351288.6 ENST00000453750.1 ENST00000452225.2 |
SMPD4 |
sphingomyelin phosphodiesterase 4, neutral membrane (neutral sphingomyelinase-3) |
| chr16_+_1359511 | 0.49 |
ENST00000397514.3 ENST00000397515.2 ENST00000567383.1 ENST00000403747.2 ENST00000566587.1 |
UBE2I |
ubiquitin-conjugating enzyme E2I |
| chr6_-_106773610 | 0.48 |
ENST00000369076.3 ENST00000369070.1 |
ATG5 |
autophagy related 5 |
| chr9_+_70856397 | 0.48 |
ENST00000360171.6 |
CBWD3 |
COBW domain containing 3 |
| chr16_+_19078911 | 0.48 |
ENST00000321998.5 |
COQ7 |
coenzyme Q7 homolog, ubiquinone (yeast) |
| chrX_-_153285251 | 0.48 |
ENST00000444230.1 ENST00000393682.1 ENST00000393687.2 ENST00000429936.2 ENST00000369974.2 |
IRAK1 |
interleukin-1 receptor-associated kinase 1 |
| chr1_-_29557383 | 0.46 |
ENST00000373791.3 ENST00000263702.6 |
MECR |
mitochondrial trans-2-enoyl-CoA reductase |
| chr11_+_64889773 | 0.46 |
ENST00000534078.1 ENST00000526171.1 ENST00000279242.2 ENST00000531705.1 ENST00000533943.1 |
MRPL49 |
mitochondrial ribosomal protein L49 |
| chr17_-_36981556 | 0.45 |
ENST00000536127.1 ENST00000225428.5 |
CWC25 |
CWC25 spliceosome-associated protein homolog (S. cerevisiae) |
| chr16_-_2318373 | 0.44 |
ENST00000566458.1 ENST00000320225.5 |
RNPS1 |
RNA binding protein S1, serine-rich domain |
| chr15_-_23034322 | 0.44 |
ENST00000539711.2 ENST00000560039.1 ENST00000398013.3 ENST00000337451.3 ENST00000359727.4 ENST00000398014.2 |
NIPA2 |
non imprinted in Prader-Willi/Angelman syndrome 2 |
| chr3_+_11314099 | 0.43 |
ENST00000446450.2 ENST00000354956.5 ENST00000354449.3 ENST00000419112.1 |
ATG7 |
autophagy related 7 |
| chr19_+_36120009 | 0.43 |
ENST00000589871.1 |
RBM42 |
RNA binding motif protein 42 |
| chr17_-_49198095 | 0.42 |
ENST00000505279.1 |
SPAG9 |
sperm associated antigen 9 |
| chr14_+_23776167 | 0.42 |
ENST00000554635.1 ENST00000557008.1 |
BCL2L2 BCL2L2-PABPN1 |
BCL2-like 2 BCL2L2-PABPN1 readthrough |
| chr12_+_69633372 | 0.41 |
ENST00000456847.3 ENST00000266679.8 |
CPSF6 |
cleavage and polyadenylation specific factor 6, 68kDa |
| chr17_-_72869086 | 0.41 |
ENST00000581530.1 ENST00000420580.2 ENST00000455107.2 ENST00000413947.2 ENST00000581219.1 ENST00000582944.1 |
FDXR |
ferredoxin reductase |
| chr11_-_10562710 | 0.41 |
ENST00000528665.1 ENST00000265981.2 |
RNF141 |
ring finger protein 141 |
| chr19_+_36119975 | 0.41 |
ENST00000589559.1 ENST00000360475.4 |
RBM42 |
RNA binding motif protein 42 |
| chr1_-_36107445 | 0.40 |
ENST00000373237.3 |
PSMB2 |
proteasome (prosome, macropain) subunit, beta type, 2 |
| chr2_+_114195268 | 0.39 |
ENST00000259199.4 ENST00000416503.2 ENST00000433343.2 |
CBWD2 |
COBW domain containing 2 |
| chr9_-_70490107 | 0.39 |
ENST00000377395.4 ENST00000429800.2 ENST00000430059.2 ENST00000377384.1 ENST00000382405.3 |
CBWD5 |
COBW domain containing 5 |
| chr19_-_5720248 | 0.39 |
ENST00000360614.3 |
LONP1 |
lon peptidase 1, mitochondrial |
| chr16_+_70380825 | 0.38 |
ENST00000417604.2 |
DDX19A |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 19A |
| chr3_-_52443799 | 0.38 |
ENST00000470173.1 ENST00000296288.5 |
BAP1 |
BRCA1 associated protein-1 (ubiquitin carboxy-terminal hydrolase) |
| chr9_+_140083099 | 0.38 |
ENST00000322310.5 |
SSNA1 |
Sjogren syndrome nuclear autoantigen 1 |
| chr16_-_2318055 | 0.37 |
ENST00000561518.1 ENST00000561718.1 ENST00000567147.1 ENST00000562690.1 ENST00000569598.2 |
RNPS1 |
RNA binding protein S1, serine-rich domain |
| chr8_+_145743360 | 0.37 |
ENST00000527730.1 ENST00000529022.1 ENST00000292524.1 |
LRRC14 |
leucine rich repeat containing 14 |
| chr1_+_186344883 | 0.36 |
ENST00000367470.3 |
C1orf27 |
chromosome 1 open reading frame 27 |
| chr17_-_8151353 | 0.36 |
ENST00000315684.8 |
CTC1 |
CTS telomere maintenance complex component 1 |
| chr11_-_64889252 | 0.36 |
ENST00000525297.1 ENST00000529259.1 |
FAU |
Finkel-Biskis-Reilly murine sarcoma virus (FBR-MuSV) ubiquitously expressed |
| chr16_+_68057179 | 0.36 |
ENST00000567100.1 ENST00000432752.1 ENST00000569289.1 ENST00000564781.1 |
DUS2 |
dihydrouridine synthase 2 |
| chr17_-_73663168 | 0.35 |
ENST00000578201.1 ENST00000423245.2 |
RECQL5 |
RecQ protein-like 5 |
| chr20_+_55043647 | 0.35 |
ENST00000023939.4 ENST00000395881.3 ENST00000357348.5 ENST00000449062.1 ENST00000435342.2 |
RTFDC1 |
replication termination factor 2 domain containing 1 |
| chrX_+_84498989 | 0.35 |
ENST00000395402.1 |
ZNF711 |
zinc finger protein 711 |
| chr1_+_186344945 | 0.35 |
ENST00000419367.3 ENST00000287859.6 |
C1orf27 |
chromosome 1 open reading frame 27 |
| chr16_+_68056844 | 0.35 |
ENST00000565263.1 |
DUS2 |
dihydrouridine synthase 2 |
| chr2_+_96068436 | 0.34 |
ENST00000445649.1 ENST00000447036.1 ENST00000233379.4 ENST00000418606.1 |
FAHD2A |
fumarylacetoacetate hydrolase domain containing 2A |
| chr22_+_50946645 | 0.33 |
ENST00000420993.2 ENST00000395698.3 ENST00000395701.3 ENST00000523045.1 ENST00000299821.11 |
NCAPH2 |
non-SMC condensin II complex, subunit H2 |
| chr17_-_72869140 | 0.33 |
ENST00000583917.1 ENST00000293195.5 ENST00000442102.2 |
FDXR |
ferredoxin reductase |
| chr16_+_68057153 | 0.32 |
ENST00000358896.6 ENST00000568099.2 |
DUS2 |
dihydrouridine synthase 2 |
| chrX_+_48660287 | 0.32 |
ENST00000444343.2 ENST00000376610.2 ENST00000334136.5 ENST00000376619.2 |
HDAC6 |
histone deacetylase 6 |
| chr6_-_114292284 | 0.32 |
ENST00000520895.1 ENST00000521163.1 ENST00000524334.1 ENST00000368632.2 ENST00000398283.2 |
HDAC2 |
histone deacetylase 2 |
| chr2_-_110962544 | 0.32 |
ENST00000355301.4 ENST00000445609.2 ENST00000417665.1 ENST00000418527.1 ENST00000316534.4 ENST00000393272.3 |
NPHP1 |
nephronophthisis 1 (juvenile) |
| chr11_+_66610883 | 0.32 |
ENST00000309657.3 ENST00000524506.1 |
RCE1 |
Ras converting CAAX endopeptidase 1 |
| chr8_-_133493200 | 0.32 |
ENST00000388996.4 |
KCNQ3 |
potassium voltage-gated channel, KQT-like subfamily, member 3 |
| chr11_+_60609537 | 0.31 |
ENST00000227520.5 |
CCDC86 |
coiled-coil domain containing 86 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 3.6 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.2 | 8.9 | REACTOME TRANSPORT OF MATURE MRNA DERIVED FROM AN INTRONLESS TRANSCRIPT | Genes involved in Transport of Mature mRNA Derived from an Intronless Transcript |
| 0.2 | 2.4 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.2 | 2.4 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.1 | 4.1 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.1 | 2.0 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.1 | 1.2 | REACTOME CDC6 ASSOCIATION WITH THE ORC ORIGIN COMPLEX | Genes involved in CDC6 association with the ORC:origin complex |
| 0.1 | 2.6 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.1 | 1.1 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.1 | 1.3 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.1 | 4.2 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.1 | 3.1 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.1 | 1.0 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.1 | 2.2 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.1 | 1.4 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 2.0 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.0 | 1.8 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 0.9 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.2 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.0 | 1.0 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 1.0 | REACTOME DEADENYLATION DEPENDENT MRNA DECAY | Genes involved in Deadenylation-dependent mRNA decay |
| 0.0 | 2.2 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 2.7 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.7 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.9 | REACTOME PIP3 ACTIVATES AKT SIGNALING | Genes involved in PIP3 activates AKT signaling |
| 0.0 | 0.8 | REACTOME IRON UPTAKE AND TRANSPORT | Genes involved in Iron uptake and transport |
| 0.0 | 1.1 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.3 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 0.6 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 1.1 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.7 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.9 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.5 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 6.3 | GO:0070283 | lipoate synthase activity(GO:0016992) radical SAM enzyme activity(GO:0070283) |
| 1.2 | 4.7 | GO:0070137 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.6 | 1.7 | GO:0070361 | mitochondrial light strand promoter anti-sense binding(GO:0070361) mitochondrial heavy strand promoter anti-sense binding(GO:0070362) mitochondrial heavy strand promoter sense binding(GO:0070364) |
| 0.4 | 1.3 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.4 | 4.9 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.4 | 3.4 | GO:0008474 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.3 | 0.9 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.3 | 0.9 | GO:0016426 | tRNA (adenine) methyltransferase activity(GO:0016426) tRNA (adenine-N1-)-methyltransferase activity(GO:0016429) |
| 0.3 | 1.4 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.2 | 1.2 | GO:0008174 | mRNA methyltransferase activity(GO:0008174) |
| 0.2 | 0.7 | GO:0004324 | ferredoxin-NADP+ reductase activity(GO:0004324) NADPH-adrenodoxin reductase activity(GO:0015039) oxidoreductase activity, acting on iron-sulfur proteins as donors(GO:0016730) oxidoreductase activity, acting on iron-sulfur proteins as donors, NAD or NADP as acceptor(GO:0016731) |
| 0.2 | 3.3 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.2 | 1.3 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 0.2 | 0.6 | GO:0051538 | 3 iron, 4 sulfur cluster binding(GO:0051538) |
| 0.2 | 2.4 | GO:0017070 | U6 snRNA binding(GO:0017070) |
| 0.2 | 0.9 | GO:0010465 | nerve growth factor receptor activity(GO:0010465) |
| 0.2 | 2.1 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.2 | 1.0 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.2 | 1.2 | GO:0030621 | U4 snRNA binding(GO:0030621) |
| 0.2 | 1.0 | GO:0050145 | nucleoside phosphate kinase activity(GO:0050145) |
| 0.2 | 0.6 | GO:0097001 | ceramide binding(GO:0097001) |
| 0.2 | 1.1 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.2 | 0.5 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 0.2 | 1.2 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.2 | 1.2 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.1 | 0.4 | GO:0034353 | RNA pyrophosphohydrolase activity(GO:0034353) |
| 0.1 | 0.7 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.1 | 0.8 | GO:0008240 | tripeptidyl-peptidase activity(GO:0008240) |
| 0.1 | 2.0 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.1 | 4.5 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.1 | 2.2 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.1 | 5.6 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 0.7 | GO:0019776 | Atg8 ligase activity(GO:0019776) |
| 0.1 | 0.6 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.1 | 1.3 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.1 | 2.0 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.1 | 0.3 | GO:0042903 | tubulin deacetylase activity(GO:0042903) |
| 0.1 | 0.6 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 0.1 | 5.2 | GO:0004004 | ATP-dependent RNA helicase activity(GO:0004004) |
| 0.1 | 1.3 | GO:0008408 | 3'-5' exonuclease activity(GO:0008408) |
| 0.1 | 2.0 | GO:0005487 | nucleocytoplasmic transporter activity(GO:0005487) |
| 0.1 | 0.3 | GO:0043398 | HLH domain binding(GO:0043398) |
| 0.1 | 0.4 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.1 | 1.3 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.1 | 0.7 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.1 | 2.1 | GO:0008308 | voltage-gated anion channel activity(GO:0008308) |
| 0.1 | 0.2 | GO:0032093 | SAM domain binding(GO:0032093) |
| 0.1 | 1.5 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 0.7 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.1 | 0.2 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.1 | 0.8 | GO:0019789 | SUMO transferase activity(GO:0019789) |
| 0.0 | 1.3 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.0 | 0.5 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.2 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.0 | 1.5 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) |
| 0.0 | 0.2 | GO:0016802 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 1.1 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.4 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 1.4 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.0 | 0.4 | GO:0043047 | single-stranded telomeric DNA binding(GO:0043047) G-rich strand telomeric DNA binding(GO:0098505) |
| 0.0 | 0.7 | GO:0031005 | filamin binding(GO:0031005) |
| 0.0 | 0.3 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 2.6 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.0 | 0.8 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.0 | 2.4 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.0 | 0.9 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.0 | 0.6 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 0.1 | GO:0030366 | molybdopterin synthase activity(GO:0030366) |
| 0.0 | 0.7 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.0 | 5.1 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 1.6 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.1 | GO:0008823 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.0 | 1.8 | GO:0002039 | p53 binding(GO:0002039) |
| 0.0 | 1.1 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 2.1 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.0 | 4.2 | GO:0061659 | ubiquitin protein ligase activity(GO:0061630) ubiquitin-like protein ligase activity(GO:0061659) |
| 0.0 | 0.3 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.0 | 3.4 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 0.8 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.0 | 0.1 | GO:0030911 | TPR domain binding(GO:0030911) |
| 0.0 | 1.1 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.0 | 2.1 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 5.3 | GO:0005525 | GTP binding(GO:0005525) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 3.4 | GO:0030689 | Noc complex(GO:0030689) |
| 0.7 | 2.2 | GO:1990876 | cytoplasmic side of nuclear pore(GO:1990876) |
| 0.6 | 3.0 | GO:0043527 | tRNA methyltransferase complex(GO:0043527) |
| 0.4 | 4.7 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.4 | 1.2 | GO:0043224 | nuclear SCF ubiquitin ligase complex(GO:0043224) |
| 0.4 | 2.7 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.3 | 1.3 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.3 | 2.4 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.3 | 1.0 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.3 | 2.2 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.3 | 1.2 | GO:0005846 | nuclear cap binding complex(GO:0005846) |
| 0.2 | 1.0 | GO:0044094 | host cell nucleus(GO:0042025) host cell nuclear part(GO:0044094) |
| 0.2 | 1.8 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.2 | 1.1 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 0.2 | 2.2 | GO:0000439 | core TFIIH complex(GO:0000439) |
| 0.2 | 1.3 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.2 | 2.4 | GO:0016602 | CCAAT-binding factor complex(GO:0016602) |
| 0.2 | 6.8 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.2 | 0.8 | GO:0032021 | NELF complex(GO:0032021) |
| 0.1 | 0.7 | GO:0089701 | U2AF(GO:0089701) |
| 0.1 | 1.2 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.1 | 1.2 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.1 | 0.8 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.1 | 0.6 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.1 | 1.2 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.1 | 3.2 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.1 | 1.4 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.1 | 1.3 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 0.1 | 0.8 | GO:0000346 | transcription export complex(GO:0000346) |
| 0.1 | 0.7 | GO:0034274 | Atg12-Atg5-Atg16 complex(GO:0034274) |
| 0.1 | 2.6 | GO:0071011 | precatalytic spliceosome(GO:0071011) |
| 0.1 | 1.0 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.1 | 0.6 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.1 | 0.5 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.1 | 2.6 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.1 | 1.4 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.1 | 1.7 | GO:0031306 | intrinsic component of mitochondrial outer membrane(GO:0031306) |
| 0.1 | 0.6 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.1 | 5.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 2.7 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 8.3 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.1 | 2.0 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.1 | 1.3 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.1 | 6.3 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.1 | 0.6 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.0 | 0.8 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 2.0 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 0.6 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 1.0 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.0 | 0.9 | GO:0000242 | pericentriolar material(GO:0000242) |
| 0.0 | 0.2 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.0 | 1.7 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 1.1 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.0 | 0.2 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.0 | 2.8 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.3 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.0 | 2.0 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.0 | 0.4 | GO:0019774 | proteasome core complex, beta-subunit complex(GO:0019774) |
| 0.0 | 0.3 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 3.4 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.0 | 0.1 | GO:0031085 | BLOC-3 complex(GO:0031085) |
| 0.0 | 0.1 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.0 | 0.4 | GO:0000782 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) |
| 0.0 | 8.0 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 0.3 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 0.3 | GO:0036020 | endolysosome membrane(GO:0036020) |
| 0.0 | 0.8 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 1.4 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 0.3 | GO:0000795 | synaptonemal complex(GO:0000795) |
| 0.0 | 1.0 | GO:0070821 | tertiary granule membrane(GO:0070821) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.9 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.1 | 3.7 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 0.8 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.1 | 0.9 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.1 | 1.0 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 1.0 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 1.1 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 5.9 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 0.6 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 6.3 | GO:0009107 | lipoate biosynthetic process(GO:0009107) |
| 1.2 | 4.7 | GO:0009386 | translational attenuation(GO:0009386) |
| 0.9 | 3.6 | GO:0034476 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 0.7 | 2.7 | GO:0045829 | negative regulation of isotype switching(GO:0045829) |
| 0.6 | 4.7 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.6 | 1.7 | GO:0070407 | oxidation-dependent protein catabolic process(GO:0070407) |
| 0.5 | 1.4 | GO:0002101 | tRNA wobble cytosine modification(GO:0002101) |
| 0.4 | 1.3 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.4 | 0.4 | GO:1902534 | single-organism membrane invagination(GO:1902534) |
| 0.4 | 3.4 | GO:0098734 | macromolecule depalmitoylation(GO:0098734) |
| 0.4 | 1.2 | GO:0000965 | mitochondrial RNA 3'-end processing(GO:0000965) |
| 0.3 | 1.4 | GO:0031081 | nuclear pore distribution(GO:0031081) nuclear pore localization(GO:0051664) |
| 0.3 | 1.3 | GO:1904502 | regulation of lipophagy(GO:1904502) positive regulation of lipophagy(GO:1904504) |
| 0.3 | 1.2 | GO:0046833 | snRNA export from nucleus(GO:0006408) positive regulation of RNA export from nucleus(GO:0046833) |
| 0.3 | 2.1 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 0.3 | 2.1 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.3 | 2.3 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 0.3 | 0.8 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.3 | 0.8 | GO:0070901 | mitochondrial tRNA methylation(GO:0070901) |
| 0.2 | 1.0 | GO:0009189 | dTTP biosynthetic process(GO:0006235) deoxyribonucleoside diphosphate biosynthetic process(GO:0009189) pyrimidine deoxyribonucleoside triphosphate biosynthetic process(GO:0009212) |
| 0.2 | 1.2 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.2 | 0.7 | GO:0039689 | negative stranded viral RNA replication(GO:0039689) multi-organism biosynthetic process(GO:0044034) |
| 0.2 | 0.6 | GO:0035621 | ER to Golgi ceramide transport(GO:0035621) |
| 0.2 | 2.0 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.2 | 1.2 | GO:0048254 | snoRNA localization(GO:0048254) |
| 0.2 | 2.1 | GO:0061734 | parkin-mediated mitophagy in response to mitochondrial depolarization(GO:0061734) |
| 0.2 | 3.3 | GO:0006744 | ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.2 | 1.0 | GO:0002943 | tRNA dihydrouridine synthesis(GO:0002943) |
| 0.1 | 0.6 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.1 | 5.6 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.1 | 0.6 | GO:0019086 | late viral transcription(GO:0019086) |
| 0.1 | 1.0 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.1 | 0.6 | GO:0035900 | response to isolation stress(GO:0035900) |
| 0.1 | 0.4 | GO:0035521 | monoubiquitinated histone deubiquitination(GO:0035521) monoubiquitinated histone H2A deubiquitination(GO:0035522) |
| 0.1 | 1.2 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.1 | 0.4 | GO:0033140 | negative regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033140) |
| 0.1 | 1.2 | GO:0000733 | DNA strand renaturation(GO:0000733) |
| 0.1 | 4.7 | GO:0071431 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.1 | 0.3 | GO:1903348 | positive regulation of bicellular tight junction assembly(GO:1903348) |
| 0.1 | 2.0 | GO:0043923 | positive regulation by host of viral transcription(GO:0043923) |
| 0.1 | 0.7 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.1 | 2.0 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.1 | 1.4 | GO:0001510 | RNA methylation(GO:0001510) |
| 0.1 | 0.4 | GO:0060830 | ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.1 | 6.8 | GO:0070126 | mitochondrial translational elongation(GO:0070125) mitochondrial translational termination(GO:0070126) |
| 0.1 | 0.9 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.1 | 0.4 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.1 | 4.8 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.1 | 0.8 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.1 | 0.6 | GO:0070862 | negative regulation of protein exit from endoplasmic reticulum(GO:0070862) negative regulation of retrograde protein transport, ER to cytosol(GO:1904153) |
| 0.1 | 1.3 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 0.2 | GO:0061034 | olfactory bulb mitral cell layer development(GO:0061034) |
| 0.1 | 1.2 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.1 | 1.0 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.1 | 0.2 | GO:1901491 | negative regulation of lymphangiogenesis(GO:1901491) |
| 0.1 | 0.2 | GO:0009304 | tRNA transcription(GO:0009304) 5S class rRNA transcription from RNA polymerase III type 1 promoter(GO:0042791) tRNA transcription from RNA polymerase III promoter(GO:0042797) |
| 0.1 | 0.3 | GO:0010793 | regulation of mRNA export from nucleus(GO:0010793) |
| 0.1 | 0.6 | GO:0000012 | single strand break repair(GO:0000012) |
| 0.1 | 0.6 | GO:2001138 | regulation of phospholipid transport(GO:2001138) positive regulation of phospholipid transport(GO:2001140) |
| 0.1 | 0.3 | GO:0099590 | neurotransmitter receptor internalization(GO:0099590) |
| 0.1 | 0.8 | GO:1900364 | negative regulation of mRNA polyadenylation(GO:1900364) |
| 0.1 | 1.7 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.1 | 0.3 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 0.1 | 0.3 | GO:0043983 | histone H4-K12 acetylation(GO:0043983) |
| 0.1 | 0.7 | GO:0015886 | heme transport(GO:0015886) |
| 0.1 | 0.7 | GO:2000210 | positive regulation of anoikis(GO:2000210) |
| 0.1 | 2.0 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.1 | 1.4 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.1 | 2.6 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.1 | 0.4 | GO:0071027 | nuclear RNA surveillance(GO:0071027) nuclear mRNA surveillance(GO:0071028) |
| 0.1 | 1.1 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 0.1 | 1.1 | GO:0034134 | toll-like receptor 2 signaling pathway(GO:0034134) |
| 0.1 | 0.4 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.1 | 0.5 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.1 | 5.2 | GO:0031124 | mRNA 3'-end processing(GO:0031124) |
| 0.1 | 2.6 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.1 | 0.2 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.0 | 0.7 | GO:0033623 | regulation of integrin activation(GO:0033623) |
| 0.0 | 2.2 | GO:0043039 | tRNA aminoacylation for protein translation(GO:0006418) amino acid activation(GO:0043038) tRNA aminoacylation(GO:0043039) |
| 0.0 | 0.1 | GO:0051461 | positive regulation of corticotropin secretion(GO:0051461) |
| 0.0 | 2.1 | GO:1903959 | regulation of anion transmembrane transport(GO:1903959) |
| 0.0 | 0.2 | GO:0048698 | negative regulation of collateral sprouting in absence of injury(GO:0048698) |
| 0.0 | 0.2 | GO:1902953 | positive regulation of ER to Golgi vesicle-mediated transport(GO:1902953) |
| 0.0 | 0.6 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.0 | 0.8 | GO:0047497 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) |
| 0.0 | 2.6 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.0 | 0.9 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.0 | 2.3 | GO:0045540 | regulation of cholesterol biosynthetic process(GO:0045540) |
| 0.0 | 0.3 | GO:1902969 | mitotic DNA replication(GO:1902969) |
| 0.0 | 1.0 | GO:0070987 | error-free translesion synthesis(GO:0070987) |
| 0.0 | 2.1 | GO:0042273 | ribosomal large subunit biogenesis(GO:0042273) |
| 0.0 | 1.3 | GO:0006363 | termination of RNA polymerase I transcription(GO:0006363) |
| 0.0 | 0.9 | GO:0045948 | positive regulation of translational initiation(GO:0045948) |
| 0.0 | 0.1 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 0.0 | 0.1 | GO:0048075 | positive regulation of eye pigmentation(GO:0048075) |
| 0.0 | 0.1 | GO:0007231 | osmosensory signaling pathway(GO:0007231) |
| 0.0 | 0.1 | GO:2000767 | positive regulation of cytoplasmic translation(GO:2000767) |
| 0.0 | 0.3 | GO:0033235 | positive regulation of protein sumoylation(GO:0033235) |
| 0.0 | 0.1 | GO:0045347 | negative regulation of MHC class II biosynthetic process(GO:0045347) |
| 0.0 | 0.3 | GO:0060081 | membrane hyperpolarization(GO:0060081) |
| 0.0 | 0.2 | GO:0070314 | G1 to G0 transition(GO:0070314) |
| 0.0 | 0.1 | GO:0097461 | ferric iron import into cell(GO:0097461) ferric iron import across plasma membrane(GO:0098706) |
| 0.0 | 1.1 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.4 | GO:0072600 | establishment of protein localization to Golgi(GO:0072600) |
| 0.0 | 0.7 | GO:0006333 | chromatin assembly or disassembly(GO:0006333) |
| 0.0 | 0.2 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 1.4 | GO:0036498 | IRE1-mediated unfolded protein response(GO:0036498) |
| 0.0 | 0.2 | GO:2001241 | positive regulation of extrinsic apoptotic signaling pathway in absence of ligand(GO:2001241) |
| 0.0 | 0.3 | GO:0032784 | regulation of DNA-templated transcription, elongation(GO:0032784) |
| 0.0 | 0.9 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 0.6 | GO:0014003 | oligodendrocyte development(GO:0014003) |
| 0.0 | 1.2 | GO:0051225 | spindle assembly(GO:0051225) |
| 0.0 | 0.3 | GO:0030261 | chromosome condensation(GO:0030261) |