Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
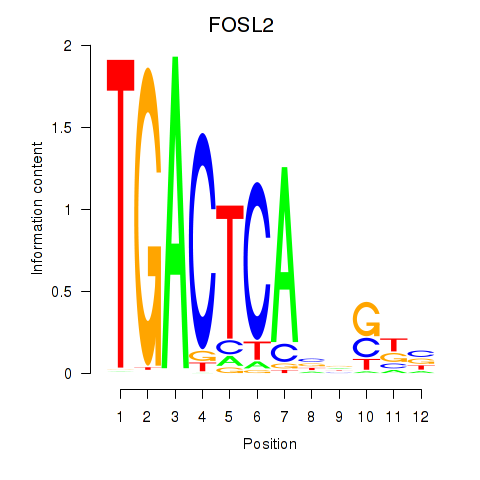
Results for FOSL2_SMARCC1
Z-value: 2.24
Transcription factors associated with FOSL2_SMARCC1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
FOSL2
|
ENSG00000075426.7 | FOSL2 |
|
SMARCC1
|
ENSG00000173473.6 | SMARCC1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SMARCC1 | hg19_v2_chr3_-_47823298_47823423 | -0.87 | 1.3e-05 | Click! |
| FOSL2 | hg19_v2_chr2_+_28615669_28615733 | 0.57 | 2.2e-02 | Click! |
Activity profile of FOSL2_SMARCC1 motif
Sorted Z-values of FOSL2_SMARCC1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of FOSL2_SMARCC1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr18_+_21452804 | 17.04 |
ENST00000269217.6 |
LAMA3 |
laminin, alpha 3 |
| chr1_-_151965048 | 16.27 |
ENST00000368809.1 |
S100A10 |
S100 calcium binding protein A10 |
| chr18_+_21452964 | 13.83 |
ENST00000587184.1 |
LAMA3 |
laminin, alpha 3 |
| chr1_+_223889285 | 12.26 |
ENST00000433674.2 |
CAPN2 |
calpain 2, (m/II) large subunit |
| chr11_-_6341844 | 10.41 |
ENST00000303927.3 |
PRKCDBP |
protein kinase C, delta binding protein |
| chr7_+_129932974 | 8.99 |
ENST00000445470.2 ENST00000222482.4 ENST00000492072.1 ENST00000473956.1 ENST00000493259.1 ENST00000486598.1 |
CPA4 |
carboxypeptidase A4 |
| chr17_-_39769005 | 7.97 |
ENST00000301653.4 ENST00000593067.1 |
KRT16 |
keratin 16 |
| chr11_-_102668879 | 7.69 |
ENST00000315274.6 |
MMP1 |
matrix metallopeptidase 1 (interstitial collagenase) |
| chr12_+_13349650 | 7.26 |
ENST00000256951.5 ENST00000431267.2 ENST00000542474.1 ENST00000544053.1 |
EMP1 |
epithelial membrane protein 1 |
| chr4_+_169753156 | 7.22 |
ENST00000393726.3 ENST00000507735.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr7_+_116312411 | 6.94 |
ENST00000456159.1 ENST00000397752.3 ENST00000318493.6 |
MET |
met proto-oncogene |
| chr11_+_35639735 | 6.54 |
ENST00000317811.4 |
FJX1 |
four jointed box 1 (Drosophila) |
| chr18_+_61442629 | 6.15 |
ENST00000398019.2 ENST00000540675.1 |
SERPINB7 |
serpin peptidase inhibitor, clade B (ovalbumin), member 7 |
| chr1_+_150480576 | 5.95 |
ENST00000346569.6 |
ECM1 |
extracellular matrix protein 1 |
| chr1_+_150480551 | 5.93 |
ENST00000369049.4 ENST00000369047.4 |
ECM1 |
extracellular matrix protein 1 |
| chr1_-_153538011 | 5.30 |
ENST00000368707.4 |
S100A2 |
S100 calcium binding protein A2 |
| chr11_+_101983176 | 5.24 |
ENST00000524575.1 |
YAP1 |
Yes-associated protein 1 |
| chr5_+_135394840 | 5.21 |
ENST00000503087.1 |
TGFBI |
transforming growth factor, beta-induced, 68kDa |
| chr10_+_17270214 | 5.16 |
ENST00000544301.1 |
VIM |
vimentin |
| chr3_-_49395705 | 4.95 |
ENST00000419349.1 |
GPX1 |
glutathione peroxidase 1 |
| chr19_+_39279838 | 4.76 |
ENST00000314980.4 |
LGALS7B |
lectin, galactoside-binding, soluble, 7B |
| chr16_-_122619 | 4.54 |
ENST00000262316.6 |
RHBDF1 |
rhomboid 5 homolog 1 (Drosophila) |
| chrX_-_10851762 | 4.43 |
ENST00000380785.1 ENST00000380787.1 |
MID1 |
midline 1 (Opitz/BBB syndrome) |
| chr13_+_102142296 | 4.41 |
ENST00000376162.3 |
ITGBL1 |
integrin, beta-like 1 (with EGF-like repeat domains) |
| chr7_+_55177416 | 4.35 |
ENST00000450046.1 ENST00000454757.2 |
EGFR |
epidermal growth factor receptor |
| chr7_+_134528635 | 4.16 |
ENST00000445569.2 |
CALD1 |
caldesmon 1 |
| chr18_+_61637159 | 3.94 |
ENST00000397985.2 ENST00000353706.2 ENST00000542677.1 ENST00000397988.3 ENST00000448851.1 |
SERPINB8 |
serpin peptidase inhibitor, clade B (ovalbumin), member 8 |
| chr15_+_67420441 | 3.71 |
ENST00000558894.1 |
SMAD3 |
SMAD family member 3 |
| chr11_-_65667997 | 3.71 |
ENST00000312562.2 ENST00000534222.1 |
FOSL1 |
FOS-like antigen 1 |
| chr2_-_175712270 | 3.71 |
ENST00000295497.7 ENST00000444394.1 |
CHN1 |
chimerin 1 |
| chr3_+_30647994 | 3.66 |
ENST00000295754.5 |
TGFBR2 |
transforming growth factor, beta receptor II (70/80kDa) |
| chr3_-_32022733 | 3.60 |
ENST00000438237.2 ENST00000396556.2 |
OSBPL10 |
oxysterol binding protein-like 10 |
| chr19_-_44174330 | 3.54 |
ENST00000340093.3 |
PLAUR |
plasminogen activator, urokinase receptor |
| chr6_+_83072923 | 3.54 |
ENST00000535040.1 |
TPBG |
trophoblast glycoprotein |
| chr19_-_44174305 | 3.52 |
ENST00000601723.1 ENST00000339082.3 |
PLAUR |
plasminogen activator, urokinase receptor |
| chr1_+_26606608 | 3.51 |
ENST00000319041.6 |
SH3BGRL3 |
SH3 domain binding glutamic acid-rich protein like 3 |
| chr10_+_75670862 | 3.47 |
ENST00000446342.1 ENST00000372764.3 ENST00000372762.4 |
PLAU |
plasminogen activator, urokinase |
| chr7_+_94023873 | 3.28 |
ENST00000297268.6 |
COL1A2 |
collagen, type I, alpha 2 |
| chr3_-_151034734 | 3.27 |
ENST00000260843.4 |
GPR87 |
G protein-coupled receptor 87 |
| chr11_+_10326612 | 3.23 |
ENST00000534464.1 ENST00000530439.1 ENST00000524948.1 ENST00000528655.1 ENST00000526492.1 ENST00000525063.1 |
ADM |
adrenomedullin |
| chr11_-_65667884 | 3.17 |
ENST00000448083.2 ENST00000531493.1 ENST00000532401.1 |
FOSL1 |
FOS-like antigen 1 |
| chr9_-_35112376 | 3.13 |
ENST00000488109.2 |
FAM214B |
family with sequence similarity 214, member B |
| chr11_-_6341724 | 3.11 |
ENST00000530979.1 |
PRKCDBP |
protein kinase C, delta binding protein |
| chr3_+_30648066 | 3.11 |
ENST00000359013.4 |
TGFBR2 |
transforming growth factor, beta receptor II (70/80kDa) |
| chr17_-_39743139 | 3.04 |
ENST00000167586.6 |
KRT14 |
keratin 14 |
| chr3_-_149095652 | 3.03 |
ENST00000305366.3 |
TM4SF1 |
transmembrane 4 L six family member 1 |
| chr6_-_84140757 | 2.93 |
ENST00000541327.1 ENST00000369705.3 ENST00000543031.1 |
ME1 |
malic enzyme 1, NADP(+)-dependent, cytosolic |
| chr10_+_121410882 | 2.91 |
ENST00000369085.3 |
BAG3 |
BCL2-associated athanogene 3 |
| chr11_-_2950642 | 2.90 |
ENST00000314222.4 |
PHLDA2 |
pleckstrin homology-like domain, family A, member 2 |
| chr17_+_35851570 | 2.88 |
ENST00000394386.1 |
DUSP14 |
dual specificity phosphatase 14 |
| chr3_+_69134124 | 2.88 |
ENST00000478935.1 |
ARL6IP5 |
ADP-ribosylation-like factor 6 interacting protein 5 |
| chr12_+_53491220 | 2.87 |
ENST00000548547.1 ENST00000301464.3 |
IGFBP6 |
insulin-like growth factor binding protein 6 |
| chr3_+_69134080 | 2.83 |
ENST00000273258.3 |
ARL6IP5 |
ADP-ribosylation-like factor 6 interacting protein 5 |
| chr6_-_138428613 | 2.80 |
ENST00000421351.3 |
PERP |
PERP, TP53 apoptosis effector |
| chr19_-_39264072 | 2.79 |
ENST00000599035.1 ENST00000378626.4 |
LGALS7 |
lectin, galactoside-binding, soluble, 7 |
| chr11_+_35198243 | 2.75 |
ENST00000528455.1 |
CD44 |
CD44 molecule (Indian blood group) |
| chr7_-_107643674 | 2.74 |
ENST00000222399.6 |
LAMB1 |
laminin, beta 1 |
| chr11_-_111794446 | 2.71 |
ENST00000527950.1 |
CRYAB |
crystallin, alpha B |
| chr12_-_91573249 | 2.70 |
ENST00000550099.1 ENST00000546391.1 ENST00000551354.1 |
DCN |
decorin |
| chr8_+_22446763 | 2.69 |
ENST00000450780.2 ENST00000430850.2 ENST00000447849.1 |
AC037459.4 |
Uncharacterized protein |
| chr9_-_35685452 | 2.68 |
ENST00000607559.1 |
TPM2 |
tropomyosin 2 (beta) |
| chrX_+_135251783 | 2.66 |
ENST00000394153.2 |
FHL1 |
four and a half LIM domains 1 |
| chr1_+_26605618 | 2.62 |
ENST00000270792.5 |
SH3BGRL3 |
SH3 domain binding glutamic acid-rich protein like 3 |
| chr11_-_66103867 | 2.59 |
ENST00000424433.2 |
RIN1 |
Ras and Rab interactor 1 |
| chr17_+_79650962 | 2.59 |
ENST00000329138.4 |
HGS |
hepatocyte growth factor-regulated tyrosine kinase substrate |
| chr12_-_91573316 | 2.57 |
ENST00000393155.1 |
DCN |
decorin |
| chr12_-_91573132 | 2.50 |
ENST00000550563.1 ENST00000546370.1 |
DCN |
decorin |
| chr18_+_61445007 | 2.50 |
ENST00000447428.1 ENST00000546027.1 |
SERPINB7 |
serpin peptidase inhibitor, clade B (ovalbumin), member 7 |
| chr11_+_35198118 | 2.47 |
ENST00000525211.1 ENST00000526000.1 ENST00000279452.6 ENST00000527889.1 |
CD44 |
CD44 molecule (Indian blood group) |
| chr1_+_36621697 | 2.43 |
ENST00000373150.4 ENST00000373151.2 |
MAP7D1 |
MAP7 domain containing 1 |
| chr1_+_156095951 | 2.41 |
ENST00000448611.2 ENST00000368297.1 |
LMNA |
lamin A/C |
| chr10_-_93392811 | 2.40 |
ENST00000238994.5 |
PPP1R3C |
protein phosphatase 1, regulatory subunit 3C |
| chr16_+_57662138 | 2.36 |
ENST00000562414.1 ENST00000561969.1 ENST00000562631.1 ENST00000563445.1 ENST00000565338.1 ENST00000567702.1 |
GPR56 |
G protein-coupled receptor 56 |
| chr3_-_114343768 | 2.30 |
ENST00000393785.2 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr2_-_163099885 | 2.28 |
ENST00000443424.1 |
FAP |
fibroblast activation protein, alpha |
| chr19_+_36630454 | 2.28 |
ENST00000246533.3 |
CAPNS1 |
calpain, small subunit 1 |
| chr1_+_169077172 | 2.24 |
ENST00000499679.3 |
ATP1B1 |
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr1_-_152009460 | 2.24 |
ENST00000271638.2 |
S100A11 |
S100 calcium binding protein A11 |
| chr12_-_52845910 | 2.23 |
ENST00000252252.3 |
KRT6B |
keratin 6B |
| chr3_+_154797877 | 2.23 |
ENST00000462745.1 ENST00000493237.1 |
MME |
membrane metallo-endopeptidase |
| chr17_+_21191341 | 2.23 |
ENST00000526076.2 ENST00000361818.5 ENST00000316920.6 |
MAP2K3 |
mitogen-activated protein kinase kinase 3 |
| chr7_+_48128194 | 2.23 |
ENST00000416681.1 ENST00000331803.4 ENST00000432131.1 |
UPP1 |
uridine phosphorylase 1 |
| chr7_+_48128316 | 2.22 |
ENST00000341253.4 |
UPP1 |
uridine phosphorylase 1 |
| chr18_+_34124507 | 2.21 |
ENST00000591635.1 |
FHOD3 |
formin homology 2 domain containing 3 |
| chr2_-_163100045 | 2.19 |
ENST00000188790.4 |
FAP |
fibroblast activation protein, alpha |
| chr3_-_48632593 | 2.18 |
ENST00000454817.1 ENST00000328333.8 |
COL7A1 |
collagen, type VII, alpha 1 |
| chr5_+_35856951 | 2.18 |
ENST00000303115.3 ENST00000343305.4 ENST00000506850.1 ENST00000511982.1 |
IL7R |
interleukin 7 receptor |
| chr16_+_57662419 | 2.18 |
ENST00000388812.4 ENST00000538815.1 ENST00000456916.1 ENST00000567154.1 ENST00000388813.5 ENST00000562558.1 ENST00000566271.2 |
GPR56 |
G protein-coupled receptor 56 |
| chr1_+_36621529 | 2.18 |
ENST00000316156.4 |
MAP7D1 |
MAP7 domain containing 1 |
| chr3_+_187930719 | 2.12 |
ENST00000312675.4 |
LPP |
LIM domain containing preferred translocation partner in lipoma |
| chr11_+_131240373 | 2.11 |
ENST00000374791.3 ENST00000436745.1 |
NTM |
neurotrimin |
| chr1_-_209824643 | 2.11 |
ENST00000391911.1 ENST00000415782.1 |
LAMB3 |
laminin, beta 3 |
| chr3_-_99833333 | 2.10 |
ENST00000354552.3 ENST00000331335.5 ENST00000398326.2 |
FILIP1L |
filamin A interacting protein 1-like |
| chr17_-_18161870 | 2.09 |
ENST00000579294.1 ENST00000545457.2 ENST00000379450.4 ENST00000578558.1 |
FLII |
flightless I homolog (Drosophila) |
| chr19_-_51523275 | 2.08 |
ENST00000309958.3 |
KLK10 |
kallikrein-related peptidase 10 |
| chr3_+_154797428 | 2.07 |
ENST00000460393.1 |
MME |
membrane metallo-endopeptidase |
| chr15_+_41136586 | 2.05 |
ENST00000431806.1 |
SPINT1 |
serine peptidase inhibitor, Kunitz type 1 |
| chrX_+_135252050 | 2.00 |
ENST00000449474.1 ENST00000345434.3 |
FHL1 |
four and a half LIM domains 1 |
| chr11_+_844406 | 2.00 |
ENST00000397404.1 |
TSPAN4 |
tetraspanin 4 |
| chr12_-_56236690 | 1.99 |
ENST00000322569.4 |
MMP19 |
matrix metallopeptidase 19 |
| chr5_+_140729649 | 1.95 |
ENST00000523390.1 |
PCDHGB1 |
protocadherin gamma subfamily B, 1 |
| chr1_-_113247543 | 1.94 |
ENST00000414971.1 ENST00000534717.1 |
RHOC |
ras homolog family member C |
| chr5_-_176923803 | 1.94 |
ENST00000506161.1 |
PDLIM7 |
PDZ and LIM domain 7 (enigma) |
| chr4_-_102268484 | 1.92 |
ENST00000394853.4 |
PPP3CA |
protein phosphatase 3, catalytic subunit, alpha isozyme |
| chr19_-_51504411 | 1.92 |
ENST00000593490.1 |
KLK8 |
kallikrein-related peptidase 8 |
| chr14_+_96722152 | 1.89 |
ENST00000216629.6 |
BDKRB1 |
bradykinin receptor B1 |
| chr5_-_176923846 | 1.87 |
ENST00000506537.1 |
PDLIM7 |
PDZ and LIM domain 7 (enigma) |
| chr1_+_183155373 | 1.86 |
ENST00000493293.1 ENST00000264144.4 |
LAMC2 |
laminin, gamma 2 |
| chr11_-_111783919 | 1.86 |
ENST00000531198.1 ENST00000533879.1 |
CRYAB |
crystallin, alpha B |
| chrX_+_49028265 | 1.86 |
ENST00000376322.3 ENST00000376327.5 |
PLP2 |
proteolipid protein 2 (colonic epithelium-enriched) |
| chr1_-_156675368 | 1.86 |
ENST00000368222.3 |
CRABP2 |
cellular retinoic acid binding protein 2 |
| chr3_-_81792780 | 1.84 |
ENST00000489715.1 |
GBE1 |
glucan (1,4-alpha-), branching enzyme 1 |
| chr16_+_15596123 | 1.82 |
ENST00000452191.2 |
C16orf45 |
chromosome 16 open reading frame 45 |
| chr16_+_89988259 | 1.82 |
ENST00000554444.1 ENST00000556565.1 |
TUBB3 |
Tubulin beta-3 chain |
| chr18_+_56338750 | 1.82 |
ENST00000345724.3 |
MALT1 |
mucosa associated lymphoid tissue lymphoma translocation gene 1 |
| chr5_-_138534071 | 1.81 |
ENST00000394817.2 |
SIL1 |
SIL1 nucleotide exchange factor |
| chr19_-_51523412 | 1.80 |
ENST00000391805.1 ENST00000599077.1 |
KLK10 |
kallikrein-related peptidase 10 |
| chr19_+_36630855 | 1.79 |
ENST00000589146.1 |
CAPNS1 |
calpain, small subunit 1 |
| chr20_-_44455976 | 1.77 |
ENST00000372555.3 |
TNNC2 |
troponin C type 2 (fast) |
| chr20_+_4666882 | 1.77 |
ENST00000379440.4 ENST00000430350.2 |
PRNP |
prion protein |
| chr3_+_105085734 | 1.76 |
ENST00000306107.5 |
ALCAM |
activated leukocyte cell adhesion molecule |
| chr17_-_27503770 | 1.73 |
ENST00000533112.1 |
MYO18A |
myosin XVIIIA |
| chr11_+_844067 | 1.72 |
ENST00000397406.1 ENST00000409543.2 ENST00000525201.1 |
TSPAN4 |
tetraspanin 4 |
| chr3_-_87040233 | 1.71 |
ENST00000398399.2 |
VGLL3 |
vestigial like 3 (Drosophila) |
| chr3_-_169587621 | 1.69 |
ENST00000523069.1 ENST00000316428.5 ENST00000264676.5 |
LRRC31 |
leucine rich repeat containing 31 |
| chr4_-_102268628 | 1.69 |
ENST00000323055.6 ENST00000512215.1 ENST00000394854.3 |
PPP3CA |
protein phosphatase 3, catalytic subunit, alpha isozyme |
| chrX_+_135251835 | 1.68 |
ENST00000456445.1 |
FHL1 |
four and a half LIM domains 1 |
| chrX_-_48901012 | 1.67 |
ENST00000315869.7 |
TFE3 |
transcription factor binding to IGHM enhancer 3 |
| chr2_+_170366203 | 1.65 |
ENST00000284669.1 |
KLHL41 |
kelch-like family member 41 |
| chr19_+_36630961 | 1.64 |
ENST00000587718.1 ENST00000592483.1 ENST00000590874.1 ENST00000588815.1 |
CAPNS1 |
calpain, small subunit 1 |
| chr5_+_138089100 | 1.62 |
ENST00000520339.1 ENST00000355078.5 ENST00000302763.7 ENST00000518910.1 |
CTNNA1 |
catenin (cadherin-associated protein), alpha 1, 102kDa |
| chr1_+_156084461 | 1.59 |
ENST00000347559.2 ENST00000361308.4 ENST00000368300.4 ENST00000368299.3 |
LMNA |
lamin A/C |
| chr5_+_140762268 | 1.58 |
ENST00000518325.1 |
PCDHGA7 |
protocadherin gamma subfamily A, 7 |
| chrX_+_47441712 | 1.58 |
ENST00000218388.4 ENST00000377018.2 ENST00000456754.2 ENST00000377017.1 ENST00000441738.1 |
TIMP1 |
TIMP metallopeptidase inhibitor 1 |
| chr18_+_56338618 | 1.58 |
ENST00000348428.3 |
MALT1 |
mucosa associated lymphoid tissue lymphoma translocation gene 1 |
| chr11_-_111784005 | 1.58 |
ENST00000527899.1 |
CRYAB |
crystallin, alpha B |
| chr5_+_150020214 | 1.58 |
ENST00000307662.4 |
SYNPO |
synaptopodin |
| chr20_+_4667094 | 1.55 |
ENST00000424424.1 ENST00000457586.1 |
PRNP |
prion protein |
| chrX_+_12993202 | 1.53 |
ENST00000451311.2 ENST00000380636.1 |
TMSB4X |
thymosin beta 4, X-linked |
| chr2_-_113542063 | 1.53 |
ENST00000263339.3 |
IL1A |
interleukin 1, alpha |
| chr1_+_156096336 | 1.52 |
ENST00000504687.1 ENST00000473598.2 |
LMNA |
lamin A/C |
| chr12_+_81110684 | 1.50 |
ENST00000228644.3 |
MYF5 |
myogenic factor 5 |
| chr8_-_145025044 | 1.50 |
ENST00000322810.4 |
PLEC |
plectin |
| chr2_-_166651191 | 1.50 |
ENST00000392701.3 |
GALNT3 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 3 (GalNAc-T3) |
| chr10_+_86088381 | 1.49 |
ENST00000224756.8 ENST00000372088.2 |
CCSER2 |
coiled-coil serine-rich protein 2 |
| chr1_+_201252580 | 1.49 |
ENST00000367324.3 ENST00000263946.3 |
PKP1 |
plakophilin 1 (ectodermal dysplasia/skin fragility syndrome) |
| chr15_+_41136216 | 1.49 |
ENST00000562057.1 ENST00000344051.4 |
SPINT1 |
serine peptidase inhibitor, Kunitz type 1 |
| chr2_-_113594279 | 1.45 |
ENST00000416750.1 ENST00000418817.1 ENST00000263341.2 |
IL1B |
interleukin 1, beta |
| chr2_+_201171064 | 1.44 |
ENST00000451764.2 |
SPATS2L |
spermatogenesis associated, serine-rich 2-like |
| chr17_-_1389419 | 1.44 |
ENST00000575158.1 |
MYO1C |
myosin IC |
| chr1_+_35220613 | 1.41 |
ENST00000338513.1 |
GJB5 |
gap junction protein, beta 5, 31.1kDa |
| chr21_-_27542972 | 1.41 |
ENST00000346798.3 ENST00000439274.2 ENST00000354192.3 ENST00000348990.5 ENST00000357903.3 ENST00000358918.3 ENST00000359726.3 |
APP |
amyloid beta (A4) precursor protein |
| chr11_-_64013663 | 1.40 |
ENST00000392210.2 |
PPP1R14B |
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr2_+_30369807 | 1.40 |
ENST00000379520.3 ENST00000379519.3 ENST00000261353.4 |
YPEL5 |
yippee-like 5 (Drosophila) |
| chr15_-_60690163 | 1.39 |
ENST00000558998.1 ENST00000560165.1 ENST00000557986.1 ENST00000559780.1 ENST00000559467.1 ENST00000559956.1 ENST00000332680.4 ENST00000396024.3 ENST00000421017.2 ENST00000560466.1 ENST00000558132.1 ENST00000559113.1 ENST00000557906.1 ENST00000558558.1 ENST00000560468.1 ENST00000559370.1 ENST00000558169.1 ENST00000559725.1 ENST00000558985.1 ENST00000451270.2 |
ANXA2 |
annexin A2 |
| chr17_-_7493390 | 1.38 |
ENST00000538513.2 ENST00000570788.1 ENST00000250055.2 |
SOX15 |
SRY (sex determining region Y)-box 15 |
| chr13_+_76334795 | 1.38 |
ENST00000526202.1 ENST00000465261.2 |
LMO7 |
LIM domain 7 |
| chr15_+_41245160 | 1.37 |
ENST00000444189.2 ENST00000446533.3 |
CHAC1 |
ChaC, cation transport regulator homolog 1 (E. coli) |
| chr1_+_27189631 | 1.37 |
ENST00000339276.4 |
SFN |
stratifin |
| chr2_+_172544182 | 1.36 |
ENST00000409197.1 ENST00000456808.1 ENST00000409317.1 ENST00000409773.1 ENST00000411953.1 ENST00000409453.1 |
DYNC1I2 |
dynein, cytoplasmic 1, intermediate chain 2 |
| chr17_-_1389228 | 1.35 |
ENST00000438665.2 |
MYO1C |
myosin IC |
| chr19_-_43702231 | 1.34 |
ENST00000597374.1 ENST00000599371.1 |
PSG4 |
pregnancy specific beta-1-glycoprotein 4 |
| chr14_+_58666824 | 1.34 |
ENST00000254286.4 |
ACTR10 |
actin-related protein 10 homolog (S. cerevisiae) |
| chr13_-_36429763 | 1.34 |
ENST00000379893.1 |
DCLK1 |
doublecortin-like kinase 1 |
| chr2_+_172543967 | 1.33 |
ENST00000534253.2 ENST00000263811.4 ENST00000397119.3 ENST00000410079.3 ENST00000438879.1 |
DYNC1I2 |
dynein, cytoplasmic 1, intermediate chain 2 |
| chr21_-_36421535 | 1.32 |
ENST00000416754.1 ENST00000437180.1 ENST00000455571.1 |
RUNX1 |
runt-related transcription factor 1 |
| chr2_+_30369859 | 1.30 |
ENST00000402003.3 |
YPEL5 |
yippee-like 5 (Drosophila) |
| chr5_+_53751445 | 1.30 |
ENST00000302005.1 |
HSPB3 |
heat shock 27kDa protein 3 |
| chr7_-_137028534 | 1.29 |
ENST00000348225.2 |
PTN |
pleiotrophin |
| chr3_+_11178779 | 1.28 |
ENST00000438284.2 |
HRH1 |
histamine receptor H1 |
| chr7_+_90338712 | 1.28 |
ENST00000265741.3 ENST00000406263.1 |
CDK14 |
cyclin-dependent kinase 14 |
| chr3_-_122512619 | 1.28 |
ENST00000383659.1 ENST00000306103.2 |
HSPBAP1 |
HSPB (heat shock 27kDa) associated protein 1 |
| chr13_+_76334567 | 1.27 |
ENST00000321797.8 |
LMO7 |
LIM domain 7 |
| chr12_-_56101647 | 1.27 |
ENST00000347027.6 ENST00000257879.6 ENST00000257880.7 ENST00000394230.2 ENST00000394229.2 |
ITGA7 |
integrin, alpha 7 |
| chr6_-_56819385 | 1.27 |
ENST00000370754.5 ENST00000449297.2 |
DST |
dystonin |
| chr5_-_176924562 | 1.24 |
ENST00000359895.2 ENST00000355572.2 ENST00000355841.2 ENST00000393551.1 ENST00000505074.1 ENST00000356618.4 ENST00000393546.4 |
PDLIM7 |
PDZ and LIM domain 7 (enigma) |
| chr12_+_53693812 | 1.24 |
ENST00000549488.1 |
C12orf10 |
chromosome 12 open reading frame 10 |
| chr12_-_56120865 | 1.22 |
ENST00000548898.1 ENST00000552067.1 |
CD63 |
CD63 molecule |
| chr6_-_56507586 | 1.22 |
ENST00000439203.1 ENST00000518935.1 ENST00000446842.2 ENST00000370765.6 ENST00000244364.6 |
DST |
dystonin |
| chr12_+_6644443 | 1.20 |
ENST00000396858.1 |
GAPDH |
glyceraldehyde-3-phosphate dehydrogenase |
| chr12_+_10365404 | 1.20 |
ENST00000266458.5 ENST00000421801.2 ENST00000544284.1 ENST00000545047.1 ENST00000543602.1 ENST00000545887.1 |
GABARAPL1 |
GABA(A) receptor-associated protein like 1 |
| chr21_-_36421626 | 1.20 |
ENST00000300305.3 |
RUNX1 |
runt-related transcription factor 1 |
| chr22_-_30642782 | 1.18 |
ENST00000249075.3 |
LIF |
leukemia inhibitory factor |
| chr1_-_154943002 | 1.17 |
ENST00000606391.1 |
SHC1 |
SHC (Src homology 2 domain containing) transforming protein 1 |
| chr2_+_172544294 | 1.17 |
ENST00000358002.6 ENST00000435234.1 ENST00000443458.1 ENST00000412370.1 |
DYNC1I2 |
dynein, cytoplasmic 1, intermediate chain 2 |
| chr21_-_27543425 | 1.17 |
ENST00000448388.2 |
APP |
amyloid beta (A4) precursor protein |
| chr2_+_172544011 | 1.17 |
ENST00000508530.1 |
DYNC1I2 |
dynein, cytoplasmic 1, intermediate chain 2 |
| chr11_-_8832182 | 1.17 |
ENST00000527510.1 ENST00000528527.1 ENST00000528523.1 ENST00000313726.6 |
ST5 |
suppression of tumorigenicity 5 |
| chr2_+_172543919 | 1.17 |
ENST00000452242.1 ENST00000340296.4 |
DYNC1I2 |
dynein, cytoplasmic 1, intermediate chain 2 |
| chr7_-_137028498 | 1.16 |
ENST00000393083.2 |
PTN |
pleiotrophin |
| chr1_-_153521597 | 1.14 |
ENST00000368712.1 |
S100A3 |
S100 calcium binding protein A3 |
| chr16_+_30386098 | 1.14 |
ENST00000322861.7 |
MYLPF |
myosin light chain, phosphorylatable, fast skeletal muscle |
| chrX_+_99899180 | 1.13 |
ENST00000373004.3 |
SRPX2 |
sushi-repeat containing protein, X-linked 2 |
| chr12_-_56122761 | 1.12 |
ENST00000552164.1 ENST00000420846.3 ENST00000257857.4 |
CD63 |
CD63 molecule |
| chr12_-_56120838 | 1.12 |
ENST00000548160.1 |
CD63 |
CD63 molecule |
| chr1_-_203155868 | 1.11 |
ENST00000255409.3 |
CHI3L1 |
chitinase 3-like 1 (cartilage glycoprotein-39) |
| chr3_+_100428188 | 1.10 |
ENST00000418917.2 ENST00000490574.1 |
TFG |
TRK-fused gene |
| chr16_-_69760409 | 1.10 |
ENST00000561500.1 ENST00000439109.2 ENST00000564043.1 ENST00000379046.2 ENST00000379047.3 |
NQO1 |
NAD(P)H dehydrogenase, quinone 1 |
| chr15_+_67458357 | 1.07 |
ENST00000537194.2 |
SMAD3 |
SMAD family member 3 |
| chr3_+_45067659 | 1.07 |
ENST00000296130.4 |
CLEC3B |
C-type lectin domain family 3, member B |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 7.1 | GO:0030377 | urokinase plasminogen activator receptor activity(GO:0030377) |
| 2.0 | 11.9 | GO:0005134 | interleukin-2 receptor binding(GO:0005134) |
| 1.7 | 6.8 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 1.4 | 8.2 | GO:1990254 | keratin filament binding(GO:1990254) |
| 1.1 | 4.4 | GO:0004850 | uridine phosphorylase activity(GO:0004850) |
| 1.1 | 5.4 | GO:0031962 | mineralocorticoid receptor binding(GO:0031962) |
| 0.9 | 6.5 | GO:0048408 | epidermal growth factor binding(GO:0048408) |
| 0.8 | 2.3 | GO:0004947 | bradykinin receptor activity(GO:0004947) |
| 0.7 | 2.2 | GO:0004917 | interleukin-7 receptor activity(GO:0004917) |
| 0.6 | 2.5 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.6 | 3.3 | GO:1903135 | cupric ion binding(GO:1903135) |
| 0.5 | 2.9 | GO:0008948 | malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) oxaloacetate decarboxylase activity(GO:0008948) |
| 0.5 | 1.4 | GO:0008426 | protein kinase C inhibitor activity(GO:0008426) |
| 0.4 | 5.7 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.4 | 4.3 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.4 | 14.3 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.4 | 3.6 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.3 | 6.2 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.3 | 7.0 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.3 | 1.2 | GO:0019828 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) aspartic-type endopeptidase inhibitor activity(GO:0019828) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 0.3 | 9.0 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.3 | 2.6 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.3 | 2.9 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.3 | 1.3 | GO:0070026 | nitric oxide binding(GO:0070026) |
| 0.3 | 0.8 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.2 | 2.9 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.2 | 1.4 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.2 | 0.7 | GO:0031811 | G-protein coupled nucleotide receptor binding(GO:0031811) P2Y1 nucleotide receptor binding(GO:0031812) |
| 0.2 | 0.9 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 0.2 | 18.7 | GO:0005518 | collagen binding(GO:0005518) |
| 0.2 | 1.0 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.2 | 1.9 | GO:0031994 | insulin-like growth factor I binding(GO:0031994) |
| 0.2 | 4.5 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.2 | 2.9 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.2 | 0.7 | GO:0004466 | long-chain-acyl-CoA dehydrogenase activity(GO:0004466) |
| 0.2 | 17.6 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.2 | 2.9 | GO:0017160 | Ral GTPase binding(GO:0017160) |
| 0.2 | 0.8 | GO:0070740 | tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.2 | 4.9 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.2 | 4.3 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 3.0 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.1 | 1.3 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.1 | 2.5 | GO:0031402 | sodium ion binding(GO:0031402) |
| 0.1 | 1.1 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.1 | 2.6 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 13.5 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 2.3 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.1 | 0.6 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.1 | 0.6 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.1 | 1.5 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.1 | 3.1 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 0.5 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.1 | 0.7 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.1 | 1.2 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.1 | 1.4 | GO:0016755 | transferase activity, transferring amino-acyl groups(GO:0016755) |
| 0.1 | 0.7 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.1 | 1.6 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.1 | 0.3 | GO:0031877 | somatostatin receptor binding(GO:0031877) |
| 0.1 | 1.5 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.1 | 1.2 | GO:0032407 | MutSalpha complex binding(GO:0032407) |
| 0.1 | 17.4 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.1 | 4.9 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 2.5 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 1.0 | GO:0004568 | chitinase activity(GO:0004568) chitin binding(GO:0008061) |
| 0.1 | 1.1 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.1 | 7.1 | GO:0004714 | transmembrane receptor protein tyrosine kinase activity(GO:0004714) |
| 0.1 | 0.9 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 1.2 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.1 | 0.6 | GO:0016167 | glial cell-derived neurotrophic factor receptor activity(GO:0016167) |
| 0.1 | 0.2 | GO:0019959 | interleukin-8 binding(GO:0019959) |
| 0.1 | 0.2 | GO:0008192 | RNA guanylyltransferase activity(GO:0008192) |
| 0.1 | 2.2 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 6.1 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.1 | 0.5 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.1 | 0.7 | GO:0000990 | transcription factor activity, core RNA polymerase binding(GO:0000990) |
| 0.1 | 0.2 | GO:0004914 | interleukin-5 receptor activity(GO:0004914) |
| 0.1 | 0.5 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.1 | 1.6 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.1 | 3.2 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 1.5 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.1 | 1.5 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 8.7 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.1 | 41.0 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.1 | 0.4 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.1 | 0.2 | GO:0034648 | histone demethylase activity (H3-dimethyl-K4 specific)(GO:0034648) |
| 0.1 | 2.9 | GO:0043621 | protein self-association(GO:0043621) |
| 0.1 | 5.2 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.1 | 1.9 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 0.6 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.3 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 2.0 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.0 | 7.2 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.9 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.0 | 0.1 | GO:0004574 | oligo-1,6-glucosidase activity(GO:0004574) |
| 0.0 | 1.1 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.0 | 0.2 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 0.0 | 0.7 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.5 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 0.2 | GO:0035368 | selenocysteine insertion sequence binding(GO:0035368) |
| 0.0 | 2.4 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.0 | 0.1 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 0.0 | 0.4 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.0 | 0.3 | GO:0003910 | DNA ligase (ATP) activity(GO:0003910) |
| 0.0 | 0.1 | GO:0070984 | SET domain binding(GO:0070984) |
| 0.0 | 0.2 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.0 | 0.1 | GO:0004878 | complement component C5a receptor activity(GO:0004878) |
| 0.0 | 0.1 | GO:0008184 | glycogen phosphorylase activity(GO:0008184) |
| 0.0 | 0.7 | GO:0031418 | L-ascorbic acid binding(GO:0031418) |
| 0.0 | 0.2 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 0.3 | GO:0038187 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.0 | 0.2 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.0 | 2.3 | GO:0019838 | growth factor binding(GO:0019838) |
| 0.0 | 2.6 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
| 0.0 | 0.5 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 0.4 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.0 | 0.7 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.0 | 0.3 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.0 | 0.1 | GO:0005057 | receptor signaling protein activity(GO:0005057) |
| 0.0 | 1.1 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.2 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.0 | 2.5 | GO:0003779 | actin binding(GO:0003779) |
| 0.0 | 0.1 | GO:0015333 | peptide:proton symporter activity(GO:0015333) proton-dependent peptide secondary active transmembrane transporter activity(GO:0022897) |
| 0.0 | 0.5 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.0 | 1.4 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 8.2 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 0.4 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 1.1 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 0.1 | GO:0004931 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.0 | 2.3 | GO:0001664 | G-protein coupled receptor binding(GO:0001664) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 4.4 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.5 | 5.0 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.4 | 10.9 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.3 | 39.5 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.3 | 4.5 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.3 | 8.0 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.3 | 6.2 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.2 | 7.2 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.2 | 6.1 | REACTOME SHC1 EVENTS IN EGFR SIGNALING | Genes involved in SHC1 events in EGFR signaling |
| 0.2 | 3.3 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.2 | 7.1 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
| 0.2 | 1.5 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.2 | 5.2 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.2 | 8.0 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.1 | 8.4 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.1 | 7.3 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 2.6 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.1 | 2.6 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.1 | 2.6 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.1 | 1.4 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.1 | 2.2 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.1 | 3.3 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.1 | 6.2 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.1 | 4.4 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 1.5 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.1 | 2.7 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.1 | 0.2 | REACTOME BINDING AND ENTRY OF HIV VIRION | Genes involved in Binding and entry of HIV virion |
| 0.1 | 0.8 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.1 | 0.6 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.1 | 4.8 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 1.7 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.1 | 2.2 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 1.8 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |
| 0.0 | 1.9 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 2.1 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 1.5 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 0.7 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.0 | 0.7 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.0 | 0.7 | REACTOME ADENYLATE CYCLASE INHIBITORY PATHWAY | Genes involved in Adenylate cyclase inhibitory pathway |
| 0.0 | 4.7 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 1.0 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 0.9 | REACTOME SIGNALING BY FGFR1 MUTANTS | Genes involved in Signaling by FGFR1 mutants |
| 0.0 | 0.7 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.0 | 2.9 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 1.1 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.9 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 3.4 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 0.7 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 1.2 | REACTOME MUSCLE CONTRACTION | Genes involved in Muscle contraction |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.4 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 2.3 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 0.5 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 0.2 | REACTOME PREFOLDIN MEDIATED TRANSFER OF SUBSTRATE TO CCT TRIC | Genes involved in Prefoldin mediated transfer of substrate to CCT/TriC |
| 0.0 | 0.5 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.2 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.2 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 39.5 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.6 | 34.9 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.3 | 6.8 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.2 | 13.1 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.2 | 15.3 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.2 | 5.4 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.2 | 9.0 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.2 | 3.7 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.2 | 8.8 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 8.0 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.1 | 0.1 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.1 | 6.0 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.1 | 3.2 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 1.6 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.1 | 8.5 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 6.2 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 3.9 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 3.1 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 13.1 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 2.2 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.1 | 0.7 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.1 | 2.2 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.1 | 14.0 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.2 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.0 | 2.8 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 2.0 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 2.3 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 1.5 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 1.1 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 1.6 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.6 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.2 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.0 | 8.9 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 1.9 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 3.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.7 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.7 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.0 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 0.5 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 1.4 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 1.1 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.3 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.0 | 0.1 | PID TRAIL PATHWAY | TRAIL signaling pathway |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.1 | 33.0 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 1.5 | 4.6 | GO:0005607 | laminin-2 complex(GO:0005607) |
| 1.5 | 4.4 | GO:0097489 | multivesicular body, internal vesicle lumen(GO:0097489) |
| 1.0 | 11.5 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 1.0 | 5.2 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) |
| 0.9 | 2.6 | GO:0033565 | ESCRT-0 complex(GO:0033565) |
| 0.8 | 6.8 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.7 | 5.2 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 0.7 | 2.2 | GO:0070435 | Shc-EGFR complex(GO:0070435) |
| 0.7 | 13.0 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.7 | 2.8 | GO:0045160 | myosin I complex(GO:0045160) |
| 0.7 | 3.4 | GO:0032449 | CBM complex(GO:0032449) |
| 0.7 | 2.0 | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex(GO:0000308) |
| 0.7 | 5.4 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.6 | 2.5 | GO:0031673 | H zone(GO:0031673) |
| 0.6 | 5.5 | GO:0005638 | lamin filament(GO:0005638) |
| 0.6 | 7.8 | GO:0098647 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.5 | 5.0 | GO:0097413 | Lewy body(GO:0097413) |
| 0.5 | 4.2 | GO:0097487 | multivesicular body, internal vesicle(GO:0097487) |
| 0.4 | 1.3 | GO:0097057 | TRAF2-GSTP1 complex(GO:0097057) |
| 0.4 | 1.2 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.4 | 3.6 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.4 | 7.0 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.4 | 6.2 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.3 | 2.9 | GO:0036454 | insulin-like growth factor binding protein complex(GO:0016942) growth factor complex(GO:0036454) |
| 0.3 | 2.6 | GO:1990761 | growth cone lamellipodium(GO:1990761) growth cone filopodium(GO:1990812) |
| 0.3 | 1.4 | GO:0044352 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.3 | 4.2 | GO:0030478 | actin cap(GO:0030478) |
| 0.3 | 3.3 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.2 | 2.2 | GO:0098644 | complex of collagen trimers(GO:0098644) |
| 0.2 | 3.4 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.2 | 6.1 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.2 | 1.2 | GO:0097452 | GAIT complex(GO:0097452) |
| 0.2 | 1.6 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.2 | 2.9 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.2 | 0.6 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.1 | 7.0 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 16.1 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 0.6 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 0.1 | 2.7 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.1 | 4.0 | GO:0005865 | striated muscle thin filament(GO:0005865) |
| 0.1 | 0.3 | GO:1990462 | omegasome(GO:1990462) |
| 0.1 | 1.1 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.1 | 3.0 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 2.2 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.1 | 2.9 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.1 | 9.2 | GO:0005604 | basement membrane(GO:0005604) |
| 0.1 | 1.3 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 2.3 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 1.4 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 2.8 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 0.7 | GO:0098839 | postsynaptic density membrane(GO:0098839) |
| 0.1 | 1.5 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.1 | 0.1 | GO:0002139 | stereocilia coupling link(GO:0002139) |
| 0.1 | 1.0 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.1 | 1.6 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.1 | 15.0 | GO:0019897 | extrinsic component of plasma membrane(GO:0019897) |
| 0.1 | 6.5 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.1 | 1.8 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.1 | 0.9 | GO:0097418 | neurofibrillary tangle(GO:0097418) |
| 0.1 | 4.4 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 1.9 | GO:0032420 | stereocilium(GO:0032420) |
| 0.1 | 2.4 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.1 | 0.9 | GO:0031094 | platelet dense tubular network(GO:0031094) |
| 0.1 | 1.0 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 0.1 | GO:0036020 | endolysosome membrane(GO:0036020) |
| 0.1 | 1.2 | GO:0008305 | integrin complex(GO:0008305) |
| 0.0 | 25.3 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 4.1 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 1.4 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 0.3 | GO:0005827 | polar microtubule(GO:0005827) |
| 0.0 | 0.3 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 1.2 | GO:0032590 | dendrite membrane(GO:0032590) |
| 0.0 | 1.8 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 10.0 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 0.2 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 2.5 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.0 | 0.2 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.0 | 0.5 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 0.7 | GO:0031105 | septin complex(GO:0031105) |
| 0.0 | 3.0 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 0.5 | GO:0010369 | chromocenter(GO:0010369) |
| 0.0 | 0.6 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 0.3 | GO:0044754 | autolysosome(GO:0044754) |
| 0.0 | 0.8 | GO:0070161 | anchoring junction(GO:0070161) |
| 0.0 | 0.3 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.0 | 0.9 | GO:0043025 | neuronal cell body(GO:0043025) |
| 0.0 | 2.1 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.6 | GO:0030315 | T-tubule(GO:0030315) |
| 0.0 | 1.4 | GO:0014069 | postsynaptic density(GO:0014069) postsynaptic specialization(GO:0099572) |
| 0.0 | 1.5 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.6 | GO:0005720 | nuclear heterochromatin(GO:0005720) |
| 0.0 | 0.7 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.7 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 0.8 | GO:0097014 | axoneme(GO:0005930) ciliary plasm(GO:0097014) |
| 0.0 | 0.6 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 0.7 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.3 | GO:0035861 | site of double-strand break(GO:0035861) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 6.8 | GO:0002514 | B cell tolerance induction(GO:0002514) regulation of B cell tolerance induction(GO:0002661) positive regulation of B cell tolerance induction(GO:0002663) |
| 2.2 | 8.6 | GO:0090362 | positive regulation of platelet-derived growth factor production(GO:0090362) |
| 1.8 | 7.1 | GO:0038195 | urokinase plasminogen activator signaling pathway(GO:0038195) |
| 1.5 | 4.4 | GO:0043006 | activation of phospholipase A2 activity by calcium-mediated signaling(GO:0043006) |
| 1.4 | 41.3 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 1.4 | 6.9 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) |
| 1.4 | 1.4 | GO:0010481 | epidermal cell division(GO:0010481) regulation of epidermal cell division(GO:0010482) |
| 1.2 | 3.6 | GO:1905205 | positive regulation of connective tissue replacement(GO:1905205) |
| 1.2 | 3.5 | GO:2000097 | regulation of smooth muscle cell-matrix adhesion(GO:2000097) |
| 1.1 | 6.9 | GO:0007296 | vitellogenesis(GO:0007296) |
| 1.1 | 3.4 | GO:0001923 | B-1 B cell differentiation(GO:0001923) |
| 1.1 | 4.5 | GO:0097325 | melanocyte proliferation(GO:0097325) |
| 1.1 | 4.4 | GO:0006218 | uridine catabolic process(GO:0006218) uridine metabolic process(GO:0046108) |
| 1.1 | 3.3 | GO:1902938 | regulation of intracellular calcium activated chloride channel activity(GO:1902938) |
| 1.1 | 5.5 | GO:0035105 | sterol regulatory element binding protein import into nucleus(GO:0035105) |
| 1.1 | 5.4 | GO:0061767 | negative regulation of lung blood pressure(GO:0061767) |
| 1.1 | 4.3 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.9 | 17.7 | GO:0001765 | membrane raft assembly(GO:0001765) |
| 0.9 | 4.3 | GO:0009609 | response to symbiont(GO:0009608) response to symbiotic bacterium(GO:0009609) |
| 0.8 | 2.5 | GO:1904395 | retinal rod cell differentiation(GO:0060221) positive regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904395) negative regulation of neuromuscular junction development(GO:1904397) |
| 0.7 | 1.4 | GO:0060355 | positive regulation of cell adhesion molecule production(GO:0060355) |
| 0.7 | 5.7 | GO:0002036 | regulation of L-glutamate transport(GO:0002036) |
| 0.7 | 2.7 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 0.7 | 5.2 | GO:1900623 | regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 0.6 | 7.8 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.6 | 2.6 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.6 | 3.2 | GO:0045906 | negative regulation of vasoconstriction(GO:0045906) |
| 0.6 | 5.7 | GO:1902459 | positive regulation of stem cell population maintenance(GO:1902459) |
| 0.6 | 3.7 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.6 | 7.7 | GO:0061436 | establishment of skin barrier(GO:0061436) |
| 0.6 | 1.7 | GO:1903028 | positive regulation of opsonization(GO:1903028) |
| 0.6 | 6.2 | GO:2000582 | positive regulation of microtubule motor activity(GO:2000576) regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.6 | 2.2 | GO:1903288 | protein localization to plasma membrane raft(GO:0044860) protein transport into plasma membrane raft(GO:0044861) positive regulation of potassium ion import(GO:1903288) |
| 0.5 | 1.6 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.5 | 2.6 | GO:0071874 | cellular response to norepinephrine stimulus(GO:0071874) |
| 0.5 | 9.9 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.5 | 2.9 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.5 | 2.8 | GO:1900748 | positive regulation of vascular endothelial growth factor signaling pathway(GO:1900748) |
| 0.5 | 1.4 | GO:0070318 | myoblast development(GO:0048627) positive regulation of G0 to G1 transition(GO:0070318) |
| 0.5 | 1.8 | GO:1903976 | negative regulation of glial cell migration(GO:1903976) |
| 0.4 | 10.7 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.4 | 1.3 | GO:0033591 | response to L-ascorbic acid(GO:0033591) |
| 0.4 | 4.2 | GO:1900746 | regulation of vascular endothelial growth factor signaling pathway(GO:1900746) |
| 0.4 | 2.5 | GO:0002248 | connective tissue replacement involved in inflammatory response wound healing(GO:0002248) |
| 0.4 | 7.3 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.4 | 1.2 | GO:0035606 | peptidyl-cysteine S-trans-nitrosylation(GO:0035606) |
| 0.4 | 0.8 | GO:0044111 | development involved in symbiotic interaction(GO:0044111) |
| 0.4 | 4.3 | GO:0060020 | Bergmann glial cell differentiation(GO:0060020) |
| 0.4 | 1.5 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.4 | 1.1 | GO:0060437 | positive regulation of metanephric mesenchymal cell migration by platelet-derived growth factor receptor-beta signaling pathway(GO:0035793) lung growth(GO:0060437) metanephric glomerulus morphogenesis(GO:0072275) metanephric glomerulus vasculature morphogenesis(GO:0072276) metanephric glomerular capillary formation(GO:0072277) regulation of metanephric mesenchymal cell migration by platelet-derived growth factor receptor-beta signaling pathway(GO:1900238) positive regulation of metanephric mesenchymal cell migration(GO:2000591) |
| 0.3 | 6.2 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.3 | 4.4 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.3 | 7.5 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.3 | 11.1 | GO:0032461 | positive regulation of protein oligomerization(GO:0032461) |
| 0.3 | 3.3 | GO:0043589 | skin morphogenesis(GO:0043589) |
| 0.3 | 2.2 | GO:0035897 | proteolysis in other organism(GO:0035897) |
| 0.3 | 1.5 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 0.3 | 2.6 | GO:0099563 | modification of synaptic structure(GO:0099563) |
| 0.3 | 0.9 | GO:2000661 | positive regulation of interleukin-1-mediated signaling pathway(GO:2000661) |
| 0.3 | 2.0 | GO:0001554 | luteolysis(GO:0001554) |
| 0.3 | 2.9 | GO:0060712 | spongiotrophoblast layer development(GO:0060712) |
| 0.3 | 0.8 | GO:0003099 | regulation of systemic arterial blood pressure by carotid sinus baroreceptor feedback(GO:0001978) baroreceptor response to increased systemic arterial blood pressure(GO:0001983) positive regulation of the force of heart contraction by chemical signal(GO:0003099) |
| 0.2 | 3.4 | GO:0090136 | epithelial cell-cell adhesion(GO:0090136) |
| 0.2 | 2.2 | GO:2000400 | positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 0.2 | 1.9 | GO:0031642 | negative regulation of myelination(GO:0031642) |
| 0.2 | 1.5 | GO:2000483 | negative regulation of interleukin-8 secretion(GO:2000483) |
| 0.2 | 0.9 | GO:0070884 | calcineurin-NFAT signaling cascade(GO:0033173) inositol phosphate-mediated signaling(GO:0048016) regulation of calcineurin-NFAT signaling cascade(GO:0070884) |
| 0.2 | 0.9 | GO:0061289 | cell-cell signaling involved in kidney development(GO:0060995) Wnt signaling pathway involved in kidney development(GO:0061289) canonical Wnt signaling pathway involved in metanephric kidney development(GO:0061290) cell-cell signaling involved in metanephros development(GO:0072204) |
| 0.2 | 0.4 | GO:0002572 | pro-T cell differentiation(GO:0002572) |
| 0.2 | 1.0 | GO:0006543 | glutamine catabolic process(GO:0006543) |
| 0.2 | 0.6 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.2 | 1.6 | GO:1901164 | negative regulation of trophoblast cell migration(GO:1901164) |
| 0.2 | 5.3 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.2 | 0.6 | GO:0002384 | hepatic immune response(GO:0002384) |
| 0.2 | 3.1 | GO:0060670 | branching involved in labyrinthine layer morphogenesis(GO:0060670) |
| 0.2 | 3.3 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.2 | 1.7 | GO:0008090 | retrograde axonal transport(GO:0008090) |
| 0.2 | 2.8 | GO:0097186 | amelogenesis(GO:0097186) |
| 0.2 | 0.7 | GO:0072674 | multinuclear osteoclast differentiation(GO:0072674) osteoclast fusion(GO:0072675) |
| 0.2 | 1.1 | GO:0010756 | positive regulation of plasminogen activation(GO:0010756) |
| 0.2 | 5.5 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.2 | 2.6 | GO:1903543 | positive regulation of exosomal secretion(GO:1903543) |
| 0.2 | 1.4 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.2 | 2.9 | GO:0035641 | locomotory exploration behavior(GO:0035641) |
| 0.2 | 3.6 | GO:0036150 | phosphatidylserine acyl-chain remodeling(GO:0036150) |
| 0.2 | 0.6 | GO:0006620 | posttranslational protein targeting to membrane(GO:0006620) |
| 0.1 | 2.2 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 0.8 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.1 | 1.5 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.1 | 1.3 | GO:0071847 | TNFSF11-mediated signaling pathway(GO:0071847) |
| 0.1 | 0.8 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.1 | 5.9 | GO:0003254 | regulation of membrane depolarization(GO:0003254) |
| 0.1 | 1.1 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.1 | 0.7 | GO:1902902 | negative regulation of autophagosome assembly(GO:1902902) |
| 0.1 | 0.6 | GO:0010807 | regulation of synaptic vesicle priming(GO:0010807) |
| 0.1 | 0.5 | GO:0021523 | somatic motor neuron differentiation(GO:0021523) protein localization to ciliary transition zone(GO:1904491) |
| 0.1 | 1.6 | GO:0022038 | corpus callosum development(GO:0022038) |
| 0.1 | 0.5 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 | 2.9 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.1 | 0.7 | GO:0045646 | regulation of erythrocyte differentiation(GO:0045646) |
| 0.1 | 2.1 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.1 | 1.7 | GO:2000291 | regulation of myoblast proliferation(GO:2000291) |
| 0.1 | 0.5 | GO:0097283 | keratinocyte apoptotic process(GO:0097283) regulation of keratinocyte apoptotic process(GO:1902172) |
| 0.1 | 1.6 | GO:0034497 | protein localization to pre-autophagosomal structure(GO:0034497) |
| 0.1 | 2.5 | GO:0097201 | negative regulation of transcription from RNA polymerase II promoter in response to stress(GO:0097201) |
| 0.1 | 1.4 | GO:0051823 | regulation of synapse structural plasticity(GO:0051823) |
| 0.1 | 1.0 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.1 | 0.3 | GO:0050894 | determination of affect(GO:0050894) |
| 0.1 | 0.4 | GO:0030822 | positive regulation of cyclic nucleotide catabolic process(GO:0030807) positive regulation of cAMP catabolic process(GO:0030822) positive regulation of purine nucleotide catabolic process(GO:0033123) |
| 0.1 | 0.5 | GO:0060356 | leucine import(GO:0060356) |
| 0.1 | 0.4 | GO:0045670 | regulation of osteoclast differentiation(GO:0045670) |
| 0.1 | 0.9 | GO:2001020 | regulation of response to DNA damage stimulus(GO:2001020) |
| 0.1 | 0.7 | GO:2000546 | positive regulation of cell chemotaxis to fibroblast growth factor(GO:1904849) positive regulation of endothelial cell chemotaxis to fibroblast growth factor(GO:2000546) |
| 0.1 | 0.7 | GO:0046322 | negative regulation of fatty acid oxidation(GO:0046322) |
| 0.1 | 1.3 | GO:0048245 | eosinophil chemotaxis(GO:0048245) |
| 0.1 | 1.0 | GO:0006030 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.1 | 0.7 | GO:0001973 | adenosine receptor signaling pathway(GO:0001973) |
| 0.1 | 0.2 | GO:1903070 | negative regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903070) |
| 0.1 | 4.9 | GO:0030834 | regulation of actin filament depolymerization(GO:0030834) |
| 0.1 | 0.2 | GO:0018874 | benzoate metabolic process(GO:0018874) butyrate metabolic process(GO:0019605) |
| 0.1 | 0.2 | GO:1903238 | positive regulation of leukocyte tethering or rolling(GO:1903238) |
| 0.1 | 0.2 | GO:2000861 | estrogen secretion(GO:0035937) estradiol secretion(GO:0035938) regulation of estrogen secretion(GO:2000861) regulation of estradiol secretion(GO:2000864) |
| 0.1 | 3.6 | GO:0035987 | endodermal cell differentiation(GO:0035987) |
| 0.1 | 2.2 | GO:0007176 | regulation of epidermal growth factor-activated receptor activity(GO:0007176) |
| 0.1 | 0.3 | GO:0002331 | pre-B cell differentiation(GO:0002329) pre-B cell allelic exclusion(GO:0002331) |
| 0.1 | 0.2 | GO:0038043 | interleukin-5-mediated signaling pathway(GO:0038043) |
| 0.1 | 1.2 | GO:0043562 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.1 | 0.3 | GO:0030239 | myofibril assembly(GO:0030239) |
| 0.1 | 0.3 | GO:0042246 | tissue regeneration(GO:0042246) |
| 0.1 | 3.6 | GO:0042058 | regulation of epidermal growth factor receptor signaling pathway(GO:0042058) |
| 0.1 | 0.1 | GO:0052047 | interaction with other organism via secreted substance involved in symbiotic interaction(GO:0052047) |
| 0.1 | 2.1 | GO:0008038 | neuron recognition(GO:0008038) |
| 0.1 | 0.6 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.1 | 1.0 | GO:0006751 | glutathione catabolic process(GO:0006751) |
| 0.1 | 1.7 | GO:0007398 | ectoderm development(GO:0007398) |
| 0.1 | 2.1 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.1 | 0.6 | GO:0035860 | glial cell-derived neurotrophic factor receptor signaling pathway(GO:0035860) |
| 0.1 | 2.3 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.1 | 5.6 | GO:0022617 | extracellular matrix disassembly(GO:0022617) |
| 0.1 | 0.1 | GO:0003011 | diaphragm contraction(GO:0002086) involuntary skeletal muscle contraction(GO:0003011) |
| 0.1 | 0.3 | GO:0050428 | purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.1 | 4.5 | GO:0009250 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.1 | 0.6 | GO:0072257 | metanephric nephron tubule epithelial cell differentiation(GO:0072257) regulation of metanephric nephron tubule epithelial cell differentiation(GO:0072307) |
| 0.1 | 0.2 | GO:0060179 | male mating behavior(GO:0060179) |
| 0.1 | 4.4 | GO:0030049 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.1 | 0.2 | GO:0098759 | response to interleukin-8(GO:0098758) cellular response to interleukin-8(GO:0098759) |
| 0.1 | 0.5 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.1 | 3.6 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.1 | 0.6 | GO:0006657 | CDP-choline pathway(GO:0006657) |
| 0.0 | 2.2 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.2 | GO:0032690 | negative regulation of interleukin-1 alpha production(GO:0032690) negative regulation of interleukin-1 alpha secretion(GO:0050712) |
| 0.0 | 0.3 | GO:0044130 | negative regulation of growth of symbiont in host(GO:0044130) |
| 0.0 | 0.6 | GO:1900745 | positive regulation of p38MAPK cascade(GO:1900745) |
| 0.0 | 8.6 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.0 | 0.1 | GO:2001206 | positive regulation of bone development(GO:1903012) positive regulation of osteoclast development(GO:2001206) |
| 0.0 | 0.2 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.0 | 0.1 | GO:1902623 | negative regulation of granulocyte chemotaxis(GO:0071623) negative regulation of neutrophil chemotaxis(GO:0090024) negative regulation of neutrophil migration(GO:1902623) |
| 0.0 | 0.3 | GO:0051106 | positive regulation of DNA ligation(GO:0051106) |
| 0.0 | 1.1 | GO:0006521 | regulation of cellular amino acid metabolic process(GO:0006521) |
| 0.0 | 0.3 | GO:0009438 | methylglyoxal metabolic process(GO:0009438) |
| 0.0 | 0.7 | GO:1902857 | positive regulation of nonmotile primary cilium assembly(GO:1902857) |
| 0.0 | 2.2 | GO:0045669 | positive regulation of osteoblast differentiation(GO:0045669) |
| 0.0 | 3.8 | GO:0002062 | chondrocyte differentiation(GO:0002062) |
| 0.0 | 0.2 | GO:0019064 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.0 | 0.2 | GO:0051402 | neuron apoptotic process(GO:0051402) |
| 0.0 | 2.3 | GO:0032755 | positive regulation of interleukin-6 production(GO:0032755) |
| 0.0 | 1.9 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.0 | 1.2 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 2.9 | GO:0048207 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.0 | 0.2 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.0 | 1.3 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.0 | 0.7 | GO:0097503 | sialylation(GO:0097503) |
| 0.0 | 1.3 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.0 | 0.1 | GO:0042938 | dipeptide transport(GO:0042938) |
| 0.0 | 0.2 | GO:1903147 | negative regulation of macromitophagy(GO:1901525) negative regulation of mitophagy(GO:1903147) |
| 0.0 | 0.1 | GO:0010286 | heat acclimation(GO:0010286) cellular heat acclimation(GO:0070370) |
| 0.0 | 0.1 | GO:0051414 | response to cortisol(GO:0051414) |
| 0.0 | 1.2 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.0 | 2.8 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.7 | GO:0042475 | odontogenesis of dentin-containing tooth(GO:0042475) |
| 0.0 | 0.1 | GO:0050910 | detection of mechanical stimulus involved in sensory perception of sound(GO:0050910) |
| 0.0 | 0.1 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.0 | 0.1 | GO:0023021 | termination of signal transduction(GO:0023021) |
| 0.0 | 0.2 | GO:0021756 | striatum development(GO:0021756) regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.0 | 0.2 | GO:1904778 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.0 | 0.3 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.0 | 0.4 | GO:0048240 | sperm capacitation(GO:0048240) |
| 0.0 | 0.1 | GO:0052565 | response to defense-related nitric oxide production by other organism involved in symbiotic interaction(GO:0052551) response to defense-related host nitric oxide production(GO:0052565) |
| 0.0 | 0.1 | GO:1904378 | maintenance of unfolded protein(GO:0036506) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) |
| 0.0 | 0.2 | GO:0035116 | embryonic hindlimb morphogenesis(GO:0035116) |
| 0.0 | 0.1 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) |
| 0.0 | 0.8 | GO:0001578 | microtubule bundle formation(GO:0001578) |
| 0.0 | 0.0 | GO:0010642 | negative regulation of platelet-derived growth factor receptor signaling pathway(GO:0010642) |
| 0.0 | 0.1 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.0 | 0.1 | GO:0035587 | purinergic receptor signaling pathway(GO:0035587) |
| 0.0 | 0.7 | GO:1902017 | regulation of cilium assembly(GO:1902017) |
| 0.0 | 0.1 | GO:0072719 | cellular response to cisplatin(GO:0072719) |
| 0.0 | 1.3 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 0.4 | GO:0010107 | potassium ion import(GO:0010107) |