Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
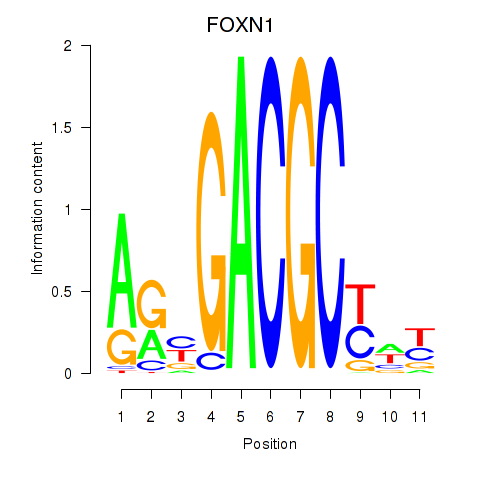
Results for FOXN1
Z-value: 0.40
Transcription factors associated with FOXN1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
FOXN1
|
ENSG00000109101.3 | FOXN1 |
Activity profile of FOXN1 motif
Sorted Z-values of FOXN1 motif
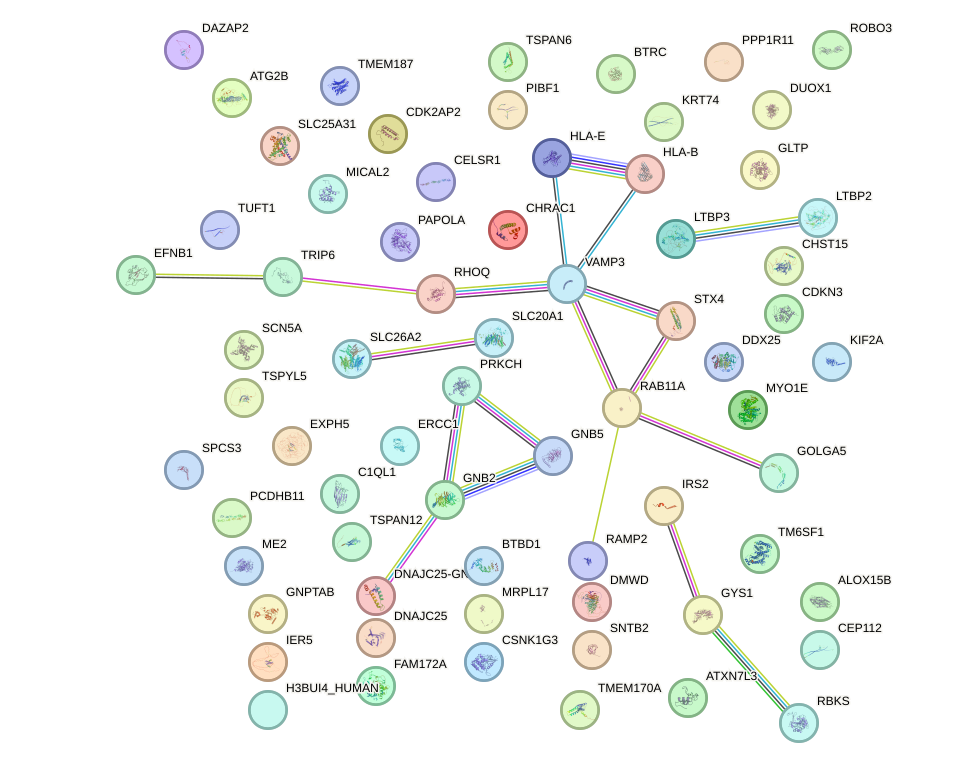
Network of associatons between targets according to the STRING database.
First level regulatory network of FOXN1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_52967600 | 0.62 |
ENST00000549343.1 ENST00000305620.2 |
KRT74 |
keratin 74 |
| chr7_-_120498357 | 0.49 |
ENST00000415871.1 ENST00000222747.3 ENST00000430985.1 |
TSPAN12 |
tetraspanin 12 |
| chr3_-_38691119 | 0.39 |
ENST00000333535.4 ENST00000413689.1 ENST00000443581.1 ENST00000425664.1 ENST00000451551.2 |
SCN5A |
sodium channel, voltage-gated, type V, alpha subunit |
| chr15_-_59665062 | 0.38 |
ENST00000288235.4 |
MYO1E |
myosin IE |
| chr15_+_71184931 | 0.37 |
ENST00000560369.1 ENST00000260382.5 |
LRRC49 |
leucine rich repeat containing 49 |
| chr15_+_71185148 | 0.35 |
ENST00000443425.2 ENST00000560755.1 |
LRRC49 |
leucine rich repeat containing 49 |
| chr13_-_110438914 | 0.34 |
ENST00000375856.3 |
IRS2 |
insulin receptor substrate 2 |
| chr3_-_49395892 | 0.32 |
ENST00000419783.1 |
GPX1 |
glutathione peroxidase 1 |
| chr17_+_7942335 | 0.32 |
ENST00000380183.4 ENST00000572022.1 ENST00000380173.2 |
ALOX15B |
arachidonate 15-lipoxygenase, type B |
| chr3_-_49395705 | 0.32 |
ENST00000419349.1 |
GPX1 |
glutathione peroxidase 1 |
| chr7_+_100464760 | 0.32 |
ENST00000200457.4 |
TRIP6 |
thyroid hormone receptor interactor 6 |
| chr15_+_83776137 | 0.31 |
ENST00000322019.9 |
TM6SF1 |
transmembrane 6 superfamily member 1 |
| chr2_+_113403434 | 0.31 |
ENST00000272542.3 |
SLC20A1 |
solute carrier family 20 (phosphate transporter), member 1 |
| chr12_-_51663728 | 0.31 |
ENST00000603864.1 ENST00000605426.1 |
SMAGP |
small cell adhesion glycoprotein |
| chr12_-_51663959 | 0.30 |
ENST00000604188.1 ENST00000398453.3 |
SMAGP |
small cell adhesion glycoprotein |
| chr14_+_61788429 | 0.30 |
ENST00000332981.5 |
PRKCH |
protein kinase C, eta |
| chr10_+_13141441 | 0.29 |
ENST00000263036.5 |
OPTN |
optineurin |
| chr15_+_83776324 | 0.29 |
ENST00000379390.6 ENST00000379386.4 ENST00000565774.1 ENST00000565982.1 |
TM6SF1 |
transmembrane 6 superfamily member 1 |
| chr1_+_7831323 | 0.29 |
ENST00000054666.6 |
VAMP3 |
vesicle-associated membrane protein 3 |
| chr5_+_149340282 | 0.28 |
ENST00000286298.4 |
SLC26A2 |
solute carrier family 26 (anion exchanger), member 2 |
| chr10_+_13141585 | 0.27 |
ENST00000378764.2 |
OPTN |
optineurin |
| chr18_+_9708162 | 0.27 |
ENST00000578921.1 |
RAB31 |
RAB31, member RAS oncogene family |
| chr11_+_124735282 | 0.25 |
ENST00000397801.1 |
ROBO3 |
roundabout, axon guidance receptor, homolog 3 (Drosophila) |
| chr19_-_45953983 | 0.25 |
ENST00000592083.1 |
ERCC1 |
excision repair cross-complementing rodent repair deficiency, complementation group 1 (includes overlapping antisense sequence) |
| chr2_-_31360887 | 0.24 |
ENST00000420311.2 ENST00000356174.3 ENST00000324589.5 |
GALNT14 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 14 (GalNAc-T14) |
| chr6_-_128841503 | 0.24 |
ENST00000368215.3 ENST00000532331.1 ENST00000368213.5 ENST00000368207.3 ENST00000525459.1 ENST00000368210.3 ENST00000368226.4 ENST00000368227.3 |
PTPRK |
protein tyrosine phosphatase, receptor type, K |
| chr8_+_31496809 | 0.24 |
ENST00000518104.1 ENST00000519301.1 |
NRG1 |
neuregulin 1 |
| chr17_-_64187973 | 0.24 |
ENST00000583358.1 ENST00000392769.2 |
CEP112 |
centrosomal protein 112kDa |
| chr11_+_12132117 | 0.23 |
ENST00000256194.4 |
MICAL2 |
microtubule associated monooxygenase, calponin and LIM domain containing 2 |
| chrX_+_153237740 | 0.23 |
ENST00000369982.4 |
TMEM187 |
transmembrane protein 187 |
| chr15_+_45422131 | 0.22 |
ENST00000321429.4 |
DUOX1 |
dual oxidase 1 |
| chr9_+_114393634 | 0.22 |
ENST00000556107.1 ENST00000374294.3 |
DNAJC25 DNAJC25-GNG10 |
DnaJ (Hsp40) homolog, subfamily C , member 25 DNAJC25-GNG10 readthrough |
| chr10_+_13142225 | 0.22 |
ENST00000378747.3 |
OPTN |
optineurin |
| chr10_+_13142075 | 0.21 |
ENST00000378757.2 ENST00000430081.1 ENST00000378752.3 ENST00000378748.3 |
OPTN |
optineurin |
| chr16_+_69221028 | 0.21 |
ENST00000336278.4 |
SNTB2 |
syntrophin, beta 2 (dystrophin-associated protein A1, 59kDa, basic component 2) |
| chr15_+_45422178 | 0.21 |
ENST00000389037.3 ENST00000558322.1 |
DUOX1 |
dual oxidase 1 |
| chr20_-_36156125 | 0.21 |
ENST00000397135.1 ENST00000397137.1 |
BLCAP |
bladder cancer associated protein |
| chr8_-_98290087 | 0.20 |
ENST00000322128.3 |
TSPYL5 |
TSPY-like 5 |
| chr11_+_1860200 | 0.20 |
ENST00000381911.1 |
TNNI2 |
troponin I type 2 (skeletal, fast) |
| chr15_+_66161802 | 0.19 |
ENST00000566233.1 ENST00000565075.1 ENST00000435304.2 |
RAB11A |
RAB11A, member RAS oncogene family |
| chr12_-_110318226 | 0.19 |
ENST00000544393.1 |
GLTP |
glycolipid transfer protein |
| chr12_+_51632638 | 0.19 |
ENST00000549732.2 |
DAZAP2 |
DAZ associated protein 2 |
| chr6_+_29910301 | 0.18 |
ENST00000376809.5 ENST00000376802.2 |
HLA-A |
major histocompatibility complex, class I, A |
| chr12_+_51632600 | 0.18 |
ENST00000549555.1 ENST00000439799.2 ENST00000425012.2 |
DAZAP2 |
DAZ associated protein 2 |
| chr1_-_115300579 | 0.18 |
ENST00000358528.4 ENST00000525132.1 |
CSDE1 |
cold shock domain containing E1, RNA-binding |
| chr15_+_66161792 | 0.17 |
ENST00000564910.1 ENST00000261890.2 |
RAB11A |
RAB11A, member RAS oncogene family |
| chr15_+_66161871 | 0.17 |
ENST00000569896.1 |
RAB11A |
RAB11A, member RAS oncogene family |
| chr1_+_181057638 | 0.16 |
ENST00000367577.4 |
IER5 |
immediate early response 5 |
| chr1_-_115300592 | 0.15 |
ENST00000261443.5 ENST00000534699.1 ENST00000339438.6 ENST00000529046.1 ENST00000525970.1 ENST00000369530.1 ENST00000530886.1 |
CSDE1 |
cold shock domain containing E1, RNA-binding |
| chr14_+_96968802 | 0.15 |
ENST00000556619.1 ENST00000392990.2 |
PAPOLA |
poly(A) polymerase alpha |
| chr2_+_10560147 | 0.15 |
ENST00000422133.1 |
HPCAL1 |
hippocalcin-like 1 |
| chr1_+_151512775 | 0.15 |
ENST00000368849.3 ENST00000392712.3 ENST00000353024.3 ENST00000368848.2 ENST00000538902.1 |
TUFT1 |
tuftelin 1 |
| chr10_+_103113802 | 0.14 |
ENST00000370187.3 |
BTRC |
beta-transducin repeat containing E3 ubiquitin protein ligase |
| chr17_+_40913264 | 0.14 |
ENST00000587142.1 ENST00000588576.1 |
RAMP2 |
receptor (G protein-coupled) activity modifying protein 2 |
| chr17_-_64188177 | 0.14 |
ENST00000535342.2 |
CEP112 |
centrosomal protein 112kDa |
| chr16_-_75498553 | 0.14 |
ENST00000569276.1 ENST00000357613.4 ENST00000561878.1 ENST00000566980.1 ENST00000567194.1 |
TMEM170A RP11-77K12.1 |
transmembrane protein 170A Uncharacterized protein |
| chr10_-_125851961 | 0.14 |
ENST00000346248.5 |
CHST15 |
carbohydrate (N-acetylgalactosamine 4-sulfate 6-O) sulfotransferase 15 |
| chr8_+_141521386 | 0.13 |
ENST00000220913.5 ENST00000519533.1 |
CHRAC1 |
chromatin accessibility complex 1 |
| chr19_-_49496557 | 0.13 |
ENST00000323798.3 ENST00000541188.1 ENST00000544287.1 ENST00000540532.1 ENST00000263276.6 |
GYS1 |
glycogen synthase 1 (muscle) |
| chr20_-_36156293 | 0.13 |
ENST00000373537.2 ENST00000414542.2 |
BLCAP |
bladder cancer associated protein |
| chr2_-_28113217 | 0.13 |
ENST00000444339.2 |
RBKS |
ribokinase |
| chr15_-_52472078 | 0.13 |
ENST00000396335.4 ENST00000560116.1 ENST00000358784.7 |
GNB5 |
guanine nucleotide binding protein (G protein), beta 5 |
| chr10_+_103113840 | 0.12 |
ENST00000393441.4 ENST00000408038.2 |
BTRC |
beta-transducin repeat containing E3 ubiquitin protein ligase |
| chr12_-_102224704 | 0.12 |
ENST00000299314.7 |
GNPTAB |
N-acetylglucosamine-1-phosphate transferase, alpha and beta subunits |
| chr12_+_51632508 | 0.12 |
ENST00000449723.3 |
DAZAP2 |
DAZ associated protein 2 |
| chrX_-_99891796 | 0.12 |
ENST00000373020.4 |
TSPAN6 |
tetraspanin 6 |
| chr16_-_75498308 | 0.12 |
ENST00000569540.1 |
TMEM170A |
transmembrane protein 170A |
| chr3_-_58572760 | 0.11 |
ENST00000447756.2 |
FAM107A |
family with sequence similarity 107, member A |
| chr16_+_31044812 | 0.11 |
ENST00000313843.3 |
STX4 |
syntaxin 4 |
| chr6_+_30457244 | 0.11 |
ENST00000376630.4 |
HLA-E |
major histocompatibility complex, class I, E |
| chr1_+_82266053 | 0.11 |
ENST00000370715.1 ENST00000370713.1 ENST00000319517.6 ENST00000370717.2 ENST00000394879.1 ENST00000271029.4 ENST00000335786.5 |
LPHN2 |
latrophilin 2 |
| chr11_-_6704513 | 0.11 |
ENST00000532203.1 ENST00000288937.6 |
MRPL17 |
mitochondrial ribosomal protein L17 |
| chr6_+_30035307 | 0.11 |
ENST00000376765.2 ENST00000376763.1 |
PPP1R11 |
protein phosphatase 1, regulatory (inhibitor) subunit 11 |
| chr18_+_48405419 | 0.10 |
ENST00000321341.5 |
ME2 |
malic enzyme 2, NAD(+)-dependent, mitochondrial |
| chr11_-_108464321 | 0.10 |
ENST00000265843.4 |
EXPH5 |
exophilin 5 |
| chr5_+_154237778 | 0.10 |
ENST00000523698.1 ENST00000517876.1 ENST00000520472.1 |
CNOT8 |
CCR4-NOT transcription complex, subunit 8 |
| chr11_-_67275542 | 0.10 |
ENST00000531506.1 |
CDK2AP2 |
cyclin-dependent kinase 2 associated protein 2 |
| chr14_+_96968707 | 0.10 |
ENST00000216277.8 ENST00000557320.1 ENST00000557471.1 |
PAPOLA |
poly(A) polymerase alpha |
| chr14_+_54863667 | 0.10 |
ENST00000335183.6 |
CDKN3 |
cyclin-dependent kinase inhibitor 3 |
| chr11_+_125774258 | 0.10 |
ENST00000263576.6 |
DDX25 |
DEAD (Asp-Glu-Ala-Asp) box helicase 25 |
| chrX_-_106959631 | 0.10 |
ENST00000486554.1 ENST00000372390.4 |
TSC22D3 |
TSC22 domain family, member 3 |
| chr14_+_54863739 | 0.10 |
ENST00000541304.1 |
CDKN3 |
cyclin-dependent kinase inhibitor 3 |
| chr3_+_137906154 | 0.10 |
ENST00000466749.1 ENST00000358441.2 ENST00000489213.1 |
ARMC8 |
armadillo repeat containing 8 |
| chr18_+_48405463 | 0.09 |
ENST00000382927.3 |
ME2 |
malic enzyme 2, NAD(+)-dependent, mitochondrial |
| chr5_-_93447333 | 0.09 |
ENST00000395965.3 ENST00000505869.1 ENST00000509163.1 |
FAM172A |
family with sequence similarity 172, member A |
| chr14_+_93260642 | 0.09 |
ENST00000355976.2 |
GOLGA5 |
golgin A5 |
| chr13_-_88323218 | 0.09 |
ENST00000436290.2 ENST00000453832.2 ENST00000606590.1 |
MIR4500HG |
MIR4500 host gene (non-protein coding) |
| chr17_+_40913210 | 0.09 |
ENST00000253796.5 |
RAMP2 |
receptor (G protein-coupled) activity modifying protein 2 |
| chr7_+_100271446 | 0.09 |
ENST00000419828.1 ENST00000427895.1 |
GNB2 |
guanine nucleotide binding protein (G protein), beta polypeptide 2 |
| chr14_+_54863682 | 0.09 |
ENST00000543789.2 ENST00000442975.2 ENST00000458126.2 ENST00000556102.2 |
CDKN3 |
cyclin-dependent kinase inhibitor 3 |
| chr22_+_20104947 | 0.09 |
ENST00000402752.1 |
RANBP1 |
RAN binding protein 1 |
| chr4_+_177241094 | 0.09 |
ENST00000503362.1 |
SPCS3 |
signal peptidase complex subunit 3 homolog (S. cerevisiae) |
| chr11_+_125774362 | 0.09 |
ENST00000530414.1 ENST00000530129.2 |
DDX25 |
DEAD (Asp-Glu-Ala-Asp) box helicase 25 |
| chr6_+_30034966 | 0.09 |
ENST00000376769.2 |
PPP1R11 |
protein phosphatase 1, regulatory (inhibitor) subunit 11 |
| chr14_+_77787227 | 0.09 |
ENST00000216465.5 ENST00000361389.4 ENST00000554279.1 ENST00000557639.1 ENST00000349555.3 ENST00000556627.1 ENST00000557053.1 |
GSTZ1 |
glutathione S-transferase zeta 1 |
| chr16_+_31044413 | 0.09 |
ENST00000394998.1 |
STX4 |
syntaxin 4 |
| chr15_-_83736091 | 0.08 |
ENST00000261721.4 |
BTBD1 |
BTB (POZ) domain containing 1 |
| chr4_+_128651530 | 0.08 |
ENST00000281154.4 |
SLC25A31 |
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 31 |
| chr5_+_61602055 | 0.08 |
ENST00000381103.2 |
KIF2A |
kinesin heavy chain member 2A |
| chr5_+_122847781 | 0.08 |
ENST00000395412.1 ENST00000395411.1 ENST00000345990.4 |
CSNK1G3 |
casein kinase 1, gamma 3 |
| chr6_+_30034865 | 0.08 |
ENST00000376772.3 |
PPP1R11 |
protein phosphatase 1, regulatory (inhibitor) subunit 11 |
| chr14_+_93260569 | 0.08 |
ENST00000163416.2 |
GOLGA5 |
golgin A5 |
| chr2_+_46769798 | 0.08 |
ENST00000238738.4 |
RHOQ |
ras homolog family member Q |
| chrX_+_68048803 | 0.08 |
ENST00000204961.4 |
EFNB1 |
ephrin-B1 |
| chr16_-_57831676 | 0.08 |
ENST00000465878.2 ENST00000539578.1 ENST00000561524.1 |
KIFC3 |
kinesin family member C3 |
| chr14_-_96830207 | 0.08 |
ENST00000359933.4 |
ATG2B |
autophagy related 2B |
| chr19_+_42772659 | 0.07 |
ENST00000572681.2 |
CIC |
capicua transcriptional repressor |
| chr17_+_40912764 | 0.07 |
ENST00000589683.1 ENST00000588928.1 |
RAMP2 |
receptor (G protein-coupled) activity modifying protein 2 |
| chr17_+_7387677 | 0.07 |
ENST00000322644.6 |
POLR2A |
polymerase (RNA) II (DNA directed) polypeptide A, 220kDa |
| chr12_-_102224457 | 0.07 |
ENST00000549165.1 ENST00000549940.1 ENST00000392919.4 |
GNPTAB |
N-acetylglucosamine-1-phosphate transferase, alpha and beta subunits |
| chr1_-_28241226 | 0.07 |
ENST00000373912.3 ENST00000373909.3 |
RPA2 |
replication protein A2, 32kDa |
| chr7_+_100271355 | 0.07 |
ENST00000436220.1 ENST00000424361.1 |
GNB2 |
guanine nucleotide binding protein (G protein), beta polypeptide 2 |
| chr1_+_172502336 | 0.06 |
ENST00000263688.3 |
SUCO |
SUN domain containing ossification factor |
| chr3_+_50654550 | 0.06 |
ENST00000430409.1 ENST00000357955.2 |
MAPKAPK3 |
mitogen-activated protein kinase-activated protein kinase 3 |
| chr20_+_3801162 | 0.06 |
ENST00000379573.2 ENST00000379567.2 ENST00000455742.1 ENST00000246041.2 |
AP5S1 |
adaptor-related protein complex 5, sigma 1 subunit |
| chr5_+_154238096 | 0.06 |
ENST00000517568.1 ENST00000524105.1 ENST00000285896.6 |
CNOT8 |
CCR4-NOT transcription complex, subunit 8 |
| chr11_-_65325430 | 0.06 |
ENST00000322147.4 |
LTBP3 |
latent transforming growth factor beta binding protein 3 |
| chr19_-_16770915 | 0.06 |
ENST00000593459.1 ENST00000358726.6 ENST00000597711.1 ENST00000487416.2 |
CTC-429P9.4 SMIM7 |
Small integral membrane protein 7; Uncharacterized protein small integral membrane protein 7 |
| chr19_-_46296011 | 0.06 |
ENST00000377735.3 ENST00000270223.6 |
DMWD |
dystrophia myotonica, WD repeat containing |
| chr13_+_73356197 | 0.06 |
ENST00000326291.6 |
PIBF1 |
progesterone immunomodulatory binding factor 1 |
| chr8_-_74884341 | 0.06 |
ENST00000284811.8 |
TCEB1 |
transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) |
| chr8_-_74884459 | 0.06 |
ENST00000522337.1 |
TCEB1 |
transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) |
| chr17_-_43045439 | 0.06 |
ENST00000253407.3 |
C1QL1 |
complement component 1, q subcomponent-like 1 |
| chr8_-_74884482 | 0.06 |
ENST00000520242.1 ENST00000519082.1 |
TCEB1 |
transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) |
| chr17_-_42277203 | 0.06 |
ENST00000587097.1 |
ATXN7L3 |
ataxin 7-like 3 |
| chr19_-_11039261 | 0.06 |
ENST00000590329.1 ENST00000587943.1 ENST00000585858.1 ENST00000586748.1 ENST00000586575.1 ENST00000253031.2 |
YIPF2 |
Yip1 domain family, member 2 |
| chr1_-_114302086 | 0.06 |
ENST00000369604.1 ENST00000357783.2 |
PHTF1 |
putative homeodomain transcription factor 1 |
| chr11_+_117014983 | 0.06 |
ENST00000527958.1 ENST00000419197.2 ENST00000304808.6 ENST00000529887.2 |
PAFAH1B2 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 2 (30kDa) |
| chr17_-_79479789 | 0.06 |
ENST00000571691.1 ENST00000571721.1 ENST00000573283.1 ENST00000575842.1 ENST00000575087.1 ENST00000570382.1 ENST00000331925.2 |
ACTG1 |
actin, gamma 1 |
| chr5_+_154238042 | 0.06 |
ENST00000519211.1 ENST00000522458.1 ENST00000519903.1 ENST00000521450.1 ENST00000403027.2 |
CNOT8 |
CCR4-NOT transcription complex, subunit 8 |
| chr1_+_171283331 | 0.06 |
ENST00000367749.3 |
FMO4 |
flavin containing monooxygenase 4 |
| chr8_-_74884511 | 0.06 |
ENST00000518127.1 |
TCEB1 |
transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) |
| chr14_-_77787198 | 0.05 |
ENST00000261534.4 |
POMT2 |
protein-O-mannosyltransferase 2 |
| chr10_+_104180580 | 0.05 |
ENST00000425536.1 |
FBXL15 |
F-box and leucine-rich repeat protein 15 |
| chrX_+_16804544 | 0.05 |
ENST00000380122.5 ENST00000398155.4 |
TXLNG |
taxilin gamma |
| chr3_+_137906353 | 0.05 |
ENST00000461822.1 ENST00000485396.1 ENST00000471453.1 ENST00000470821.1 ENST00000471709.1 ENST00000538260.1 ENST00000393058.3 ENST00000463485.1 |
ARMC8 |
armadillo repeat containing 8 |
| chr12_+_32260085 | 0.05 |
ENST00000548411.1 ENST00000281474.5 ENST00000551086.1 |
BICD1 |
bicaudal D homolog 1 (Drosophila) |
| chr1_-_8939265 | 0.05 |
ENST00000489867.1 |
ENO1 |
enolase 1, (alpha) |
| chr8_-_74884399 | 0.05 |
ENST00000520210.1 ENST00000602840.1 |
TCEB1 |
transcription elongation factor B (SIII), polypeptide 1 (15kDa, elongin C) |
| chr1_-_114301960 | 0.05 |
ENST00000369598.1 ENST00000369600.1 |
PHTF1 |
putative homeodomain transcription factor 1 |
| chr12_+_120933859 | 0.05 |
ENST00000242577.6 ENST00000548214.1 ENST00000392508.2 |
DYNLL1 |
dynein, light chain, LC8-type 1 |
| chr1_-_114301755 | 0.05 |
ENST00000393357.2 ENST00000369596.2 ENST00000446739.1 |
PHTF1 |
putative homeodomain transcription factor 1 |
| chr20_-_57607347 | 0.05 |
ENST00000395663.1 ENST00000395659.1 ENST00000243997.3 |
ATP5E |
ATP synthase, H+ transporting, mitochondrial F1 complex, epsilon subunit |
| chr15_+_90777424 | 0.05 |
ENST00000561433.1 ENST00000559204.1 ENST00000558291.1 |
GDPGP1 |
GDP-D-glucose phosphorylase 1 |
| chr10_-_104179682 | 0.05 |
ENST00000406432.1 |
PSD |
pleckstrin and Sec7 domain containing |
| chr17_+_33914460 | 0.05 |
ENST00000537622.2 |
AP2B1 |
adaptor-related protein complex 2, beta 1 subunit |
| chr10_+_14880157 | 0.05 |
ENST00000378372.3 |
HSPA14 |
heat shock 70kDa protein 14 |
| chr3_-_27525865 | 0.05 |
ENST00000445684.1 |
SLC4A7 |
solute carrier family 4, sodium bicarbonate cotransporter, member 7 |
| chr19_+_48867652 | 0.05 |
ENST00000344846.2 |
SYNGR4 |
synaptogyrin 4 |
| chr19_+_39109735 | 0.05 |
ENST00000593149.1 ENST00000248342.4 ENST00000538434.1 ENST00000588934.1 ENST00000545173.2 ENST00000589307.1 ENST00000586513.1 ENST00000591409.1 ENST00000592558.1 |
EIF3K |
eukaryotic translation initiation factor 3, subunit K |
| chr5_+_140019079 | 0.05 |
ENST00000252100.6 |
TMCO6 |
transmembrane and coiled-coil domains 6 |
| chr2_-_20251744 | 0.05 |
ENST00000175091.4 |
LAPTM4A |
lysosomal protein transmembrane 4 alpha |
| chr13_-_95364389 | 0.05 |
ENST00000376945.2 |
SOX21 |
SRY (sex determining region Y)-box 21 |
| chr4_-_10023095 | 0.05 |
ENST00000264784.3 |
SLC2A9 |
solute carrier family 2 (facilitated glucose transporter), member 9 |
| chr19_-_42806919 | 0.04 |
ENST00000595530.1 ENST00000538771.1 ENST00000601865.1 |
PAFAH1B3 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr5_-_131892501 | 0.04 |
ENST00000450655.1 |
IL5 |
interleukin 5 (colony-stimulating factor, eosinophil) |
| chr4_+_110354928 | 0.04 |
ENST00000504968.2 ENST00000399100.2 ENST00000265175.5 |
SEC24B |
SEC24 family member B |
| chr18_+_3448455 | 0.04 |
ENST00000549780.1 |
TGIF1 |
TGFB-induced factor homeobox 1 |
| chr12_+_70760056 | 0.04 |
ENST00000258111.4 |
KCNMB4 |
potassium large conductance calcium-activated channel, subfamily M, beta member 4 |
| chr9_+_34458771 | 0.04 |
ENST00000437363.1 ENST00000242317.4 |
DNAI1 |
dynein, axonemal, intermediate chain 1 |
| chr5_+_140019004 | 0.04 |
ENST00000394671.3 ENST00000511410.1 ENST00000537378.1 |
TMCO6 |
transmembrane and coiled-coil domains 6 |
| chr20_-_58515344 | 0.04 |
ENST00000370996.3 |
PPP1R3D |
protein phosphatase 1, regulatory subunit 3D |
| chr2_+_28113583 | 0.04 |
ENST00000344773.2 ENST00000379624.1 ENST00000342045.2 ENST00000379632.2 ENST00000361704.2 |
BRE |
brain and reproductive organ-expressed (TNFRSF1A modulator) |
| chr15_-_40213080 | 0.04 |
ENST00000561100.1 |
GPR176 |
G protein-coupled receptor 176 |
| chr1_-_78444738 | 0.04 |
ENST00000436586.2 ENST00000370768.2 |
FUBP1 |
far upstream element (FUSE) binding protein 1 |
| chr5_-_150138061 | 0.04 |
ENST00000521533.1 ENST00000424236.1 |
DCTN4 |
dynactin 4 (p62) |
| chr3_+_14989186 | 0.04 |
ENST00000435454.1 ENST00000323373.6 |
NR2C2 |
nuclear receptor subfamily 2, group C, member 2 |
| chr16_-_28937027 | 0.04 |
ENST00000358201.4 |
RABEP2 |
rabaptin, RAB GTPase binding effector protein 2 |
| chr21_+_17102311 | 0.04 |
ENST00000285679.6 ENST00000351097.5 ENST00000285681.2 ENST00000400183.2 |
USP25 |
ubiquitin specific peptidase 25 |
| chr19_+_12902289 | 0.04 |
ENST00000302754.4 |
JUNB |
jun B proto-oncogene |
| chr1_+_153606513 | 0.04 |
ENST00000368694.3 |
CHTOP |
chromatin target of PRMT1 |
| chr12_+_120933904 | 0.04 |
ENST00000550178.1 ENST00000550845.1 ENST00000549989.1 ENST00000552870.1 |
DYNLL1 |
dynein, light chain, LC8-type 1 |
| chr1_-_28241024 | 0.04 |
ENST00000313433.7 ENST00000444045.1 |
RPA2 |
replication protein A2, 32kDa |
| chr5_-_100238956 | 0.04 |
ENST00000231461.5 |
ST8SIA4 |
ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 4 |
| chrY_+_22737678 | 0.04 |
ENST00000382772.3 |
EIF1AY |
eukaryotic translation initiation factor 1A, Y-linked |
| chr15_+_99791567 | 0.03 |
ENST00000558879.1 ENST00000301981.3 ENST00000422500.2 ENST00000447360.2 ENST00000442993.2 |
LRRC28 |
leucine rich repeat containing 28 |
| chr12_+_50451331 | 0.03 |
ENST00000228468.4 |
ASIC1 |
acid-sensing (proton-gated) ion channel 1 |
| chr5_-_150138551 | 0.03 |
ENST00000446090.2 ENST00000447998.2 |
DCTN4 |
dynactin 4 (p62) |
| chr1_-_114301503 | 0.03 |
ENST00000447664.2 |
PHTF1 |
putative homeodomain transcription factor 1 |
| chr19_-_42806723 | 0.03 |
ENST00000262890.3 |
PAFAH1B3 |
platelet-activating factor acetylhydrolase 1b, catalytic subunit 3 (29kDa) |
| chr1_+_38478432 | 0.03 |
ENST00000537711.1 |
UTP11L |
UTP11-like, U3 small nucleolar ribonucleoprotein, (yeast) |
| chr15_+_49447947 | 0.03 |
ENST00000327171.3 ENST00000560654.1 |
GALK2 |
galactokinase 2 |
| chr10_+_18629628 | 0.03 |
ENST00000377329.4 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr20_-_25062767 | 0.03 |
ENST00000429762.3 ENST00000444511.2 ENST00000376707.3 |
VSX1 |
visual system homeobox 1 |
| chr4_-_103746924 | 0.03 |
ENST00000505207.1 ENST00000502404.1 ENST00000507845.1 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chr13_+_27131798 | 0.03 |
ENST00000361042.4 |
WASF3 |
WAS protein family, member 3 |
| chr8_-_100905363 | 0.03 |
ENST00000524245.1 |
COX6C |
cytochrome c oxidase subunit VIc |
| chr12_+_3600356 | 0.03 |
ENST00000382622.3 |
PRMT8 |
protein arginine methyltransferase 8 |
| chr13_-_73356009 | 0.03 |
ENST00000377780.4 ENST00000377767.4 |
DIS3 |
DIS3 mitotic control homolog (S. cerevisiae) |
| chr15_-_49447771 | 0.03 |
ENST00000558843.1 ENST00000542928.1 ENST00000561248.1 |
COPS2 |
COP9 signalosome subunit 2 |
| chr17_+_36508111 | 0.03 |
ENST00000331159.5 ENST00000577233.1 |
SOCS7 |
suppressor of cytokine signaling 7 |
| chr5_+_76506706 | 0.02 |
ENST00000340978.3 ENST00000346042.3 ENST00000264917.5 ENST00000342343.4 ENST00000333194.4 |
PDE8B |
phosphodiesterase 8B |
| chr8_-_100905850 | 0.02 |
ENST00000520271.1 ENST00000522940.1 ENST00000523016.1 ENST00000517682.2 ENST00000297564.2 |
COX6C |
cytochrome c oxidase subunit VIc |
| chr14_-_92588013 | 0.02 |
ENST00000553514.1 ENST00000605997.1 |
NDUFB1 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 1, 7kDa |
| chr3_+_53528659 | 0.02 |
ENST00000350061.5 |
CACNA1D |
calcium channel, voltage-dependent, L type, alpha 1D subunit |
| chr19_-_58662139 | 0.02 |
ENST00000598312.1 |
ZNF329 |
zinc finger protein 329 |
| chr15_-_55700457 | 0.02 |
ENST00000442196.3 ENST00000563171.1 ENST00000425574.3 |
CCPG1 |
cell cycle progression 1 |
| chrX_-_83442915 | 0.02 |
ENST00000262752.2 ENST00000543399.1 |
RPS6KA6 |
ribosomal protein S6 kinase, 90kDa, polypeptide 6 |
| chr3_-_62358690 | 0.02 |
ENST00000475839.1 |
FEZF2 |
FEZ family zinc finger 2 |
| chr13_+_27131887 | 0.02 |
ENST00000335327.5 |
WASF3 |
WAS protein family, member 3 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.0 | 0.2 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.0 | 0.3 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.0 | 0.2 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.3 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.0 | 0.4 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.5 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.0 | 0.4 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.3 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 0.3 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0086062 | voltage-gated sodium channel activity involved in Purkinje myocyte action potential(GO:0086062) |
| 0.1 | 0.6 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.1 | 0.3 | GO:0050473 | arachidonate 15-lipoxygenase activity(GO:0050473) |
| 0.1 | 0.3 | GO:0015319 | sodium:inorganic phosphate symporter activity(GO:0015319) |
| 0.1 | 0.2 | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity(GO:0003976) |
| 0.1 | 0.3 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.0 | 0.2 | GO:0016230 | sphingomyelin phosphodiesterase activator activity(GO:0016230) |
| 0.0 | 0.3 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.0 | 0.2 | GO:0017089 | glycolipid transporter activity(GO:0017089) |
| 0.0 | 0.9 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 0.3 | GO:0097643 | amylin receptor activity(GO:0097643) |
| 0.0 | 0.2 | GO:1990599 | 3' overhang single-stranded DNA endodeoxyribonuclease activity(GO:1990599) |
| 0.0 | 0.3 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.0 | 0.2 | GO:0004473 | malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) oxaloacetate decarboxylase activity(GO:0008948) |
| 0.0 | 0.2 | GO:0047179 | platelet-activating factor acetyltransferase activity(GO:0047179) |
| 0.0 | 0.4 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 0.0 | 0.1 | GO:0032427 | GBD domain binding(GO:0032427) |
| 0.0 | 0.2 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.0 | 0.7 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.0 | 0.2 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 0.0 | 0.1 | GO:0050656 | 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 0.0 | 0.5 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.1 | GO:0030021 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.0 | 0.5 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.0 | 0.1 | GO:0003968 | RNA-directed RNA polymerase activity(GO:0003968) |
| 0.0 | 0.3 | GO:0030881 | beta-2-microglobulin binding(GO:0030881) |
| 0.0 | 0.2 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.3 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.0 | GO:0005137 | interleukin-5 receptor binding(GO:0005137) |
| 0.0 | 0.2 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.0 | 0.0 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 0.0 | 0.1 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.0 | 0.2 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.0 | 0.2 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.1 | GO:0035251 | UDP-glucosyltransferase activity(GO:0035251) |
| 0.0 | 0.1 | GO:0043426 | MRF binding(GO:0043426) |
| 0.0 | 0.1 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.3 | GO:0043548 | phosphatidylinositol 3-kinase binding(GO:0043548) |
| 0.0 | 0.5 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.0 | GO:0033858 | N-acetylgalactosamine kinase activity(GO:0033858) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 0.1 | 0.7 | GO:0097413 | Lewy body(GO:0097413) |
| 0.1 | 0.3 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 0.0 | 0.3 | GO:1903440 | calcitonin family receptor complex(GO:1903439) amylin receptor complex(GO:1903440) |
| 0.0 | 0.2 | GO:0070522 | nucleotide-excision repair factor 1 complex(GO:0000110) ERCC4-ERCC1 complex(GO:0070522) |
| 0.0 | 0.4 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.0 | 0.3 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.0 | 0.1 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.0 | 0.2 | GO:0000322 | storage vacuole(GO:0000322) |
| 0.0 | 0.2 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.0 | 0.4 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.0 | 1.2 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.2 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 0.3 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.6 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.1 | GO:0033648 | host intracellular organelle(GO:0033647) host intracellular membrane-bounded organelle(GO:0033648) |
| 0.0 | 0.1 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.0 | 0.1 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 0.0 | GO:0042587 | glycogen granule(GO:0042587) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0010796 | regulation of multivesicular body size(GO:0010796) |
| 0.1 | 0.6 | GO:0009609 | response to symbiont(GO:0009608) response to symbiotic bacterium(GO:0009609) |
| 0.1 | 1.0 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.1 | 0.4 | GO:0042335 | cuticle development(GO:0042335) |
| 0.1 | 0.6 | GO:0035166 | post-embryonic hemopoiesis(GO:0035166) |
| 0.1 | 0.2 | GO:0019417 | sulfur oxidation(GO:0019417) |
| 0.1 | 0.4 | GO:0060373 | regulation of ventricular cardiac muscle cell membrane depolarization(GO:0060373) |
| 0.1 | 0.2 | GO:0043311 | positive regulation of eosinophil degranulation(GO:0043311) positive regulation of eosinophil activation(GO:1902568) |
| 0.1 | 0.3 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 0.1 | 0.3 | GO:0034036 | purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.1 | 0.3 | GO:0010746 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) |
| 0.1 | 0.3 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.1 | 0.3 | GO:0051121 | hepoxilin metabolic process(GO:0051121) hepoxilin biosynthetic process(GO:0051122) |
| 0.0 | 0.3 | GO:1903593 | regulation of histamine secretion by mast cell(GO:1903593) |
| 0.0 | 0.3 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.0 | 0.1 | GO:0032679 | antigen processing and presentation of endogenous peptide antigen via MHC class Ib(GO:0002476) TRAIL production(GO:0032639) regulation of TRAIL production(GO:0032679) positive regulation of TRAIL production(GO:0032759) protection from natural killer cell mediated cytotoxicity(GO:0042270) positive regulation of CD8-positive, alpha-beta T cell proliferation(GO:2000566) |
| 0.0 | 0.3 | GO:0016199 | axon midline choice point recognition(GO:0016199) |
| 0.0 | 0.2 | GO:0021840 | directional guidance of interneurons involved in migration from the subpallium to the cortex(GO:0021840) chemorepulsion involved in interneuron migration from the subpallium to the cortex(GO:0021842) |
| 0.0 | 0.3 | GO:0097646 | calcitonin family receptor signaling pathway(GO:0097646) amylin receptor signaling pathway(GO:0097647) |
| 0.0 | 0.1 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.0 | 0.1 | GO:0006014 | D-ribose metabolic process(GO:0006014) |
| 0.0 | 0.3 | GO:0044341 | sodium-dependent phosphate transport(GO:0044341) |
| 0.0 | 0.2 | GO:1902031 | regulation of NADP metabolic process(GO:1902031) |
| 0.0 | 0.2 | GO:0046836 | glycolipid transport(GO:0046836) |
| 0.0 | 0.2 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 0.0 | 0.1 | GO:2000035 | regulation of stem cell division(GO:2000035) |
| 0.0 | 0.2 | GO:0033299 | secretion of lysosomal enzymes(GO:0033299) |
| 0.0 | 0.1 | GO:0001172 | transcription, RNA-templated(GO:0001172) |
| 0.0 | 0.5 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 0.0 | GO:0051083 | 'de novo' cotranslational protein folding(GO:0051083) |
| 0.0 | 0.2 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.0 | 0.1 | GO:1902460 | regulation of mesenchymal stem cell proliferation(GO:1902460) positive regulation of mesenchymal stem cell proliferation(GO:1902462) |
| 0.0 | 0.1 | GO:0032815 | negative regulation of natural killer cell activation(GO:0032815) |
| 0.0 | 0.0 | GO:0045643 | regulation of eosinophil differentiation(GO:0045643) positive regulation of eosinophil differentiation(GO:0045645) |
| 0.0 | 0.0 | GO:1901301 | regulation of cargo loading into COPII-coated vesicle(GO:1901301) |
| 0.0 | 0.1 | GO:0061743 | motor learning(GO:0061743) |
| 0.0 | 0.6 | GO:0045104 | intermediate filament cytoskeleton organization(GO:0045104) |
| 0.0 | 0.2 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.0 | 0.1 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.0 | 0.3 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.0 | 0.0 | GO:0099590 | neurotransmitter receptor internalization(GO:0099590) |
| 0.0 | 0.1 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.0 | 0.1 | GO:0070235 | regulation of activation-induced cell death of T cells(GO:0070235) negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.0 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.0 | 0.4 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 0.0 | 0.1 | GO:1904100 | regulation of protein O-linked glycosylation(GO:1904098) positive regulation of protein O-linked glycosylation(GO:1904100) |
| 0.0 | 0.3 | GO:0042753 | positive regulation of circadian rhythm(GO:0042753) |
| 0.0 | 0.6 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.1 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.0 | 0.1 | GO:0072385 | minus-end-directed organelle transport along microtubule(GO:0072385) |