Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
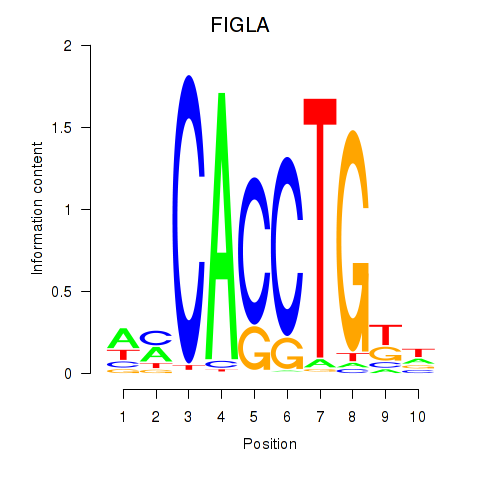
Results for FIGLA
Z-value: 1.04
Transcription factors associated with FIGLA
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
FIGLA
|
ENSG00000183733.6 | FIGLA |
Activity profile of FIGLA motif
Sorted Z-values of FIGLA motif
Network of associatons between targets according to the STRING database.
First level regulatory network of FIGLA
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chrX_-_11445856 | 3.25 |
ENST00000380736.1 |
ARHGAP6 |
Rho GTPase activating protein 6 |
| chr7_-_150675372 | 2.85 |
ENST00000262186.5 |
KCNH2 |
potassium voltage-gated channel, subfamily H (eag-related), member 2 |
| chrX_-_137793826 | 2.72 |
ENST00000315930.6 |
FGF13 |
fibroblast growth factor 13 |
| chr11_-_33913708 | 2.67 |
ENST00000257818.2 |
LMO2 |
LIM domain only 2 (rhombotin-like 1) |
| chr7_+_12727250 | 1.78 |
ENST00000404894.1 |
ARL4A |
ADP-ribosylation factor-like 4A |
| chr7_-_99698338 | 1.70 |
ENST00000354230.3 ENST00000425308.1 |
MCM7 |
minichromosome maintenance complex component 7 |
| chr17_+_45286387 | 1.60 |
ENST00000572316.1 ENST00000354968.1 ENST00000576874.1 ENST00000536623.2 |
MYL4 |
myosin, light chain 4, alkali; atrial, embryonic |
| chr6_-_42016385 | 1.50 |
ENST00000502771.1 ENST00000508143.1 ENST00000514588.1 ENST00000510503.1 ENST00000415497.2 ENST00000372988.4 |
CCND3 |
cyclin D3 |
| chr16_-_103572 | 1.42 |
ENST00000293860.5 |
POLR3K |
polymerase (RNA) III (DNA directed) polypeptide K, 12.3 kDa |
| chr8_-_91095099 | 1.29 |
ENST00000265431.3 |
CALB1 |
calbindin 1, 28kDa |
| chr17_+_76164639 | 1.23 |
ENST00000225777.3 ENST00000585591.1 ENST00000589711.1 ENST00000588282.1 ENST00000589168.1 |
SYNGR2 |
synaptogyrin 2 |
| chr1_+_45478568 | 1.19 |
ENST00000428106.1 |
UROD |
uroporphyrinogen decarboxylase |
| chr17_+_76165213 | 1.15 |
ENST00000590201.1 |
SYNGR2 |
synaptogyrin 2 |
| chr7_+_128470431 | 1.13 |
ENST00000325888.8 ENST00000346177.6 |
FLNC |
filamin C, gamma |
| chr5_-_54281407 | 1.12 |
ENST00000381403.4 |
ESM1 |
endothelial cell-specific molecule 1 |
| chr14_+_31343747 | 1.10 |
ENST00000216361.4 ENST00000396618.3 ENST00000475087.1 |
COCH |
cochlin |
| chr11_-_118972575 | 1.09 |
ENST00000432443.2 |
DPAGT1 |
dolichyl-phosphate (UDP-N-acetylglucosamine) N-acetylglucosaminephosphotransferase 1 (GlcNAc-1-P transferase) |
| chr11_+_118958689 | 1.04 |
ENST00000535253.1 ENST00000392841.1 |
HMBS |
hydroxymethylbilane synthase |
| chr6_+_34204642 | 0.90 |
ENST00000347617.6 ENST00000401473.3 ENST00000311487.5 ENST00000447654.1 ENST00000395004.3 |
HMGA1 |
high mobility group AT-hook 1 |
| chr6_-_39197226 | 0.90 |
ENST00000359534.3 |
KCNK5 |
potassium channel, subfamily K, member 5 |
| chr6_-_31697255 | 0.89 |
ENST00000436437.1 |
DDAH2 |
dimethylarginine dimethylaminohydrolase 2 |
| chr4_-_90757364 | 0.89 |
ENST00000508895.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr5_-_94620239 | 0.88 |
ENST00000515393.1 |
MCTP1 |
multiple C2 domains, transmembrane 1 |
| chr5_-_54281491 | 0.85 |
ENST00000381405.4 |
ESM1 |
endothelial cell-specific molecule 1 |
| chr11_-_122931881 | 0.85 |
ENST00000526110.1 ENST00000227378.3 |
HSPA8 |
heat shock 70kDa protein 8 |
| chr1_-_39339777 | 0.84 |
ENST00000397572.2 |
MYCBP |
MYC binding protein |
| chr17_-_7165662 | 0.83 |
ENST00000571881.2 ENST00000360325.7 |
CLDN7 |
claudin 7 |
| chr7_-_8301768 | 0.83 |
ENST00000265577.7 |
ICA1 |
islet cell autoantigen 1, 69kDa |
| chr2_+_170590321 | 0.82 |
ENST00000392647.2 |
KLHL23 |
kelch-like family member 23 |
| chr7_-_8301682 | 0.78 |
ENST00000396675.3 ENST00000430867.1 |
ICA1 |
islet cell autoantigen 1, 69kDa |
| chr1_-_27216729 | 0.75 |
ENST00000431781.2 ENST00000374135.4 |
GPN2 |
GPN-loop GTPase 2 |
| chr12_+_58176525 | 0.72 |
ENST00000543727.1 ENST00000540550.1 ENST00000323833.8 ENST00000350762.5 ENST00000550559.1 ENST00000548851.1 ENST00000434359.1 ENST00000457189.1 |
TSFM |
Ts translation elongation factor, mitochondrial |
| chr6_+_31783291 | 0.72 |
ENST00000375651.5 ENST00000608703.1 ENST00000458062.2 |
HSPA1A |
heat shock 70kDa protein 1A |
| chrX_-_153714994 | 0.70 |
ENST00000369660.4 |
UBL4A |
ubiquitin-like 4A |
| chr6_-_27440460 | 0.69 |
ENST00000377419.1 |
ZNF184 |
zinc finger protein 184 |
| chr1_-_11120057 | 0.68 |
ENST00000376957.2 |
SRM |
spermidine synthase |
| chr8_+_145438870 | 0.68 |
ENST00000527931.1 |
FAM203B |
family with sequence similarity 203, member B |
| chr1_-_160990886 | 0.67 |
ENST00000537746.1 |
F11R |
F11 receptor |
| chr6_-_27440837 | 0.67 |
ENST00000211936.6 |
ZNF184 |
zinc finger protein 184 |
| chr7_-_8302164 | 0.66 |
ENST00000447326.1 ENST00000406470.2 |
ICA1 |
islet cell autoantigen 1, 69kDa |
| chr1_+_66258846 | 0.66 |
ENST00000341517.4 |
PDE4B |
phosphodiesterase 4B, cAMP-specific |
| chr7_-_8301869 | 0.65 |
ENST00000402384.3 |
ICA1 |
islet cell autoantigen 1, 69kDa |
| chr6_+_41888926 | 0.63 |
ENST00000230340.4 |
BYSL |
bystin-like |
| chr6_-_31697563 | 0.62 |
ENST00000375789.2 ENST00000416410.1 |
DDAH2 |
dimethylarginine dimethylaminohydrolase 2 |
| chr4_-_76598544 | 0.62 |
ENST00000515457.1 ENST00000357854.3 |
G3BP2 |
GTPase activating protein (SH3 domain) binding protein 2 |
| chr1_+_100315613 | 0.61 |
ENST00000361915.3 |
AGL |
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase |
| chr6_+_96463840 | 0.60 |
ENST00000302103.5 |
FUT9 |
fucosyltransferase 9 (alpha (1,3) fucosyltransferase) |
| chr16_+_103816 | 0.60 |
ENST00000383018.3 ENST00000417493.1 |
SNRNP25 |
small nuclear ribonucleoprotein 25kDa (U11/U12) |
| chr5_+_34656569 | 0.59 |
ENST00000428746.2 |
RAI14 |
retinoic acid induced 14 |
| chr18_+_48405419 | 0.58 |
ENST00000321341.5 |
ME2 |
malic enzyme 2, NAD(+)-dependent, mitochondrial |
| chr17_-_31204124 | 0.58 |
ENST00000579584.1 ENST00000318217.5 ENST00000583621.1 |
MYO1D |
myosin ID |
| chr12_+_32655048 | 0.57 |
ENST00000427716.2 ENST00000266482.3 |
FGD4 |
FYVE, RhoGEF and PH domain containing 4 |
| chr7_+_130794846 | 0.57 |
ENST00000421797.2 |
MKLN1 |
muskelin 1, intracellular mediator containing kelch motifs |
| chr19_-_3786253 | 0.56 |
ENST00000585778.1 |
MATK |
megakaryocyte-associated tyrosine kinase |
| chr1_-_167905225 | 0.55 |
ENST00000367846.4 |
MPC2 |
mitochondrial pyruvate carrier 2 |
| chr1_+_100316041 | 0.53 |
ENST00000370165.3 ENST00000370163.3 ENST00000294724.4 |
AGL |
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase |
| chr7_-_11871815 | 0.53 |
ENST00000423059.4 |
THSD7A |
thrombospondin, type I, domain containing 7A |
| chr21_-_39870339 | 0.52 |
ENST00000429727.2 ENST00000398905.1 ENST00000398907.1 ENST00000453032.2 ENST00000288319.7 |
ERG |
v-ets avian erythroblastosis virus E26 oncogene homolog |
| chr1_-_150669500 | 0.52 |
ENST00000271732.3 |
GOLPH3L |
golgi phosphoprotein 3-like |
| chr22_-_37882395 | 0.52 |
ENST00000416983.3 ENST00000424765.2 ENST00000356998.3 |
MFNG |
MFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase |
| chr17_+_40985407 | 0.50 |
ENST00000586114.1 ENST00000590720.1 ENST00000585805.1 ENST00000541124.1 ENST00000441946.2 ENST00000591152.1 ENST00000589469.1 ENST00000293362.3 ENST00000592169.1 |
PSME3 |
proteasome (prosome, macropain) activator subunit 3 (PA28 gamma; Ki) |
| chr8_-_130951940 | 0.50 |
ENST00000522250.1 ENST00000522941.1 ENST00000522746.1 ENST00000520204.1 ENST00000519070.1 ENST00000520254.1 ENST00000519824.2 ENST00000519540.1 |
FAM49B |
family with sequence similarity 49, member B |
| chr8_-_99837856 | 0.49 |
ENST00000518165.1 ENST00000419617.2 |
STK3 |
serine/threonine kinase 3 |
| chr6_-_35888824 | 0.48 |
ENST00000361690.3 ENST00000512445.1 |
SRPK1 |
SRSF protein kinase 1 |
| chr16_-_2314222 | 0.48 |
ENST00000566397.1 |
RNPS1 |
RNA binding protein S1, serine-rich domain |
| chr8_-_8318847 | 0.48 |
ENST00000521218.1 |
CTA-398F10.2 |
CTA-398F10.2 |
| chr17_+_39411636 | 0.48 |
ENST00000394008.1 |
KRTAP9-9 |
keratin associated protein 9-9 |
| chr11_+_128634589 | 0.47 |
ENST00000281428.8 |
FLI1 |
Fli-1 proto-oncogene, ETS transcription factor |
| chr9_+_90112767 | 0.45 |
ENST00000408954.3 |
DAPK1 |
death-associated protein kinase 1 |
| chr18_+_48405463 | 0.45 |
ENST00000382927.3 |
ME2 |
malic enzyme 2, NAD(+)-dependent, mitochondrial |
| chr3_-_69435224 | 0.43 |
ENST00000398540.3 |
FRMD4B |
FERM domain containing 4B |
| chr5_-_177423243 | 0.42 |
ENST00000308304.2 |
PROP1 |
PROP paired-like homeobox 1 |
| chr16_-_30006922 | 0.42 |
ENST00000564026.1 |
HIRIP3 |
HIRA interacting protein 3 |
| chr3_-_46904946 | 0.40 |
ENST00000292327.4 |
MYL3 |
myosin, light chain 3, alkali; ventricular, skeletal, slow |
| chr14_-_20923195 | 0.40 |
ENST00000206542.4 |
OSGEP |
O-sialoglycoprotein endopeptidase |
| chr6_-_31938700 | 0.40 |
ENST00000495340.1 |
DXO |
decapping exoribonuclease |
| chr5_+_122110691 | 0.40 |
ENST00000379516.2 ENST00000505934.1 ENST00000514949.1 |
SNX2 |
sorting nexin 2 |
| chr1_+_26737253 | 0.39 |
ENST00000326279.6 |
LIN28A |
lin-28 homolog A (C. elegans) |
| chr1_-_111991850 | 0.39 |
ENST00000411751.2 |
WDR77 |
WD repeat domain 77 |
| chr2_+_228029281 | 0.39 |
ENST00000396578.3 |
COL4A3 |
collagen, type IV, alpha 3 (Goodpasture antigen) |
| chr10_-_30638090 | 0.39 |
ENST00000421701.1 ENST00000263063.4 |
MTPAP |
mitochondrial poly(A) polymerase |
| chr1_-_154842741 | 0.38 |
ENST00000271915.4 |
KCNN3 |
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 |
| chr3_-_46904918 | 0.38 |
ENST00000395869.1 |
MYL3 |
myosin, light chain 3, alkali; ventricular, skeletal, slow |
| chr8_-_102216925 | 0.38 |
ENST00000517844.1 |
ZNF706 |
zinc finger protein 706 |
| chr11_-_65626753 | 0.38 |
ENST00000526975.1 ENST00000531413.1 |
CFL1 |
cofilin 1 (non-muscle) |
| chr6_-_133055815 | 0.38 |
ENST00000509351.1 ENST00000417437.2 ENST00000414302.2 ENST00000423615.2 ENST00000427187.2 ENST00000275223.3 ENST00000519686.2 |
VNN3 |
vanin 3 |
| chr19_-_46272106 | 0.38 |
ENST00000560168.1 |
SIX5 |
SIX homeobox 5 |
| chr5_-_132113083 | 0.37 |
ENST00000296873.7 |
SEPT8 |
septin 8 |
| chr13_+_26828275 | 0.37 |
ENST00000381527.3 |
CDK8 |
cyclin-dependent kinase 8 |
| chr9_+_140172200 | 0.37 |
ENST00000357503.2 |
TOR4A |
torsin family 4, member A |
| chr5_-_137071756 | 0.37 |
ENST00000394937.3 ENST00000309755.4 |
KLHL3 |
kelch-like family member 3 |
| chr6_-_41888843 | 0.36 |
ENST00000434077.1 ENST00000409312.1 |
MED20 |
mediator complex subunit 20 |
| chr9_+_133710453 | 0.36 |
ENST00000318560.5 |
ABL1 |
c-abl oncogene 1, non-receptor tyrosine kinase |
| chr11_-_123525289 | 0.35 |
ENST00000392770.2 ENST00000299333.3 ENST00000530277.1 |
SCN3B |
sodium channel, voltage-gated, type III, beta subunit |
| chr4_-_139163491 | 0.35 |
ENST00000280612.5 |
SLC7A11 |
solute carrier family 7 (anionic amino acid transporter light chain, xc- system), member 11 |
| chr9_+_90112117 | 0.34 |
ENST00000358077.5 |
DAPK1 |
death-associated protein kinase 1 |
| chr6_-_31550192 | 0.32 |
ENST00000429299.2 ENST00000446745.2 |
LTB |
lymphotoxin beta (TNF superfamily, member 3) |
| chr6_-_133055896 | 0.32 |
ENST00000367927.5 ENST00000425515.2 ENST00000207771.3 ENST00000392393.3 ENST00000450865.2 ENST00000392394.2 |
VNN3 |
vanin 3 |
| chr13_+_112721913 | 0.31 |
ENST00000330949.1 |
SOX1 |
SRY (sex determining region Y)-box 1 |
| chr6_-_35888905 | 0.31 |
ENST00000510290.1 ENST00000423325.2 ENST00000373822.1 |
SRPK1 |
SRSF protein kinase 1 |
| chr19_-_42916499 | 0.31 |
ENST00000601189.1 ENST00000599211.1 |
LIPE |
lipase, hormone-sensitive |
| chr13_-_41635512 | 0.31 |
ENST00000405737.2 |
ELF1 |
E74-like factor 1 (ets domain transcription factor) |
| chr4_-_140223614 | 0.31 |
ENST00000394223.1 |
NDUFC1 |
NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 1, 6kDa |
| chr19_+_50180317 | 0.31 |
ENST00000534465.1 |
PRMT1 |
protein arginine methyltransferase 1 |
| chr4_-_140223670 | 0.30 |
ENST00000394228.1 ENST00000539387.1 |
NDUFC1 |
NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 1, 6kDa |
| chr17_+_68165657 | 0.29 |
ENST00000243457.3 |
KCNJ2 |
potassium inwardly-rectifying channel, subfamily J, member 2 |
| chr7_+_6629621 | 0.29 |
ENST00000344417.5 |
C7orf26 |
chromosome 7 open reading frame 26 |
| chr1_-_40367668 | 0.28 |
ENST00000397332.2 ENST00000429311.1 |
MYCL |
v-myc avian myelocytomatosis viral oncogene lung carcinoma derived homolog |
| chr5_-_132948216 | 0.28 |
ENST00000265342.7 |
FSTL4 |
follistatin-like 4 |
| chr4_-_151936865 | 0.28 |
ENST00000535741.1 |
LRBA |
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr3_+_148545586 | 0.27 |
ENST00000282957.4 ENST00000468341.1 |
CPB1 |
carboxypeptidase B1 (tissue) |
| chr11_+_71249071 | 0.27 |
ENST00000398534.3 |
KRTAP5-8 |
keratin associated protein 5-8 |
| chr5_-_141257954 | 0.27 |
ENST00000456271.1 ENST00000394536.3 ENST00000503492.1 ENST00000287008.3 |
PCDH1 |
protocadherin 1 |
| chr17_-_34890665 | 0.26 |
ENST00000586007.1 |
MYO19 |
myosin XIX |
| chr4_-_164534657 | 0.26 |
ENST00000339875.5 |
MARCH1 |
membrane-associated ring finger (C3HC4) 1, E3 ubiquitin protein ligase |
| chr10_+_18689637 | 0.26 |
ENST00000377315.4 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr11_-_108093329 | 0.26 |
ENST00000278612.8 |
NPAT |
nuclear protein, ataxia-telangiectasia locus |
| chr6_-_33160231 | 0.26 |
ENST00000395194.1 ENST00000457788.1 ENST00000341947.2 ENST00000357486.1 ENST00000374714.1 ENST00000374713.1 ENST00000395197.1 ENST00000374712.1 ENST00000361917.1 ENST00000374708.4 |
COL11A2 |
collagen, type XI, alpha 2 |
| chr14_-_50999307 | 0.25 |
ENST00000013125.4 |
MAP4K5 |
mitogen-activated protein kinase kinase kinase kinase 5 |
| chr4_-_103749205 | 0.25 |
ENST00000508249.1 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chr11_-_19223523 | 0.25 |
ENST00000265968.3 |
CSRP3 |
cysteine and glycine-rich protein 3 (cardiac LIM protein) |
| chr6_-_33281979 | 0.24 |
ENST00000426633.2 ENST00000467025.1 |
TAPBP |
TAP binding protein (tapasin) |
| chr6_+_63921351 | 0.24 |
ENST00000370659.1 |
FKBP1C |
FK506 binding protein 1C |
| chr1_-_201391149 | 0.24 |
ENST00000555948.1 ENST00000556362.1 |
TNNI1 |
troponin I type 1 (skeletal, slow) |
| chr17_-_39222131 | 0.24 |
ENST00000394015.2 |
KRTAP2-4 |
keratin associated protein 2-4 |
| chr13_-_113862948 | 0.24 |
ENST00000375457.2 ENST00000375477.1 ENST00000246505.5 ENST00000337344.4 ENST00000375479.2 |
PCID2 |
PCI domain containing 2 |
| chr19_-_10230540 | 0.24 |
ENST00000589454.1 |
EIF3G |
eukaryotic translation initiation factor 3, subunit G |
| chr1_+_154244987 | 0.23 |
ENST00000328703.7 ENST00000457918.2 ENST00000483970.2 ENST00000435087.1 ENST00000532105.1 |
HAX1 |
HCLS1 associated protein X-1 |
| chr6_-_41888814 | 0.23 |
ENST00000409060.1 ENST00000265350.4 |
MED20 |
mediator complex subunit 20 |
| chr12_+_50497784 | 0.23 |
ENST00000548814.1 |
GPD1 |
glycerol-3-phosphate dehydrogenase 1 (soluble) |
| chr19_-_10230562 | 0.23 |
ENST00000587146.1 ENST00000588709.1 ENST00000253108.4 |
EIF3G |
eukaryotic translation initiation factor 3, subunit G |
| chr7_+_50348268 | 0.23 |
ENST00000438033.1 ENST00000439701.1 |
IKZF1 |
IKAROS family zinc finger 1 (Ikaros) |
| chr3_+_35721106 | 0.22 |
ENST00000474696.1 ENST00000412048.1 ENST00000396482.2 ENST00000432682.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr15_-_67546963 | 0.22 |
ENST00000561452.1 ENST00000261880.5 |
AAGAB |
alpha- and gamma-adaptin binding protein |
| chr7_+_20370746 | 0.22 |
ENST00000222573.4 |
ITGB8 |
integrin, beta 8 |
| chr12_+_79258547 | 0.22 |
ENST00000457153.2 |
SYT1 |
synaptotagmin I |
| chrX_+_151806637 | 0.22 |
ENST00000370306.2 |
GABRQ |
gamma-aminobutyric acid (GABA) A receptor, theta |
| chr1_-_230561475 | 0.21 |
ENST00000391860.1 |
PGBD5 |
piggyBac transposable element derived 5 |
| chr15_+_91411810 | 0.21 |
ENST00000268171.3 |
FURIN |
furin (paired basic amino acid cleaving enzyme) |
| chr19_+_50936142 | 0.21 |
ENST00000357701.5 |
MYBPC2 |
myosin binding protein C, fast type |
| chr8_-_103424986 | 0.21 |
ENST00000521922.1 |
UBR5 |
ubiquitin protein ligase E3 component n-recognin 5 |
| chr6_-_36953833 | 0.21 |
ENST00000538808.1 ENST00000460219.1 ENST00000373616.5 ENST00000373627.5 |
MTCH1 |
mitochondrial carrier 1 |
| chr5_+_34656331 | 0.20 |
ENST00000265109.3 |
RAI14 |
retinoic acid induced 14 |
| chrX_-_153775426 | 0.20 |
ENST00000393562.2 |
G6PD |
glucose-6-phosphate dehydrogenase |
| chr1_-_111991908 | 0.20 |
ENST00000235090.5 |
WDR77 |
WD repeat domain 77 |
| chr4_-_103749179 | 0.20 |
ENST00000502690.1 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chr8_-_103424916 | 0.19 |
ENST00000220959.4 |
UBR5 |
ubiquitin protein ligase E3 component n-recognin 5 |
| chr16_+_77224732 | 0.19 |
ENST00000569610.1 ENST00000248248.3 ENST00000567291.1 ENST00000320859.6 ENST00000563612.1 ENST00000563279.1 |
MON1B |
MON1 secretory trafficking family member B |
| chr4_+_81951957 | 0.19 |
ENST00000282701.2 |
BMP3 |
bone morphogenetic protein 3 |
| chr1_+_110754094 | 0.19 |
ENST00000369787.3 ENST00000413138.3 ENST00000438661.2 |
KCNC4 |
potassium voltage-gated channel, Shaw-related subfamily, member 4 |
| chr15_+_92937058 | 0.19 |
ENST00000268164.3 |
ST8SIA2 |
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 2 |
| chr16_-_46865047 | 0.19 |
ENST00000394806.2 |
C16orf87 |
chromosome 16 open reading frame 87 |
| chr6_-_114292449 | 0.18 |
ENST00000519065.1 |
HDAC2 |
histone deacetylase 2 |
| chrX_-_6146876 | 0.18 |
ENST00000381095.3 |
NLGN4X |
neuroligin 4, X-linked |
| chr5_+_52856456 | 0.18 |
ENST00000296684.5 ENST00000506765.1 |
NDUFS4 |
NADH dehydrogenase (ubiquinone) Fe-S protein 4, 18kDa (NADH-coenzyme Q reductase) |
| chr1_-_171621815 | 0.18 |
ENST00000037502.6 |
MYOC |
myocilin, trabecular meshwork inducible glucocorticoid response |
| chr1_+_168250194 | 0.18 |
ENST00000367821.3 |
TBX19 |
T-box 19 |
| chrX_-_118827333 | 0.18 |
ENST00000360156.7 ENST00000354228.4 ENST00000489216.1 ENST00000354416.3 ENST00000394610.1 ENST00000343984.5 |
SEPT6 |
septin 6 |
| chr11_-_84634447 | 0.18 |
ENST00000532653.1 |
DLG2 |
discs, large homolog 2 (Drosophila) |
| chr20_-_1974692 | 0.18 |
ENST00000217305.2 ENST00000539905.1 |
PDYN |
prodynorphin |
| chr22_+_31090793 | 0.18 |
ENST00000332585.6 ENST00000382310.3 ENST00000446658.2 |
OSBP2 |
oxysterol binding protein 2 |
| chr2_+_136343820 | 0.18 |
ENST00000410054.1 |
R3HDM1 |
R3H domain containing 1 |
| chr2_-_175629164 | 0.18 |
ENST00000409323.1 ENST00000261007.5 ENST00000348749.5 |
CHRNA1 |
cholinergic receptor, nicotinic, alpha 1 (muscle) |
| chr2_+_233390863 | 0.17 |
ENST00000449596.1 ENST00000543200.1 |
CHRND |
cholinergic receptor, nicotinic, delta (muscle) |
| chr15_+_45694523 | 0.17 |
ENST00000305560.6 |
SPATA5L1 |
spermatogenesis associated 5-like 1 |
| chr6_+_33172407 | 0.17 |
ENST00000374662.3 |
HSD17B8 |
hydroxysteroid (17-beta) dehydrogenase 8 |
| chr14_-_106586656 | 0.17 |
ENST00000390602.2 |
IGHV3-13 |
immunoglobulin heavy variable 3-13 |
| chrX_-_16887963 | 0.16 |
ENST00000380084.4 |
RBBP7 |
retinoblastoma binding protein 7 |
| chr7_-_37488834 | 0.16 |
ENST00000310758.4 |
ELMO1 |
engulfment and cell motility 1 |
| chr2_+_37571845 | 0.15 |
ENST00000537448.1 |
QPCT |
glutaminyl-peptide cyclotransferase |
| chr7_-_44229022 | 0.15 |
ENST00000403799.3 |
GCK |
glucokinase (hexokinase 4) |
| chrX_+_128872998 | 0.15 |
ENST00000371106.3 |
XPNPEP2 |
X-prolyl aminopeptidase (aminopeptidase P) 2, membrane-bound |
| chr14_-_55369525 | 0.14 |
ENST00000543643.2 ENST00000536224.2 ENST00000395514.1 ENST00000491895.2 |
GCH1 |
GTP cyclohydrolase 1 |
| chr2_+_173600514 | 0.14 |
ENST00000264111.6 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr3_-_49851313 | 0.14 |
ENST00000333486.3 |
UBA7 |
ubiquitin-like modifier activating enzyme 7 |
| chr1_-_42384343 | 0.14 |
ENST00000372584.1 |
HIVEP3 |
human immunodeficiency virus type I enhancer binding protein 3 |
| chr9_+_136399929 | 0.14 |
ENST00000393060.1 |
ADAMTSL2 |
ADAMTS-like 2 |
| chr17_+_7462031 | 0.14 |
ENST00000380535.4 |
TNFSF13 |
tumor necrosis factor (ligand) superfamily, member 13 |
| chr2_+_176987088 | 0.14 |
ENST00000249499.6 |
HOXD9 |
homeobox D9 |
| chr11_-_84634217 | 0.14 |
ENST00000524982.1 |
DLG2 |
discs, large homolog 2 (Drosophila) |
| chr12_+_122150646 | 0.14 |
ENST00000449592.2 |
TMEM120B |
transmembrane protein 120B |
| chr11_-_2182388 | 0.13 |
ENST00000421783.1 ENST00000397262.1 ENST00000250971.3 ENST00000381330.4 ENST00000397270.1 |
INS INS-IGF2 |
insulin INS-IGF2 readthrough |
| chr7_+_77428066 | 0.13 |
ENST00000422959.2 ENST00000307305.8 ENST00000424760.1 |
PHTF2 |
putative homeodomain transcription factor 2 |
| chr2_+_37571717 | 0.13 |
ENST00000338415.3 ENST00000404976.1 |
QPCT |
glutaminyl-peptide cyclotransferase |
| chr2_-_175629135 | 0.13 |
ENST00000409542.1 ENST00000409219.1 |
CHRNA1 |
cholinergic receptor, nicotinic, alpha 1 (muscle) |
| chr1_+_22307592 | 0.13 |
ENST00000400277.2 |
CELA3B |
chymotrypsin-like elastase family, member 3B |
| chr16_+_30383613 | 0.13 |
ENST00000568749.1 |
MYLPF |
myosin light chain, phosphorylatable, fast skeletal muscle |
| chr1_+_10509971 | 0.13 |
ENST00000320498.4 |
CORT |
cortistatin |
| chr14_-_106967788 | 0.12 |
ENST00000390622.2 |
IGHV1-46 |
immunoglobulin heavy variable 1-46 |
| chr13_+_115000339 | 0.12 |
ENST00000360383.3 ENST00000375312.3 |
CDC16 |
cell division cycle 16 |
| chr16_-_12009833 | 0.12 |
ENST00000420576.2 |
GSPT1 |
G1 to S phase transition 1 |
| chr3_+_182511266 | 0.12 |
ENST00000323116.5 ENST00000493826.1 |
ATP11B |
ATPase, class VI, type 11B |
| chr12_+_48357340 | 0.12 |
ENST00000256686.6 ENST00000549288.1 ENST00000552561.1 ENST00000546749.1 ENST00000552546.1 ENST00000550552.1 |
TMEM106C |
transmembrane protein 106C |
| chr12_-_48119340 | 0.11 |
ENST00000229003.3 |
ENDOU |
endonuclease, polyU-specific |
| chr17_-_56429500 | 0.11 |
ENST00000225504.3 |
SUPT4H1 |
suppressor of Ty 4 homolog 1 (S. cerevisiae) |
| chr8_-_103425047 | 0.11 |
ENST00000520539.1 |
UBR5 |
ubiquitin protein ligase E3 component n-recognin 5 |
| chr7_+_93535817 | 0.11 |
ENST00000248572.5 |
GNGT1 |
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr3_-_49066811 | 0.11 |
ENST00000442157.1 ENST00000326739.4 |
IMPDH2 |
IMP (inosine 5'-monophosphate) dehydrogenase 2 |
| chr2_+_233390890 | 0.11 |
ENST00000258385.3 ENST00000536614.1 ENST00000457943.2 |
CHRND |
cholinergic receptor, nicotinic, delta (muscle) |
| chr1_-_55266865 | 0.11 |
ENST00000371274.4 |
TTC22 |
tetratricopeptide repeat domain 22 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.7 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.1 | 0.9 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.1 | 2.2 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 3.1 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.3 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.1 | 0.8 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 0.1 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.0 | 1.1 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 1.1 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 1.1 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.6 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.0 | 1.1 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.0 | 0.7 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 1.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 0.7 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 4.4 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 2.3 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.3 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.2 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.4 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.0 | 0.2 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.0 | 0.3 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.0 | 0.7 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.4 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 0.1 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.4 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.8 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.5 | REACTOME MRNA 3 END PROCESSING | Genes involved in mRNA 3'-end processing |
| 0.0 | 0.2 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 2.9 | GO:1902303 | regulation of heart rate by hormone(GO:0003064) negative regulation of potassium ion export(GO:1902303) |
| 0.5 | 2.7 | GO:1990834 | response to odorant(GO:1990834) |
| 0.3 | 2.0 | GO:1902202 | regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 0.3 | 2.6 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.2 | 0.7 | GO:1990022 | RNA polymerase II complex import to nucleus(GO:0044376) RNA polymerase III complex localization to nucleus(GO:1990022) |
| 0.2 | 1.1 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.2 | 0.7 | GO:0070426 | positive regulation of nucleotide-binding oligomerization domain containing signaling pathway(GO:0070426) positive regulation of nucleotide-binding oligomerization domain containing 2 signaling pathway(GO:0070434) |
| 0.2 | 0.9 | GO:0051621 | negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) regulation of peroxidase activity(GO:2000468) positive regulation of peroxidase activity(GO:2000470) |
| 0.2 | 2.2 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) |
| 0.2 | 0.9 | GO:1902904 | negative regulation of fibril organization(GO:1902904) chaperone-mediated autophagy translocation complex disassembly(GO:1904764) |
| 0.1 | 1.0 | GO:1902031 | regulation of NADP metabolic process(GO:1902031) |
| 0.1 | 1.3 | GO:0035502 | metanephric part of ureteric bud development(GO:0035502) |
| 0.1 | 0.5 | GO:0006850 | mitochondrial pyruvate transport(GO:0006850) mitochondrial pyruvate transmembrane transport(GO:1902361) |
| 0.1 | 0.4 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.1 | 0.4 | GO:0001922 | B-1 B cell homeostasis(GO:0001922) B cell proliferation involved in immune response(GO:0002322) |
| 0.1 | 0.7 | GO:0071816 | tail-anchored membrane protein insertion into ER membrane(GO:0071816) |
| 0.1 | 2.7 | GO:0042789 | mRNA transcription from RNA polymerase II promoter(GO:0042789) |
| 0.1 | 3.3 | GO:0051895 | negative regulation of focal adhesion assembly(GO:0051895) |
| 0.1 | 0.5 | GO:2000504 | positive regulation of blood vessel remodeling(GO:2000504) |
| 0.1 | 0.5 | GO:0060800 | regulation of cell differentiation involved in embryonic placenta development(GO:0060800) |
| 0.1 | 0.6 | GO:0060528 | secretory columnal luminar epithelial cell differentiation involved in prostate glandular acinus development(GO:0060528) |
| 0.1 | 0.3 | GO:0098746 | fast, calcium ion-dependent exocytosis of neurotransmitter(GO:0098746) |
| 0.1 | 0.7 | GO:0008295 | spermidine biosynthetic process(GO:0008295) |
| 0.1 | 0.5 | GO:2000769 | regulation of unidimensional cell growth(GO:0051510) negative regulation of unidimensional cell growth(GO:0051511) establishment of cell polarity regulating cell shape(GO:0071964) regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000769) positive regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000771) regulation of establishment of cell polarity regulating cell shape(GO:2000782) positive regulation of establishment of cell polarity regulating cell shape(GO:2000784) regulation of barbed-end actin filament capping(GO:2000812) positive regulation of barbed-end actin filament capping(GO:2000814) |
| 0.1 | 0.6 | GO:0061502 | early endosome to recycling endosome transport(GO:0061502) |
| 0.1 | 0.3 | GO:0017186 | peptidyl-pyroglutamic acid biosynthetic process, using glutaminyl-peptide cyclotransferase(GO:0017186) |
| 0.1 | 0.4 | GO:0014859 | negative regulation of skeletal muscle cell proliferation(GO:0014859) negative regulation of skeletal muscle satellite cell proliferation(GO:1902723) |
| 0.1 | 1.5 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.1 | 0.3 | GO:0031548 | regulation of brain-derived neurotrophic factor receptor signaling pathway(GO:0031548) |
| 0.1 | 0.4 | GO:0048852 | hypophysis morphogenesis(GO:0048850) diencephalon morphogenesis(GO:0048852) |
| 0.1 | 0.6 | GO:0048630 | skeletal muscle tissue growth(GO:0048630) |
| 0.1 | 0.5 | GO:0035709 | memory T cell activation(GO:0035709) regulation of memory T cell activation(GO:2000567) positive regulation of memory T cell activation(GO:2000568) |
| 0.1 | 0.3 | GO:0035995 | detection of muscle stretch(GO:0035995) |
| 0.1 | 0.8 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.1 | 0.4 | GO:0010587 | miRNA catabolic process(GO:0010587) |
| 0.1 | 0.5 | GO:1904879 | positive regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1904879) |
| 0.1 | 0.2 | GO:0006072 | glycerol-3-phosphate metabolic process(GO:0006072) |
| 0.1 | 2.4 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 0.1 | 0.5 | GO:1901314 | negative regulation of histone ubiquitination(GO:0033183) histone H2A K63-linked ubiquitination(GO:0070535) negative regulation of protein K63-linked ubiquitination(GO:1900045) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.1 | 0.3 | GO:0090076 | relaxation of skeletal muscle(GO:0090076) |
| 0.1 | 0.2 | GO:0032900 | negative regulation of low-density lipoprotein particle receptor catabolic process(GO:0032804) negative regulation of neurotrophin production(GO:0032900) negative regulation of transforming growth factor beta1 production(GO:0032911) |
| 0.1 | 0.4 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.1 | 0.7 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.1 | 0.7 | GO:0070129 | regulation of mitochondrial translation(GO:0070129) |
| 0.1 | 0.7 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.1 | 0.3 | GO:0060023 | soft palate development(GO:0060023) |
| 0.1 | 0.3 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.1 | 0.3 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.1 | 0.2 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.1 | 0.3 | GO:2000980 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 0.1 | 0.7 | GO:1901898 | negative regulation of relaxation of cardiac muscle(GO:1901898) |
| 0.1 | 0.2 | GO:0010286 | heat acclimation(GO:0010286) cellular heat acclimation(GO:0070370) |
| 0.1 | 0.4 | GO:0070294 | renal sodium ion absorption(GO:0070294) |
| 0.1 | 0.3 | GO:0046985 | positive regulation of hemoglobin biosynthetic process(GO:0046985) |
| 0.1 | 0.6 | GO:0042354 | fucose catabolic process(GO:0019317) L-fucose metabolic process(GO:0042354) L-fucose catabolic process(GO:0042355) |
| 0.0 | 0.1 | GO:0051066 | dihydrobiopterin metabolic process(GO:0051066) |
| 0.0 | 0.2 | GO:0019086 | late viral transcription(GO:0019086) |
| 0.0 | 0.2 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.0 | 0.2 | GO:0014734 | skeletal muscle hypertrophy(GO:0014734) |
| 0.0 | 0.3 | GO:0060573 | ventral spinal cord interneuron specification(GO:0021521) cell fate specification involved in pattern specification(GO:0060573) |
| 0.0 | 0.1 | GO:0033861 | negative regulation of NAD(P)H oxidase activity(GO:0033861) |
| 0.0 | 0.8 | GO:0071447 | cellular response to hydroperoxide(GO:0071447) |
| 0.0 | 1.4 | GO:0006379 | mRNA cleavage(GO:0006379) |
| 0.0 | 1.8 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.0 | 0.4 | GO:1902455 | negative regulation of stem cell population maintenance(GO:1902455) |
| 0.0 | 0.7 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 0.2 | GO:0061198 | fungiform papilla formation(GO:0061198) |
| 0.0 | 0.6 | GO:0007253 | cytoplasmic sequestering of NF-kappaB(GO:0007253) stress granule assembly(GO:0034063) |
| 0.0 | 0.2 | GO:0061635 | regulation of protein complex stability(GO:0061635) |
| 0.0 | 1.1 | GO:0005980 | glycogen catabolic process(GO:0005980) |
| 0.0 | 0.1 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 0.0 | 0.8 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.0 | 0.2 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.0 | 0.4 | GO:0038063 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) |
| 0.0 | 0.4 | GO:0071044 | histone mRNA catabolic process(GO:0071044) |
| 0.0 | 0.1 | GO:0015917 | aminophospholipid transport(GO:0015917) |
| 0.0 | 0.4 | GO:0071028 | nuclear RNA surveillance(GO:0071027) nuclear mRNA surveillance(GO:0071028) |
| 0.0 | 0.4 | GO:0034498 | early endosome to Golgi transport(GO:0034498) |
| 0.0 | 0.3 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.0 | 0.2 | GO:0090394 | negative regulation of excitatory postsynaptic potential(GO:0090394) |
| 0.0 | 0.2 | GO:0075525 | viral translational termination-reinitiation(GO:0075525) |
| 0.0 | 0.5 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.0 | 1.5 | GO:0046626 | regulation of insulin receptor signaling pathway(GO:0046626) |
| 0.0 | 0.2 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.0 | 0.1 | GO:0060481 | lobar bronchus epithelium development(GO:0060481) |
| 0.0 | 0.2 | GO:0001573 | ganglioside metabolic process(GO:0001573) |
| 0.0 | 0.4 | GO:0001829 | trophectodermal cell differentiation(GO:0001829) |
| 0.0 | 0.1 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.0 | 0.1 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.0 | 0.2 | GO:0014883 | transition between fast and slow fiber(GO:0014883) |
| 0.0 | 0.4 | GO:0050860 | negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.0 | 0.1 | GO:0006701 | progesterone biosynthetic process(GO:0006701) |
| 0.0 | 0.5 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.0 | 0.2 | GO:0045161 | neuronal ion channel clustering(GO:0045161) |
| 0.0 | 0.0 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 0.0 | 0.7 | GO:0048747 | muscle fiber development(GO:0048747) |
| 0.0 | 0.2 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.0 | 0.1 | GO:0000712 | resolution of meiotic recombination intermediates(GO:0000712) |
| 0.0 | 0.1 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.0 | 0.3 | GO:0046710 | GDP metabolic process(GO:0046710) |
| 0.0 | 0.0 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.0 | 0.2 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.0 | 0.0 | GO:0098907 | T-tubule organization(GO:0033292) protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) regulation of SA node cell action potential(GO:0098907) |
| 0.0 | 0.3 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.0 | 0.1 | GO:0044320 | cellular response to leptin stimulus(GO:0044320) |
| 0.0 | 2.8 | GO:0006836 | neurotransmitter transport(GO:0006836) |
| 0.0 | 0.7 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 3.2 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 2.0 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.8 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.0 | 1.8 | PID ATR PATHWAY | ATR signaling pathway |
| 0.0 | 0.7 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 0.3 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.0 | 0.8 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 1.0 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.7 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.0 | 0.6 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 0.8 | PID IFNG PATHWAY | IFN-gamma pathway |
| 0.0 | 0.9 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.9 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.0 | 1.1 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 0.4 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.9 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.2 | 0.7 | GO:0071818 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 1.7 | GO:0042555 | MCM complex(GO:0042555) |
| 0.1 | 0.9 | GO:0098575 | lumenal side of lysosomal membrane(GO:0098575) |
| 0.1 | 0.7 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.1 | 0.5 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.1 | 0.9 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.1 | 0.4 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.1 | 0.2 | GO:0034686 | integrin alphav-beta8 complex(GO:0034686) |
| 0.1 | 0.4 | GO:0030905 | retromer, tubulation complex(GO:0030905) |
| 0.1 | 5.6 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.1 | 0.2 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.1 | 0.5 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.1 | 1.3 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.0 | 0.9 | GO:0034709 | methylosome(GO:0034709) |
| 0.0 | 0.4 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.0 | 0.2 | GO:0042825 | TAP complex(GO:0042825) |
| 0.0 | 0.2 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.0 | 2.8 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.0 | 0.4 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.0 | 0.6 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.0 | 0.8 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.0 | 0.3 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.0 | 2.4 | GO:0031672 | A band(GO:0031672) |
| 0.0 | 0.9 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 0.9 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 0.1 | GO:0032044 | DSIF complex(GO:0032044) |
| 0.0 | 0.3 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.6 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 0.7 | GO:0000930 | gamma-tubulin complex(GO:0000930) |
| 0.0 | 2.1 | GO:0016528 | sarcoplasm(GO:0016528) |
| 0.0 | 0.9 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.3 | GO:0098643 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.0 | 0.8 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.2 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.0 | 0.5 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 1.2 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 0.1 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 1.3 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.2 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.3 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 0.5 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.0 | 0.5 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.0 | 0.7 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.2 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.0 | 0.0 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.0 | 0.2 | GO:0097227 | sperm annulus(GO:0097227) |
| 0.0 | 1.4 | GO:0001726 | ruffle(GO:0001726) |
| 0.0 | 0.3 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.0 | 0.7 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.2 | GO:0005861 | troponin complex(GO:0005861) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.0 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 0.6 | 2.4 | GO:0032038 | myosin II heavy chain binding(GO:0032038) |
| 0.5 | 2.9 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.4 | 1.2 | GO:0004853 | uroporphyrinogen decarboxylase activity(GO:0004853) |
| 0.4 | 1.1 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.3 | 1.5 | GO:0016403 | dimethylargininase activity(GO:0016403) |
| 0.2 | 0.7 | GO:0004766 | spermidine synthase activity(GO:0004766) |
| 0.2 | 1.3 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.2 | 0.9 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.2 | 1.6 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.2 | 1.0 | GO:0004471 | malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) oxaloacetate decarboxylase activity(GO:0008948) |
| 0.1 | 1.1 | GO:0090599 | alpha-glucosidase activity(GO:0090599) |
| 0.1 | 0.7 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.1 | 0.4 | GO:0034353 | RNA pyrophosphohydrolase activity(GO:0034353) |
| 0.1 | 0.5 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.1 | 0.4 | GO:0002134 | pyrimidine nucleoside binding(GO:0001884) UTP binding(GO:0002134) pyrimidine ribonucleoside binding(GO:0032551) |
| 0.1 | 0.4 | GO:0004515 | nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.1 | 0.3 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.1 | 0.3 | GO:0016603 | glutaminyl-peptide cyclotransferase activity(GO:0016603) |
| 0.1 | 0.5 | GO:0023025 | MHC class Ib protein complex binding(GO:0023025) MHC class Ib protein binding, via antigen binding groove(GO:0023030) |
| 0.1 | 0.4 | GO:1990460 | leptin receptor binding(GO:1990460) |
| 0.1 | 0.4 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.1 | 0.6 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.1 | 0.5 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.1 | 0.7 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.1 | 0.4 | GO:0086006 | voltage-gated sodium channel activity involved in cardiac muscle cell action potential(GO:0086006) |
| 0.1 | 0.4 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) |
| 0.1 | 0.5 | GO:0050833 | pyruvate transmembrane transporter activity(GO:0050833) |
| 0.1 | 0.2 | GO:0004345 | glucose-6-phosphate dehydrogenase activity(GO:0004345) |
| 0.1 | 0.9 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 0.2 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) |
| 0.1 | 0.9 | GO:0016274 | arginine N-methyltransferase activity(GO:0016273) protein-arginine N-methyltransferase activity(GO:0016274) |
| 0.1 | 0.3 | GO:0030348 | syntaxin-3 binding(GO:0030348) |
| 0.1 | 1.3 | GO:0001056 | RNA polymerase III activity(GO:0001056) |
| 0.1 | 2.5 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.0 | 2.7 | GO:0043425 | bHLH transcription factor binding(GO:0043425) |
| 0.0 | 0.2 | GO:0016901 | glycerol-3-phosphate dehydrogenase activity(GO:0004368) oxidoreductase activity, acting on the CH-OH group of donors, quinone or similar compound as acceptor(GO:0016901) |
| 0.0 | 0.2 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.0 | 0.3 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 0.0 | 1.8 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.0 | 0.4 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.0 | 0.4 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.0 | 0.4 | GO:0002151 | G-quadruplex RNA binding(GO:0002151) |
| 0.0 | 0.2 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.0 | 0.2 | GO:0070404 | NADH binding(GO:0070404) |
| 0.0 | 0.5 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.0 | 0.3 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.0 | 0.3 | GO:0086056 | voltage-gated calcium channel activity involved in AV node cell action potential(GO:0086056) |
| 0.0 | 1.1 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.8 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.2 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.4 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.1 | GO:0050436 | microfibril binding(GO:0050436) |
| 0.0 | 3.3 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 0.7 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.1 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.0 | 0.7 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.0 | 0.2 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.0 | 0.6 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.0 | 0.1 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.0 | 0.1 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 0.3 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 0.3 | GO:0086008 | voltage-gated potassium channel activity involved in cardiac muscle cell action potential repolarization(GO:0086008) |
| 0.0 | 0.2 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.0 | 0.6 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) |
| 0.0 | 0.1 | GO:0004396 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.0 | 0.4 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.0 | 0.5 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.0 | 0.6 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.4 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.2 | GO:0019966 | interleukin-1 binding(GO:0019966) |
| 0.0 | 0.2 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.0 | 0.3 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.1 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.0 | 0.0 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.0 | 0.2 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 0.5 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.7 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.2 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 1.0 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.0 | 0.2 | GO:0005326 | neurotransmitter transporter activity(GO:0005326) |