Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
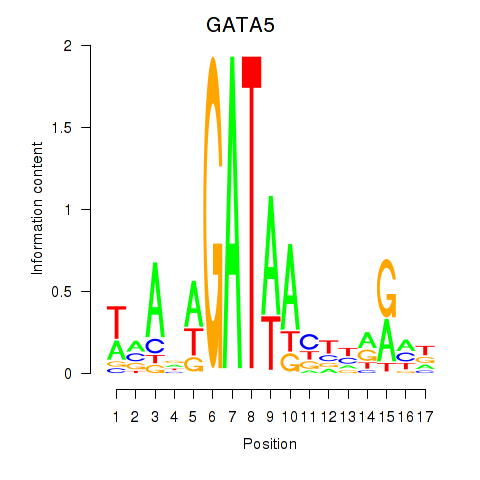
Results for GATA5
Z-value: 1.18
Transcription factors associated with GATA5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
GATA5
|
ENSG00000130700.6 | GATA5 |
Activity profile of GATA5 motif
Sorted Z-values of GATA5 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of GATA5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_160681593 | 3.23 |
ENST00000368045.3 ENST00000368046.3 |
CD48 |
CD48 molecule |
| chr13_-_46756351 | 3.13 |
ENST00000323076.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chr9_-_6645628 | 3.13 |
ENST00000321612.6 |
GLDC |
glycine dehydrogenase (decarboxylating) |
| chr4_-_110723335 | 2.69 |
ENST00000394634.2 |
CFI |
complement factor I |
| chr7_+_26332645 | 2.63 |
ENST00000396376.1 |
SNX10 |
sorting nexin 10 |
| chr4_-_110723134 | 2.52 |
ENST00000510800.1 ENST00000512148.1 |
CFI |
complement factor I |
| chr4_-_110723194 | 2.52 |
ENST00000394635.3 |
CFI |
complement factor I |
| chr6_-_32557610 | 2.45 |
ENST00000360004.5 |
HLA-DRB1 |
major histocompatibility complex, class II, DR beta 1 |
| chr6_+_32709119 | 2.42 |
ENST00000374940.3 |
HLA-DQA2 |
major histocompatibility complex, class II, DQ alpha 2 |
| chr17_+_53344945 | 2.36 |
ENST00000575345.1 |
HLF |
hepatic leukemia factor |
| chr10_+_7745232 | 2.36 |
ENST00000358415.4 |
ITIH2 |
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr10_+_7745303 | 2.35 |
ENST00000429820.1 ENST00000379587.4 |
ITIH2 |
inter-alpha-trypsin inhibitor heavy chain 2 |
| chr11_+_121461097 | 2.05 |
ENST00000527934.1 |
SORL1 |
sortilin-related receptor, L(DLR class) A repeats containing |
| chr10_-_98031265 | 2.01 |
ENST00000224337.5 ENST00000371176.2 |
BLNK |
B-cell linker |
| chr2_+_143635067 | 1.71 |
ENST00000264170.4 |
KYNU |
kynureninase |
| chr19_+_35820064 | 1.70 |
ENST00000341773.6 ENST00000600131.1 ENST00000270311.6 ENST00000595780.1 ENST00000597916.1 ENST00000593867.1 ENST00000600424.1 ENST00000599811.1 ENST00000536635.2 ENST00000085219.5 ENST00000544992.2 ENST00000419549.2 |
CD22 |
CD22 molecule |
| chr4_+_155484155 | 1.60 |
ENST00000509493.1 |
FGB |
fibrinogen beta chain |
| chr22_-_29107919 | 1.47 |
ENST00000434810.1 ENST00000456369.1 |
CHEK2 |
checkpoint kinase 2 |
| chr8_+_28196157 | 1.47 |
ENST00000522209.1 |
PNOC |
prepronociceptin |
| chr6_+_33043703 | 1.43 |
ENST00000418931.2 ENST00000535465.1 |
HLA-DPB1 |
major histocompatibility complex, class II, DP beta 1 |
| chr9_-_35650900 | 1.41 |
ENST00000259608.3 |
SIT1 |
signaling threshold regulating transmembrane adaptor 1 |
| chr3_+_178276488 | 1.33 |
ENST00000432997.1 ENST00000455865.1 |
KCNMB2 |
potassium large conductance calcium-activated channel, subfamily M, beta member 2 |
| chr1_+_174844645 | 1.25 |
ENST00000486220.1 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr15_-_20193370 | 1.17 |
ENST00000558565.2 |
IGHV3OR15-7 |
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr12_-_9268707 | 1.16 |
ENST00000318602.7 |
A2M |
alpha-2-macroglobulin |
| chr6_+_31916733 | 1.15 |
ENST00000483004.1 |
CFB |
complement factor B |
| chr8_+_86099884 | 1.12 |
ENST00000517476.1 ENST00000521429.1 |
E2F5 |
E2F transcription factor 5, p130-binding |
| chr19_-_10450328 | 1.08 |
ENST00000160262.5 |
ICAM3 |
intercellular adhesion molecule 3 |
| chr8_+_40010989 | 1.04 |
ENST00000315792.3 |
C8orf4 |
chromosome 8 open reading frame 4 |
| chr6_-_154677900 | 1.03 |
ENST00000265198.4 ENST00000520261.1 |
IPCEF1 |
interaction protein for cytohesin exchange factors 1 |
| chr11_+_128563652 | 1.01 |
ENST00000527786.2 |
FLI1 |
Fli-1 proto-oncogene, ETS transcription factor |
| chr4_+_155484103 | 1.00 |
ENST00000302068.4 |
FGB |
fibrinogen beta chain |
| chr1_+_207277590 | 0.99 |
ENST00000367070.3 |
C4BPA |
complement component 4 binding protein, alpha |
| chr12_+_54892550 | 0.95 |
ENST00000545638.2 |
NCKAP1L |
NCK-associated protein 1-like |
| chr2_+_102615416 | 0.90 |
ENST00000393414.2 |
IL1R2 |
interleukin 1 receptor, type II |
| chr6_-_32908765 | 0.84 |
ENST00000416244.2 |
HLA-DMB |
major histocompatibility complex, class II, DM beta |
| chr3_-_197300194 | 0.77 |
ENST00000358186.2 ENST00000431056.1 |
BDH1 |
3-hydroxybutyrate dehydrogenase, type 1 |
| chr11_-_33891362 | 0.76 |
ENST00000395833.3 |
LMO2 |
LIM domain only 2 (rhombotin-like 1) |
| chr2_+_69201705 | 0.76 |
ENST00000377938.2 |
GKN1 |
gastrokine 1 |
| chrX_+_30261847 | 0.74 |
ENST00000378981.3 ENST00000397550.1 |
MAGEB1 |
melanoma antigen family B, 1 |
| chr8_+_117778736 | 0.70 |
ENST00000309822.2 ENST00000357148.3 ENST00000517814.1 ENST00000517820.1 |
UTP23 |
UTP23, small subunit (SSU) processome component, homolog (yeast) |
| chr18_+_13611763 | 0.70 |
ENST00000585931.1 |
LDLRAD4 |
low density lipoprotein receptor class A domain containing 4 |
| chr14_+_39735411 | 0.69 |
ENST00000603904.1 |
RP11-407N17.3 |
cTAGE family member 5 isoform 4 |
| chr1_+_156123318 | 0.68 |
ENST00000368285.3 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr1_+_57320437 | 0.67 |
ENST00000361249.3 |
C8A |
complement component 8, alpha polypeptide |
| chrX_+_52235228 | 0.67 |
ENST00000518075.1 |
XAGE1B |
X antigen family, member 1B |
| chr13_-_99910673 | 0.66 |
ENST00000397473.2 ENST00000397470.2 |
GPR18 |
G protein-coupled receptor 18 |
| chr11_+_101785727 | 0.66 |
ENST00000263468.8 |
KIAA1377 |
KIAA1377 |
| chr15_-_34628951 | 0.65 |
ENST00000397707.2 ENST00000560611.1 |
SLC12A6 |
solute carrier family 12 (potassium/chloride transporter), member 6 |
| chr1_+_79115503 | 0.64 |
ENST00000370747.4 ENST00000438486.1 ENST00000545124.1 |
IFI44 |
interferon-induced protein 44 |
| chr6_-_43027105 | 0.64 |
ENST00000230413.5 ENST00000487429.1 ENST00000489623.1 ENST00000468957.1 |
MRPL2 |
mitochondrial ribosomal protein L2 |
| chr3_-_178969403 | 0.62 |
ENST00000314235.5 ENST00000392685.2 |
KCNMB3 |
potassium large conductance calcium-activated channel, subfamily M beta member 3 |
| chr7_-_123197733 | 0.62 |
ENST00000470123.1 ENST00000471770.1 |
NDUFA5 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 5 |
| chr10_+_35484053 | 0.58 |
ENST00000487763.1 ENST00000473940.1 ENST00000488328.1 ENST00000356917.5 |
CREM |
cAMP responsive element modulator |
| chr11_+_124824000 | 0.57 |
ENST00000529051.1 ENST00000344762.5 |
CCDC15 |
coiled-coil domain containing 15 |
| chr6_-_86099898 | 0.56 |
ENST00000455071.1 |
RP11-30P6.6 |
RP11-30P6.6 |
| chr2_+_201997595 | 0.53 |
ENST00000470178.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr3_+_184080387 | 0.53 |
ENST00000455712.1 |
POLR2H |
polymerase (RNA) II (DNA directed) polypeptide H |
| chr5_+_140261703 | 0.51 |
ENST00000409494.1 ENST00000289272.2 |
PCDHA13 |
protocadherin alpha 13 |
| chr3_-_182817297 | 0.50 |
ENST00000539926.1 ENST00000476176.1 |
MCCC1 |
methylcrotonoyl-CoA carboxylase 1 (alpha) |
| chr1_+_150954493 | 0.50 |
ENST00000368947.4 |
ANXA9 |
annexin A9 |
| chr1_-_11918988 | 0.50 |
ENST00000376468.3 |
NPPB |
natriuretic peptide B |
| chr6_-_53409890 | 0.49 |
ENST00000229416.6 |
GCLC |
glutamate-cysteine ligase, catalytic subunit |
| chr6_-_55739542 | 0.48 |
ENST00000446683.2 |
BMP5 |
bone morphogenetic protein 5 |
| chr2_-_191115229 | 0.46 |
ENST00000409820.2 ENST00000410045.1 |
HIBCH |
3-hydroxyisobutyryl-CoA hydrolase |
| chr2_+_201997492 | 0.45 |
ENST00000494258.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr22_+_23412479 | 0.44 |
ENST00000248996.4 |
GNAZ |
guanine nucleotide binding protein (G protein), alpha z polypeptide |
| chr1_+_10292308 | 0.43 |
ENST00000377081.1 |
KIF1B |
kinesin family member 1B |
| chr6_-_135271260 | 0.42 |
ENST00000265605.2 |
ALDH8A1 |
aldehyde dehydrogenase 8 family, member A1 |
| chr1_-_150669500 | 0.42 |
ENST00000271732.3 |
GOLPH3L |
golgi phosphoprotein 3-like |
| chr11_-_72070206 | 0.40 |
ENST00000544382.1 |
CLPB |
ClpB caseinolytic peptidase B homolog (E. coli) |
| chr3_-_137851220 | 0.37 |
ENST00000236709.3 |
A4GNT |
alpha-1,4-N-acetylglucosaminyltransferase |
| chr1_+_150229554 | 0.37 |
ENST00000369111.4 |
CA14 |
carbonic anhydrase XIV |
| chr14_+_61995722 | 0.35 |
ENST00000556347.1 |
RP11-47I22.4 |
RP11-47I22.4 |
| chr19_-_52133588 | 0.34 |
ENST00000570106.2 |
SIGLEC5 |
sialic acid binding Ig-like lectin 5 |
| chr6_-_133079022 | 0.34 |
ENST00000525289.1 ENST00000326499.6 |
VNN2 |
vanin 2 |
| chr16_+_77224732 | 0.34 |
ENST00000569610.1 ENST00000248248.3 ENST00000567291.1 ENST00000320859.6 ENST00000563612.1 ENST00000563279.1 |
MON1B |
MON1 secretory trafficking family member B |
| chr16_-_33647696 | 0.33 |
ENST00000558425.1 ENST00000569103.2 |
RP11-812E19.9 |
Uncharacterized protein |
| chr3_-_178984759 | 0.33 |
ENST00000349697.2 ENST00000497599.1 |
KCNMB3 |
potassium large conductance calcium-activated channel, subfamily M beta member 3 |
| chr1_-_158656488 | 0.32 |
ENST00000368147.4 |
SPTA1 |
spectrin, alpha, erythrocytic 1 (elliptocytosis 2) |
| chr6_-_135271219 | 0.31 |
ENST00000367847.2 ENST00000367845.2 |
ALDH8A1 |
aldehyde dehydrogenase 8 family, member A1 |
| chr4_+_110834033 | 0.30 |
ENST00000509793.1 ENST00000265171.5 |
EGF |
epidermal growth factor |
| chr6_+_118869452 | 0.29 |
ENST00000357525.5 |
PLN |
phospholamban |
| chr5_-_75919253 | 0.29 |
ENST00000296641.4 |
F2RL2 |
coagulation factor II (thrombin) receptor-like 2 |
| chr5_-_75919217 | 0.29 |
ENST00000504899.1 |
F2RL2 |
coagulation factor II (thrombin) receptor-like 2 |
| chr7_+_136553370 | 0.28 |
ENST00000445907.2 |
CHRM2 |
cholinergic receptor, muscarinic 2 |
| chr4_+_175204818 | 0.28 |
ENST00000503780.1 |
CEP44 |
centrosomal protein 44kDa |
| chr16_+_31366536 | 0.28 |
ENST00000562522.1 |
ITGAX |
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chr2_-_109605663 | 0.28 |
ENST00000409271.1 ENST00000258443.2 ENST00000376651.1 |
EDAR |
ectodysplasin A receptor |
| chr17_-_41277467 | 0.27 |
ENST00000494123.1 ENST00000346315.3 ENST00000309486.4 ENST00000468300.1 ENST00000354071.3 ENST00000352993.3 ENST00000471181.2 |
BRCA1 |
breast cancer 1, early onset |
| chr8_-_91095099 | 0.26 |
ENST00000265431.3 |
CALB1 |
calbindin 1, 28kDa |
| chr9_+_106856831 | 0.26 |
ENST00000303219.8 ENST00000374787.3 |
SMC2 |
structural maintenance of chromosomes 2 |
| chr3_+_108541545 | 0.25 |
ENST00000295756.6 |
TRAT1 |
T cell receptor associated transmembrane adaptor 1 |
| chr9_-_21166659 | 0.25 |
ENST00000380225.1 |
IFNA21 |
interferon, alpha 21 |
| chr3_+_141144954 | 0.25 |
ENST00000441582.2 ENST00000321464.5 |
ZBTB38 |
zinc finger and BTB domain containing 38 |
| chr1_-_182921119 | 0.25 |
ENST00000423786.1 |
SHCBP1L |
SHC SH2-domain binding protein 1-like |
| chr10_+_15085936 | 0.25 |
ENST00000378217.3 |
OLAH |
oleoyl-ACP hydrolase |
| chr4_-_100356551 | 0.24 |
ENST00000209665.4 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr10_+_62538089 | 0.24 |
ENST00000519078.2 ENST00000395284.3 ENST00000316629.4 |
CDK1 |
cyclin-dependent kinase 1 |
| chr17_-_61996136 | 0.23 |
ENST00000342364.4 |
GH1 |
growth hormone 1 |
| chr3_+_108541608 | 0.23 |
ENST00000426646.1 |
TRAT1 |
T cell receptor associated transmembrane adaptor 1 |
| chr19_-_14952689 | 0.23 |
ENST00000248058.1 |
OR7A10 |
olfactory receptor, family 7, subfamily A, member 10 |
| chrX_+_100645812 | 0.22 |
ENST00000427805.2 ENST00000553110.3 ENST00000392994.3 ENST00000409338.1 ENST00000409170.3 |
RPL36A RPL36A-HNRNPH2 |
ribosomal protein L36a RPL36A-HNRNPH2 readthrough |
| chr1_+_62439037 | 0.22 |
ENST00000545929.1 |
INADL |
InaD-like (Drosophila) |
| chr12_+_101988627 | 0.21 |
ENST00000547405.1 ENST00000452455.2 ENST00000441232.1 ENST00000360610.2 ENST00000392934.3 ENST00000547509.1 ENST00000361685.2 ENST00000549145.1 ENST00000553190.1 |
MYBPC1 |
myosin binding protein C, slow type |
| chr16_-_28503357 | 0.20 |
ENST00000333496.9 ENST00000561505.1 ENST00000567963.1 ENST00000354630.5 ENST00000355477.5 ENST00000357076.5 ENST00000565688.1 ENST00000359984.7 |
CLN3 |
ceroid-lipofuscinosis, neuronal 3 |
| chr1_-_89357179 | 0.20 |
ENST00000448623.1 ENST00000418217.1 ENST00000370500.5 |
GTF2B |
general transcription factor IIB |
| chr15_+_99791567 | 0.20 |
ENST00000558879.1 ENST00000301981.3 ENST00000422500.2 ENST00000447360.2 ENST00000442993.2 |
LRRC28 |
leucine rich repeat containing 28 |
| chr6_-_33290580 | 0.20 |
ENST00000446511.1 ENST00000446403.1 ENST00000414083.2 ENST00000266000.6 ENST00000374542.5 |
DAXX |
death-domain associated protein |
| chr17_-_61996160 | 0.20 |
ENST00000458650.2 ENST00000351388.4 ENST00000323322.5 |
GH1 |
growth hormone 1 |
| chrX_+_46771711 | 0.19 |
ENST00000424392.1 ENST00000397189.1 |
PHF16 |
jade family PHD finger 3 |
| chr14_+_21785693 | 0.19 |
ENST00000382933.4 ENST00000557351.1 |
RPGRIP1 |
retinitis pigmentosa GTPase regulator interacting protein 1 |
| chr5_-_34916871 | 0.19 |
ENST00000382038.2 |
RAD1 |
RAD1 homolog (S. pombe) |
| chr12_-_45270151 | 0.19 |
ENST00000429094.2 |
NELL2 |
NEL-like 2 (chicken) |
| chr1_+_171454639 | 0.19 |
ENST00000392078.3 ENST00000426496.2 |
PRRC2C |
proline-rich coiled-coil 2C |
| chr12_-_45270077 | 0.19 |
ENST00000551601.1 ENST00000549027.1 ENST00000452445.2 |
NELL2 |
NEL-like 2 (chicken) |
| chr1_+_115572415 | 0.18 |
ENST00000256592.1 |
TSHB |
thyroid stimulating hormone, beta |
| chr17_-_72772462 | 0.18 |
ENST00000582870.1 ENST00000581136.1 ENST00000357814.3 ENST00000579218.1 ENST00000583476.1 ENST00000580301.1 ENST00000583757.1 ENST00000582524.1 |
NAT9 |
N-acetyltransferase 9 (GCN5-related, putative) |
| chr14_+_76618242 | 0.16 |
ENST00000557542.1 ENST00000557263.1 ENST00000557207.1 ENST00000312858.5 ENST00000261530.7 |
GPATCH2L |
G patch domain containing 2-like |
| chr2_-_40006289 | 0.15 |
ENST00000260619.6 ENST00000454352.2 |
THUMPD2 |
THUMP domain containing 2 |
| chr10_+_15085895 | 0.14 |
ENST00000378228.3 |
OLAH |
oleoyl-ACP hydrolase |
| chr12_-_68619586 | 0.14 |
ENST00000229134.4 |
IL26 |
interleukin 26 |
| chr6_-_29395509 | 0.13 |
ENST00000377147.2 |
OR11A1 |
olfactory receptor, family 11, subfamily A, member 1 |
| chr1_-_157014865 | 0.12 |
ENST00000361409.2 |
ARHGEF11 |
Rho guanine nucleotide exchange factor (GEF) 11 |
| chr16_+_31366455 | 0.11 |
ENST00000268296.4 |
ITGAX |
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chr12_-_45269430 | 0.11 |
ENST00000395487.2 |
NELL2 |
NEL-like 2 (chicken) |
| chr17_-_61996192 | 0.11 |
ENST00000392824.4 |
CSHL1 |
chorionic somatomammotropin hormone-like 1 |
| chr6_-_90529418 | 0.10 |
ENST00000439638.1 ENST00000369393.3 ENST00000428876.1 |
MDN1 |
MDN1, midasin homolog (yeast) |
| chr18_+_13612613 | 0.10 |
ENST00000586765.1 |
LDLRAD4 |
low density lipoprotein receptor class A domain containing 4 |
| chr6_+_46761118 | 0.10 |
ENST00000230588.4 |
MEP1A |
meprin A, alpha (PABA peptide hydrolase) |
| chr9_-_123239632 | 0.10 |
ENST00000416449.1 |
CDK5RAP2 |
CDK5 regulatory subunit associated protein 2 |
| chr15_+_66874502 | 0.10 |
ENST00000558797.1 |
RP11-321F6.1 |
HCG2003567; Uncharacterized protein |
| chr14_-_24553834 | 0.10 |
ENST00000397002.2 |
NRL |
neural retina leucine zipper |
| chr2_-_49381572 | 0.10 |
ENST00000454032.1 ENST00000304421.4 |
FSHR |
follicle stimulating hormone receptor |
| chr11_+_30252549 | 0.10 |
ENST00000254122.3 ENST00000417547.1 |
FSHB |
follicle stimulating hormone, beta polypeptide |
| chr16_+_31539183 | 0.09 |
ENST00000302312.4 |
AHSP |
alpha hemoglobin stabilizing protein |
| chr2_+_201997676 | 0.08 |
ENST00000462763.1 ENST00000479953.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr1_-_27226928 | 0.08 |
ENST00000361720.5 |
GPATCH3 |
G patch domain containing 3 |
| chr4_-_47983519 | 0.08 |
ENST00000358519.4 ENST00000544810.1 ENST00000402813.3 |
CNGA1 |
cyclic nucleotide gated channel alpha 1 |
| chr15_+_100106155 | 0.07 |
ENST00000557785.1 ENST00000558049.1 ENST00000449277.2 |
MEF2A |
myocyte enhancer factor 2A |
| chr20_+_43211149 | 0.06 |
ENST00000372886.1 |
PKIG |
protein kinase (cAMP-dependent, catalytic) inhibitor gamma |
| chr6_-_24936170 | 0.06 |
ENST00000538035.1 |
FAM65B |
family with sequence similarity 65, member B |
| chr19_-_14992264 | 0.06 |
ENST00000327462.2 |
OR7A17 |
olfactory receptor, family 7, subfamily A, member 17 |
| chr4_+_71570430 | 0.06 |
ENST00000417478.2 |
RUFY3 |
RUN and FYVE domain containing 3 |
| chr12_+_62860581 | 0.05 |
ENST00000393632.2 ENST00000393630.3 ENST00000280379.6 ENST00000546600.1 ENST00000552738.1 ENST00000393629.2 ENST00000552115.1 |
MON2 |
MON2 homolog (S. cerevisiae) |
| chr2_-_49381646 | 0.05 |
ENST00000346173.3 ENST00000406846.2 |
FSHR |
follicle stimulating hormone receptor |
| chr6_+_126102292 | 0.05 |
ENST00000368357.3 |
NCOA7 |
nuclear receptor coactivator 7 |
| chr1_-_203274418 | 0.05 |
ENST00000457348.1 |
RP11-134P9.1 |
long intergenic non-protein coding RNA 1136 |
| chr18_-_3874752 | 0.03 |
ENST00000534970.1 |
DLGAP1 |
discs, large (Drosophila) homolog-associated protein 1 |
| chr10_-_114206649 | 0.03 |
ENST00000369404.3 ENST00000369405.3 |
ZDHHC6 |
zinc finger, DHHC-type containing 6 |
| chr9_-_215744 | 0.03 |
ENST00000382387.2 |
C9orf66 |
chromosome 9 open reading frame 66 |
| chr12_+_20968608 | 0.03 |
ENST00000381541.3 ENST00000540229.1 ENST00000553473.1 ENST00000554957.1 |
LST3 SLCO1B3 SLCO1B7 |
Putative solute carrier organic anion transporter family member 1B7; Uncharacterized protein solute carrier organic anion transporter family, member 1B3 solute carrier organic anion transporter family, member 1B7 (non-functional) |
| chr10_+_60028818 | 0.02 |
ENST00000333926.5 |
CISD1 |
CDGSH iron sulfur domain 1 |
| chr3_+_155860751 | 0.02 |
ENST00000471742.1 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr12_-_54582655 | 0.02 |
ENST00000504338.1 ENST00000514685.1 ENST00000504797.1 ENST00000513838.1 ENST00000505128.1 ENST00000337581.3 ENST00000503306.1 ENST00000243112.5 ENST00000514196.1 ENST00000506169.1 ENST00000507904.1 ENST00000508394.2 |
SMUG1 |
single-strand-selective monofunctional uracil-DNA glycosylase 1 |
| chr9_-_70490107 | 0.01 |
ENST00000377395.4 ENST00000429800.2 ENST00000430059.2 ENST00000377384.1 ENST00000382405.3 |
CBWD5 |
COBW domain containing 5 |
| chr12_+_57881740 | 0.00 |
ENST00000262027.5 ENST00000315473.5 |
MARS |
methionyl-tRNA synthetase |
| chr11_+_118754475 | 0.00 |
ENST00000292174.4 |
CXCR5 |
chemokine (C-X-C motif) receptor 5 |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.7 | GO:0016823 | hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 0.4 | 1.2 | GO:0019959 | interleukin-8 binding(GO:0019959) |
| 0.3 | 0.8 | GO:0003858 | 3-hydroxybutyrate dehydrogenase activity(GO:0003858) |
| 0.2 | 3.1 | GO:0016594 | glycine binding(GO:0016594) |
| 0.2 | 1.5 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.2 | 0.9 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 0.2 | 2.4 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.1 | 0.4 | GO:0005148 | prolactin receptor binding(GO:0005148) |
| 0.1 | 2.5 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.1 | 3.9 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.1 | 2.1 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.1 | 0.4 | GO:0004320 | oleoyl-[acyl-carrier-protein] hydrolase activity(GO:0004320) myristoyl-[acyl-carrier-protein] hydrolase activity(GO:0016295) palmitoyl-[acyl-carrier-protein] hydrolase activity(GO:0016296) acyl-[acyl-carrier-protein] hydrolase activity(GO:0016297) |
| 0.1 | 2.6 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 7.7 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 0.5 | GO:0004485 | methylcrotonoyl-CoA carboxylase activity(GO:0004485) |
| 0.1 | 0.6 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.1 | 0.7 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.1 | 2.3 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.1 | 1.0 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.1 | 0.3 | GO:0042030 | ATPase inhibitor activity(GO:0042030) |
| 0.1 | 0.7 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.1 | 0.2 | GO:0035276 | aldehyde oxidase activity(GO:0004031) ethanol binding(GO:0035276) |
| 0.1 | 0.6 | GO:0015379 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 0.1 | 0.3 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.1 | 1.8 | GO:0001848 | complement binding(GO:0001848) |
| 0.0 | 4.6 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.5 | GO:0016595 | glutamate binding(GO:0016595) |
| 0.0 | 0.7 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.0 | 0.6 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.0 | 0.1 | GO:0004963 | follicle-stimulating hormone receptor activity(GO:0004963) |
| 0.0 | 0.4 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.0 | 0.5 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.0 | 1.1 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.2 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.0 | 0.8 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.0 | 0.5 | GO:0001054 | RNA polymerase I activity(GO:0001054) |
| 0.0 | 0.5 | GO:0016289 | CoA hydrolase activity(GO:0016289) |
| 0.0 | 2.6 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 0.3 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.0 | 0.1 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.0 | 0.4 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.0 | 0.5 | GO:0015464 | acetylcholine receptor activity(GO:0015464) |
| 0.0 | 1.1 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 0.3 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.0 | 0.2 | GO:0031432 | titin binding(GO:0031432) |
| 0.0 | 0.8 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.0 | 0.8 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 0.3 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 1.2 | GO:0030295 | protein kinase activator activity(GO:0030295) |
| 0.0 | 2.9 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.1 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.0 | 0.6 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 0.6 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.3 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.0 | 0.2 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.0 | 1.0 | GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding(GO:0000980) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 4.1 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 2.0 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.0 | 1.0 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.0 | 2.3 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 1.1 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.0 | 1.7 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.0 | 1.1 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 0.6 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.0 | 0.4 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.0 | 1.7 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 1.0 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 1.2 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 0.9 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 0.2 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.0 | 4.2 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.3 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.0 | 0.5 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.0 | 0.8 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.1 | PID E2F PATHWAY | E2F transcription factor network |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.1 | GO:1902769 | regulation of choline O-acetyltransferase activity(GO:1902769) positive regulation of choline O-acetyltransferase activity(GO:1902771) negative regulation of tau-protein kinase activity(GO:1902948) positive regulation of early endosome to recycling endosome transport(GO:1902955) negative regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902960) negative regulation of neurofibrillary tangle assembly(GO:1902997) negative regulation of aspartic-type peptidase activity(GO:1905246) |
| 0.7 | 3.3 | GO:0002399 | MHC class II protein complex assembly(GO:0002399) |
| 0.6 | 3.1 | GO:0006546 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 0.6 | 1.7 | GO:0097052 | tryptophan catabolic process to acetyl-CoA(GO:0019442) L-kynurenine metabolic process(GO:0097052) |
| 0.5 | 1.5 | GO:1903925 | signal transduction involved in intra-S DNA damage checkpoint(GO:0072428) response to bisphenol A(GO:1903925) cellular response to bisphenol A(GO:1903926) |
| 0.3 | 2.6 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.2 | 1.0 | GO:0043376 | regulation of CD8-positive, alpha-beta T cell differentiation(GO:0043376) actin polymerization-dependent cell motility(GO:0070358) |
| 0.2 | 0.9 | GO:0032690 | negative regulation of interleukin-1 alpha production(GO:0032690) negative regulation of interleukin-1 alpha secretion(GO:0050712) |
| 0.2 | 3.1 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.2 | 0.5 | GO:0071676 | negative regulation of mononuclear cell migration(GO:0071676) |
| 0.2 | 0.8 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.1 | 1.0 | GO:1900020 | regulation of protein kinase C activity(GO:1900019) positive regulation of protein kinase C activity(GO:1900020) |
| 0.1 | 0.4 | GO:1904647 | response to rotenone(GO:1904647) |
| 0.1 | 0.6 | GO:0071477 | hypotonic salinity response(GO:0042539) cellular hypotonic salinity response(GO:0071477) |
| 0.1 | 0.5 | GO:0097068 | response to thyroxine(GO:0097068) response to L-phenylalanine derivative(GO:1904386) |
| 0.1 | 2.3 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.1 | 2.1 | GO:2000258 | negative regulation of complement activation(GO:0045916) negative regulation of protein activation cascade(GO:2000258) |
| 0.1 | 0.7 | GO:0000480 | endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000480) |
| 0.1 | 1.1 | GO:1903944 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 2.6 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.1 | 0.3 | GO:1901876 | regulation of calcium ion binding(GO:1901876) negative regulation of calcium ion binding(GO:1901877) regulation of calcium ion import into sarcoplasmic reticulum(GO:1902080) negative regulation of calcium ion import into sarcoplasmic reticulum(GO:1902081) |
| 0.1 | 0.5 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.1 | 0.3 | GO:0070512 | regulation of histone H4-K20 methylation(GO:0070510) positive regulation of histone H4-K20 methylation(GO:0070512) |
| 0.1 | 0.3 | GO:0019086 | late viral transcription(GO:0019086) |
| 0.1 | 1.8 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.1 | 0.2 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.1 | 0.3 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.1 | 0.7 | GO:0042492 | gamma-delta T cell differentiation(GO:0042492) |
| 0.1 | 4.2 | GO:0030212 | hyaluronan metabolic process(GO:0030212) |
| 0.1 | 7.5 | GO:0030449 | regulation of complement activation(GO:0030449) |
| 0.1 | 1.0 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.1 | 0.7 | GO:0042905 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.1 | 0.8 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.1 | 0.5 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.1 | 0.3 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.1 | 0.5 | GO:0006552 | leucine catabolic process(GO:0006552) |
| 0.1 | 0.4 | GO:0010536 | positive regulation of activation of Janus kinase activity(GO:0010536) |
| 0.1 | 0.8 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.1 | 0.2 | GO:0035752 | lysosomal lumen pH elevation(GO:0035752) |
| 0.0 | 0.2 | GO:1903935 | response to sodium arsenite(GO:1903935) cellular response to sodium arsenite(GO:1903936) |
| 0.0 | 0.2 | GO:0033313 | meiotic cell cycle checkpoint(GO:0033313) |
| 0.0 | 1.0 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.0 | 0.5 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.0 | 0.2 | GO:0014038 | regulation of Schwann cell differentiation(GO:0014038) |
| 0.0 | 0.1 | GO:0001545 | primary ovarian follicle growth(GO:0001545) |
| 0.0 | 0.4 | GO:0048194 | Golgi vesicle budding(GO:0048194) |
| 0.0 | 0.8 | GO:0042789 | mRNA transcription from RNA polymerase II promoter(GO:0042789) |
| 0.0 | 0.2 | GO:1904798 | positive regulation of core promoter binding(GO:1904798) |
| 0.0 | 1.4 | GO:0043029 | T cell homeostasis(GO:0043029) |
| 0.0 | 3.8 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.0 | 0.3 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.0 | 2.4 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.0 | 0.6 | GO:0070493 | thrombin receptor signaling pathway(GO:0070493) |
| 0.0 | 0.3 | GO:0060662 | tube lumen cavitation(GO:0060605) salivary gland cavitation(GO:0060662) |
| 0.0 | 1.1 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.0 | 0.1 | GO:0045872 | positive regulation of rhodopsin gene expression(GO:0045872) |
| 0.0 | 0.3 | GO:0002260 | lymphocyte homeostasis(GO:0002260) |
| 0.0 | 0.7 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.0 | 2.4 | GO:0002819 | regulation of adaptive immune response(GO:0002819) |
| 0.0 | 2.3 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.0 | 0.1 | GO:0060011 | Sertoli cell proliferation(GO:0060011) |
| 0.0 | 1.5 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 1.3 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 0.2 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.0 | 0.5 | GO:0006363 | termination of RNA polymerase I transcription(GO:0006363) |
| 0.0 | 0.1 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 9.5 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.3 | 6.3 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.2 | 2.6 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.1 | 1.7 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.1 | 1.7 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.1 | 1.2 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.1 | 2.0 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.0 | 1.1 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 1.6 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 1.1 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 1.0 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 3.6 | REACTOME CELL SURFACE INTERACTIONS AT THE VASCULAR WALL | Genes involved in Cell surface interactions at the vascular wall |
| 0.0 | 0.7 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.7 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.0 | 0.2 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.0 | 0.4 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.0 | 0.4 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.4 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.0 | 0.3 | REACTOME SHC1 EVENTS IN EGFR SIGNALING | Genes involved in SHC1 events in EGFR signaling |
| 0.0 | 0.5 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |
| 0.0 | 0.9 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.5 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |
| 0.0 | 0.6 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |
| 0.0 | 1.1 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.5 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 0.3 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 3.1 | GO:0005960 | glycine cleavage complex(GO:0005960) |
| 0.4 | 2.6 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.3 | 7.1 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.2 | 2.6 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.2 | 3.1 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 0.4 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.1 | 2.1 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.1 | 0.5 | GO:0002169 | 3-methylcrotonyl-CoA carboxylase complex, mitochondrial(GO:0002169) methylcrotonoyl-CoA carboxylase complex(GO:1905202) |
| 0.1 | 0.7 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 0.3 | GO:0032279 | asymmetric synapse(GO:0032279) |
| 0.1 | 0.3 | GO:0031436 | BRCA1-BARD1 complex(GO:0031436) |
| 0.1 | 1.0 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.1 | 1.1 | GO:0097342 | ripoptosome(GO:0097342) |
| 0.0 | 0.2 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 0.0 | 0.2 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.0 | 0.3 | GO:0032437 | cuticular plate(GO:0032437) |
| 0.0 | 7.8 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 0.2 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.5 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 0.7 | GO:0098576 | lumenal side of membrane(GO:0098576) |
| 0.0 | 0.3 | GO:0000796 | condensin complex(GO:0000796) |
| 0.0 | 0.7 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.0 | 3.2 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 2.3 | GO:0034705 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.0 | 0.2 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.0 | 0.6 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.5 | GO:0042101 | T cell receptor complex(GO:0042101) |
| 0.0 | 1.7 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 1.7 | GO:0043679 | axon terminus(GO:0043679) |
| 0.0 | 0.1 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.0 | 0.6 | GO:0005747 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |