Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
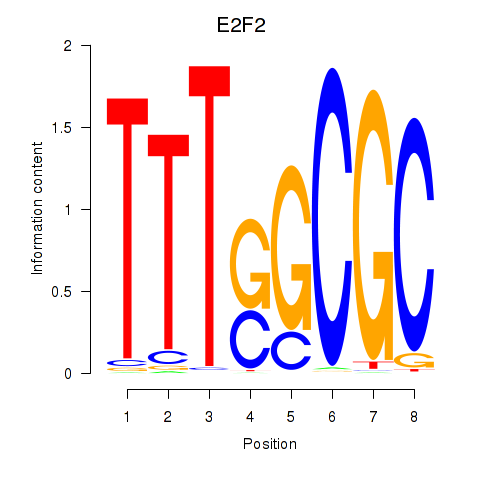
Results for E2F2_E2F5
Z-value: 2.27
Transcription factors associated with E2F2_E2F5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
E2F2
|
ENSG00000007968.6 | E2F2 |
|
E2F5
|
ENSG00000133740.6 | E2F5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| E2F5 | hg19_v2_chr8_+_86121448_86121494 | 0.71 | 2.0e-03 | Click! |
| E2F2 | hg19_v2_chr1_-_23857698_23857733 | -0.12 | 6.6e-01 | Click! |
Activity profile of E2F2_E2F5 motif
Sorted Z-values of E2F2_E2F5 motif
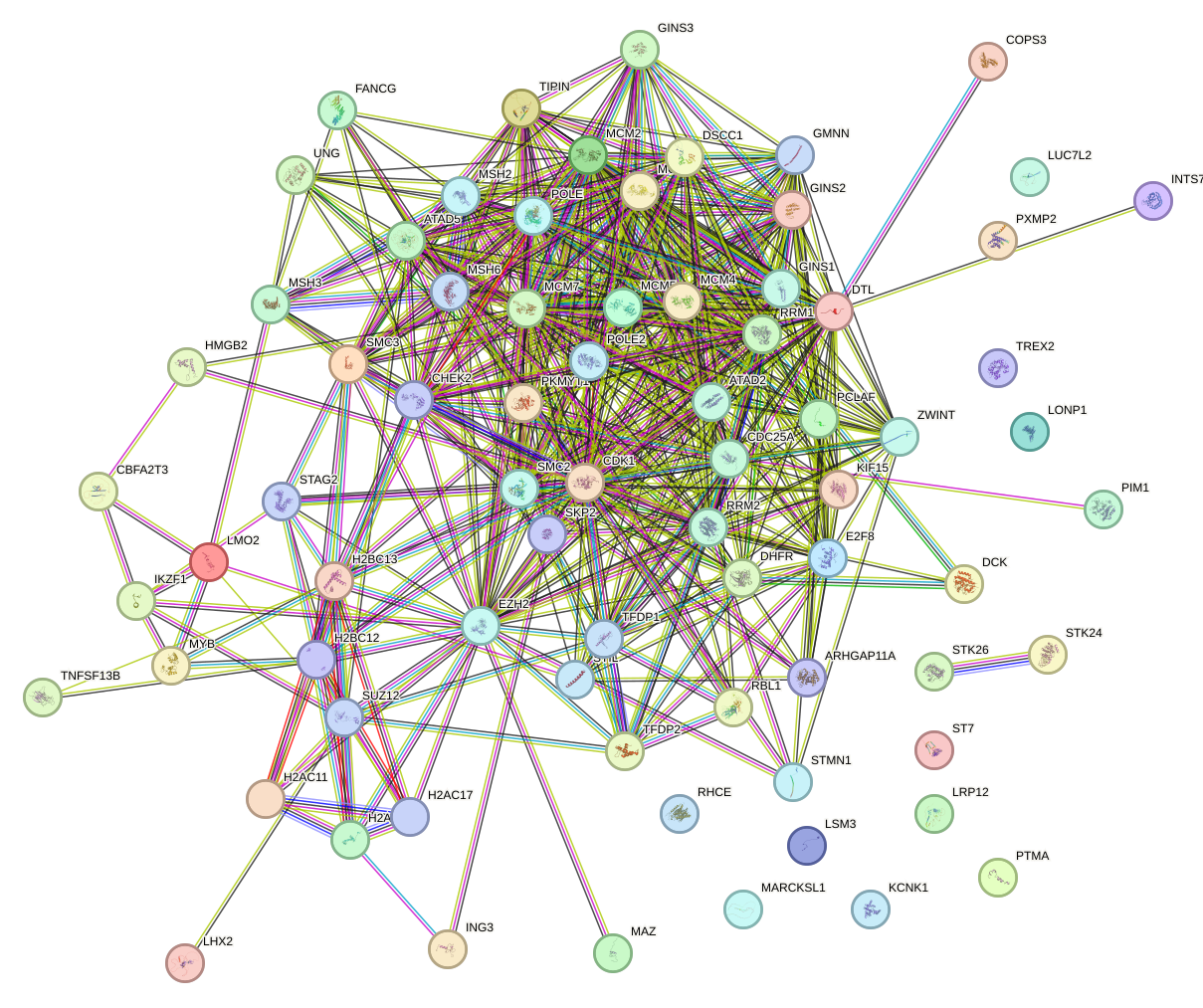
Network of associatons between targets according to the STRING database.
First level regulatory network of E2F2_E2F5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_19263145 | 5.16 |
ENST00000532666.1 ENST00000527884.1 |
E2F8 |
E2F transcription factor 8 |
| chr7_-_99699538 | 5.06 |
ENST00000343023.6 ENST00000303887.5 |
MCM7 |
minichromosome maintenance complex component 7 |
| chr6_-_27114577 | 4.90 |
ENST00000356950.1 ENST00000396891.4 |
HIST1H2BK |
histone cluster 1, H2bk |
| chr10_+_62538089 | 4.83 |
ENST00000519078.2 ENST00000395284.3 ENST00000316629.4 |
CDK1 |
cyclin-dependent kinase 1 |
| chr10_+_62538248 | 4.72 |
ENST00000448257.2 |
CDK1 |
cyclin-dependent kinase 1 |
| chr6_+_24775641 | 4.62 |
ENST00000378054.2 ENST00000476555.1 |
GMNN |
geminin, DNA replication inhibitor |
| chr6_+_135502408 | 4.47 |
ENST00000341911.5 ENST00000442647.2 ENST00000316528.8 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr6_+_135502466 | 4.35 |
ENST00000367814.4 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr2_-_136633940 | 4.19 |
ENST00000264156.2 |
MCM6 |
minichromosome maintenance complex component 6 |
| chr5_-_79950371 | 4.18 |
ENST00000511032.1 ENST00000504396.1 ENST00000505337.1 |
DHFR |
dihydrofolate reductase |
| chr11_+_125495862 | 3.96 |
ENST00000428830.2 ENST00000544373.1 ENST00000527013.1 ENST00000526937.1 ENST00000534685.1 |
CHEK1 |
checkpoint kinase 1 |
| chr8_-_124408652 | 3.83 |
ENST00000287394.5 |
ATAD2 |
ATPase family, AAA domain containing 2 |
| chr7_-_148581251 | 3.75 |
ENST00000478654.1 ENST00000460911.1 ENST00000350995.2 |
EZH2 |
enhancer of zeste homolog 2 (Drosophila) |
| chr10_+_13203543 | 3.53 |
ENST00000378714.3 ENST00000479669.1 ENST00000484800.2 |
MCM10 |
minichromosome maintenance complex component 10 |
| chr14_-_50154921 | 3.50 |
ENST00000553805.2 ENST00000554396.1 ENST00000216367.5 ENST00000539565.2 |
POLE2 |
polymerase (DNA directed), epsilon 2, accessory subunit |
| chr3_+_127317066 | 3.43 |
ENST00000265056.7 |
MCM2 |
minichromosome maintenance complex component 2 |
| chr22_+_35796056 | 3.34 |
ENST00000216122.4 |
MCM5 |
minichromosome maintenance complex component 5 |
| chr7_-_148581360 | 3.29 |
ENST00000320356.2 ENST00000541220.1 ENST00000483967.1 ENST00000536783.1 |
EZH2 |
enhancer of zeste homolog 2 (Drosophila) |
| chr22_+_35796108 | 3.21 |
ENST00000382011.5 ENST00000416905.1 |
MCM5 |
minichromosome maintenance complex component 5 |
| chr8_-_120868078 | 3.15 |
ENST00000313655.4 |
DSCC1 |
DNA replication and sister chromatid cohesion 1 |
| chr1_-_28241024 | 3.14 |
ENST00000313433.7 ENST00000444045.1 |
RPA2 |
replication protein A2, 32kDa |
| chr16_-_89007491 | 3.09 |
ENST00000327483.5 ENST00000564416.1 |
CBFA2T3 |
core-binding factor, runt domain, alpha subunit 2; translocated to, 3 |
| chr20_+_25388293 | 3.08 |
ENST00000262460.4 ENST00000429262.2 |
GINS1 |
GINS complex subunit 1 (Psf1 homolog) |
| chr5_-_79950775 | 3.08 |
ENST00000439211.2 |
DHFR |
dihydrofolate reductase |
| chr6_+_24775153 | 3.00 |
ENST00000356509.3 ENST00000230056.3 |
GMNN |
geminin, DNA replication inhibitor |
| chr15_-_64673630 | 2.87 |
ENST00000558008.1 ENST00000559519.1 ENST00000380258.2 |
KIAA0101 |
KIAA0101 |
| chr7_+_120591170 | 2.86 |
ENST00000431467.1 |
ING3 |
inhibitor of growth family, member 3 |
| chr16_-_3030283 | 2.86 |
ENST00000572619.1 ENST00000574415.1 ENST00000440027.2 ENST00000572059.1 |
PKMYT1 |
protein kinase, membrane associated tyrosine/threonine 1 |
| chr2_+_232573208 | 2.81 |
ENST00000409115.3 |
PTMA |
prothymosin, alpha |
| chr7_+_120590803 | 2.79 |
ENST00000315870.5 ENST00000339121.5 ENST00000445699.1 |
ING3 |
inhibitor of growth family, member 3 |
| chr16_-_3030407 | 2.79 |
ENST00000431515.2 ENST00000574385.1 ENST00000576268.1 ENST00000574730.1 ENST00000575632.1 ENST00000573944.1 ENST00000262300.8 |
PKMYT1 |
protein kinase, membrane associated tyrosine/threonine 1 |
| chr15_-_66649010 | 2.78 |
ENST00000367709.4 ENST00000261881.4 |
TIPIN |
TIMELESS interacting protein |
| chrX_+_24711997 | 2.77 |
ENST00000379068.3 ENST00000379059.3 |
POLA1 |
polymerase (DNA directed), alpha 1, catalytic subunit |
| chr7_+_139044621 | 2.59 |
ENST00000354926.4 |
C7orf55-LUC7L2 |
C7orf55-LUC7L2 readthrough |
| chrX_-_152736013 | 2.57 |
ENST00000330912.2 ENST00000338525.2 ENST00000334497.2 ENST00000370232.1 ENST00000370212.3 ENST00000370211.4 |
TREX2 HAUS7 |
three prime repair exonuclease 2 HAUS augmin-like complex, subunit 7 |
| chr15_+_32907691 | 2.55 |
ENST00000361627.3 ENST00000567348.1 ENST00000563864.1 ENST00000543522.1 |
ARHGAP11A |
Rho GTPase activating protein 11A |
| chr2_+_232573222 | 2.46 |
ENST00000341369.7 ENST00000409683.1 |
PTMA |
prothymosin, alpha |
| chr15_-_64673665 | 2.45 |
ENST00000300035.4 |
KIAA0101 |
KIAA0101 |
| chr1_+_212208919 | 2.42 |
ENST00000366991.4 ENST00000542077.1 |
DTL |
denticleless E3 ubiquitin protein ligase homolog (Drosophila) |
| chr1_-_26232951 | 2.39 |
ENST00000426559.2 ENST00000455785.2 |
STMN1 |
stathmin 1 |
| chr2_+_10262857 | 2.39 |
ENST00000304567.5 |
RRM2 |
ribonucleotide reductase M2 |
| chr11_-_33891362 | 2.35 |
ENST00000395833.3 |
LMO2 |
LIM domain only 2 (rhombotin-like 1) |
| chr6_+_27806319 | 2.34 |
ENST00000606613.1 ENST00000396980.3 |
HIST1H2BN |
histone cluster 1, H2bn |
| chr6_-_52149475 | 2.23 |
ENST00000419835.2 ENST00000229854.7 ENST00000596288.1 |
MCM3 |
minichromosome maintenance complex component 3 |
| chr1_+_228645796 | 2.09 |
ENST00000369160.2 |
HIST3H2BB |
histone cluster 3, H2bb |
| chr7_-_158497431 | 2.04 |
ENST00000449727.2 ENST00000409339.3 ENST00000409423.1 ENST00000356309.3 |
NCAPG2 |
non-SMC condensin II complex, subunit G2 |
| chr6_+_135502501 | 2.02 |
ENST00000527615.1 ENST00000420123.2 ENST00000525369.1 ENST00000528774.1 ENST00000534121.1 ENST00000534044.1 ENST00000533624.1 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr2_+_48010312 | 2.02 |
ENST00000540021.1 |
MSH6 |
mutS homolog 6 |
| chr1_-_47779762 | 2.00 |
ENST00000371877.3 ENST00000360380.3 ENST00000337817.5 ENST00000447475.2 |
STIL |
SCL/TAL1 interrupting locus |
| chr20_-_35724388 | 1.90 |
ENST00000344359.3 ENST00000373664.3 |
RBL1 |
retinoblastoma-like 1 (p107) |
| chr1_-_26232522 | 1.89 |
ENST00000399728.1 |
STMN1 |
stathmin 1 |
| chr8_+_48873479 | 1.84 |
ENST00000262105.2 |
MCM4 |
minichromosome maintenance complex component 4 |
| chr8_+_48873453 | 1.83 |
ENST00000523944.1 |
MCM4 |
minichromosome maintenance complex component 4 |
| chr1_+_25598989 | 1.81 |
ENST00000454452.2 |
RHD |
Rh blood group, D antigen |
| chr15_+_40987327 | 1.81 |
ENST00000423169.2 ENST00000267868.3 ENST00000557850.1 ENST00000532743.1 ENST00000382643.3 |
RAD51 |
RAD51 recombinase |
| chr3_-_48229846 | 1.79 |
ENST00000302506.3 ENST00000351231.3 ENST00000437972.1 |
CDC25A |
cell division cycle 25A |
| chr2_+_48010221 | 1.77 |
ENST00000234420.5 |
MSH6 |
mutS homolog 6 |
| chr16_-_85722530 | 1.64 |
ENST00000253462.3 |
GINS2 |
GINS complex subunit 2 (Psf2 homolog) |
| chr19_+_50887585 | 1.61 |
ENST00000440232.2 ENST00000601098.1 ENST00000599857.1 ENST00000593887.1 |
POLD1 |
polymerase (DNA directed), delta 1, catalytic subunit |
| chr17_+_30264014 | 1.60 |
ENST00000322652.5 ENST00000580398.1 |
SUZ12 |
SUZ12 polycomb repressive complex 2 subunit |
| chrX_+_123095546 | 1.57 |
ENST00000371157.3 ENST00000371145.3 ENST00000371144.3 |
STAG2 |
stromal antigen 2 |
| chr3_+_44803209 | 1.53 |
ENST00000326047.4 |
KIF15 |
kinesin family member 15 |
| chr12_-_133263893 | 1.53 |
ENST00000535270.1 ENST00000320574.5 |
POLE |
polymerase (DNA directed), epsilon, catalytic subunit |
| chr5_+_43121698 | 1.51 |
ENST00000505606.2 ENST00000509634.1 ENST00000509341.1 |
ZNF131 |
zinc finger protein 131 |
| chr7_+_116593568 | 1.43 |
ENST00000446490.1 |
ST7 |
suppression of tumorigenicity 7 |
| chr1_-_32801825 | 1.42 |
ENST00000329421.7 |
MARCKSL1 |
MARCKS-like 1 |
| chr1_-_28241226 | 1.42 |
ENST00000373912.3 ENST00000373909.3 |
RPA2 |
replication protein A2, 32kDa |
| chr12_+_133264156 | 1.38 |
ENST00000317479.3 ENST00000543589.1 |
PXMP2 |
peroxisomal membrane protein 2, 22kDa |
| chr6_-_35656712 | 1.38 |
ENST00000357266.4 ENST00000542713.1 |
FKBP5 |
FK506 binding protein 5 |
| chr22_+_20105012 | 1.36 |
ENST00000331821.3 ENST00000411892.1 |
RANBP1 |
RAN binding protein 1 |
| chr1_-_25747283 | 1.35 |
ENST00000346452.4 ENST00000340849.4 ENST00000349438.4 ENST00000294413.7 ENST00000413854.1 ENST00000455194.1 ENST00000243186.6 ENST00000425135.1 |
RHCE |
Rh blood group, CcEe antigens |
| chr8_+_128748308 | 1.34 |
ENST00000377970.2 |
MYC |
v-myc avian myelocytomatosis viral oncogene homolog |
| chrX_+_131157290 | 1.33 |
ENST00000394334.2 |
MST4 |
Serine/threonine-protein kinase MST4 |
| chr1_+_25598872 | 1.32 |
ENST00000328664.4 |
RHD |
Rh blood group, D antigen |
| chr6_-_27775694 | 1.29 |
ENST00000377401.2 |
HIST1H2BL |
histone cluster 1, H2bl |
| chr6_+_37137939 | 1.29 |
ENST00000373509.5 |
PIM1 |
pim-1 oncogene |
| chrX_+_131157322 | 1.29 |
ENST00000481105.1 ENST00000354719.6 ENST00000394335.2 |
MST4 |
Serine/threonine-protein kinase MST4 |
| chr1_+_233749739 | 1.28 |
ENST00000366621.3 |
KCNK1 |
potassium channel, subfamily K, member 1 |
| chr11_-_19262486 | 1.25 |
ENST00000250024.4 |
E2F8 |
E2F transcription factor 8 |
| chr7_+_116593433 | 1.25 |
ENST00000323984.3 ENST00000393449.1 |
ST7 |
suppression of tumorigenicity 7 |
| chr6_+_27775899 | 1.24 |
ENST00000358739.3 |
HIST1H2AI |
histone cluster 1, H2ai |
| chr5_+_43120985 | 1.23 |
ENST00000515326.1 |
ZNF131 |
zinc finger protein 131 |
| chr5_+_36152179 | 1.22 |
ENST00000508514.1 ENST00000513151.1 ENST00000546211.1 |
SKP2 |
S-phase kinase-associated protein 2, E3 ubiquitin protein ligase |
| chr10_-_58120996 | 1.21 |
ENST00000361148.6 ENST00000395405.1 ENST00000373944.3 |
ZWINT |
ZW10 interacting kinetochore protein |
| chr3_-_53080047 | 1.21 |
ENST00000482396.1 ENST00000358080.2 ENST00000296295.6 ENST00000394752.3 |
SFMBT1 |
Scm-like with four mbt domains 1 |
| chr6_+_27100811 | 1.20 |
ENST00000359193.2 |
HIST1H2AG |
histone cluster 1, H2ag |
| chr7_+_50344289 | 1.19 |
ENST00000413698.1 ENST00000359197.5 ENST00000331340.3 ENST00000357364.4 ENST00000343574.5 ENST00000349824.4 ENST00000346667.4 ENST00000440768.2 |
IKZF1 |
IKAROS family zinc finger 1 (Ikaros) |
| chr1_-_212208842 | 1.19 |
ENST00000366992.3 ENST00000366993.3 ENST00000440600.2 ENST00000366994.3 |
INTS7 |
integrator complex subunit 7 |
| chr6_-_27100529 | 1.17 |
ENST00000607124.1 ENST00000339812.2 ENST00000541790.1 |
HIST1H2BJ |
histone cluster 1, H2bj |
| chr9_+_126773880 | 1.15 |
ENST00000373615.4 |
LHX2 |
LIM homeobox 2 |
| chr22_-_29137771 | 1.13 |
ENST00000439200.1 ENST00000405598.1 ENST00000398017.2 ENST00000425190.2 ENST00000348295.3 ENST00000382578.1 ENST00000382565.1 ENST00000382566.1 ENST00000382580.2 ENST00000328354.6 |
CHEK2 |
checkpoint kinase 2 |
| chr5_+_43121607 | 1.12 |
ENST00000509156.1 ENST00000508259.1 ENST00000306938.4 ENST00000399534.1 |
ZNF131 |
zinc finger protein 131 |
| chr9_+_106856831 | 1.11 |
ENST00000303219.8 ENST00000374787.3 |
SMC2 |
structural maintenance of chromosomes 2 |
| chr10_+_112327425 | 1.11 |
ENST00000361804.4 |
SMC3 |
structural maintenance of chromosomes 3 |
| chr12_+_109535923 | 1.10 |
ENST00000336865.2 |
UNG |
uracil-DNA glycosylase |
| chr9_-_35080013 | 1.10 |
ENST00000378643.3 |
FANCG |
Fanconi anemia, complementation group G |
| chr11_+_4116054 | 1.09 |
ENST00000423050.2 |
RRM1 |
ribonucleotide reductase M1 |
| chr5_+_36152091 | 1.08 |
ENST00000274254.5 |
SKP2 |
S-phase kinase-associated protein 2, E3 ubiquitin protein ligase |
| chr3_-_141868293 | 1.07 |
ENST00000317104.7 ENST00000494358.1 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr1_+_25599018 | 1.05 |
ENST00000417538.2 ENST00000357542.4 ENST00000568195.1 ENST00000342055.5 ENST00000423810.2 |
RHD |
Rh blood group, D antigen |
| chr13_+_114238997 | 1.05 |
ENST00000538138.1 ENST00000375370.5 |
TFDP1 |
transcription factor Dp-1 |
| chr4_+_71859156 | 1.04 |
ENST00000286648.5 ENST00000504730.1 ENST00000504952.1 |
DCK |
deoxycytidine kinase |
| chr16_+_58426296 | 1.03 |
ENST00000426538.2 ENST00000328514.7 ENST00000318129.5 |
GINS3 |
GINS complex subunit 3 (Psf3 homolog) |
| chr4_-_174256276 | 1.01 |
ENST00000296503.5 |
HMGB2 |
high mobility group box 2 |
| chr8_+_128748466 | 1.01 |
ENST00000524013.1 ENST00000520751.1 |
MYC |
v-myc avian myelocytomatosis viral oncogene homolog |
| chr16_+_29817399 | 1.01 |
ENST00000545521.1 |
MAZ |
MYC-associated zinc finger protein (purine-binding transcription factor) |
| chr19_+_36706024 | 1.00 |
ENST00000443387.2 |
ZNF146 |
zinc finger protein 146 |
| chr22_-_20104700 | 0.98 |
ENST00000439169.2 ENST00000445045.1 ENST00000404751.3 ENST00000252136.7 ENST00000403707.3 |
TRMT2A |
tRNA methyltransferase 2 homolog A (S. cerevisiae) |
| chr17_+_29158962 | 0.97 |
ENST00000321990.4 |
ATAD5 |
ATPase family, AAA domain containing 5 |
| chr17_-_17184605 | 0.96 |
ENST00000268717.5 |
COPS3 |
COP9 signalosome subunit 3 |
| chr3_-_141868357 | 0.96 |
ENST00000489671.1 ENST00000475734.1 ENST00000467072.1 ENST00000499676.2 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr2_+_47630108 | 0.95 |
ENST00000233146.2 ENST00000454849.1 ENST00000543555.1 |
MSH2 |
mutS homolog 2 |
| chr5_+_79950463 | 0.94 |
ENST00000265081.6 |
MSH3 |
mutS homolog 3 |
| chr3_+_14219858 | 0.92 |
ENST00000306024.3 |
LSM3 |
LSM3 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr1_+_66458072 | 0.92 |
ENST00000423207.2 |
PDE4B |
phosphodiesterase 4B, cAMP-specific |
| chr5_+_36876833 | 0.92 |
ENST00000282516.8 ENST00000448238.2 |
NIPBL |
Nipped-B homolog (Drosophila) |
| chr7_-_44887620 | 0.91 |
ENST00000349299.3 ENST00000521529.1 ENST00000308153.4 ENST00000350771.3 ENST00000222690.6 ENST00000381124.5 ENST00000437072.1 ENST00000446531.1 |
H2AFV |
H2A histone family, member V |
| chr17_+_58677539 | 0.91 |
ENST00000305921.3 |
PPM1D |
protein phosphatase, Mg2+/Mn2+ dependent, 1D |
| chr11_+_64808368 | 0.88 |
ENST00000531072.1 ENST00000398846.1 |
SAC3D1 |
SAC3 domain containing 1 |
| chr14_-_69864993 | 0.88 |
ENST00000555373.1 |
ERH |
enhancer of rudimentary homolog (Drosophila) |
| chr19_+_10765699 | 0.87 |
ENST00000590009.1 |
ILF3 |
interleukin enhancer binding factor 3, 90kDa |
| chr15_-_70388943 | 0.87 |
ENST00000559048.1 ENST00000560939.1 ENST00000440567.3 ENST00000557907.1 ENST00000558379.1 ENST00000451782.2 ENST00000559929.1 |
TLE3 |
transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) |
| chr6_-_144329531 | 0.86 |
ENST00000429150.1 ENST00000392309.1 ENST00000416623.1 ENST00000392307.1 |
PLAGL1 |
pleiomorphic adenoma gene-like 1 |
| chr13_+_32889605 | 0.85 |
ENST00000380152.3 ENST00000544455.1 ENST00000530893.2 |
BRCA2 |
breast cancer 2, early onset |
| chrX_+_48554986 | 0.82 |
ENST00000376687.3 ENST00000453214.2 |
SUV39H1 |
suppressor of variegation 3-9 homolog 1 (Drosophila) |
| chr20_-_32274179 | 0.82 |
ENST00000343380.5 |
E2F1 |
E2F transcription factor 1 |
| chr6_-_27782548 | 0.81 |
ENST00000333151.3 |
HIST1H2AJ |
histone cluster 1, H2aj |
| chr9_+_106856541 | 0.78 |
ENST00000286398.7 ENST00000440179.1 ENST00000374793.3 |
SMC2 |
structural maintenance of chromosomes 2 |
| chr6_+_35995552 | 0.75 |
ENST00000468133.1 |
MAPK14 |
mitogen-activated protein kinase 14 |
| chrX_-_135849484 | 0.75 |
ENST00000370620.1 ENST00000535227.1 |
ARHGEF6 |
Rac/Cdc42 guanine nucleotide exchange factor (GEF) 6 |
| chr1_-_54303934 | 0.73 |
ENST00000537333.1 |
NDC1 |
NDC1 transmembrane nucleoporin |
| chr1_+_100598691 | 0.72 |
ENST00000370143.1 ENST00000370141.2 |
TRMT13 |
tRNA methyltransferase 13 homolog (S. cerevisiae) |
| chr11_+_74303575 | 0.71 |
ENST00000263681.2 |
POLD3 |
polymerase (DNA-directed), delta 3, accessory subunit |
| chr13_-_79177673 | 0.71 |
ENST00000377208.5 |
POU4F1 |
POU class 4 homeobox 1 |
| chr11_+_64808675 | 0.71 |
ENST00000529996.1 |
SAC3D1 |
SAC3 domain containing 1 |
| chr14_+_51706886 | 0.70 |
ENST00000457354.2 |
TMX1 |
thioredoxin-related transmembrane protein 1 |
| chr19_-_10305752 | 0.69 |
ENST00000540357.1 ENST00000359526.4 ENST00000340748.4 |
DNMT1 |
DNA (cytosine-5-)-methyltransferase 1 |
| chr18_+_19192228 | 0.68 |
ENST00000300413.5 ENST00000579618.1 ENST00000582475.1 |
SNRPD1 |
small nuclear ribonucleoprotein D1 polypeptide 16kDa |
| chr16_-_23607598 | 0.66 |
ENST00000562133.1 ENST00000570319.1 ENST00000007516.3 |
NDUFAB1 |
NADH dehydrogenase (ubiquinone) 1, alpha/beta subcomplex, 1, 8kDa |
| chr7_+_26241325 | 0.66 |
ENST00000456948.1 ENST00000409747.1 |
CBX3 |
chromobox homolog 3 |
| chr9_+_86595626 | 0.65 |
ENST00000445877.1 ENST00000325875.3 |
RMI1 |
RecQ mediated genome instability 1 |
| chr10_-_21786179 | 0.64 |
ENST00000377113.5 |
CASC10 |
cancer susceptibility candidate 10 |
| chr3_-_133380731 | 0.64 |
ENST00000260810.5 |
TOPBP1 |
topoisomerase (DNA) II binding protein 1 |
| chr4_+_129730779 | 0.64 |
ENST00000226319.6 |
PHF17 |
jade family PHD finger 1 |
| chr10_-_13390021 | 0.62 |
ENST00000537130.1 |
SEPHS1 |
selenophosphate synthetase 1 |
| chr4_+_178230985 | 0.62 |
ENST00000264596.3 |
NEIL3 |
nei endonuclease VIII-like 3 (E. coli) |
| chr17_-_74733404 | 0.61 |
ENST00000508921.3 ENST00000583836.1 ENST00000358156.6 ENST00000392485.2 ENST00000359995.5 |
SRSF2 |
serine/arginine-rich splicing factor 2 |
| chr6_+_30312908 | 0.59 |
ENST00000433076.2 ENST00000442966.2 ENST00000428040.2 ENST00000436442.2 |
RPP21 |
ribonuclease P/MRP 21kDa subunit |
| chr20_-_5100591 | 0.59 |
ENST00000379143.5 |
PCNA |
proliferating cell nuclear antigen |
| chr6_+_84569359 | 0.59 |
ENST00000369681.5 ENST00000369679.4 |
CYB5R4 |
cytochrome b5 reductase 4 |
| chr3_-_169381183 | 0.58 |
ENST00000494292.1 |
MECOM |
MDS1 and EVI1 complex locus |
| chr1_+_33116743 | 0.58 |
ENST00000414241.3 ENST00000373493.5 |
RBBP4 |
retinoblastoma binding protein 4 |
| chr19_+_1383890 | 0.58 |
ENST00000539480.1 ENST00000313408.7 ENST00000414651.2 |
NDUFS7 |
NADH dehydrogenase (ubiquinone) Fe-S protein 7, 20kDa (NADH-coenzyme Q reductase) |
| chr12_-_31479045 | 0.57 |
ENST00000539409.1 ENST00000395766.1 |
FAM60A |
family with sequence similarity 60, member A |
| chr6_+_35995488 | 0.56 |
ENST00000229795.3 |
MAPK14 |
mitogen-activated protein kinase 14 |
| chr6_-_17706618 | 0.54 |
ENST00000262077.2 ENST00000537253.1 |
NUP153 |
nucleoporin 153kDa |
| chr19_+_36705504 | 0.54 |
ENST00000456324.1 |
ZNF146 |
zinc finger protein 146 |
| chr6_+_35995531 | 0.53 |
ENST00000229794.4 |
MAPK14 |
mitogen-activated protein kinase 14 |
| chr7_-_105516923 | 0.51 |
ENST00000478915.1 |
ATXN7L1 |
ataxin 7-like 1 |
| chr3_-_185655795 | 0.50 |
ENST00000342294.4 ENST00000382191.4 ENST00000453386.2 |
TRA2B |
transformer 2 beta homolog (Drosophila) |
| chr1_-_222885770 | 0.50 |
ENST00000355727.2 ENST00000340020.6 |
AIDA |
axin interactor, dorsalization associated |
| chr4_+_129730947 | 0.50 |
ENST00000452328.2 ENST00000504089.1 |
PHF17 |
jade family PHD finger 1 |
| chrX_+_106871713 | 0.50 |
ENST00000372435.4 ENST00000372428.4 ENST00000372419.3 ENST00000543248.1 |
PRPS1 |
phosphoribosyl pyrophosphate synthetase 1 |
| chr19_+_50432400 | 0.49 |
ENST00000423777.2 ENST00000600336.1 ENST00000597227.1 |
ATF5 |
activating transcription factor 5 |
| chr19_+_49588677 | 0.49 |
ENST00000598984.1 ENST00000598441.1 |
SNRNP70 |
small nuclear ribonucleoprotein 70kDa (U1) |
| chr22_-_38966172 | 0.49 |
ENST00000216024.2 |
DMC1 |
DNA meiotic recombinase 1 |
| chr5_+_133861790 | 0.48 |
ENST00000395003.1 |
PHF15 |
jade family PHD finger 2 |
| chr7_+_26241310 | 0.48 |
ENST00000396386.2 |
CBX3 |
chromobox homolog 3 |
| chr3_-_51975942 | 0.47 |
ENST00000232888.6 |
RRP9 |
ribosomal RNA processing 9, small subunit (SSU) processome component, homolog (yeast) |
| chr3_+_16926441 | 0.47 |
ENST00000418129.2 ENST00000396755.2 |
PLCL2 |
phospholipase C-like 2 |
| chr6_+_27114861 | 0.47 |
ENST00000377459.1 |
HIST1H2AH |
histone cluster 1, H2ah |
| chr4_-_39979576 | 0.47 |
ENST00000303538.8 ENST00000503396.1 |
PDS5A |
PDS5, regulator of cohesion maintenance, homolog A (S. cerevisiae) |
| chr6_+_88299833 | 0.46 |
ENST00000392844.3 ENST00000257789.4 ENST00000546266.1 ENST00000417380.2 |
ORC3 |
origin recognition complex, subunit 3 |
| chr2_+_61108771 | 0.45 |
ENST00000394479.3 |
REL |
v-rel avian reticuloendotheliosis viral oncogene homolog |
| chr7_-_105517021 | 0.43 |
ENST00000318724.4 ENST00000419735.3 |
ATXN7L1 |
ataxin 7-like 1 |
| chr3_+_134514093 | 0.43 |
ENST00000398015.3 |
EPHB1 |
EPH receptor B1 |
| chr1_-_26233423 | 0.42 |
ENST00000357865.2 |
STMN1 |
stathmin 1 |
| chr12_-_49245936 | 0.42 |
ENST00000308025.3 |
DDX23 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 23 |
| chr12_-_51611477 | 0.41 |
ENST00000389243.4 |
POU6F1 |
POU class 6 homeobox 1 |
| chr5_-_140998616 | 0.41 |
ENST00000389054.3 ENST00000398562.2 ENST00000389057.5 ENST00000398566.3 ENST00000398557.4 ENST00000253811.6 |
DIAPH1 |
diaphanous-related formin 1 |
| chr16_+_3019552 | 0.39 |
ENST00000572687.1 |
PAQR4 |
progestin and adipoQ receptor family member IV |
| chr19_-_50432654 | 0.39 |
ENST00000596680.1 ENST00000594673.1 ENST00000597029.1 |
NUP62 |
nucleoporin 62kDa |
| chr1_-_149982624 | 0.38 |
ENST00000417191.1 ENST00000369135.4 |
OTUD7B |
OTU domain containing 7B |
| chr7_-_150924121 | 0.38 |
ENST00000441774.1 ENST00000222388.2 ENST00000287844.2 |
ABCF2 |
ATP-binding cassette, sub-family F (GCN20), member 2 |
| chr16_-_30102547 | 0.37 |
ENST00000279386.2 |
TBX6 |
T-box 6 |
| chr15_+_66994561 | 0.37 |
ENST00000288840.5 |
SMAD6 |
SMAD family member 6 |
| chr3_+_182511266 | 0.37 |
ENST00000323116.5 ENST00000493826.1 |
ATP11B |
ATPase, class VI, type 11B |
| chr5_+_36152163 | 0.37 |
ENST00000274255.6 |
SKP2 |
S-phase kinase-associated protein 2, E3 ubiquitin protein ligase |
| chr11_+_66384053 | 0.36 |
ENST00000310137.4 ENST00000393979.3 ENST00000409372.1 ENST00000443702.1 ENST00000409738.4 ENST00000514361.3 ENST00000503028.2 ENST00000412278.2 ENST00000500635.2 |
RBM14 RBM4 RBM14-RBM4 |
RNA binding motif protein 14 RNA binding motif protein 4 RBM14-RBM4 readthrough |
| chr10_-_13390270 | 0.36 |
ENST00000378614.4 ENST00000545675.1 ENST00000327347.5 |
SEPHS1 |
selenophosphate synthetase 1 |
| chr6_+_31633011 | 0.36 |
ENST00000375885.4 |
CSNK2B |
casein kinase 2, beta polypeptide |
| chr16_+_67596310 | 0.36 |
ENST00000264010.4 ENST00000401394.1 |
CTCF |
CCCTC-binding factor (zinc finger protein) |
| chr5_+_75378997 | 0.35 |
ENST00000502798.2 |
SV2C |
synaptic vesicle glycoprotein 2C |
| chr9_-_120177216 | 0.35 |
ENST00000373996.3 ENST00000313400.4 ENST00000361477.3 |
ASTN2 |
astrotactin 2 |
| chr1_-_151119087 | 0.35 |
ENST00000341697.3 ENST00000368914.3 |
SEMA6C |
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6C |
| chr1_-_23670752 | 0.35 |
ENST00000302271.6 ENST00000426846.2 ENST00000427764.2 ENST00000606561.1 ENST00000374616.3 |
HNRNPR |
heterogeneous nuclear ribonucleoprotein R |
| chr6_-_5260963 | 0.34 |
ENST00000464010.1 ENST00000468929.1 ENST00000480566.1 |
LYRM4 |
LYR motif containing 4 |
| chr22_+_31031639 | 0.34 |
ENST00000343605.4 ENST00000300385.8 |
SLC35E4 |
solute carrier family 35, member E4 |
| chr19_-_50432711 | 0.34 |
ENST00000597723.1 ENST00000599788.1 ENST00000596217.1 ENST00000593652.1 ENST00000599567.1 ENST00000600935.1 ENST00000596011.1 ENST00000596022.1 ENST00000597295.1 |
NUP62 IL4I1 |
nucleoporin 62kDa interleukin 4 induced 1 |
| chr5_+_140019079 | 0.33 |
ENST00000252100.6 |
TMCO6 |
transmembrane and coiled-coil domains 6 |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 7.3 | GO:0051870 | methotrexate binding(GO:0051870) |
| 1.2 | 4.7 | GO:0032143 | single thymine insertion binding(GO:0032143) |
| 1.0 | 8.6 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.9 | 0.9 | GO:0000404 | heteroduplex DNA loop binding(GO:0000404) double-strand/single-strand DNA junction binding(GO:0000406) dinucleotide repeat insertion binding(GO:0032181) |
| 0.9 | 3.5 | GO:0016728 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.8 | 14.0 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.8 | 9.0 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.7 | 4.0 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.6 | 5.0 | GO:0043142 | single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.5 | 1.9 | GO:0036033 | mediator complex binding(GO:0036033) |
| 0.5 | 5.7 | GO:0008296 | 3'-5'-exodeoxyribonuclease activity(GO:0008296) |
| 0.4 | 4.7 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 0.4 | 4.6 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.4 | 0.8 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.4 | 12.9 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.4 | 9.5 | GO:0035173 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) histone kinase activity(GO:0035173) |
| 0.3 | 1.0 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) |
| 0.3 | 1.0 | GO:0004756 | selenide, water dikinase activity(GO:0004756) phosphotransferase activity, paired acceptors(GO:0016781) |
| 0.3 | 0.6 | GO:0032139 | dinucleotide insertion or deletion binding(GO:0032139) |
| 0.3 | 7.0 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.3 | 1.0 | GO:0044378 | non-sequence-specific DNA binding, bending(GO:0044378) |
| 0.2 | 0.7 | GO:0000035 | acyl binding(GO:0000035) phosphopantetheine binding(GO:0031177) |
| 0.2 | 5.5 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.2 | 0.8 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) |
| 0.2 | 1.8 | GO:0051525 | NFAT protein binding(GO:0051525) |
| 0.2 | 0.5 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.2 | 1.1 | GO:0097506 | uracil DNA N-glycosylase activity(GO:0004844) deaminated base DNA N-glycosylase activity(GO:0097506) |
| 0.1 | 1.3 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.1 | 1.3 | GO:1990446 | U1 snRNP binding(GO:1990446) |
| 0.1 | 0.4 | GO:0016262 | protein N-acetylglucosaminyltransferase activity(GO:0016262) |
| 0.1 | 1.3 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.1 | 0.6 | GO:0000405 | bubble DNA binding(GO:0000405) |
| 0.1 | 0.4 | GO:0097158 | pre-mRNA intronic pyrimidine-rich binding(GO:0097158) |
| 0.1 | 0.7 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 7.5 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 1.3 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.1 | 8.6 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.1 | 0.9 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.1 | 0.5 | GO:0004749 | ribose phosphate diphosphokinase activity(GO:0004749) |
| 0.1 | 0.4 | GO:0070697 | activin receptor binding(GO:0070697) |
| 0.1 | 7.1 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.1 | 0.4 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.1 | 0.5 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.1 | 0.1 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.1 | 1.9 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 0.3 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.1 | 1.4 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.1 | 1.3 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.6 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.0 | 1.4 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.3 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.0 | 7.0 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 1.3 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 0.3 | GO:0043533 | inositol 1,3,4,5 tetrakisphosphate binding(GO:0043533) |
| 0.0 | 0.9 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 0.4 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.0 | 1.1 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 5.3 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 0.4 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.0 | 1.2 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.0 | 0.6 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 0.3 | GO:0099583 | neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration(GO:0099583) |
| 0.0 | 1.7 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.0 | 0.7 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 0.9 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 0.6 | GO:0048038 | quinone binding(GO:0048038) |
| 0.0 | 0.2 | GO:0004793 | glycine hydroxymethyltransferase activity(GO:0004372) threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.0 | 0.1 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.0 | 1.0 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.0 | 0.0 | GO:0032135 | DNA insertion or deletion binding(GO:0032135) |
| 0.0 | 0.5 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 12.3 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.0 | 0.6 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.0 | 0.1 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 2.2 | GO:0004386 | helicase activity(GO:0004386) |
| 0.0 | 1.0 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.0 | 1.5 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 5.9 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 0.4 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.0 | 0.8 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.0 | 0.5 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 1.8 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 0.3 | GO:0019903 | protein phosphatase binding(GO:0019903) |
| 0.0 | 0.1 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.0 | 0.1 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 1.2 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 0.3 | GO:0070566 | adenylyltransferase activity(GO:0070566) |
| 0.0 | 5.4 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.0 | 0.1 | GO:0016018 | cyclosporin A binding(GO:0016018) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 29.9 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 1.0 | 26.2 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.8 | 11.9 | REACTOME REPAIR SYNTHESIS FOR GAP FILLING BY DNA POL IN TC NER | Genes involved in Repair synthesis for gap-filling by DNA polymerase in TC-NER |
| 0.4 | 6.1 | REACTOME ASSOCIATION OF LICENSING FACTORS WITH THE PRE REPLICATIVE COMPLEX | Genes involved in Association of licensing factors with the pre-replicative complex |
| 0.4 | 7.5 | REACTOME ACTIVATION OF ATR IN RESPONSE TO REPLICATION STRESS | Genes involved in Activation of ATR in response to replication stress |
| 0.3 | 5.4 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.2 | 12.4 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.2 | 1.1 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.2 | 1.3 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.2 | 2.5 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.2 | 2.7 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.1 | 1.8 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.1 | 12.9 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 0.9 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.1 | 5.1 | REACTOME CYCLIN E ASSOCIATED EVENTS DURING G1 S TRANSITION | Genes involved in Cyclin E associated events during G1/S transition |
| 0.0 | 1.0 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 1.4 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.0 | 1.1 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 1.9 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 1.3 | REACTOME RNA POL I TRANSCRIPTION | Genes involved in RNA Polymerase I Transcription |
| 0.0 | 0.9 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 2.6 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.0 | 2.4 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 0.7 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.4 | REACTOME MEIOTIC RECOMBINATION | Genes involved in Meiotic Recombination |
| 0.0 | 0.4 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 0.2 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.0 | 0.3 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.0 | 1.4 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.5 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.9 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.0 | 2.2 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.3 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 25.1 | GO:0042555 | MCM complex(GO:0042555) |
| 1.9 | 9.5 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 1.4 | 5.7 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 1.2 | 4.7 | GO:0032301 | MutSalpha complex(GO:0032301) |
| 1.2 | 4.7 | GO:0000811 | GINS complex(GO:0000811) |
| 1.1 | 6.3 | GO:0031298 | replication fork protection complex(GO:0031298) |
| 1.0 | 5.0 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.9 | 0.9 | GO:0032302 | MutSbeta complex(GO:0032302) |
| 0.9 | 3.5 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 0.6 | 2.3 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.6 | 2.8 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.5 | 3.1 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.5 | 1.5 | GO:0005873 | plus-end kinesin complex(GO:0005873) |
| 0.5 | 7.0 | GO:0045120 | pronucleus(GO:0045120) |
| 0.5 | 0.5 | GO:0036387 | nuclear pre-replicative complex(GO:0005656) pre-replicative complex(GO:0036387) |
| 0.4 | 3.9 | GO:0000796 | condensin complex(GO:0000796) |
| 0.4 | 1.1 | GO:0034991 | nuclear meiotic cohesin complex(GO:0034991) |
| 0.3 | 4.6 | GO:0043601 | nuclear replisome(GO:0043601) |
| 0.3 | 0.6 | GO:0030894 | replisome(GO:0030894) |
| 0.3 | 2.6 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.3 | 0.8 | GO:0035189 | Rb-E2F complex(GO:0035189) |
| 0.3 | 1.6 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.3 | 2.7 | GO:0000800 | lateral element(GO:0000800) |
| 0.3 | 1.3 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.2 | 0.7 | GO:0070762 | nuclear pore transmembrane ring(GO:0070762) |
| 0.2 | 13.8 | GO:0044815 | nucleosome(GO:0000786) DNA packaging complex(GO:0044815) |
| 0.2 | 2.5 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.2 | 2.4 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.2 | 0.8 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.2 | 0.5 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.1 | 0.6 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.1 | 1.2 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 0.9 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.1 | 0.5 | GO:0002189 | ribose phosphate diphosphokinase complex(GO:0002189) |
| 0.1 | 0.6 | GO:0035061 | interchromatin granule(GO:0035061) |
| 0.1 | 0.4 | GO:0060187 | cell pole(GO:0060187) |
| 0.1 | 1.3 | GO:0000243 | commitment complex(GO:0000243) |
| 0.1 | 0.3 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.1 | 0.7 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.1 | 1.4 | GO:0044613 | nuclear pore central transport channel(GO:0044613) |
| 0.1 | 11.5 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.1 | 0.3 | GO:0038038 | G-protein coupled receptor homodimeric complex(GO:0038038) |
| 0.1 | 4.9 | GO:0000794 | condensed nuclear chromosome(GO:0000794) |
| 0.1 | 0.5 | GO:0005720 | nuclear heterochromatin(GO:0005720) |
| 0.1 | 0.3 | GO:0033503 | HULC complex(GO:0033503) |
| 0.1 | 0.6 | GO:0005655 | nucleolar ribonuclease P complex(GO:0005655) |
| 0.1 | 0.9 | GO:0034709 | methylosome(GO:0034709) |
| 0.1 | 1.4 | GO:0031231 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.1 | 0.1 | GO:0000444 | MIS12/MIND type complex(GO:0000444) |
| 0.0 | 2.7 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 1.1 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.0 | 0.1 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.0 | 0.4 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.0 | 0.5 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.0 | 0.7 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 0.2 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.0 | 0.4 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 0.0 | 0.6 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.0 | 0.2 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 10.4 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.3 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.0 | 3.9 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
| 0.0 | 0.4 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 0.1 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.0 | 0.9 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 1.0 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 1.2 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.3 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 2.2 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.0 | 0.1 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 2.0 | GO:0000922 | spindle pole(GO:0000922) |
| 0.0 | 0.0 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.1 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.0 | 1.1 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.0 | 0.5 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 0.2 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.0 | 0.1 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.0 | 1.8 | GO:0005819 | spindle(GO:0005819) |
| 0.0 | 0.9 | GO:0000785 | chromatin(GO:0000785) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 15.0 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.3 | 17.9 | PID ATR PATHWAY | ATR signaling pathway |
| 0.3 | 4.7 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.2 | 23.8 | PID E2F PATHWAY | E2F transcription factor network |
| 0.2 | 10.8 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.1 | 6.0 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.1 | 1.3 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.1 | 2.0 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 2.0 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 3.1 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 1.6 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 2.6 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 1.8 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 0.8 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.0 | 2.4 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.5 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 1.8 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 1.4 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 1.3 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 1.0 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.3 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 0.4 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.6 | 10.8 | GO:1904897 | regulation of hepatic stellate cell proliferation(GO:1904897) positive regulation of hepatic stellate cell proliferation(GO:1904899) hepatic stellate cell proliferation(GO:1990922) |
| 2.3 | 7.0 | GO:1904772 | hepatocyte homeostasis(GO:0036333) response to tetrachloromethane(GO:1904772) |
| 1.8 | 18.2 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 1.8 | 7.3 | GO:0046452 | dihydrofolate metabolic process(GO:0046452) |
| 1.6 | 9.5 | GO:0014038 | regulation of Schwann cell differentiation(GO:0014038) |
| 1.3 | 6.4 | GO:0060718 | chorionic trophoblast cell differentiation(GO:0060718) |
| 1.3 | 7.6 | GO:0071163 | DNA replication preinitiation complex assembly(GO:0071163) |
| 1.2 | 4.9 | GO:0006272 | leading strand elongation(GO:0006272) |
| 0.9 | 4.7 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) |
| 0.7 | 3.6 | GO:0048478 | replication fork protection(GO:0048478) |
| 0.7 | 4.0 | GO:0035407 | histone H3-T11 phosphorylation(GO:0035407) |
| 0.6 | 5.7 | GO:0043570 | maintenance of DNA repeat elements(GO:0043570) |
| 0.6 | 3.1 | GO:1900264 | regulation of DNA-directed DNA polymerase activity(GO:1900262) positive regulation of DNA-directed DNA polymerase activity(GO:1900264) |
| 0.6 | 5.1 | GO:0001833 | inner cell mass cell proliferation(GO:0001833) |
| 0.5 | 2.7 | GO:0071460 | cellular response to cell-matrix adhesion(GO:0071460) |
| 0.5 | 18.6 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.4 | 6.9 | GO:0032201 | telomere maintenance via semi-conservative replication(GO:0032201) |
| 0.4 | 3.8 | GO:0031936 | negative regulation of chromatin silencing(GO:0031936) |
| 0.4 | 2.3 | GO:0072131 | kidney mesenchyme morphogenesis(GO:0072131) metanephric mesenchyme morphogenesis(GO:0072133) |
| 0.4 | 3.1 | GO:1903715 | regulation of aerobic respiration(GO:1903715) |
| 0.4 | 1.1 | GO:1903926 | signal transduction involved in intra-S DNA damage checkpoint(GO:0072428) response to bisphenol A(GO:1903925) cellular response to bisphenol A(GO:1903926) |
| 0.4 | 1.8 | GO:0014835 | myoblast differentiation involved in skeletal muscle regeneration(GO:0014835) |
| 0.3 | 2.0 | GO:0033504 | floor plate development(GO:0033504) |
| 0.3 | 1.0 | GO:0016260 | selenocysteine biosynthetic process(GO:0016260) |
| 0.3 | 6.1 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.3 | 0.9 | GO:0061010 | gall bladder development(GO:0061010) |
| 0.3 | 1.0 | GO:0010216 | maintenance of DNA methylation(GO:0010216) |
| 0.2 | 0.7 | GO:0071790 | spindle pole body duplication(GO:0030474) spindle pole body organization(GO:0051300) spindle pole body localization(GO:0070631) establishment of spindle pole body localization(GO:0070632) spindle pole body localization to nuclear envelope(GO:0071789) establishment of spindle pole body localization to nuclear envelope(GO:0071790) |
| 0.2 | 0.7 | GO:0021986 | epithalamus development(GO:0021538) habenula development(GO:0021986) |
| 0.2 | 1.9 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.2 | 2.6 | GO:0032876 | negative regulation of DNA endoreduplication(GO:0032876) |
| 0.2 | 0.9 | GO:1903438 | regulation of cytokinetic process(GO:0032954) regulation of mitotic cytokinetic process(GO:1903436) positive regulation of mitotic cytokinetic process(GO:1903438) positive regulation of mitotic cytokinesis(GO:1903490) |
| 0.2 | 1.9 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.2 | 1.4 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.2 | 0.4 | GO:0090526 | regulation of gluconeogenesis involved in cellular glucose homeostasis(GO:0090526) |
| 0.2 | 5.7 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.2 | 1.7 | GO:0021978 | telencephalon regionalization(GO:0021978) |
| 0.2 | 1.0 | GO:0009157 | deoxyribonucleoside monophosphate biosynthetic process(GO:0009157) |
| 0.2 | 5.5 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.2 | 1.3 | GO:0060075 | regulation of resting membrane potential(GO:0060075) |
| 0.2 | 0.5 | GO:0001923 | B-1 B cell differentiation(GO:0001923) |
| 0.2 | 3.5 | GO:0009263 | deoxyribonucleotide biosynthetic process(GO:0009263) |
| 0.1 | 0.4 | GO:0099545 | trans-synaptic signaling by trans-synaptic complex(GO:0099545) |
| 0.1 | 1.3 | GO:2000138 | positive regulation of cell proliferation involved in heart morphogenesis(GO:2000138) |
| 0.1 | 0.4 | GO:0000354 | cis assembly of pre-catalytic spliceosome(GO:0000354) |
| 0.1 | 0.3 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.1 | 1.0 | GO:0045654 | positive regulation of megakaryocyte differentiation(GO:0045654) |
| 0.1 | 0.5 | GO:0046101 | hypoxanthine metabolic process(GO:0046100) hypoxanthine biosynthetic process(GO:0046101) |
| 0.1 | 0.6 | GO:0061083 | regulation of protein refolding(GO:0061083) negative regulation of protein refolding(GO:0061084) |
| 0.1 | 0.8 | GO:0036123 | histone H3-K9 dimethylation(GO:0036123) |
| 0.1 | 2.3 | GO:0042789 | mRNA transcription from RNA polymerase II promoter(GO:0042789) |
| 0.1 | 0.3 | GO:0042727 | flavin-containing compound biosynthetic process(GO:0042727) |
| 0.1 | 1.2 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.1 | 1.0 | GO:0048304 | positive regulation of isotype switching to IgG isotypes(GO:0048304) |
| 0.1 | 0.4 | GO:0015917 | aminophospholipid transport(GO:0015917) |
| 0.1 | 1.2 | GO:0016446 | somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.1 | 0.3 | GO:0006597 | spermine biosynthetic process(GO:0006597) |
| 0.1 | 1.6 | GO:0070734 | histone H3-K27 methylation(GO:0070734) |
| 0.1 | 1.0 | GO:0006271 | DNA strand elongation involved in DNA replication(GO:0006271) |
| 0.1 | 2.0 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.1 | 0.9 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.1 | 0.3 | GO:0035822 | gene conversion(GO:0035822) |
| 0.1 | 0.5 | GO:0032525 | somite rostral/caudal axis specification(GO:0032525) |
| 0.1 | 7.8 | GO:0000079 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) |
| 0.1 | 0.7 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.1 | 0.4 | GO:0007352 | zygotic specification of dorsal/ventral axis(GO:0007352) |
| 0.1 | 0.6 | GO:0006335 | DNA replication-dependent nucleosome assembly(GO:0006335) DNA replication-dependent nucleosome organization(GO:0034723) |
| 0.1 | 0.5 | GO:0042148 | strand invasion(GO:0042148) |
| 0.1 | 1.0 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.1 | 0.4 | GO:0097167 | circadian regulation of translation(GO:0097167) |
| 0.1 | 0.3 | GO:0060266 | negative regulation of respiratory burst involved in inflammatory response(GO:0060266) |
| 0.1 | 0.4 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.1 | 3.2 | GO:0006342 | chromatin silencing(GO:0006342) |
| 0.1 | 0.4 | GO:0032927 | positive regulation of activin receptor signaling pathway(GO:0032927) |
| 0.1 | 0.5 | GO:0090666 | scaRNA localization to Cajal body(GO:0090666) |
| 0.1 | 0.4 | GO:0048104 | establishment of body hair or bristle planar orientation(GO:0048104) establishment of body hair planar orientation(GO:0048105) |
| 0.1 | 0.2 | GO:0019264 | glycine biosynthetic process from serine(GO:0019264) |
| 0.0 | 0.7 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.0 | 0.5 | GO:0046831 | regulation of RNA export from nucleus(GO:0046831) |
| 0.0 | 0.1 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.0 | 0.8 | GO:0032688 | negative regulation of interferon-beta production(GO:0032688) |
| 0.0 | 1.4 | GO:0019985 | translesion synthesis(GO:0019985) |
| 0.0 | 0.5 | GO:0021891 | olfactory bulb interneuron development(GO:0021891) |
| 0.0 | 0.7 | GO:0000732 | strand displacement(GO:0000732) |
| 0.0 | 0.9 | GO:0033962 | cytoplasmic mRNA processing body assembly(GO:0033962) |
| 0.0 | 0.5 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 0.3 | GO:0099566 | regulation of postsynaptic cytosolic calcium ion concentration(GO:0099566) |
| 0.0 | 0.6 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.0 | 0.6 | GO:0033197 | response to vitamin E(GO:0033197) |
| 0.0 | 1.7 | GO:0001510 | RNA methylation(GO:0001510) |
| 0.0 | 0.3 | GO:0098789 | pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.0 | 0.1 | GO:1900186 | negative regulation of clathrin-mediated endocytosis(GO:1900186) |
| 0.0 | 1.5 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.2 | GO:0045903 | positive regulation of translational fidelity(GO:0045903) |
| 0.0 | 0.4 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.0 | 0.1 | GO:0001757 | somite specification(GO:0001757) |
| 0.0 | 0.9 | GO:0006221 | pyrimidine nucleotide biosynthetic process(GO:0006221) |
| 0.0 | 0.3 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.0 | 0.2 | GO:0051415 | interphase microtubule nucleation by interphase microtubule organizing center(GO:0051415) microtubule nucleation by microtubule organizing center(GO:0051418) |
| 0.0 | 0.3 | GO:0048262 | determination of dorsal/ventral asymmetry(GO:0048262) |
| 0.0 | 1.8 | GO:0043550 | regulation of lipid kinase activity(GO:0043550) |
| 0.0 | 2.6 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 0.2 | GO:1904814 | regulation of protein localization to chromosome, telomeric region(GO:1904814) positive regulation of protein localization to chromosome, telomeric region(GO:1904816) |
| 0.0 | 0.5 | GO:0061157 | mRNA destabilization(GO:0061157) |
| 0.0 | 0.2 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.0 | 0.2 | GO:1903800 | positive regulation of production of miRNAs involved in gene silencing by miRNA(GO:1903800) |
| 0.0 | 0.1 | GO:0070682 | proteasome regulatory particle assembly(GO:0070682) |
| 0.0 | 1.1 | GO:0048635 | negative regulation of muscle organ development(GO:0048635) |
| 0.0 | 0.6 | GO:0071425 | hematopoietic stem cell proliferation(GO:0071425) |
| 0.0 | 0.9 | GO:0006977 | DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest(GO:0006977) |
| 0.0 | 0.8 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 2.0 | GO:0051225 | spindle assembly(GO:0051225) |
| 0.0 | 1.5 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 1.1 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.0 | 0.4 | GO:0006836 | neurotransmitter transport(GO:0006836) |
| 0.0 | 1.1 | GO:0001541 | ovarian follicle development(GO:0001541) |
| 0.0 | 0.5 | GO:0006284 | base-excision repair(GO:0006284) |
| 0.0 | 1.3 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.0 | 0.4 | GO:1904837 | beta-catenin-TCF complex assembly(GO:1904837) |
| 0.0 | 0.4 | GO:0005980 | glycogen catabolic process(GO:0005980) |
| 0.0 | 1.1 | GO:0030218 | erythrocyte differentiation(GO:0030218) |
| 0.0 | 0.6 | GO:0046677 | response to antibiotic(GO:0046677) |