Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
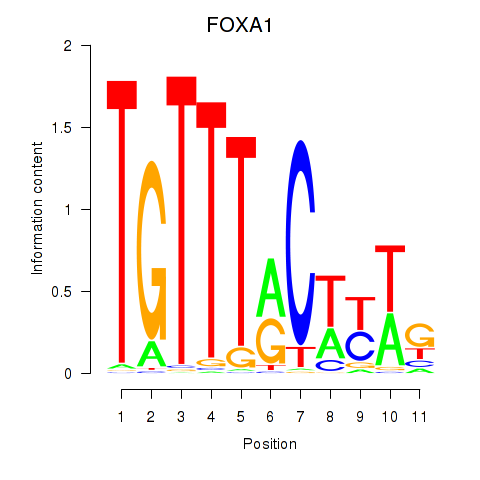
Results for FOXA1
Z-value: 1.10
Transcription factors associated with FOXA1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
FOXA1
|
ENSG00000129514.4 | FOXA1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| FOXA1 | hg19_v2_chr14_-_38064198_38064239 | 0.91 | 1.2e-06 | Click! |
Activity profile of FOXA1 motif
Sorted Z-values of FOXA1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of FOXA1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_26695013 | 6.62 |
ENST00000555059.2 |
CTB-96E2.2 |
Homeobox protein SEBOX |
| chr17_-_26694979 | 6.53 |
ENST00000438614.1 |
VTN |
vitronectin |
| chr2_-_21266935 | 4.85 |
ENST00000233242.1 |
APOB |
apolipoprotein B |
| chr3_+_186330712 | 4.59 |
ENST00000411641.2 ENST00000273784.5 |
AHSG |
alpha-2-HS-glycoprotein |
| chr17_+_79953310 | 4.28 |
ENST00000582355.2 |
ASPSCR1 |
alveolar soft part sarcoma chromosome region, candidate 1 |
| chr14_-_94789663 | 3.99 |
ENST00000557225.1 ENST00000341584.3 |
SERPINA6 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 6 |
| chr3_-_148939835 | 3.94 |
ENST00000264613.6 |
CP |
ceruloplasmin (ferroxidase) |
| chr22_+_21128167 | 3.87 |
ENST00000215727.5 |
SERPIND1 |
serpin peptidase inhibitor, clade D (heparin cofactor), member 1 |
| chr4_-_70080449 | 2.87 |
ENST00000446444.1 |
UGT2B11 |
UDP glucuronosyltransferase 2 family, polypeptide B11 |
| chr4_-_155533787 | 2.74 |
ENST00000407946.1 ENST00000405164.1 ENST00000336098.3 ENST00000393846.2 ENST00000404648.3 ENST00000443553.1 |
FGG |
fibrinogen gamma chain |
| chr17_-_39684550 | 2.20 |
ENST00000455635.1 ENST00000361566.3 |
KRT19 |
keratin 19 |
| chr6_-_41715128 | 2.12 |
ENST00000356667.4 ENST00000373025.3 ENST00000425343.2 |
PGC |
progastricsin (pepsinogen C) |
| chr17_+_72426891 | 1.91 |
ENST00000392627.1 |
GPRC5C |
G protein-coupled receptor, family C, group 5, member C |
| chr8_+_126442563 | 1.88 |
ENST00000311922.3 |
TRIB1 |
tribbles pseudokinase 1 |
| chr1_-_27240455 | 1.84 |
ENST00000254227.3 |
NR0B2 |
nuclear receptor subfamily 0, group B, member 2 |
| chr1_+_227127981 | 1.82 |
ENST00000366778.1 ENST00000366777.3 ENST00000458507.2 |
ADCK3 |
aarF domain containing kinase 3 |
| chr19_+_18496957 | 1.78 |
ENST00000252809.3 |
GDF15 |
growth differentiation factor 15 |
| chrX_-_20236970 | 1.58 |
ENST00000379548.4 |
RPS6KA3 |
ribosomal protein S6 kinase, 90kDa, polypeptide 3 |
| chr14_-_38064198 | 1.57 |
ENST00000250448.2 |
FOXA1 |
forkhead box A1 |
| chr17_+_72427477 | 1.51 |
ENST00000342648.5 ENST00000481232.1 |
GPRC5C |
G protein-coupled receptor, family C, group 5, member C |
| chr14_-_21493884 | 1.27 |
ENST00000556974.1 ENST00000554419.1 ENST00000298687.5 ENST00000397858.1 ENST00000360463.3 ENST00000350792.3 ENST00000397847.2 |
NDRG2 |
NDRG family member 2 |
| chr17_+_74372662 | 1.27 |
ENST00000591651.1 ENST00000545180.1 |
SPHK1 |
sphingosine kinase 1 |
| chr12_-_48398104 | 1.26 |
ENST00000337299.6 ENST00000380518.3 |
COL2A1 |
collagen, type II, alpha 1 |
| chr20_-_45984401 | 1.21 |
ENST00000311275.7 |
ZMYND8 |
zinc finger, MYND-type containing 8 |
| chr18_+_60382672 | 1.17 |
ENST00000400316.4 ENST00000262719.5 |
PHLPP1 |
PH domain and leucine rich repeat protein phosphatase 1 |
| chr1_+_214161854 | 1.16 |
ENST00000435016.1 |
PROX1 |
prospero homeobox 1 |
| chr3_+_142315225 | 1.14 |
ENST00000457734.2 ENST00000483373.1 ENST00000475296.1 ENST00000495744.1 ENST00000476044.1 ENST00000461644.1 |
PLS1 |
plastin 1 |
| chr3_-_129407535 | 1.13 |
ENST00000432054.2 |
TMCC1 |
transmembrane and coiled-coil domain family 1 |
| chr16_-_87970122 | 1.08 |
ENST00000309893.2 |
CA5A |
carbonic anhydrase VA, mitochondrial |
| chr13_-_46716969 | 1.07 |
ENST00000435666.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chr12_-_71551868 | 1.06 |
ENST00000247829.3 |
TSPAN8 |
tetraspanin 8 |
| chr2_+_58655461 | 1.03 |
ENST00000429095.1 ENST00000429664.1 ENST00000452840.1 |
AC007092.1 |
long intergenic non-protein coding RNA 1122 |
| chr12_-_53343602 | 1.00 |
ENST00000546897.1 ENST00000552551.1 |
KRT8 |
keratin 8 |
| chr1_-_57431679 | 0.90 |
ENST00000371237.4 ENST00000535057.1 ENST00000543257.1 |
C8B |
complement component 8, beta polypeptide |
| chr6_+_10528560 | 0.88 |
ENST00000379597.3 |
GCNT2 |
glucosaminyl (N-acetyl) transferase 2, I-branching enzyme (I blood group) |
| chr7_+_134832808 | 0.85 |
ENST00000275767.3 |
TMEM140 |
transmembrane protein 140 |
| chrX_+_9431324 | 0.85 |
ENST00000407597.2 ENST00000424279.1 ENST00000536365.1 ENST00000441088.1 ENST00000380961.1 ENST00000415293.1 |
TBL1X |
transducin (beta)-like 1X-linked |
| chr20_+_306177 | 0.84 |
ENST00000544632.1 |
SOX12 |
SRY (sex determining region Y)-box 12 |
| chr12_+_121416489 | 0.83 |
ENST00000541395.1 ENST00000544413.1 |
HNF1A |
HNF1 homeobox A |
| chr21_-_43735446 | 0.80 |
ENST00000398431.2 |
TFF3 |
trefoil factor 3 (intestinal) |
| chr12_-_71551652 | 0.73 |
ENST00000546561.1 |
TSPAN8 |
tetraspanin 8 |
| chr7_+_73242490 | 0.71 |
ENST00000431918.1 |
CLDN4 |
claudin 4 |
| chr5_+_156712372 | 0.70 |
ENST00000541131.1 |
CYFIP2 |
cytoplasmic FMR1 interacting protein 2 |
| chr10_-_73848086 | 0.69 |
ENST00000536168.1 |
SPOCK2 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 2 |
| chr14_-_21493649 | 0.68 |
ENST00000553442.1 ENST00000555869.1 ENST00000556457.1 ENST00000397844.2 ENST00000554415.1 |
NDRG2 |
NDRG family member 2 |
| chr6_+_7107999 | 0.65 |
ENST00000491191.1 ENST00000379938.2 ENST00000471433.1 |
RREB1 |
ras responsive element binding protein 1 |
| chr11_+_120973375 | 0.61 |
ENST00000264037.2 |
TECTA |
tectorin alpha |
| chr20_+_306221 | 0.60 |
ENST00000342665.2 |
SOX12 |
SRY (sex determining region Y)-box 12 |
| chr2_-_172017393 | 0.59 |
ENST00000442919.2 |
TLK1 |
tousled-like kinase 1 |
| chr1_+_174933899 | 0.57 |
ENST00000367688.3 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr2_-_172290482 | 0.55 |
ENST00000442541.1 ENST00000392599.2 ENST00000375258.4 |
METTL8 |
methyltransferase like 8 |
| chr17_-_19651668 | 0.55 |
ENST00000494157.2 ENST00000225740.6 |
ALDH3A1 |
aldehyde dehydrogenase 3 family, member A1 |
| chr17_+_4613776 | 0.54 |
ENST00000269260.2 |
ARRB2 |
arrestin, beta 2 |
| chr14_-_64971288 | 0.54 |
ENST00000394715.1 |
ZBTB25 |
zinc finger and BTB domain containing 25 |
| chr6_+_135502501 | 0.51 |
ENST00000527615.1 ENST00000420123.2 ENST00000525369.1 ENST00000528774.1 ENST00000534121.1 ENST00000534044.1 ENST00000533624.1 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr2_-_172017343 | 0.51 |
ENST00000431350.2 ENST00000360843.3 |
TLK1 |
tousled-like kinase 1 |
| chr2_-_165424973 | 0.49 |
ENST00000543549.1 |
GRB14 |
growth factor receptor-bound protein 14 |
| chr5_-_115872142 | 0.48 |
ENST00000510263.1 |
SEMA6A |
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6A |
| chr4_+_187187098 | 0.46 |
ENST00000403665.2 ENST00000264692.4 |
F11 |
coagulation factor XI |
| chr4_+_68424434 | 0.46 |
ENST00000265404.2 ENST00000396225.1 |
STAP1 |
signal transducing adaptor family member 1 |
| chr18_-_52626622 | 0.45 |
ENST00000591504.1 |
CCDC68 |
coiled-coil domain containing 68 |
| chr17_+_4613918 | 0.45 |
ENST00000574954.1 ENST00000346341.2 ENST00000572457.1 ENST00000381488.6 ENST00000412477.3 ENST00000571428.1 ENST00000575877.1 |
ARRB2 |
arrestin, beta 2 |
| chr14_-_24911448 | 0.44 |
ENST00000555355.1 ENST00000553343.1 ENST00000556523.1 ENST00000556249.1 ENST00000538105.2 ENST00000555225.1 |
SDR39U1 |
short chain dehydrogenase/reductase family 39U, member 1 |
| chr13_+_76334567 | 0.44 |
ENST00000321797.8 |
LMO7 |
LIM domain 7 |
| chr19_-_15443318 | 0.42 |
ENST00000360016.5 |
BRD4 |
bromodomain containing 4 |
| chr6_+_135502466 | 0.42 |
ENST00000367814.4 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chrX_+_37639302 | 0.42 |
ENST00000545017.1 ENST00000536160.1 |
CYBB |
cytochrome b-245, beta polypeptide |
| chrX_+_37639264 | 0.41 |
ENST00000378588.4 |
CYBB |
cytochrome b-245, beta polypeptide |
| chr6_+_135502408 | 0.40 |
ENST00000341911.5 ENST00000442647.2 ENST00000316528.8 |
MYB |
v-myb avian myeloblastosis viral oncogene homolog |
| chr2_-_158345462 | 0.40 |
ENST00000439355.1 ENST00000540637.1 |
CYTIP |
cytohesin 1 interacting protein |
| chr3_+_52811596 | 0.40 |
ENST00000542827.1 ENST00000273283.2 |
ITIH1 |
inter-alpha-trypsin inhibitor heavy chain 1 |
| chr12_+_121416340 | 0.40 |
ENST00000257555.6 ENST00000400024.2 |
HNF1A |
HNF1 homeobox A |
| chr7_+_116654935 | 0.40 |
ENST00000432298.1 ENST00000422922.1 |
ST7 |
suppression of tumorigenicity 7 |
| chr20_+_52105495 | 0.38 |
ENST00000439873.2 |
AL354993.1 |
Cell growth-inhibiting protein 7; HCG1784586; Uncharacterized protein |
| chr8_-_71316021 | 0.37 |
ENST00000452400.2 |
NCOA2 |
nuclear receptor coactivator 2 |
| chr6_+_106546808 | 0.36 |
ENST00000369089.3 |
PRDM1 |
PR domain containing 1, with ZNF domain |
| chr13_+_76334795 | 0.36 |
ENST00000526202.1 ENST00000465261.2 |
LMO7 |
LIM domain 7 |
| chr7_+_28452130 | 0.36 |
ENST00000357727.2 |
CREB5 |
cAMP responsive element binding protein 5 |
| chr3_-_120068143 | 0.35 |
ENST00000295628.3 |
LRRC58 |
leucine rich repeat containing 58 |
| chr4_+_170581213 | 0.34 |
ENST00000507875.1 |
CLCN3 |
chloride channel, voltage-sensitive 3 |
| chrX_+_24167746 | 0.34 |
ENST00000428571.1 ENST00000539115.1 |
ZFX |
zinc finger protein, X-linked |
| chrX_+_22050546 | 0.34 |
ENST00000379374.4 |
PHEX |
phosphate regulating endopeptidase homolog, X-linked |
| chr7_+_112063192 | 0.33 |
ENST00000005558.4 |
IFRD1 |
interferon-related developmental regulator 1 |
| chr7_+_95115210 | 0.33 |
ENST00000428113.1 ENST00000325885.5 |
ASB4 |
ankyrin repeat and SOCS box containing 4 |
| chr14_-_31926701 | 0.33 |
ENST00000310850.4 |
DTD2 |
D-tyrosyl-tRNA deacylase 2 (putative) |
| chr9_-_20622478 | 0.31 |
ENST00000355930.6 ENST00000380338.4 |
MLLT3 |
myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 |
| chr6_+_31582961 | 0.31 |
ENST00000376059.3 ENST00000337917.7 |
AIF1 |
allograft inflammatory factor 1 |
| chr1_+_12538594 | 0.29 |
ENST00000543710.1 |
VPS13D |
vacuolar protein sorting 13 homolog D (S. cerevisiae) |
| chr18_-_25616519 | 0.29 |
ENST00000399380.3 |
CDH2 |
cadherin 2, type 1, N-cadherin (neuronal) |
| chr6_+_33589161 | 0.29 |
ENST00000605930.1 |
ITPR3 |
inositol 1,4,5-trisphosphate receptor, type 3 |
| chr19_-_48894104 | 0.28 |
ENST00000597017.1 |
KDELR1 |
KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 1 |
| chr6_-_89927151 | 0.28 |
ENST00000454853.2 |
GABRR1 |
gamma-aminobutyric acid (GABA) A receptor, rho 1 |
| chr6_+_108977520 | 0.28 |
ENST00000540898.1 |
FOXO3 |
forkhead box O3 |
| chrX_-_24665208 | 0.28 |
ENST00000356768.4 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr2_+_47630108 | 0.27 |
ENST00000233146.2 ENST00000454849.1 ENST00000543555.1 |
MSH2 |
mutS homolog 2 |
| chr4_-_100356844 | 0.27 |
ENST00000437033.2 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr8_+_119294456 | 0.26 |
ENST00000366457.2 |
AC023590.1 |
Uncharacterized protein |
| chr3_+_148545586 | 0.25 |
ENST00000282957.4 ENST00000468341.1 |
CPB1 |
carboxypeptidase B1 (tissue) |
| chr9_-_98279241 | 0.25 |
ENST00000437951.1 ENST00000375274.2 ENST00000430669.2 ENST00000468211.2 |
PTCH1 |
patched 1 |
| chr2_-_220042825 | 0.24 |
ENST00000409789.1 |
CNPPD1 |
cyclin Pas1/PHO80 domain containing 1 |
| chr9_-_123239632 | 0.24 |
ENST00000416449.1 |
CDK5RAP2 |
CDK5 regulatory subunit associated protein 2 |
| chr12_+_10658489 | 0.23 |
ENST00000538173.1 |
EIF2S3L |
Putative eukaryotic translation initiation factor 2 subunit 3-like protein |
| chr11_+_71249071 | 0.23 |
ENST00000398534.3 |
KRTAP5-8 |
keratin associated protein 5-8 |
| chr2_-_214016314 | 0.22 |
ENST00000434687.1 ENST00000374319.4 |
IKZF2 |
IKAROS family zinc finger 2 (Helios) |
| chr6_+_112375462 | 0.22 |
ENST00000361714.1 |
WISP3 |
WNT1 inducible signaling pathway protein 3 |
| chr2_+_47630255 | 0.22 |
ENST00000406134.1 |
MSH2 |
mutS homolog 2 |
| chr1_-_216596738 | 0.21 |
ENST00000307340.3 ENST00000366943.2 ENST00000366942.3 |
USH2A |
Usher syndrome 2A (autosomal recessive, mild) |
| chr14_+_37126765 | 0.21 |
ENST00000402703.2 |
PAX9 |
paired box 9 |
| chr12_-_772901 | 0.21 |
ENST00000305108.4 |
NINJ2 |
ninjurin 2 |
| chrX_+_135730373 | 0.20 |
ENST00000370628.2 |
CD40LG |
CD40 ligand |
| chr7_+_106505912 | 0.20 |
ENST00000359195.3 |
PIK3CG |
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit gamma |
| chrX_+_24167828 | 0.20 |
ENST00000379188.3 ENST00000419690.1 ENST00000379177.1 ENST00000304543.5 |
ZFX |
zinc finger protein, X-linked |
| chr8_+_77593448 | 0.20 |
ENST00000521891.2 |
ZFHX4 |
zinc finger homeobox 4 |
| chr9_-_95186739 | 0.20 |
ENST00000375550.4 |
OMD |
osteomodulin |
| chr7_+_106505696 | 0.19 |
ENST00000440650.2 ENST00000496166.1 ENST00000473541.1 |
PIK3CG |
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit gamma |
| chrX_-_24665353 | 0.19 |
ENST00000379144.2 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr3_+_148447887 | 0.19 |
ENST00000475347.1 ENST00000474935.1 ENST00000461609.1 |
AGTR1 |
angiotensin II receptor, type 1 |
| chr22_-_40929812 | 0.19 |
ENST00000422851.1 |
MKL1 |
megakaryoblastic leukemia (translocation) 1 |
| chr3_-_113897899 | 0.18 |
ENST00000383673.2 ENST00000295881.7 |
DRD3 |
dopamine receptor D3 |
| chrX_+_135730297 | 0.18 |
ENST00000370629.2 |
CD40LG |
CD40 ligand |
| chr9_+_136287444 | 0.18 |
ENST00000355699.2 ENST00000356589.2 ENST00000371911.3 |
ADAMTS13 |
ADAM metallopeptidase with thrombospondin type 1 motif, 13 |
| chr5_+_161494770 | 0.17 |
ENST00000414552.2 ENST00000361925.4 |
GABRG2 |
gamma-aminobutyric acid (GABA) A receptor, gamma 2 |
| chr2_+_204801471 | 0.16 |
ENST00000316386.6 ENST00000435193.1 |
ICOS |
inducible T-cell co-stimulator |
| chr16_-_67517716 | 0.16 |
ENST00000290953.2 |
AGRP |
agouti related protein homolog (mouse) |
| chr12_-_89746173 | 0.16 |
ENST00000308385.6 |
DUSP6 |
dual specificity phosphatase 6 |
| chr3_+_137728842 | 0.16 |
ENST00000183605.5 |
CLDN18 |
claudin 18 |
| chrX_+_99839799 | 0.14 |
ENST00000373031.4 |
TNMD |
tenomodulin |
| chr6_+_7107830 | 0.14 |
ENST00000379933.3 |
RREB1 |
ras responsive element binding protein 1 |
| chr20_+_62795827 | 0.14 |
ENST00000328439.1 ENST00000536311.1 |
MYT1 |
myelin transcription factor 1 |
| chr14_+_23299088 | 0.13 |
ENST00000355151.5 ENST00000397496.3 ENST00000555345.1 ENST00000432849.3 ENST00000553711.1 ENST00000556465.1 ENST00000397505.2 ENST00000557221.1 ENST00000311892.6 ENST00000556840.1 ENST00000555536.1 |
MRPL52 |
mitochondrial ribosomal protein L52 |
| chr17_-_48546324 | 0.13 |
ENST00000508540.1 |
CHAD |
chondroadherin |
| chr2_-_219850277 | 0.13 |
ENST00000295727.1 |
FEV |
FEV (ETS oncogene family) |
| chr4_+_165675269 | 0.13 |
ENST00000507311.1 |
RP11-294O2.2 |
RP11-294O2.2 |
| chr12_-_21487829 | 0.13 |
ENST00000445053.1 ENST00000452078.1 ENST00000458504.1 ENST00000422327.1 ENST00000421294.1 |
SLCO1A2 |
solute carrier organic anion transporter family, member 1A2 |
| chr8_+_77593474 | 0.12 |
ENST00000455469.2 ENST00000050961.6 |
ZFHX4 |
zinc finger homeobox 4 |
| chr8_+_133879193 | 0.12 |
ENST00000377869.1 ENST00000220616.4 |
TG |
thyroglobulin |
| chr6_+_7108210 | 0.11 |
ENST00000467782.1 ENST00000334984.6 ENST00000349384.6 |
RREB1 |
ras responsive element binding protein 1 |
| chr1_+_74701062 | 0.11 |
ENST00000326637.3 |
TNNI3K |
TNNI3 interacting kinase |
| chr4_-_111119804 | 0.11 |
ENST00000394607.3 ENST00000302274.3 |
ELOVL6 |
ELOVL fatty acid elongase 6 |
| chr3_+_141106643 | 0.11 |
ENST00000514251.1 |
ZBTB38 |
zinc finger and BTB domain containing 38 |
| chr5_+_161494521 | 0.11 |
ENST00000356592.3 |
GABRG2 |
gamma-aminobutyric acid (GABA) A receptor, gamma 2 |
| chr17_+_67410832 | 0.10 |
ENST00000590474.1 |
MAP2K6 |
mitogen-activated protein kinase kinase 6 |
| chr21_-_31588365 | 0.10 |
ENST00000399899.1 |
CLDN8 |
claudin 8 |
| chr6_+_161123270 | 0.10 |
ENST00000366924.2 ENST00000308192.9 ENST00000418964.1 |
PLG |
plasminogen |
| chr6_-_31550192 | 0.10 |
ENST00000429299.2 ENST00000446745.2 |
LTB |
lymphotoxin beta (TNF superfamily, member 3) |
| chr3_-_186080012 | 0.10 |
ENST00000544847.1 ENST00000265022.3 |
DGKG |
diacylglycerol kinase, gamma 90kDa |
| chr2_+_103089756 | 0.09 |
ENST00000295269.4 |
SLC9A4 |
solute carrier family 9, subfamily A (NHE4, cation proton antiporter 4), member 4 |
| chr1_+_87797351 | 0.09 |
ENST00000370542.1 |
LMO4 |
LIM domain only 4 |
| chr1_-_111743285 | 0.09 |
ENST00000357640.4 |
DENND2D |
DENN/MADD domain containing 2D |
| chr5_-_137674000 | 0.09 |
ENST00000510119.1 ENST00000513970.1 |
CDC25C |
cell division cycle 25C |
| chr5_+_32531893 | 0.08 |
ENST00000512913.1 |
SUB1 |
SUB1 homolog (S. cerevisiae) |
| chr3_+_160394940 | 0.08 |
ENST00000320767.2 |
ARL14 |
ADP-ribosylation factor-like 14 |
| chr14_+_56127989 | 0.08 |
ENST00000555573.1 |
KTN1 |
kinectin 1 (kinesin receptor) |
| chr4_-_87028478 | 0.08 |
ENST00000515400.1 ENST00000395157.3 |
MAPK10 |
mitogen-activated protein kinase 10 |
| chr3_-_105588231 | 0.08 |
ENST00000545639.1 ENST00000394027.3 ENST00000438603.1 ENST00000447441.1 ENST00000443752.1 |
CBLB |
Cbl proto-oncogene B, E3 ubiquitin protein ligase |
| chr2_+_162101247 | 0.08 |
ENST00000439050.1 ENST00000436506.1 |
AC009299.3 |
AC009299.3 |
| chr10_-_32217717 | 0.08 |
ENST00000396144.4 ENST00000375245.4 ENST00000344936.2 ENST00000375250.5 |
ARHGAP12 |
Rho GTPase activating protein 12 |
| chr5_-_127674883 | 0.08 |
ENST00000507835.1 |
FBN2 |
fibrillin 2 |
| chr10_+_118350468 | 0.07 |
ENST00000358834.4 ENST00000528052.1 ENST00000442761.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr2_-_3521518 | 0.07 |
ENST00000382093.5 |
ADI1 |
acireductone dioxygenase 1 |
| chr5_+_161495038 | 0.07 |
ENST00000393933.4 |
GABRG2 |
gamma-aminobutyric acid (GABA) A receptor, gamma 2 |
| chr17_-_48546232 | 0.07 |
ENST00000258969.4 |
CHAD |
chondroadherin |
| chr17_-_17485731 | 0.07 |
ENST00000395783.1 |
PEMT |
phosphatidylethanolamine N-methyltransferase |
| chr4_-_174256276 | 0.07 |
ENST00000296503.5 |
HMGB2 |
high mobility group box 2 |
| chr13_-_99910673 | 0.07 |
ENST00000397473.2 ENST00000397470.2 |
GPR18 |
G protein-coupled receptor 18 |
| chr7_-_99698338 | 0.07 |
ENST00000354230.3 ENST00000425308.1 |
MCM7 |
minichromosome maintenance complex component 7 |
| chr21_-_31588338 | 0.06 |
ENST00000286809.1 |
CLDN8 |
claudin 8 |
| chr9_+_105757590 | 0.06 |
ENST00000374798.3 ENST00000487798.1 |
CYLC2 |
cylicin, basic protein of sperm head cytoskeleton 2 |
| chr15_+_75335604 | 0.06 |
ENST00000563393.1 |
PPCDC |
phosphopantothenoylcysteine decarboxylase |
| chr10_-_81708854 | 0.05 |
ENST00000372292.3 |
SFTPD |
surfactant protein D |
| chr17_+_8924837 | 0.05 |
ENST00000173229.2 |
NTN1 |
netrin 1 |
| chr6_-_87804815 | 0.05 |
ENST00000369582.2 |
CGA |
glycoprotein hormones, alpha polypeptide |
| chr7_-_107443652 | 0.05 |
ENST00000340010.5 ENST00000422236.2 ENST00000453332.1 |
SLC26A3 |
solute carrier family 26 (anion exchanger), member 3 |
| chr5_-_148758839 | 0.05 |
ENST00000261796.3 |
IL17B |
interleukin 17B |
| chr21_-_38445011 | 0.05 |
ENST00000464265.1 ENST00000399102.1 |
PIGP |
phosphatidylinositol glycan anchor biosynthesis, class P |
| chrX_-_124097620 | 0.04 |
ENST00000371130.3 ENST00000422452.2 |
TENM1 |
teneurin transmembrane protein 1 |
| chr20_-_55841398 | 0.04 |
ENST00000395864.3 |
BMP7 |
bone morphogenetic protein 7 |
| chr17_+_9728828 | 0.04 |
ENST00000262441.5 |
GLP2R |
glucagon-like peptide 2 receptor |
| chr6_+_146350267 | 0.04 |
ENST00000355289.4 ENST00000282753.1 |
GRM1 |
glutamate receptor, metabotropic 1 |
| chr12_-_102872317 | 0.03 |
ENST00000424202.2 |
IGF1 |
insulin-like growth factor 1 (somatomedin C) |
| chr2_+_169757750 | 0.03 |
ENST00000375363.3 ENST00000429379.2 ENST00000421979.1 |
G6PC2 |
glucose-6-phosphatase, catalytic, 2 |
| chr7_+_107384142 | 0.02 |
ENST00000440859.3 |
CBLL1 |
Cbl proto-oncogene-like 1, E3 ubiquitin protein ligase |
| chr16_-_68034470 | 0.01 |
ENST00000412757.2 |
DPEP2 |
dipeptidase 2 |
| chr20_-_55841662 | 0.01 |
ENST00000395863.3 ENST00000450594.2 |
BMP7 |
bone morphogenetic protein 7 |
| chr7_+_129906660 | 0.01 |
ENST00000222481.4 |
CPA2 |
carboxypeptidase A2 (pancreatic) |
| chr12_-_68647281 | 0.01 |
ENST00000328087.4 ENST00000538666.1 |
IL22 |
interleukin 22 |
| chr16_-_86542455 | 0.01 |
ENST00000595886.1 ENST00000597578.1 ENST00000593604.1 |
FENDRR |
FOXF1 adjacent non-coding developmental regulatory RNA |
| chr13_-_49975632 | 0.01 |
ENST00000457041.1 ENST00000355854.4 |
CAB39L |
calcium binding protein 39-like |
| chr7_+_31003621 | 0.01 |
ENST00000326139.2 |
GHRHR |
growth hormone releasing hormone receptor |
| chr12_+_21525818 | 0.00 |
ENST00000240652.3 ENST00000542023.1 ENST00000537593.1 |
IAPP |
islet amyloid polypeptide |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 4.9 | GO:0035473 | lipase binding(GO:0035473) |
| 0.4 | 3.9 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.4 | 6.5 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.3 | 1.0 | GO:0031859 | platelet activating factor receptor binding(GO:0031859) |
| 0.3 | 1.3 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.2 | 1.3 | GO:0017050 | sphinganine kinase activity(GO:0008481) D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.2 | 0.5 | GO:0032181 | heteroduplex DNA loop binding(GO:0000404) double-strand/single-strand DNA junction binding(GO:0000406) dinucleotide repeat insertion binding(GO:0032181) |
| 0.1 | 1.2 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 1.8 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 0.4 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.1 | 0.9 | GO:0008109 | N-acetyllactosaminide beta-1,6-N-acetylglucosaminyltransferase activity(GO:0008109) |
| 0.1 | 0.4 | GO:0010861 | thyroid hormone receptor activator activity(GO:0010861) thyroid hormone receptor coactivator activity(GO:0030375) |
| 0.1 | 1.3 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.1 | 1.9 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.1 | 0.5 | GO:0018479 | benzaldehyde dehydrogenase (NAD+) activity(GO:0018479) |
| 0.1 | 2.1 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.1 | 0.5 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.1 | 0.5 | GO:0005157 | macrophage colony-stimulating factor receptor binding(GO:0005157) |
| 0.1 | 8.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 2.9 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 5.6 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.1 | 0.3 | GO:0000822 | inositol hexakisphosphate binding(GO:0000822) inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.1 | 0.3 | GO:0005046 | KDEL sequence binding(GO:0005046) |
| 0.1 | 0.3 | GO:0035276 | aldehyde oxidase activity(GO:0004031) ethanol binding(GO:0035276) |
| 0.1 | 1.1 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.1 | 0.2 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.1 | 0.3 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.1 | 0.4 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.1 | 0.3 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 0.0 | 1.6 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.8 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.0 | 0.7 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.3 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.0 | 1.5 | GO:0001105 | RNA polymerase II transcription coactivator activity(GO:0001105) |
| 0.0 | 1.8 | GO:0005160 | transforming growth factor beta receptor binding(GO:0005160) |
| 0.0 | 0.3 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) |
| 0.0 | 0.2 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.1 | GO:0080101 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 0.0 | 0.2 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 0.3 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.0 | 0.5 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.0 | 3.4 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.1 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.1 | GO:0044378 | non-sequence-specific DNA binding, bending(GO:0044378) |
| 0.0 | 0.0 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 0.0 | 0.1 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.0 | 1.0 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 1.2 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.0 | 0.1 | GO:1990405 | protein antigen binding(GO:1990405) |
| 0.0 | 0.2 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.0 | 0.4 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.1 | GO:0031013 | troponin I binding(GO:0031013) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 6.5 | GO:0071062 | rough endoplasmic reticulum lumen(GO:0048237) alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 0.6 | 4.9 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 0.6 | 3.3 | GO:1990357 | terminal web(GO:1990357) |
| 0.2 | 2.7 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.2 | 0.5 | GO:0032302 | MutSbeta complex(GO:0032302) |
| 0.1 | 0.4 | GO:0005944 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) |
| 0.1 | 1.3 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.1 | 0.8 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 0.2 | GO:0002139 | stereocilia coupling link(GO:0002139) |
| 0.1 | 1.4 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 1.9 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 0.3 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.0 | 4.6 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.8 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.0 | 13.0 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.3 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.0 | 0.1 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.1 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.0 | 1.2 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 0.1 | GO:0033011 | perinuclear theca(GO:0033011) cytoskeletal calyx(GO:0033150) |
| 0.0 | 0.2 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.0 | 0.3 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.0 | 0.2 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.5 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 2.5 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.0 | 0.9 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.0 | 1.9 | GO:0030426 | growth cone(GO:0030426) |
| 0.0 | 1.0 | GO:0005905 | clathrin-coated pit(GO:0005905) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.9 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.2 | 2.7 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.2 | 3.9 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.1 | 1.2 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.1 | 2.9 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.1 | 1.1 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.1 | 7.8 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 1.7 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.0 | 1.2 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 1.3 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.9 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.3 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.0 | 1.0 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.4 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 0.5 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 1.7 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.7 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.4 | REACTOME G BETA GAMMA SIGNALLING THROUGH PI3KGAMMA | Genes involved in G beta:gamma signalling through PI3Kgamma |
| 0.0 | 0.6 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.0 | 0.7 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.0 | 0.5 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.0 | 0.1 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 6.5 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.7 | 4.9 | GO:0006642 | triglyceride mobilization(GO:0006642) response to selenium ion(GO:0010269) |
| 0.6 | 1.9 | GO:0045659 | regulation of eosinophil differentiation(GO:0045643) positive regulation of eosinophil differentiation(GO:0045645) regulation of neutrophil differentiation(GO:0045658) negative regulation of neutrophil differentiation(GO:0045659) |
| 0.6 | 3.9 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.4 | 1.3 | GO:1904897 | regulation of hepatic stellate cell proliferation(GO:1904897) positive regulation of hepatic stellate cell proliferation(GO:1904899) hepatic stellate cell proliferation(GO:1990922) |
| 0.4 | 1.3 | GO:0046521 | sphingoid catabolic process(GO:0046521) |
| 0.4 | 1.6 | GO:0060743 | epithelial cell maturation involved in prostate gland development(GO:0060743) |
| 0.4 | 1.2 | GO:0046619 | optic placode formation involved in camera-type eye formation(GO:0046619) |
| 0.4 | 2.1 | GO:0002760 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antimicrobial humoral response(GO:0002760) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.3 | 1.0 | GO:0035565 | regulation of pronephros size(GO:0035565) renal glucose absorption(GO:0035623) |
| 0.2 | 1.1 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.2 | 0.9 | GO:1903691 | positive regulation of wound healing, spreading of epidermal cells(GO:1903691) |
| 0.2 | 1.5 | GO:1902897 | regulation of postsynaptic density protein 95 clustering(GO:1902897) |
| 0.2 | 1.0 | GO:0002032 | desensitization of G-protein coupled receptor protein signaling pathway by arrestin(GO:0002032) |
| 0.2 | 4.6 | GO:0030502 | negative regulation of bone mineralization(GO:0030502) |
| 0.2 | 0.8 | GO:1904844 | response to L-glutamine(GO:1904844) cellular response to L-glutamine(GO:1904845) |
| 0.2 | 0.5 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) negative regulation of macrophage colony-stimulating factor signaling pathway(GO:1902227) negative regulation of response to macrophage colony-stimulating factor(GO:1903970) negative regulation of cellular response to macrophage colony-stimulating factor stimulus(GO:1903973) |
| 0.2 | 3.9 | GO:0006825 | copper ion transport(GO:0006825) |
| 0.1 | 2.7 | GO:0090331 | negative regulation of platelet aggregation(GO:0090331) |
| 0.1 | 0.5 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 0.1 | 4.0 | GO:0008211 | glucocorticoid metabolic process(GO:0008211) |
| 0.1 | 1.2 | GO:0002667 | lymphocyte anergy(GO:0002249) regulation of T cell anergy(GO:0002667) T cell anergy(GO:0002870) regulation of lymphocyte anergy(GO:0002911) |
| 0.1 | 0.9 | GO:0036438 | maintenance of lens transparency(GO:0036438) |
| 0.1 | 2.0 | GO:0090361 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 0.1 | 1.1 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.1 | 1.3 | GO:0060174 | limb bud formation(GO:0060174) |
| 0.1 | 0.3 | GO:0007518 | myoblast fate determination(GO:0007518) |
| 0.1 | 0.2 | GO:1900155 | regulation of bone trabecula formation(GO:1900154) negative regulation of bone trabecula formation(GO:1900155) |
| 0.1 | 0.5 | GO:0010520 | meiotic gene conversion(GO:0006311) regulation of reciprocal meiotic recombination(GO:0010520) |
| 0.1 | 0.3 | GO:0050917 | sensory perception of sweet taste(GO:0050916) sensory perception of umami taste(GO:0050917) |
| 0.1 | 0.3 | GO:0001544 | initiation of primordial ovarian follicle growth(GO:0001544) |
| 0.1 | 0.3 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.1 | 0.7 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.1 | 0.3 | GO:1904383 | response to sodium phosphate(GO:1904383) |
| 0.1 | 1.8 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.1 | 1.1 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 0.4 | GO:1904016 | response to Thyroglobulin triiodothyronine(GO:1904016) |
| 0.1 | 0.4 | GO:0033078 | extrathymic T cell differentiation(GO:0033078) |
| 0.1 | 1.0 | GO:0051599 | response to hydrostatic pressure(GO:0051599) |
| 0.1 | 0.3 | GO:0048388 | endosomal lumen acidification(GO:0048388) |
| 0.1 | 0.4 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) secretory granule localization(GO:0032252) |
| 0.1 | 0.3 | GO:0021997 | response to chlorate(GO:0010157) neural plate axis specification(GO:0021997) |
| 0.1 | 0.3 | GO:0014738 | regulation of muscle hyperplasia(GO:0014738) |
| 0.1 | 1.6 | GO:0043555 | regulation of translation in response to stress(GO:0043555) |
| 0.1 | 0.2 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.0 | 0.2 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.0 | 0.2 | GO:0050883 | negative regulation of sodium:proton antiporter activity(GO:0032416) gastric motility(GO:0035482) gastric emptying(GO:0035483) musculoskeletal movement, spinal reflex action(GO:0050883) |
| 0.0 | 0.3 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.0 | 1.8 | GO:0030195 | negative regulation of blood coagulation(GO:0030195) negative regulation of hemostasis(GO:1900047) |
| 0.0 | 4.3 | GO:0046324 | regulation of glucose import(GO:0046324) |
| 0.0 | 0.2 | GO:1903224 | regulation of endodermal cell differentiation(GO:1903224) |
| 0.0 | 0.5 | GO:2001224 | positive regulation of neuron migration(GO:2001224) |
| 0.0 | 0.5 | GO:0006657 | CDP-choline pathway(GO:0006657) |
| 0.0 | 0.9 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.0 | 0.2 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.0 | 0.3 | GO:2001214 | positive regulation of vasculogenesis(GO:2001214) |
| 0.0 | 1.5 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 0.0 | 0.1 | GO:0035990 | tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) |
| 0.0 | 2.2 | GO:0045214 | sarcomere organization(GO:0045214) |
| 0.0 | 0.4 | GO:1901407 | regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901407) |
| 0.0 | 0.4 | GO:2000353 | positive regulation of endothelial cell apoptotic process(GO:2000353) |
| 0.0 | 0.1 | GO:0072709 | cellular response to sorbitol(GO:0072709) |
| 0.0 | 0.1 | GO:0021502 | neural fold elevation formation(GO:0021502) allantois development(GO:1905069) |
| 0.0 | 0.7 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.0 | 0.3 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.0 | 1.8 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.0 | 0.1 | GO:0015705 | iodide transport(GO:0015705) |
| 0.0 | 1.1 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.0 | 0.2 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.0 | 1.1 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.0 | 0.2 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.0 | 0.1 | GO:0034625 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.2 | GO:0008343 | adult feeding behavior(GO:0008343) |
| 0.0 | 0.3 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.0 | 0.1 | GO:0045085 | negative regulation of interleukin-2 biosynthetic process(GO:0045085) |
| 0.0 | 0.3 | GO:0006450 | regulation of translational fidelity(GO:0006450) |
| 0.0 | 0.2 | GO:0002517 | T cell tolerance induction(GO:0002517) |
| 0.0 | 0.1 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.0 | 0.5 | GO:1900077 | negative regulation of insulin receptor signaling pathway(GO:0046627) negative regulation of cellular response to insulin stimulus(GO:1900077) |
| 0.0 | 1.3 | GO:0021510 | spinal cord development(GO:0021510) |
| 0.0 | 0.1 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 6.5 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 2.2 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 4.3 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.1 | 3.1 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 4.2 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 1.3 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.0 | 1.6 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.0 | 4.7 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.4 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.0 | 2.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 2.4 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 1.6 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.0 | 1.3 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 6.1 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.6 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 0.5 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 1.0 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.3 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.0 | 0.3 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.0 | 1.1 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.5 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |