Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
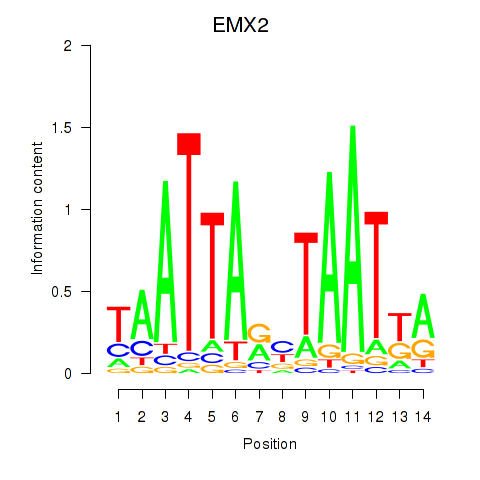
Results for EMX2
Z-value: 0.74
Transcription factors associated with EMX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
EMX2
|
ENSG00000170370.10 | EMX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| EMX2 | hg19_v2_chr10_+_119301928_119301955 | -0.15 | 5.9e-01 | Click! |
Activity profile of EMX2 motif
Sorted Z-values of EMX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of EMX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_99279928 | 2.36 |
ENST00000414521.2 |
MGAT4A |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme A |
| chr1_-_173174681 | 1.00 |
ENST00000367718.1 |
TNFSF4 |
tumor necrosis factor (ligand) superfamily, member 4 |
| chr6_+_13272904 | 0.90 |
ENST00000379335.3 ENST00000379329.1 |
PHACTR1 |
phosphatase and actin regulator 1 |
| chr12_-_25055177 | 0.88 |
ENST00000538118.1 |
BCAT1 |
branched chain amino-acid transaminase 1, cytosolic |
| chr4_-_101439148 | 0.82 |
ENST00000511970.1 ENST00000502569.1 ENST00000305864.3 |
EMCN |
endomucin |
| chr4_-_101439242 | 0.81 |
ENST00000296420.4 |
EMCN |
endomucin |
| chr6_+_26251835 | 0.79 |
ENST00000356350.2 |
HIST1H2BH |
histone cluster 1, H2bh |
| chr5_-_94417339 | 0.77 |
ENST00000429576.2 ENST00000508509.1 ENST00000510732.1 |
MCTP1 |
multiple C2 domains, transmembrane 1 |
| chr6_-_33041378 | 0.74 |
ENST00000428995.1 |
HLA-DPA1 |
major histocompatibility complex, class II, DP alpha 1 |
| chr6_+_28092338 | 0.61 |
ENST00000340487.4 |
ZSCAN16 |
zinc finger and SCAN domain containing 16 |
| chr16_+_31885079 | 0.60 |
ENST00000300870.10 ENST00000394846.3 |
ZNF267 |
zinc finger protein 267 |
| chr12_+_29302119 | 0.58 |
ENST00000536681.3 |
FAR2 |
fatty acyl CoA reductase 2 |
| chr3_-_141747950 | 0.56 |
ENST00000497579.1 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr5_-_111093167 | 0.56 |
ENST00000446294.2 ENST00000419114.2 |
NREP |
neuronal regeneration related protein |
| chr4_-_25865159 | 0.55 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr19_+_54466179 | 0.50 |
ENST00000270458.2 |
CACNG8 |
calcium channel, voltage-dependent, gamma subunit 8 |
| chr12_-_112279694 | 0.50 |
ENST00000443596.1 ENST00000442119.1 |
MAPKAPK5-AS1 |
MAPKAPK5 antisense RNA 1 |
| chr8_-_57123815 | 0.48 |
ENST00000316981.3 ENST00000423799.2 ENST00000429357.2 |
PLAG1 |
pleiomorphic adenoma gene 1 |
| chr4_-_100356291 | 0.46 |
ENST00000476959.1 ENST00000482593.1 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr5_+_112312416 | 0.46 |
ENST00000389063.2 |
DCP2 |
decapping mRNA 2 |
| chr11_-_63376013 | 0.46 |
ENST00000540943.1 |
PLA2G16 |
phospholipase A2, group XVI |
| chr6_-_26235206 | 0.46 |
ENST00000244534.5 |
HIST1H1D |
histone cluster 1, H1d |
| chr5_-_111093081 | 0.45 |
ENST00000453526.2 ENST00000509427.1 |
NREP |
neuronal regeneration related protein |
| chr11_-_107729887 | 0.42 |
ENST00000525815.1 |
SLC35F2 |
solute carrier family 35, member F2 |
| chr7_+_138145076 | 0.41 |
ENST00000343526.4 |
TRIM24 |
tripartite motif containing 24 |
| chr4_-_71532339 | 0.41 |
ENST00000254801.4 |
IGJ |
immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides |
| chr5_-_111092930 | 0.41 |
ENST00000257435.7 |
NREP |
neuronal regeneration related protein |
| chr3_+_113616317 | 0.40 |
ENST00000440446.2 ENST00000488680.1 |
GRAMD1C |
GRAM domain containing 1C |
| chr6_+_6588316 | 0.40 |
ENST00000379953.2 |
LY86 |
lymphocyte antigen 86 |
| chr14_+_24099318 | 0.37 |
ENST00000432832.2 |
DHRS2 |
dehydrogenase/reductase (SDR family) member 2 |
| chr18_-_5419797 | 0.37 |
ENST00000542146.1 ENST00000427684.2 |
EPB41L3 |
erythrocyte membrane protein band 4.1-like 3 |
| chr12_-_10007448 | 0.37 |
ENST00000538152.1 |
CLEC2B |
C-type lectin domain family 2, member B |
| chr15_+_77712993 | 0.36 |
ENST00000336216.4 ENST00000381714.3 ENST00000558651.1 |
HMG20A |
high mobility group 20A |
| chr19_+_50016610 | 0.36 |
ENST00000596975.1 |
FCGRT |
Fc fragment of IgG, receptor, transporter, alpha |
| chr4_-_100356551 | 0.35 |
ENST00000209665.4 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr6_-_26197478 | 0.35 |
ENST00000356476.2 |
HIST1H3D |
histone cluster 1, H3d |
| chr7_-_140482926 | 0.34 |
ENST00000496384.2 |
BRAF |
v-raf murine sarcoma viral oncogene homolog B |
| chr6_+_44187242 | 0.33 |
ENST00000393844.1 |
SLC29A1 |
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr6_+_63921351 | 0.33 |
ENST00000370659.1 |
FKBP1C |
FK506 binding protein 1C |
| chrX_+_77166172 | 0.32 |
ENST00000343533.5 ENST00000350425.4 ENST00000341514.6 |
ATP7A |
ATPase, Cu++ transporting, alpha polypeptide |
| chr6_+_44187334 | 0.32 |
ENST00000313248.7 ENST00000427851.2 |
SLC29A1 |
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr6_+_34725263 | 0.32 |
ENST00000374018.1 ENST00000374017.3 |
SNRPC |
small nuclear ribonucleoprotein polypeptide C |
| chr6_+_63921399 | 0.31 |
ENST00000356170.3 |
FKBP1C |
FK506 binding protein 1C |
| chr5_-_111312622 | 0.30 |
ENST00000395634.3 |
NREP |
neuronal regeneration related protein |
| chr15_-_20193370 | 0.29 |
ENST00000558565.2 |
IGHV3OR15-7 |
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chrX_-_9734004 | 0.28 |
ENST00000467482.1 ENST00000380929.2 |
GPR143 |
G protein-coupled receptor 143 |
| chr1_+_174844645 | 0.28 |
ENST00000486220.1 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr1_+_160709055 | 0.28 |
ENST00000368043.3 ENST00000368042.3 ENST00000458602.2 ENST00000458104.2 |
SLAMF7 |
SLAM family member 7 |
| chr1_+_28764653 | 0.27 |
ENST00000373836.3 |
PHACTR4 |
phosphatase and actin regulator 4 |
| chr12_-_22063787 | 0.25 |
ENST00000544039.1 |
ABCC9 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 9 |
| chr16_+_446713 | 0.25 |
ENST00000397722.1 ENST00000454619.1 |
NME4 |
NME/NM23 nucleoside diphosphate kinase 4 |
| chr6_+_34725181 | 0.24 |
ENST00000244520.5 |
SNRPC |
small nuclear ribonucleoprotein polypeptide C |
| chr4_-_186877502 | 0.24 |
ENST00000431902.1 ENST00000284776.7 ENST00000415274.1 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr4_-_186877806 | 0.24 |
ENST00000355634.5 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr4_-_68411275 | 0.24 |
ENST00000273853.6 |
CENPC |
centromere protein C |
| chr12_-_54652060 | 0.23 |
ENST00000552562.1 |
CBX5 |
chromobox homolog 5 |
| chr5_-_55412774 | 0.23 |
ENST00000434982.2 |
ANKRD55 |
ankyrin repeat domain 55 |
| chr8_-_101571933 | 0.23 |
ENST00000520311.1 |
ANKRD46 |
ankyrin repeat domain 46 |
| chr1_-_119682812 | 0.22 |
ENST00000537870.1 |
WARS2 |
tryptophanyl tRNA synthetase 2, mitochondrial |
| chr16_+_69599861 | 0.22 |
ENST00000354436.2 |
NFAT5 |
nuclear factor of activated T-cells 5, tonicity-responsive |
| chr8_-_93029865 | 0.21 |
ENST00000422361.2 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr8_+_110552337 | 0.21 |
ENST00000337573.5 |
EBAG9 |
estrogen receptor binding site associated, antigen, 9 |
| chr1_-_161147275 | 0.21 |
ENST00000319769.5 ENST00000367998.1 |
B4GALT3 |
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 3 |
| chr1_+_78470530 | 0.21 |
ENST00000370763.5 |
DNAJB4 |
DnaJ (Hsp40) homolog, subfamily B, member 4 |
| chr3_-_186288097 | 0.20 |
ENST00000446782.1 |
TBCCD1 |
TBCC domain containing 1 |
| chr4_-_185275104 | 0.19 |
ENST00000317596.3 |
RP11-290F5.2 |
RP11-290F5.2 |
| chr17_-_42100474 | 0.19 |
ENST00000585950.1 ENST00000592127.1 ENST00000589334.1 |
TMEM101 |
transmembrane protein 101 |
| chr15_-_65426174 | 0.19 |
ENST00000204549.4 |
PDCD7 |
programmed cell death 7 |
| chr4_+_74702214 | 0.19 |
ENST00000226317.5 ENST00000515050.1 |
CXCL6 |
chemokine (C-X-C motif) ligand 6 |
| chr9_+_135854091 | 0.19 |
ENST00000450530.1 ENST00000534944.1 |
GFI1B |
growth factor independent 1B transcription repressor |
| chr15_-_65903407 | 0.18 |
ENST00000395644.4 ENST00000567744.1 ENST00000568573.1 ENST00000562830.1 ENST00000569491.1 ENST00000561769.1 |
VWA9 |
von Willebrand factor A domain containing 9 |
| chr17_-_67138015 | 0.18 |
ENST00000284425.2 ENST00000590645.1 |
ABCA6 |
ATP-binding cassette, sub-family A (ABC1), member 6 |
| chr11_+_119019722 | 0.18 |
ENST00000307417.3 |
ABCG4 |
ATP-binding cassette, sub-family G (WHITE), member 4 |
| chr5_+_129083772 | 0.18 |
ENST00000564719.1 |
KIAA1024L |
KIAA1024-like |
| chr12_-_68696652 | 0.18 |
ENST00000539972.1 |
MDM1 |
Mdm1 nuclear protein homolog (mouse) |
| chr8_+_110552831 | 0.18 |
ENST00000530629.1 |
EBAG9 |
estrogen receptor binding site associated, antigen, 9 |
| chr17_+_7239821 | 0.18 |
ENST00000158762.3 ENST00000570457.2 |
ACAP1 |
ArfGAP with coiled-coil, ankyrin repeat and PH domains 1 |
| chrX_-_23926004 | 0.17 |
ENST00000379226.4 ENST00000379220.3 |
APOO |
apolipoprotein O |
| chr4_-_168155169 | 0.16 |
ENST00000534949.1 ENST00000535728.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr2_+_196313239 | 0.16 |
ENST00000413290.1 |
AC064834.1 |
AC064834.1 |
| chr12_-_53715328 | 0.16 |
ENST00000547757.1 ENST00000394384.3 ENST00000209873.4 |
AAAS |
achalasia, adrenocortical insufficiency, alacrimia |
| chr15_+_75970672 | 0.16 |
ENST00000435356.1 |
AC105020.1 |
Uncharacterized protein; cDNA FLJ12988 fis, clone NT2RP3000080 |
| chr12_-_58220078 | 0.16 |
ENST00000549039.1 |
CTDSP2 |
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 2 |
| chr14_+_102276132 | 0.15 |
ENST00000350249.3 ENST00000557621.1 ENST00000556946.1 |
PPP2R5C |
protein phosphatase 2, regulatory subunit B', gamma |
| chr11_+_92085262 | 0.15 |
ENST00000298047.6 ENST00000409404.2 ENST00000541502.1 |
FAT3 |
FAT atypical cadherin 3 |
| chr15_-_57025759 | 0.15 |
ENST00000267807.7 |
ZNF280D |
zinc finger protein 280D |
| chr15_+_64680003 | 0.15 |
ENST00000261884.3 |
TRIP4 |
thyroid hormone receptor interactor 4 |
| chr7_+_107224364 | 0.15 |
ENST00000491150.1 |
BCAP29 |
B-cell receptor-associated protein 29 |
| chr1_-_45452240 | 0.15 |
ENST00000372183.3 ENST00000372182.4 ENST00000360403.2 |
EIF2B3 |
eukaryotic translation initiation factor 2B, subunit 3 gamma, 58kDa |
| chr3_-_130745571 | 0.14 |
ENST00000514044.1 ENST00000264992.3 |
ASTE1 |
asteroid homolog 1 (Drosophila) |
| chr1_+_79115503 | 0.14 |
ENST00000370747.4 ENST00000438486.1 ENST00000545124.1 |
IFI44 |
interferon-induced protein 44 |
| chr3_-_141719195 | 0.14 |
ENST00000397991.4 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr12_+_10365404 | 0.14 |
ENST00000266458.5 ENST00000421801.2 ENST00000544284.1 ENST00000545047.1 ENST00000543602.1 ENST00000545887.1 |
GABARAPL1 |
GABA(A) receptor-associated protein like 1 |
| chr2_+_11682790 | 0.14 |
ENST00000389825.3 ENST00000381483.2 |
GREB1 |
growth regulation by estrogen in breast cancer 1 |
| chr1_+_10292308 | 0.14 |
ENST00000377081.1 |
KIF1B |
kinesin family member 1B |
| chr6_+_26204825 | 0.13 |
ENST00000360441.4 |
HIST1H4E |
histone cluster 1, H4e |
| chr1_-_190446759 | 0.13 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr6_+_28317685 | 0.13 |
ENST00000252211.2 ENST00000341464.5 ENST00000377255.3 |
ZKSCAN3 |
zinc finger with KRAB and SCAN domains 3 |
| chr10_-_115904361 | 0.12 |
ENST00000428953.1 ENST00000543782.1 |
C10orf118 |
chromosome 10 open reading frame 118 |
| chr17_+_5031687 | 0.12 |
ENST00000250066.6 ENST00000304328.5 |
USP6 |
ubiquitin specific peptidase 6 (Tre-2 oncogene) |
| chr8_+_26150628 | 0.12 |
ENST00000523925.1 ENST00000315985.7 |
PPP2R2A |
protein phosphatase 2, regulatory subunit B, alpha |
| chr1_+_235530675 | 0.12 |
ENST00000366601.3 ENST00000406207.1 ENST00000543662.1 |
TBCE |
tubulin folding cofactor E |
| chr12_+_21207503 | 0.12 |
ENST00000545916.1 |
SLCO1B7 |
solute carrier organic anion transporter family, member 1B7 (non-functional) |
| chr5_+_54455946 | 0.12 |
ENST00000503787.1 ENST00000296734.6 ENST00000515370.1 |
GPX8 |
glutathione peroxidase 8 (putative) |
| chr18_+_21693306 | 0.12 |
ENST00000540918.2 |
TTC39C |
tetratricopeptide repeat domain 39C |
| chr22_+_45725524 | 0.12 |
ENST00000405548.3 |
FAM118A |
family with sequence similarity 118, member A |
| chr8_-_13134045 | 0.12 |
ENST00000512044.2 |
DLC1 |
deleted in liver cancer 1 |
| chr5_+_178977546 | 0.11 |
ENST00000319449.4 ENST00000377001.2 |
RUFY1 |
RUN and FYVE domain containing 1 |
| chr11_-_59950622 | 0.11 |
ENST00000323961.3 ENST00000412309.2 |
MS4A6A |
membrane-spanning 4-domains, subfamily A, member 6A |
| chr2_+_32502952 | 0.11 |
ENST00000238831.4 |
YIPF4 |
Yip1 domain family, member 4 |
| chr13_-_70682590 | 0.11 |
ENST00000377844.4 |
KLHL1 |
kelch-like family member 1 |
| chr1_-_35450897 | 0.11 |
ENST00000373337.3 |
ZMYM6NB |
ZMYM6 neighbor |
| chr16_-_28634874 | 0.11 |
ENST00000395609.1 ENST00000350842.4 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr9_-_21368075 | 0.10 |
ENST00000449498.1 |
IFNA13 |
interferon, alpha 13 |
| chr1_-_67142710 | 0.10 |
ENST00000502413.2 |
AL139147.1 |
Uncharacterized protein |
| chr1_-_110283660 | 0.10 |
ENST00000361066.2 |
GSTM3 |
glutathione S-transferase mu 3 (brain) |
| chr19_+_58144529 | 0.10 |
ENST00000347302.3 ENST00000254182.7 ENST00000391703.3 ENST00000541801.1 ENST00000299871.5 ENST00000544273.1 |
ZNF211 |
zinc finger protein 211 |
| chr11_-_59950486 | 0.10 |
ENST00000426738.2 ENST00000533023.1 ENST00000420732.2 |
MS4A6A |
membrane-spanning 4-domains, subfamily A, member 6A |
| chr8_-_38008783 | 0.10 |
ENST00000276449.4 |
STAR |
steroidogenic acute regulatory protein |
| chr3_-_108248169 | 0.10 |
ENST00000273353.3 |
MYH15 |
myosin, heavy chain 15 |
| chr3_+_52245458 | 0.10 |
ENST00000459884.1 |
ALAS1 |
aminolevulinate, delta-, synthase 1 |
| chr6_-_74231444 | 0.10 |
ENST00000331523.2 ENST00000356303.2 |
EEF1A1 |
eukaryotic translation elongation factor 1 alpha 1 |
| chr4_+_80584903 | 0.10 |
ENST00000506460.1 |
RP11-452C8.1 |
RP11-452C8.1 |
| chr5_+_161495038 | 0.10 |
ENST00000393933.4 |
GABRG2 |
gamma-aminobutyric acid (GABA) A receptor, gamma 2 |
| chr11_-_59950519 | 0.10 |
ENST00000528851.1 |
MS4A6A |
membrane-spanning 4-domains, subfamily A, member 6A |
| chr16_+_56995854 | 0.09 |
ENST00000566128.1 |
CETP |
cholesteryl ester transfer protein, plasma |
| chr1_-_39347255 | 0.09 |
ENST00000454994.2 ENST00000357771.3 |
GJA9 |
gap junction protein, alpha 9, 59kDa |
| chr2_+_122494676 | 0.09 |
ENST00000455432.1 |
TSN |
translin |
| chr5_-_75919253 | 0.09 |
ENST00000296641.4 |
F2RL2 |
coagulation factor II (thrombin) receptor-like 2 |
| chr2_+_157330081 | 0.09 |
ENST00000409674.1 |
GPD2 |
glycerol-3-phosphate dehydrogenase 2 (mitochondrial) |
| chr5_-_111754948 | 0.08 |
ENST00000261486.5 |
EPB41L4A |
erythrocyte membrane protein band 4.1 like 4A |
| chr2_+_67624430 | 0.08 |
ENST00000272342.5 |
ETAA1 |
Ewing tumor-associated antigen 1 |
| chr3_+_11314099 | 0.08 |
ENST00000446450.2 ENST00000354956.5 ENST00000354449.3 ENST00000419112.1 |
ATG7 |
autophagy related 7 |
| chr11_+_74303575 | 0.08 |
ENST00000263681.2 |
POLD3 |
polymerase (DNA-directed), delta 3, accessory subunit |
| chr8_-_121457608 | 0.08 |
ENST00000306185.3 |
MRPL13 |
mitochondrial ribosomal protein L13 |
| chr1_-_20250110 | 0.08 |
ENST00000375116.3 |
PLA2G2E |
phospholipase A2, group IIE |
| chr2_-_89619904 | 0.08 |
ENST00000498574.1 |
IGKV1-39 |
immunoglobulin kappa variable 1-39 (gene/pseudogene) |
| chr5_-_88119580 | 0.08 |
ENST00000539796.1 |
MEF2C |
myocyte enhancer factor 2C |
| chr8_+_12803176 | 0.08 |
ENST00000524591.2 |
KIAA1456 |
KIAA1456 |
| chr12_+_9980113 | 0.08 |
ENST00000537723.1 |
KLRF1 |
killer cell lectin-like receptor subfamily F, member 1 |
| chr3_+_132316081 | 0.08 |
ENST00000249887.2 |
ACKR4 |
atypical chemokine receptor 4 |
| chr12_-_122985067 | 0.08 |
ENST00000540586.1 ENST00000543897.1 |
ZCCHC8 |
zinc finger, CCHC domain containing 8 |
| chr3_+_121774202 | 0.07 |
ENST00000469710.1 ENST00000493101.1 ENST00000330540.2 ENST00000264468.5 |
CD86 |
CD86 molecule |
| chr10_+_51187938 | 0.07 |
ENST00000311663.5 |
FAM21D |
family with sequence similarity 21, member D |
| chr11_+_4116054 | 0.07 |
ENST00000423050.2 |
RRM1 |
ribonucleotide reductase M1 |
| chr12_+_9980069 | 0.07 |
ENST00000354855.3 ENST00000324214.4 ENST00000279544.3 |
KLRF1 |
killer cell lectin-like receptor subfamily F, member 1 |
| chr1_-_13390765 | 0.07 |
ENST00000357367.2 |
PRAMEF8 |
PRAME family member 8 |
| chr11_-_327537 | 0.07 |
ENST00000602735.1 |
IFITM3 |
interferon induced transmembrane protein 3 |
| chr5_+_147774275 | 0.07 |
ENST00000513826.1 |
FBXO38 |
F-box protein 38 |
| chr6_+_27833034 | 0.07 |
ENST00000357320.2 |
HIST1H2AL |
histone cluster 1, H2al |
| chr15_+_84115868 | 0.07 |
ENST00000427482.2 |
SH3GL3 |
SH3-domain GRB2-like 3 |
| chr7_-_44887620 | 0.07 |
ENST00000349299.3 ENST00000521529.1 ENST00000308153.4 ENST00000350771.3 ENST00000222690.6 ENST00000381124.5 ENST00000437072.1 ENST00000446531.1 |
H2AFV |
H2A histone family, member V |
| chr8_+_21823726 | 0.07 |
ENST00000433566.4 |
XPO7 |
exportin 7 |
| chr16_-_46655538 | 0.07 |
ENST00000303383.3 |
SHCBP1 |
SHC SH2-domain binding protein 1 |
| chr1_+_206858232 | 0.07 |
ENST00000294981.4 |
MAPKAPK2 |
mitogen-activated protein kinase-activated protein kinase 2 |
| chr6_-_29343068 | 0.07 |
ENST00000396806.3 |
OR12D3 |
olfactory receptor, family 12, subfamily D, member 3 |
| chr11_+_66276550 | 0.07 |
ENST00000419755.3 |
CTD-3074O7.11 |
Bardet-Biedl syndrome 1 protein |
| chr11_-_13517565 | 0.07 |
ENST00000282091.1 ENST00000529816.1 |
PTH |
parathyroid hormone |
| chr14_+_39644387 | 0.06 |
ENST00000553331.1 ENST00000216832.4 |
PNN |
pinin, desmosome associated protein |
| chr20_+_30697298 | 0.06 |
ENST00000398022.2 |
TM9SF4 |
transmembrane 9 superfamily protein member 4 |
| chr20_+_9494987 | 0.06 |
ENST00000427562.2 ENST00000246070.2 |
LAMP5 |
lysosomal-associated membrane protein family, member 5 |
| chr4_+_156824840 | 0.06 |
ENST00000536354.2 |
TDO2 |
tryptophan 2,3-dioxygenase |
| chr6_+_110501621 | 0.06 |
ENST00000368930.1 ENST00000307731.1 |
CDC40 |
cell division cycle 40 |
| chr5_+_81575281 | 0.06 |
ENST00000380167.4 |
ATP6AP1L |
ATPase, H+ transporting, lysosomal accessory protein 1-like |
| chr9_+_111624577 | 0.06 |
ENST00000333999.3 |
ACTL7A |
actin-like 7A |
| chr7_-_16505440 | 0.06 |
ENST00000307068.4 |
SOSTDC1 |
sclerostin domain containing 1 |
| chr6_-_74233480 | 0.06 |
ENST00000455918.1 |
EEF1A1 |
eukaryotic translation elongation factor 1 alpha 1 |
| chr6_+_110501344 | 0.06 |
ENST00000368932.1 |
CDC40 |
cell division cycle 40 |
| chr10_-_47239738 | 0.05 |
ENST00000413193.2 |
AGAP10 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 10 |
| chr15_-_66816853 | 0.05 |
ENST00000568588.1 |
RPL4 |
ribosomal protein L4 |
| chr6_+_150690028 | 0.05 |
ENST00000229447.5 ENST00000344419.3 |
IYD |
iodotyrosine deiodinase |
| chr2_+_11674213 | 0.05 |
ENST00000381486.2 |
GREB1 |
growth regulation by estrogen in breast cancer 1 |
| chr5_-_43557791 | 0.05 |
ENST00000338972.4 ENST00000511321.1 ENST00000515338.1 |
PAIP1 |
poly(A) binding protein interacting protein 1 |
| chr10_-_75008620 | 0.05 |
ENST00000372950.4 |
DNAJC9 |
DnaJ (Hsp40) homolog, subfamily C, member 9 |
| chr15_+_59903975 | 0.05 |
ENST00000560585.1 ENST00000396065.1 |
GCNT3 |
glucosaminyl (N-acetyl) transferase 3, mucin type |
| chr22_+_24407642 | 0.05 |
ENST00000454754.1 ENST00000263119.5 |
CABIN1 |
calcineurin binding protein 1 |
| chr17_-_15244894 | 0.05 |
ENST00000338696.2 ENST00000543896.1 ENST00000539245.1 ENST00000539316.1 ENST00000395930.1 |
TEKT3 |
tektin 3 |
| chr1_+_146714291 | 0.05 |
ENST00000431239.1 ENST00000369259.3 ENST00000369258.4 ENST00000361293.5 |
CHD1L |
chromodomain helicase DNA binding protein 1-like |
| chr17_+_39240459 | 0.05 |
ENST00000391417.4 |
KRTAP4-7 |
keratin associated protein 4-7 |
| chr2_-_62733476 | 0.05 |
ENST00000335390.5 |
TMEM17 |
transmembrane protein 17 |
| chr17_-_34503867 | 0.05 |
ENST00000589443.1 ENST00000454519.3 ENST00000586369.1 |
TBC1D3B |
TBC1 domain family, member 3B |
| chr1_-_182921119 | 0.05 |
ENST00000423786.1 |
SHCBP1L |
SHC SH2-domain binding protein 1-like |
| chr5_+_118812237 | 0.05 |
ENST00000513628.1 |
HSD17B4 |
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr5_-_58882219 | 0.05 |
ENST00000505453.1 ENST00000360047.5 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr11_-_62609281 | 0.05 |
ENST00000525239.1 ENST00000538098.2 |
WDR74 |
WD repeat domain 74 |
| chr2_+_120687335 | 0.05 |
ENST00000544261.1 |
PTPN4 |
protein tyrosine phosphatase, non-receptor type 4 (megakaryocyte) |
| chr11_-_125773085 | 0.05 |
ENST00000227474.3 ENST00000534158.1 ENST00000529801.1 |
PUS3 |
pseudouridylate synthase 3 |
| chr2_-_207023918 | 0.05 |
ENST00000455934.2 ENST00000449699.1 ENST00000454195.1 |
NDUFS1 |
NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) |
| chr12_+_58176525 | 0.05 |
ENST00000543727.1 ENST00000540550.1 ENST00000323833.8 ENST00000350762.5 ENST00000550559.1 ENST00000548851.1 ENST00000434359.1 ENST00000457189.1 |
TSFM |
Ts translation elongation factor, mitochondrial |
| chr6_+_72926145 | 0.04 |
ENST00000425662.2 ENST00000453976.2 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr5_+_118812294 | 0.04 |
ENST00000509514.1 |
HSD17B4 |
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr12_-_71533055 | 0.04 |
ENST00000552128.1 |
TSPAN8 |
tetraspanin 8 |
| chr8_-_25281747 | 0.04 |
ENST00000421054.2 |
GNRH1 |
gonadotropin-releasing hormone 1 (luteinizing-releasing hormone) |
| chr1_-_19615744 | 0.04 |
ENST00000361640.4 |
AKR7A3 |
aldo-keto reductase family 7, member A3 (aflatoxin aldehyde reductase) |
| chr1_+_115572415 | 0.04 |
ENST00000256592.1 |
TSHB |
thyroid stimulating hormone, beta |
| chrX_+_135730373 | 0.04 |
ENST00000370628.2 |
CD40LG |
CD40 ligand |
| chr17_-_10276319 | 0.04 |
ENST00000252172.4 ENST00000418404.3 |
MYH13 |
myosin, heavy chain 13, skeletal muscle |
| chr12_-_24102576 | 0.04 |
ENST00000537393.1 ENST00000309359.1 ENST00000381381.2 ENST00000451604.2 |
SOX5 |
SRY (sex determining region Y)-box 5 |
| chr8_+_70404996 | 0.04 |
ENST00000402687.4 ENST00000419716.3 |
SULF1 |
sulfatase 1 |
| chr21_-_43735446 | 0.04 |
ENST00000398431.2 |
TFF3 |
trefoil factor 3 (intestinal) |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.2 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.2 | 0.8 | GO:0035276 | aldehyde oxidase activity(GO:0004031) ethanol binding(GO:0035276) |
| 0.2 | 0.6 | GO:0080019 | fatty-acyl-CoA reductase (alcohol-forming) activity(GO:0080019) |
| 0.2 | 0.6 | GO:0030627 | pre-mRNA 5'-splice site binding(GO:0030627) |
| 0.1 | 0.9 | GO:0052655 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.1 | 0.3 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 0.1 | 0.5 | GO:0052740 | 1-acyl-2-lysophosphatidylserine acylhydrolase activity(GO:0052740) |
| 0.1 | 0.4 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 0.1 | 0.4 | GO:0004090 | carbonyl reductase (NADPH) activity(GO:0004090) |
| 0.1 | 0.2 | GO:0034041 | sterol-transporting ATPase activity(GO:0034041) |
| 0.1 | 0.7 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.1 | 0.5 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.0 | 0.2 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.0 | 0.3 | GO:0072542 | protein phosphatase activator activity(GO:0072542) |
| 0.0 | 0.4 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.0 | 0.9 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.2 | GO:0003945 | N-acetyllactosamine synthase activity(GO:0003945) |
| 0.0 | 0.1 | GO:0003870 | 5-aminolevulinate synthase activity(GO:0003870) N-succinyltransferase activity(GO:0016749) |
| 0.0 | 0.1 | GO:0017129 | triglyceride binding(GO:0017129) |
| 0.0 | 0.2 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.0 | 0.6 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.0 | 0.3 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.0 | 0.1 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.0 | 0.2 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.0 | 0.3 | GO:0035240 | dopamine binding(GO:0035240) |
| 0.0 | 0.6 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.1 | GO:0052591 | sn-glycerol-3-phosphate:ubiquinone oxidoreductase activity(GO:0052590) sn-glycerol-3-phosphate:ubiquinone-8 oxidoreductase activity(GO:0052591) |
| 0.0 | 1.0 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.1 | GO:0032393 | MHC class I receptor activity(GO:0032393) |
| 0.0 | 0.1 | GO:0004618 | phosphoglycerate kinase activity(GO:0004618) |
| 0.0 | 0.1 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.0 | 0.5 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.1 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.0 | 0.1 | GO:0003829 | beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase activity(GO:0003829) |
| 0.0 | 0.2 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.0 | 0.1 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) steroid sulfotransferase activity(GO:0050294) |
| 0.0 | 0.1 | GO:0044594 | 3alpha,7alpha,12alpha-trihydroxy-5beta-cholest-24-enoyl-CoA hydratase activity(GO:0033989) 17-beta-hydroxysteroid dehydrogenase (NAD+) activity(GO:0044594) |
| 0.0 | 0.1 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.0 | 0.1 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 0.0 | GO:0015403 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) |
| 0.0 | 0.1 | GO:0030957 | Tat protein binding(GO:0030957) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.1 | 2.4 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.5 | REACTOME ACYL CHAIN REMODELLING OF PS | Genes involved in Acyl chain remodelling of PS |
| 0.0 | 0.9 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.5 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 0.7 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.0 | 0.5 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 1.3 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.0 | 0.6 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 0.4 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.3 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.0 | 0.6 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.2 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.4 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 0.2 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:2000525 | regulation of T cell costimulation(GO:2000523) positive regulation of T cell costimulation(GO:2000525) |
| 0.3 | 0.8 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.2 | 0.6 | GO:0010025 | wax biosynthetic process(GO:0010025) wax metabolic process(GO:0010166) |
| 0.1 | 0.9 | GO:0009099 | branched-chain amino acid biosynthetic process(GO:0009082) leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.1 | 0.6 | GO:0015862 | uridine transport(GO:0015862) |
| 0.1 | 0.3 | GO:0071284 | cellular response to lead ion(GO:0071284) |
| 0.1 | 0.3 | GO:2001045 | closure of optic fissure(GO:0061386) negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.1 | 2.4 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.1 | 0.3 | GO:0043324 | eye pigment biosynthetic process(GO:0006726) eye pigment metabolic process(GO:0042441) pigment metabolic process involved in developmental pigmentation(GO:0043324) pigment metabolic process involved in pigmentation(GO:0043474) regulation of melanosome transport(GO:1902908) |
| 0.1 | 0.3 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.1 | 0.2 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.1 | 0.5 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.1 | 0.5 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.1 | 0.6 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.1 | 0.4 | GO:0070562 | regulation of vitamin D receptor signaling pathway(GO:0070562) |
| 0.1 | 0.1 | GO:0090662 | energy coupled proton transmembrane transport, against electrochemical gradient(GO:0015988) ATP hydrolysis coupled proton transport(GO:0015991) ATP hydrolysis coupled transmembrane transport(GO:0090662) |
| 0.1 | 0.5 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.1 | 0.4 | GO:0002175 | protein localization to paranode region of axon(GO:0002175) |
| 0.1 | 0.4 | GO:0002414 | immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 0.0 | 0.1 | GO:1904647 | response to rotenone(GO:1904647) |
| 0.0 | 0.2 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.0 | 0.4 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.0 | 0.1 | GO:0048936 | peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.0 | 0.0 | GO:0051031 | tRNA export from nucleus(GO:0006409) tRNA transport(GO:0051031) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.0 | 0.1 | GO:0018894 | dibenzo-p-dioxin metabolic process(GO:0018894) |
| 0.0 | 0.0 | GO:0051180 | vitamin transport(GO:0051180) |
| 0.0 | 0.1 | GO:0090298 | late nucleophagy(GO:0044805) negative regulation of mitochondrial DNA replication(GO:0090298) negative regulation of mitochondrial DNA metabolic process(GO:1901859) single-organism membrane invagination(GO:1902534) |
| 0.0 | 0.5 | GO:0060252 | positive regulation of glial cell proliferation(GO:0060252) |
| 0.0 | 0.6 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.0 | 0.2 | GO:0019375 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.0 | 0.1 | GO:0008065 | establishment of blood-nerve barrier(GO:0008065) |
| 0.0 | 0.1 | GO:0032641 | negative regulation of tolerance induction(GO:0002644) lymphotoxin A production(GO:0032641) interleukin-4 biosynthetic process(GO:0042097) lymphotoxin A biosynthetic process(GO:0042109) regulation of interleukin-4 biosynthetic process(GO:0045402) positive regulation of interleukin-4 biosynthetic process(GO:0045404) |
| 0.0 | 0.3 | GO:0046051 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 0.0 | 0.1 | GO:0090290 | positive regulation of osteoclast proliferation(GO:0090290) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.0 | 0.1 | GO:0019442 | tryptophan catabolic process to acetyl-CoA(GO:0019442) |
| 0.0 | 0.1 | GO:2000015 | regulation of determination of dorsal identity(GO:2000015) |
| 0.0 | 0.2 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.0 | 0.1 | GO:0032057 | negative regulation of translational initiation in response to stress(GO:0032057) |
| 0.0 | 0.4 | GO:0033234 | negative regulation of protein sumoylation(GO:0033234) |
| 0.0 | 0.1 | GO:0006127 | glycerophosphate shuttle(GO:0006127) |
| 0.0 | 0.7 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.0 | 0.1 | GO:0036111 | very long-chain fatty-acyl-CoA metabolic process(GO:0036111) |
| 0.0 | 0.4 | GO:0043011 | myeloid dendritic cell differentiation(GO:0043011) |
| 0.0 | 0.2 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 0.1 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.0 | 0.0 | GO:1990637 | response to prolactin(GO:1990637) |
| 0.0 | 0.5 | GO:0003301 | physiological muscle hypertrophy(GO:0003298) physiological cardiac muscle hypertrophy(GO:0003301) cell growth involved in cardiac muscle cell development(GO:0061049) |
| 0.0 | 0.2 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.0 | 0.0 | GO:0034505 | tooth mineralization(GO:0034505) |
| 0.0 | 0.6 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.1 | GO:2000773 | negative regulation of cellular senescence(GO:2000773) |
| 0.0 | 0.0 | GO:0015888 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 0.0 | 0.2 | GO:0034374 | low-density lipoprotein particle remodeling(GO:0034374) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.1 | 0.4 | GO:0005726 | perichromatin fibrils(GO:0005726) |
| 0.0 | 0.6 | GO:0000243 | commitment complex(GO:0000243) |
| 0.0 | 0.7 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.0 | 0.5 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.5 | GO:0031332 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.0 | 0.2 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 1.4 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.6 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 0.4 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.0 | 0.1 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.0 | 0.1 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 0.0 | 0.1 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.0 | 0.5 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.3 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.0 | 0.1 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.0 | 0.1 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.0 | 0.2 | GO:0098574 | cytoplasmic side of lysosomal membrane(GO:0098574) |
| 0.0 | 0.2 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 0.6 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |