Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
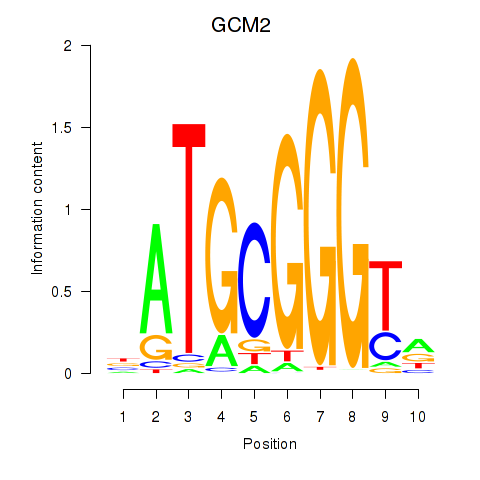
Results for GCM2
Z-value: 0.74
Transcription factors associated with GCM2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
GCM2
|
ENSG00000124827.6 | GCM2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| GCM2 | hg19_v2_chr6_-_10882174_10882274 | 0.08 | 7.8e-01 | Click! |
Activity profile of GCM2 motif
Sorted Z-values of GCM2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of GCM2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_116139744 | 1.88 |
ENST00000343213.2 |
CAV2 |
caveolin 2 |
| chr11_+_86511569 | 1.72 |
ENST00000441050.1 |
PRSS23 |
protease, serine, 23 |
| chr9_-_130635741 | 1.50 |
ENST00000223836.10 |
AK1 |
adenylate kinase 1 |
| chr5_+_82767284 | 1.43 |
ENST00000265077.3 |
VCAN |
versican |
| chr7_+_12727250 | 1.22 |
ENST00000404894.1 |
ARL4A |
ADP-ribosylation factor-like 4A |
| chr5_+_82767487 | 1.20 |
ENST00000343200.5 ENST00000342785.4 |
VCAN |
versican |
| chr1_-_159893507 | 1.07 |
ENST00000368096.1 |
TAGLN2 |
transgelin 2 |
| chr5_+_82767583 | 1.01 |
ENST00000512590.2 ENST00000513960.1 ENST00000513984.1 ENST00000502527.2 |
VCAN |
versican |
| chr21_-_28338732 | 0.96 |
ENST00000284987.5 |
ADAMTS5 |
ADAM metallopeptidase with thrombospondin type 1 motif, 5 |
| chr18_+_3449695 | 0.93 |
ENST00000343820.5 |
TGIF1 |
TGFB-induced factor homeobox 1 |
| chr6_-_41909191 | 0.91 |
ENST00000512426.1 ENST00000372987.4 |
CCND3 |
cyclin D3 |
| chr11_-_6341724 | 0.90 |
ENST00000530979.1 |
PRKCDBP |
protein kinase C, delta binding protein |
| chr12_+_81110684 | 0.89 |
ENST00000228644.3 |
MYF5 |
myogenic factor 5 |
| chr14_-_105635090 | 0.87 |
ENST00000331782.3 ENST00000347004.2 |
JAG2 |
jagged 2 |
| chr22_-_20307532 | 0.74 |
ENST00000405465.3 ENST00000248879.3 |
DGCR6L |
DiGeorge syndrome critical region gene 6-like |
| chr7_-_151329416 | 0.74 |
ENST00000418337.2 |
PRKAG2 |
protein kinase, AMP-activated, gamma 2 non-catalytic subunit |
| chr5_-_54281407 | 0.71 |
ENST00000381403.4 |
ESM1 |
endothelial cell-specific molecule 1 |
| chr13_-_45010939 | 0.66 |
ENST00000261489.2 |
TSC22D1 |
TSC22 domain family, member 1 |
| chr15_+_43809797 | 0.62 |
ENST00000399453.1 ENST00000300231.5 |
MAP1A |
microtubule-associated protein 1A |
| chr22_+_31489344 | 0.60 |
ENST00000404574.1 |
SMTN |
smoothelin |
| chr8_+_20054878 | 0.59 |
ENST00000276390.2 ENST00000519667.1 |
ATP6V1B2 |
ATPase, H+ transporting, lysosomal 56/58kDa, V1 subunit B2 |
| chr18_+_3449821 | 0.59 |
ENST00000407501.2 ENST00000405385.3 ENST00000546979.1 |
TGIF1 |
TGFB-induced factor homeobox 1 |
| chr1_+_156105878 | 0.59 |
ENST00000508500.1 |
LMNA |
lamin A/C |
| chr9_-_13279589 | 0.57 |
ENST00000319217.7 |
MPDZ |
multiple PDZ domain protein |
| chr3_+_126707437 | 0.52 |
ENST00000393409.2 ENST00000251772.4 |
PLXNA1 |
plexin A1 |
| chr10_+_102790980 | 0.48 |
ENST00000393459.1 ENST00000224807.5 |
SFXN3 |
sideroflexin 3 |
| chr9_-_13279563 | 0.48 |
ENST00000541718.1 |
MPDZ |
multiple PDZ domain protein |
| chr3_+_35681081 | 0.47 |
ENST00000428373.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr22_+_19710468 | 0.46 |
ENST00000366425.3 |
GP1BB |
glycoprotein Ib (platelet), beta polypeptide |
| chr19_-_12886327 | 0.45 |
ENST00000397668.3 ENST00000587178.1 ENST00000264827.5 |
HOOK2 |
hook microtubule-tethering protein 2 |
| chr3_+_30647994 | 0.43 |
ENST00000295754.5 |
TGFBR2 |
transforming growth factor, beta receptor II (70/80kDa) |
| chr16_+_69221028 | 0.42 |
ENST00000336278.4 |
SNTB2 |
syntrophin, beta 2 (dystrophin-associated protein A1, 59kDa, basic component 2) |
| chr15_+_89010923 | 0.42 |
ENST00000353598.6 |
MRPS11 |
mitochondrial ribosomal protein S11 |
| chr2_+_12857043 | 0.42 |
ENST00000381465.2 |
TRIB2 |
tribbles pseudokinase 2 |
| chr1_+_65886326 | 0.41 |
ENST00000371059.3 ENST00000371060.3 ENST00000349533.6 ENST00000406510.3 |
LEPR |
leptin receptor |
| chr7_+_6048856 | 0.40 |
ENST00000223029.3 ENST00000400479.2 ENST00000395236.2 |
AIMP2 |
aminoacyl tRNA synthetase complex-interacting multifunctional protein 2 |
| chr2_+_12857015 | 0.38 |
ENST00000155926.4 |
TRIB2 |
tribbles pseudokinase 2 |
| chr11_+_4116005 | 0.38 |
ENST00000300738.5 |
RRM1 |
ribonucleotide reductase M1 |
| chr11_+_4116054 | 0.37 |
ENST00000423050.2 |
RRM1 |
ribonucleotide reductase M1 |
| chr8_-_98290087 | 0.36 |
ENST00000322128.3 |
TSPYL5 |
TSPY-like 5 |
| chr17_-_76899275 | 0.35 |
ENST00000322630.2 ENST00000586713.1 |
DDC8 |
Protein DDC8 homolog |
| chr8_+_66556936 | 0.34 |
ENST00000262146.4 |
MTFR1 |
mitochondrial fission regulator 1 |
| chr12_-_56122124 | 0.34 |
ENST00000552754.1 |
CD63 |
CD63 molecule |
| chr10_-_70092671 | 0.33 |
ENST00000358769.2 ENST00000432941.1 ENST00000495025.2 |
PBLD |
phenazine biosynthesis-like protein domain containing |
| chr3_+_186383741 | 0.33 |
ENST00000232003.4 |
HRG |
histidine-rich glycoprotein |
| chr1_+_178062855 | 0.32 |
ENST00000448150.3 |
RASAL2 |
RAS protein activator like 2 |
| chr5_+_140743859 | 0.31 |
ENST00000518069.1 |
PCDHGA5 |
protocadherin gamma subfamily A, 5 |
| chr6_-_110500905 | 0.31 |
ENST00000392587.2 |
WASF1 |
WAS protein family, member 1 |
| chr5_-_54281491 | 0.31 |
ENST00000381405.4 |
ESM1 |
endothelial cell-specific molecule 1 |
| chr7_+_148395733 | 0.31 |
ENST00000602748.1 |
CUL1 |
cullin 1 |
| chr1_-_53686261 | 0.31 |
ENST00000294360.4 |
C1orf123 |
chromosome 1 open reading frame 123 |
| chr12_+_124069070 | 0.31 |
ENST00000262225.3 ENST00000438031.2 |
TMED2 |
transmembrane emp24 domain trafficking protein 2 |
| chr1_-_167906277 | 0.30 |
ENST00000271373.4 |
MPC2 |
mitochondrial pyruvate carrier 2 |
| chr1_+_203595903 | 0.30 |
ENST00000367218.3 ENST00000367219.3 ENST00000391954.2 |
ATP2B4 |
ATPase, Ca++ transporting, plasma membrane 4 |
| chr17_+_7358889 | 0.30 |
ENST00000575379.1 |
CHRNB1 |
cholinergic receptor, nicotinic, beta 1 (muscle) |
| chr2_-_33824336 | 0.29 |
ENST00000431950.1 ENST00000403368.1 ENST00000441530.2 |
FAM98A |
family with sequence similarity 98, member A |
| chr8_-_145559943 | 0.29 |
ENST00000332135.4 |
SCRT1 |
scratch family zinc finger 1 |
| chr5_+_176560007 | 0.28 |
ENST00000510954.1 ENST00000354179.4 |
NSD1 |
nuclear receptor binding SET domain protein 1 |
| chr2_-_33824382 | 0.28 |
ENST00000238823.8 |
FAM98A |
family with sequence similarity 98, member A |
| chr1_+_64239657 | 0.27 |
ENST00000371080.1 ENST00000371079.1 |
ROR1 |
receptor tyrosine kinase-like orphan receptor 1 |
| chr12_-_49582593 | 0.27 |
ENST00000295766.5 |
TUBA1A |
tubulin, alpha 1a |
| chr7_-_6048650 | 0.27 |
ENST00000382321.4 ENST00000406569.3 |
PMS2 |
PMS2 postmeiotic segregation increased 2 (S. cerevisiae) |
| chr11_+_73882144 | 0.27 |
ENST00000328257.8 |
PPME1 |
protein phosphatase methylesterase 1 |
| chr7_+_106809406 | 0.26 |
ENST00000468410.1 ENST00000478930.1 ENST00000464009.1 ENST00000222574.4 |
HBP1 |
HMG-box transcription factor 1 |
| chr11_+_66406088 | 0.25 |
ENST00000310092.7 ENST00000396053.4 ENST00000408993.2 |
RBM4 |
RNA binding motif protein 4 |
| chr7_-_44365020 | 0.25 |
ENST00000395747.2 ENST00000347193.4 ENST00000346990.4 ENST00000258682.6 ENST00000353625.4 ENST00000421607.1 ENST00000424197.1 ENST00000502837.2 ENST00000350811.3 ENST00000395749.2 |
CAMK2B |
calcium/calmodulin-dependent protein kinase II beta |
| chr15_+_74466744 | 0.25 |
ENST00000560862.1 ENST00000395118.1 |
ISLR |
immunoglobulin superfamily containing leucine-rich repeat |
| chr1_+_203595689 | 0.25 |
ENST00000357681.5 |
ATP2B4 |
ATPase, Ca++ transporting, plasma membrane 4 |
| chr8_-_144897138 | 0.25 |
ENST00000377533.3 |
SCRIB |
scribbled planar cell polarity protein |
| chr8_-_144897549 | 0.23 |
ENST00000356994.2 ENST00000320476.3 |
SCRIB |
scribbled planar cell polarity protein |
| chr1_+_46640750 | 0.23 |
ENST00000372003.1 |
TSPAN1 |
tetraspanin 1 |
| chr2_-_44588624 | 0.21 |
ENST00000438314.1 ENST00000409936.1 |
PREPL |
prolyl endopeptidase-like |
| chr5_-_137878887 | 0.21 |
ENST00000507939.1 ENST00000572514.1 ENST00000499810.2 ENST00000360541.5 |
ETF1 |
eukaryotic translation termination factor 1 |
| chr11_+_66036004 | 0.21 |
ENST00000311481.6 ENST00000527397.1 |
RAB1B |
RAB1B, member RAS oncogene family |
| chr2_-_179315490 | 0.21 |
ENST00000487082.1 |
PRKRA |
protein kinase, interferon-inducible double stranded RNA dependent activator |
| chr2_-_44588694 | 0.21 |
ENST00000409957.1 |
PREPL |
prolyl endopeptidase-like |
| chr7_-_6048702 | 0.21 |
ENST00000265849.7 |
PMS2 |
PMS2 postmeiotic segregation increased 2 (S. cerevisiae) |
| chr17_-_41132010 | 0.21 |
ENST00000409103.1 ENST00000360221.4 |
PTGES3L-AARSD1 |
PTGES3L-AARSD1 readthrough |
| chr20_+_42086525 | 0.21 |
ENST00000244020.3 |
SRSF6 |
serine/arginine-rich splicing factor 6 |
| chr2_-_44588679 | 0.20 |
ENST00000409411.1 |
PREPL |
prolyl endopeptidase-like |
| chr22_+_37447771 | 0.20 |
ENST00000402077.3 ENST00000403888.3 ENST00000456470.1 |
KCTD17 |
potassium channel tetramerization domain containing 17 |
| chr15_-_75248954 | 0.20 |
ENST00000499788.2 |
RPP25 |
ribonuclease P/MRP 25kDa subunit |
| chrX_-_119763835 | 0.20 |
ENST00000371313.2 ENST00000304661.5 |
C1GALT1C1 |
C1GALT1-specific chaperone 1 |
| chr3_+_15247686 | 0.19 |
ENST00000253693.2 |
CAPN7 |
calpain 7 |
| chr17_-_42907564 | 0.19 |
ENST00000592524.1 |
GJC1 |
gap junction protein, gamma 1, 45kDa |
| chr7_+_148395959 | 0.18 |
ENST00000325222.4 |
CUL1 |
cullin 1 |
| chr4_-_74864386 | 0.18 |
ENST00000296027.4 |
CXCL5 |
chemokine (C-X-C motif) ligand 5 |
| chr6_-_52859968 | 0.18 |
ENST00000370959.1 |
GSTA4 |
glutathione S-transferase alpha 4 |
| chr16_+_22825475 | 0.18 |
ENST00000261374.3 |
HS3ST2 |
heparan sulfate (glucosamine) 3-O-sulfotransferase 2 |
| chr13_-_47471155 | 0.18 |
ENST00000543956.1 ENST00000542664.1 |
HTR2A |
5-hydroxytryptamine (serotonin) receptor 2A, G protein-coupled |
| chr13_-_44735393 | 0.18 |
ENST00000400419.1 |
SMIM2 |
small integral membrane protein 2 |
| chr14_-_75536182 | 0.18 |
ENST00000555463.1 |
ACYP1 |
acylphosphatase 1, erythrocyte (common) type |
| chr7_-_44365216 | 0.17 |
ENST00000358707.3 ENST00000457475.1 ENST00000440254.2 |
CAMK2B |
calcium/calmodulin-dependent protein kinase II beta |
| chr19_+_35783047 | 0.17 |
ENST00000595791.1 ENST00000597035.1 ENST00000537831.2 |
MAG |
myelin associated glycoprotein |
| chr11_+_63448918 | 0.17 |
ENST00000341307.2 ENST00000356000.3 ENST00000542238.1 |
RTN3 |
reticulon 3 |
| chr17_+_79989500 | 0.16 |
ENST00000306897.4 |
RAC3 |
ras-related C3 botulinum toxin substrate 3 (rho family, small GTP binding protein Rac3) |
| chr11_-_64410787 | 0.16 |
ENST00000301894.2 |
NRXN2 |
neurexin 2 |
| chr3_+_156008809 | 0.16 |
ENST00000302490.8 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr19_+_35783037 | 0.15 |
ENST00000361922.4 |
MAG |
myelin associated glycoprotein |
| chr14_+_21359558 | 0.15 |
ENST00000304639.3 |
RNASE3 |
ribonuclease, RNase A family, 3 |
| chr2_-_179315453 | 0.15 |
ENST00000432031.2 |
PRKRA |
protein kinase, interferon-inducible double stranded RNA dependent activator |
| chr3_-_12705600 | 0.15 |
ENST00000542177.1 ENST00000442415.2 ENST00000251849.4 |
RAF1 |
v-raf-1 murine leukemia viral oncogene homolog 1 |
| chr16_+_2564254 | 0.15 |
ENST00000565223.1 |
ATP6V0C |
ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c |
| chr12_-_96794330 | 0.15 |
ENST00000261211.3 |
CDK17 |
cyclin-dependent kinase 17 |
| chr11_+_63448955 | 0.15 |
ENST00000377819.5 ENST00000339997.4 ENST00000540798.1 ENST00000545432.1 ENST00000543552.1 ENST00000537981.1 |
RTN3 |
reticulon 3 |
| chr19_+_35783028 | 0.14 |
ENST00000600291.1 ENST00000392213.3 |
MAG |
myelin associated glycoprotein |
| chr3_-_50605150 | 0.14 |
ENST00000357203.3 |
C3orf18 |
chromosome 3 open reading frame 18 |
| chr17_-_42908155 | 0.14 |
ENST00000426548.1 ENST00000590758.1 ENST00000591424.1 |
GJC1 |
gap junction protein, gamma 1, 45kDa |
| chr12_-_96794143 | 0.14 |
ENST00000543119.2 |
CDK17 |
cyclin-dependent kinase 17 |
| chr3_-_50605077 | 0.13 |
ENST00000426034.1 ENST00000441239.1 |
C3orf18 |
chromosome 3 open reading frame 18 |
| chr1_-_27226928 | 0.13 |
ENST00000361720.5 |
GPATCH3 |
G patch domain containing 3 |
| chr3_+_184032919 | 0.13 |
ENST00000427845.1 ENST00000342981.4 ENST00000319274.6 |
EIF4G1 |
eukaryotic translation initiation factor 4 gamma, 1 |
| chrX_+_12156582 | 0.12 |
ENST00000380682.1 |
FRMPD4 |
FERM and PDZ domain containing 4 |
| chr22_-_29457832 | 0.12 |
ENST00000216071.4 |
C22orf31 |
chromosome 22 open reading frame 31 |
| chr19_-_14316980 | 0.12 |
ENST00000361434.3 ENST00000340736.6 |
LPHN1 |
latrophilin 1 |
| chr6_+_31633902 | 0.12 |
ENST00000375865.2 ENST00000375866.2 |
CSNK2B |
casein kinase 2, beta polypeptide |
| chr11_+_64002292 | 0.12 |
ENST00000426086.2 |
VEGFB |
vascular endothelial growth factor B |
| chr9_-_99417562 | 0.11 |
ENST00000375234.3 ENST00000446045.1 |
AAED1 |
AhpC/TSA antioxidant enzyme domain containing 1 |
| chr6_-_32077100 | 0.11 |
ENST00000375244.3 ENST00000375247.2 |
TNXB |
tenascin XB |
| chr1_+_43824669 | 0.11 |
ENST00000372462.1 |
CDC20 |
cell division cycle 20 |
| chr7_-_150754935 | 0.10 |
ENST00000297518.4 |
CDK5 |
cyclin-dependent kinase 5 |
| chr3_+_50712672 | 0.10 |
ENST00000266037.9 |
DOCK3 |
dedicator of cytokinesis 3 |
| chr5_+_177027101 | 0.10 |
ENST00000029410.5 ENST00000510761.1 ENST00000505468.1 |
B4GALT7 |
xylosylprotein beta 1,4-galactosyltransferase, polypeptide 7 |
| chr19_+_6361440 | 0.10 |
ENST00000245816.4 |
CLPP |
caseinolytic mitochondrial matrix peptidase proteolytic subunit |
| chr7_-_36764004 | 0.10 |
ENST00000431169.1 |
AOAH |
acyloxyacyl hydrolase (neutrophil) |
| chr6_-_150039249 | 0.10 |
ENST00000543571.1 |
LATS1 |
large tumor suppressor kinase 1 |
| chr15_+_40861487 | 0.10 |
ENST00000315616.7 ENST00000559271.1 |
RPUSD2 |
RNA pseudouridylate synthase domain containing 2 |
| chr7_-_8301768 | 0.10 |
ENST00000265577.7 |
ICA1 |
islet cell autoantigen 1, 69kDa |
| chr1_-_156698591 | 0.10 |
ENST00000368219.1 |
ISG20L2 |
interferon stimulated exonuclease gene 20kDa-like 2 |
| chr7_-_36764142 | 0.09 |
ENST00000258749.5 ENST00000535891.1 |
AOAH |
acyloxyacyl hydrolase (neutrophil) |
| chr5_-_141257954 | 0.09 |
ENST00000456271.1 ENST00000394536.3 ENST00000503492.1 ENST00000287008.3 |
PCDH1 |
protocadherin 1 |
| chr2_+_234627424 | 0.09 |
ENST00000373409.3 |
UGT1A4 |
UDP glucuronosyltransferase 1 family, polypeptide A4 |
| chr19_+_42829702 | 0.09 |
ENST00000334370.4 |
MEGF8 |
multiple EGF-like-domains 8 |
| chr4_+_26321284 | 0.08 |
ENST00000506956.1 ENST00000512671.1 ENST00000345843.3 ENST00000342295.1 |
RBPJ |
recombination signal binding protein for immunoglobulin kappa J region |
| chr11_+_64001962 | 0.08 |
ENST00000309422.2 |
VEGFB |
vascular endothelial growth factor B |
| chr17_-_7197881 | 0.08 |
ENST00000007699.5 |
YBX2 |
Y box binding protein 2 |
| chr12_+_5541267 | 0.08 |
ENST00000423158.3 |
NTF3 |
neurotrophin 3 |
| chr6_-_142409936 | 0.08 |
ENST00000258042.1 |
NMBR |
neuromedin B receptor |
| chr15_+_40731920 | 0.08 |
ENST00000561234.1 |
BAHD1 |
bromo adjacent homology domain containing 1 |
| chr1_+_29063271 | 0.07 |
ENST00000373812.3 |
YTHDF2 |
YTH domain family, member 2 |
| chr16_+_28996364 | 0.07 |
ENST00000564277.1 |
LAT |
linker for activation of T cells |
| chr3_+_130650738 | 0.07 |
ENST00000504612.1 |
ATP2C1 |
ATPase, Ca++ transporting, type 2C, member 1 |
| chr15_-_64665911 | 0.07 |
ENST00000606793.1 ENST00000561349.1 ENST00000560278.1 |
CTD-2116N17.1 |
Uncharacterized protein |
| chr1_-_228613026 | 0.07 |
ENST00000366696.1 |
HIST3H3 |
histone cluster 3, H3 |
| chr1_+_43824577 | 0.07 |
ENST00000310955.6 |
CDC20 |
cell division cycle 20 |
| chr14_-_59932044 | 0.07 |
ENST00000395116.1 |
GPR135 |
G protein-coupled receptor 135 |
| chr6_+_110012462 | 0.07 |
ENST00000441478.2 ENST00000230124.3 |
FIG4 |
FIG4 homolog, SAC1 lipid phosphatase domain containing (S. cerevisiae) |
| chr11_-_83393457 | 0.07 |
ENST00000404783.3 |
DLG2 |
discs, large homolog 2 (Drosophila) |
| chr19_-_51530916 | 0.07 |
ENST00000594768.1 |
KLK11 |
kallikrein-related peptidase 11 |
| chr17_-_46703826 | 0.07 |
ENST00000550387.1 ENST00000311177.5 |
HOXB9 |
homeobox B9 |
| chr3_-_45837959 | 0.06 |
ENST00000353278.4 ENST00000456124.2 |
SLC6A20 |
solute carrier family 6 (proline IMINO transporter), member 20 |
| chr17_-_7123021 | 0.06 |
ENST00000399510.2 |
DLG4 |
discs, large homolog 4 (Drosophila) |
| chr6_+_31633833 | 0.06 |
ENST00000375882.2 ENST00000375880.2 |
CSNK2B CSNK2B-LY6G5B-1181 |
casein kinase 2, beta polypeptide Uncharacterized protein |
| chr15_+_40733387 | 0.06 |
ENST00000416165.1 |
BAHD1 |
bromo adjacent homology domain containing 1 |
| chr11_+_76778033 | 0.06 |
ENST00000456580.2 |
CAPN5 |
calpain 5 |
| chr1_+_150521876 | 0.06 |
ENST00000369041.5 ENST00000271643.4 ENST00000538795.1 |
ADAMTSL4 AL356356.1 |
ADAMTS-like 4 Protein LOC100996516 |
| chr11_-_83393429 | 0.05 |
ENST00000426717.2 |
DLG2 |
discs, large homolog 2 (Drosophila) |
| chr7_-_100808394 | 0.05 |
ENST00000445482.2 |
VGF |
VGF nerve growth factor inducible |
| chrX_-_111326004 | 0.05 |
ENST00000262839.2 |
TRPC5 |
transient receptor potential cation channel, subfamily C, member 5 |
| chr20_-_23731569 | 0.05 |
ENST00000304749.2 |
CST1 |
cystatin SN |
| chr19_+_41222998 | 0.05 |
ENST00000263370.2 |
ITPKC |
inositol-trisphosphate 3-kinase C |
| chr1_-_223537401 | 0.04 |
ENST00000343846.3 ENST00000454695.2 ENST00000484758.2 |
SUSD4 |
sushi domain containing 4 |
| chrX_-_71526813 | 0.04 |
ENST00000246139.5 |
CITED1 |
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 1 |
| chr10_-_21786179 | 0.04 |
ENST00000377113.5 |
CASC10 |
cancer susceptibility candidate 10 |
| chr15_+_78632666 | 0.04 |
ENST00000299529.6 |
CRABP1 |
cellular retinoic acid binding protein 1 |
| chr21_+_27011584 | 0.04 |
ENST00000400532.1 ENST00000480456.1 ENST00000312957.5 |
JAM2 |
junctional adhesion molecule 2 |
| chr11_+_64009072 | 0.04 |
ENST00000535135.1 ENST00000394540.3 |
FKBP2 |
FK506 binding protein 2, 13kDa |
| chr5_+_139493665 | 0.04 |
ENST00000331327.3 |
PURA |
purine-rich element binding protein A |
| chr1_+_44444865 | 0.04 |
ENST00000372324.1 |
B4GALT2 |
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 2 |
| chr19_+_22469236 | 0.04 |
ENST00000357491.6 |
ZNF729 |
zinc finger protein 729 |
| chr11_+_76777979 | 0.04 |
ENST00000531028.1 ENST00000278559.3 ENST00000527066.1 ENST00000529629.1 |
CAPN5 |
calpain 5 |
| chr2_+_233243233 | 0.03 |
ENST00000392027.2 |
ALPP |
alkaline phosphatase, placental |
| chr6_-_138539627 | 0.03 |
ENST00000527246.2 |
PBOV1 |
prostate and breast cancer overexpressed 1 |
| chrX_-_63005405 | 0.03 |
ENST00000374878.1 ENST00000437457.2 |
ARHGEF9 |
Cdc42 guanine nucleotide exchange factor (GEF) 9 |
| chr2_-_179315786 | 0.03 |
ENST00000457633.1 ENST00000438687.3 ENST00000325748.4 |
PRKRA |
protein kinase, interferon-inducible double stranded RNA dependent activator |
| chr14_+_75536280 | 0.03 |
ENST00000238686.8 |
ZC2HC1C |
zinc finger, C2HC-type containing 1C |
| chr17_+_7517264 | 0.03 |
ENST00000593717.1 ENST00000572182.1 ENST00000574539.1 ENST00000576728.1 ENST00000575314.1 ENST00000570547.1 ENST00000572262.1 ENST00000576478.1 |
AC007421.1 SHBG |
AC007421.1 sex hormone-binding globulin |
| chr9_+_15422702 | 0.03 |
ENST00000380821.3 ENST00000421710.1 |
SNAPC3 |
small nuclear RNA activating complex, polypeptide 3, 50kDa |
| chr1_+_29063439 | 0.03 |
ENST00000541996.1 ENST00000496288.1 |
YTHDF2 |
YTH domain family, member 2 |
| chr12_+_56618102 | 0.03 |
ENST00000267023.4 ENST00000380198.2 ENST00000341463.5 |
NABP2 |
nucleic acid binding protein 2 |
| chr1_-_223537475 | 0.03 |
ENST00000344029.6 ENST00000494793.2 ENST00000366878.4 ENST00000366877.3 |
SUSD4 |
sushi domain containing 4 |
| chr15_+_44069276 | 0.03 |
ENST00000381359.1 |
SERF2 |
small EDRK-rich factor 2 |
| chr12_+_56360605 | 0.03 |
ENST00000553376.1 ENST00000440311.2 ENST00000354056.4 |
CDK2 |
cyclin-dependent kinase 2 |
| chr1_+_237205476 | 0.03 |
ENST00000366574.2 |
RYR2 |
ryanodine receptor 2 (cardiac) |
| chr14_-_77737543 | 0.03 |
ENST00000298352.4 |
NGB |
neuroglobin |
| chr14_+_75536335 | 0.03 |
ENST00000554763.1 ENST00000439583.2 ENST00000526130.1 ENST00000525046.1 |
ZC2HC1C |
zinc finger, C2HC-type containing 1C |
| chr1_-_21503337 | 0.03 |
ENST00000400422.1 ENST00000602326.1 ENST00000411888.1 ENST00000438975.1 |
EIF4G3 |
eukaryotic translation initiation factor 4 gamma, 3 |
| chr12_+_50344516 | 0.03 |
ENST00000199280.3 ENST00000550862.1 |
AQP2 |
aquaporin 2 (collecting duct) |
| chr5_+_122847781 | 0.02 |
ENST00000395412.1 ENST00000395411.1 ENST00000345990.4 |
CSNK1G3 |
casein kinase 1, gamma 3 |
| chr10_-_102790852 | 0.02 |
ENST00000470414.1 ENST00000370215.3 |
PDZD7 |
PDZ domain containing 7 |
| chr19_-_2236290 | 0.02 |
ENST00000591099.2 ENST00000586608.2 ENST00000326631.2 ENST00000587962.2 |
PLEKHJ1 |
pleckstrin homology domain containing, family J member 1 |
| chr5_-_32313019 | 0.02 |
ENST00000280285.5 ENST00000264934.5 |
MTMR12 |
myotubularin related protein 12 |
| chrX_+_128674213 | 0.01 |
ENST00000371113.4 ENST00000357121.5 |
OCRL |
oculocerebrorenal syndrome of Lowe |
| chr7_+_99933730 | 0.01 |
ENST00000610247.1 |
PILRB |
paired immunoglobin-like type 2 receptor beta |
| chrX_-_71526999 | 0.01 |
ENST00000453707.2 ENST00000373619.3 |
CITED1 |
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 1 |
| chr2_-_40739501 | 0.01 |
ENST00000403092.1 ENST00000402441.1 ENST00000448531.1 |
SLC8A1 |
solute carrier family 8 (sodium/calcium exchanger), member 1 |
| chr19_+_19626531 | 0.01 |
ENST00000507754.4 |
NDUFA13 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 13 |
| chr8_+_134125727 | 0.01 |
ENST00000521107.1 |
TG |
thyroglobulin |
| chr19_-_51531210 | 0.01 |
ENST00000391804.3 |
KLK11 |
kallikrein-related peptidase 11 |
| chr15_-_48470558 | 0.00 |
ENST00000324324.7 |
MYEF2 |
myelin expression factor 2 |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0005171 | hepatocyte growth factor receptor binding(GO:0005171) |
| 0.2 | 0.8 | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor(GO:0004748) oxidoreductase activity, acting on CH or CH2 groups, disulfide as acceptor(GO:0016728) ribonucleoside-diphosphate reductase activity(GO:0061731) |
| 0.2 | 0.5 | GO:0036487 | nitric-oxide synthase inhibitor activity(GO:0036487) |
| 0.1 | 1.5 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.1 | 3.6 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 0.4 | GO:0005026 | transforming growth factor beta receptor activity, type II(GO:0005026) |
| 0.1 | 0.3 | GO:0042799 | histone methyltransferase activity (H4-K20 specific)(GO:0042799) |
| 0.1 | 0.3 | GO:0097158 | pre-mRNA intronic pyrimidine-rich binding(GO:0097158) |
| 0.1 | 0.3 | GO:0086020 | gap junction channel activity involved in SA node cell-atrial cardiac muscle cell electrical coupling(GO:0086020) gap junction channel activity involved in AV node cell-bundle of His cell electrical coupling(GO:0086077) |
| 0.1 | 0.7 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.1 | 0.5 | GO:0032138 | single base insertion or deletion binding(GO:0032138) |
| 0.1 | 0.2 | GO:0050528 | acyloxyacyl hydrolase activity(GO:0050528) |
| 0.1 | 1.6 | GO:0070410 | co-SMAD binding(GO:0070410) |
| 0.1 | 0.4 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.0 | 0.3 | GO:1904929 | coreceptor activity involved in Wnt signaling pathway, planar cell polarity pathway(GO:1904929) |
| 0.0 | 0.4 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 0.0 | 0.9 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 0.3 | GO:0050833 | pyruvate transmembrane transporter activity(GO:0050833) |
| 0.0 | 0.6 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.0 | 0.2 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.5 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 0.2 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.0 | 0.5 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.2 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.0 | 0.2 | GO:0003998 | acylphosphatase activity(GO:0003998) |
| 0.0 | 0.6 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.0 | 0.7 | GO:0046961 | proton-transporting ATPase activity, rotational mechanism(GO:0046961) |
| 0.0 | 1.9 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 1.3 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.0 | 0.1 | GO:0005166 | neurotrophin p75 receptor binding(GO:0005166) |
| 0.0 | 0.1 | GO:0043812 | phosphatidylinositol-4-phosphate phosphatase activity(GO:0043812) |
| 0.0 | 0.4 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.0 | 0.6 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.0 | 0.9 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.2 | GO:0051378 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 1.0 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.3 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.0 | 0.2 | GO:0048531 | beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.0 | 0.0 | GO:0032422 | purine-rich negative regulatory element binding(GO:0032422) |
| 0.0 | 0.2 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 0.2 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 0.0 | 0.3 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.0 | 0.2 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.1 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.0 | 0.2 | GO:0097602 | cullin family protein binding(GO:0097602) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.6 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 1.9 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 1.0 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 2.2 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 0.6 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.0 | 1.1 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 1.6 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 0.4 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.0 | 0.2 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.0 | 0.4 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.6 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 0.9 | REACTOME RECEPTOR LIGAND BINDING INITIATES THE SECOND PROTEOLYTIC CLEAVAGE OF NOTCH RECEPTOR | Genes involved in Receptor-ligand binding initiates the second proteolytic cleavage of Notch receptor |
| 0.1 | 2.3 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.7 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 1.5 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 0.5 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.5 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.0 | 0.7 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.0 | 1.4 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.4 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.0 | 0.3 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 0.9 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.3 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.2 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.3 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.0 | 0.2 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.0 | 0.5 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.4 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 0.1 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.0 | 0.3 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0005971 | ribonucleoside-diphosphate reductase complex(GO:0005971) |
| 0.2 | 0.5 | GO:0097453 | mesaxon(GO:0097453) ensheathing process(GO:1990015) |
| 0.1 | 0.4 | GO:0070695 | FHF complex(GO:0070695) |
| 0.1 | 0.5 | GO:0034750 | Scrib-APC-beta-catenin complex(GO:0034750) |
| 0.1 | 0.6 | GO:0072669 | tRNA-splicing ligase complex(GO:0072669) |
| 0.1 | 1.9 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.1 | 0.6 | GO:0005638 | lamin filament(GO:0005638) |
| 0.1 | 1.1 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.1 | 0.5 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.4 | GO:0070022 | transforming growth factor beta receptor homodimeric complex(GO:0070022) |
| 0.0 | 0.5 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 0.6 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.0 | 0.7 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 0.2 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 0.4 | GO:0070578 | RISC-loading complex(GO:0070578) |
| 0.0 | 0.3 | GO:0097487 | multivesicular body, internal vesicle(GO:0097487) |
| 0.0 | 3.6 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 0.1 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.0 | 0.2 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.0 | 0.2 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.0 | 0.2 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.0 | 0.3 | GO:0030663 | COPI-coated vesicle membrane(GO:0030663) |
| 0.0 | 0.5 | GO:0097228 | sperm principal piece(GO:0097228) |
| 0.0 | 0.3 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.4 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.0 | 0.3 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.1 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.0 | 0.1 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.0 | 0.1 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.0 | 0.4 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.1 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.0 | 0.9 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 0.4 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 0.3 | GO:0036019 | endolysosome(GO:0036019) |
| 0.0 | 0.3 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 0.3 | GO:0031304 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.9 | GO:0044791 | modulation by host of viral release from host cell(GO:0044789) positive regulation by host of viral release from host cell(GO:0044791) |
| 0.2 | 0.5 | GO:1903249 | regulation of cellular amine catabolic process(GO:0033241) negative regulation of cellular amine catabolic process(GO:0033242) negative regulation of the force of heart contraction(GO:0098736) regulation of arginine catabolic process(GO:1900081) negative regulation of arginine catabolic process(GO:1900082) regulation of citrulline biosynthetic process(GO:1903248) negative regulation of citrulline biosynthetic process(GO:1903249) negative regulation of cellular amino acid biosynthetic process(GO:2000283) |
| 0.2 | 0.3 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.2 | 0.8 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.2 | 0.5 | GO:0098968 | neurotransmitter receptor transport postsynaptic membrane to endosome(GO:0098968) |
| 0.1 | 3.6 | GO:0030208 | dermatan sulfate biosynthetic process(GO:0030208) |
| 0.1 | 1.0 | GO:1902202 | regulation of hepatocyte growth factor receptor signaling pathway(GO:1902202) |
| 0.1 | 0.4 | GO:0002663 | B cell tolerance induction(GO:0002514) regulation of B cell tolerance induction(GO:0002661) positive regulation of B cell tolerance induction(GO:0002663) |
| 0.1 | 1.0 | GO:0009912 | auditory receptor cell fate commitment(GO:0009912) inner ear receptor cell fate commitment(GO:0060120) |
| 0.1 | 0.6 | GO:0035105 | sterol regulatory element binding protein import into nucleus(GO:0035105) |
| 0.1 | 1.0 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.1 | 0.3 | GO:0086053 | SA node cell to atrial cardiac muscle cell communication by electrical coupling(GO:0086021) AV node cell to bundle of His cell communication by electrical coupling(GO:0086053) |
| 0.1 | 0.9 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.1 | 0.2 | GO:0097116 | gephyrin clustering involved in postsynaptic density assembly(GO:0097116) |
| 0.1 | 0.3 | GO:0048200 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 0.1 | 0.3 | GO:0006850 | mitochondrial pyruvate transport(GO:0006850) mitochondrial pyruvate transmembrane transport(GO:1902361) |
| 0.1 | 0.2 | GO:0060392 | negative regulation of SMAD protein import into nucleus(GO:0060392) |
| 0.1 | 0.5 | GO:0060666 | dichotomous subdivision of terminal units involved in salivary gland branching(GO:0060666) |
| 0.0 | 0.5 | GO:0010265 | SCF complex assembly(GO:0010265) |
| 0.0 | 0.4 | GO:0060510 | Type II pneumocyte differentiation(GO:0060510) |
| 0.0 | 2.3 | GO:0015949 | nucleobase-containing small molecule interconversion(GO:0015949) |
| 0.0 | 0.5 | GO:0048711 | positive regulation of astrocyte differentiation(GO:0048711) |
| 0.0 | 0.3 | GO:0000414 | regulation of histone H3-K36 methylation(GO:0000414) |
| 0.0 | 0.3 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.0 | 0.7 | GO:0006853 | carnitine shuttle(GO:0006853) |
| 0.0 | 0.2 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.0 | 0.6 | GO:0090129 | positive regulation of synapse maturation(GO:0090129) |
| 0.0 | 0.2 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.0 | 0.3 | GO:0071787 | endoplasmic reticulum tubular network assembly(GO:0071787) |
| 0.0 | 0.2 | GO:0060501 | positive regulation of epithelial cell proliferation involved in lung morphogenesis(GO:0060501) |
| 0.0 | 0.4 | GO:0033210 | leptin-mediated signaling pathway(GO:0033210) |
| 0.0 | 0.1 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.0 | 0.4 | GO:0030422 | production of siRNA involved in RNA interference(GO:0030422) |
| 0.0 | 1.2 | GO:0050873 | brown fat cell differentiation(GO:0050873) |
| 0.0 | 0.3 | GO:0097167 | circadian regulation of translation(GO:0097167) |
| 0.0 | 0.0 | GO:0098904 | regulation of AV node cell action potential(GO:0098904) |
| 0.0 | 0.3 | GO:0035095 | behavioral response to nicotine(GO:0035095) |
| 0.0 | 0.5 | GO:0016446 | somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.0 | 0.3 | GO:2000680 | rubidium ion transport(GO:0035826) regulation of rubidium ion transport(GO:2000680) |
| 0.0 | 0.1 | GO:0097155 | fasciculation of sensory neuron axon(GO:0097155) |
| 0.0 | 0.2 | GO:0060754 | positive regulation of mast cell chemotaxis(GO:0060754) |
| 0.0 | 0.7 | GO:0090383 | phagosome acidification(GO:0090383) |
| 0.0 | 0.5 | GO:0007597 | blood coagulation, intrinsic pathway(GO:0007597) |
| 0.0 | 0.1 | GO:0001826 | inner cell mass cell differentiation(GO:0001826) |
| 0.0 | 0.1 | GO:0009386 | translational attenuation(GO:0009386) |
| 0.0 | 0.1 | GO:2000507 | positive regulation of energy homeostasis(GO:2000507) |
| 0.0 | 0.1 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.0 | 0.4 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.0 | 0.0 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 0.0 | 0.1 | GO:0006789 | bilirubin conjugation(GO:0006789) |
| 0.0 | 0.2 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.0 | 0.1 | GO:1903679 | regulation of cap-independent translational initiation(GO:1903677) positive regulation of cap-independent translational initiation(GO:1903679) regulation of cytoplasmic translational initiation(GO:1904688) positive regulation of cytoplasmic translational initiation(GO:1904690) |
| 0.0 | 0.2 | GO:0014041 | regulation of neuron maturation(GO:0014041) |
| 0.0 | 0.6 | GO:2000300 | regulation of synaptic vesicle exocytosis(GO:2000300) |
| 0.0 | 0.2 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 0.0 | 0.1 | GO:0031118 | rRNA pseudouridine synthesis(GO:0031118) |
| 0.0 | 0.0 | GO:0034653 | diterpenoid catabolic process(GO:0016103) retinoic acid catabolic process(GO:0034653) |
| 0.0 | 0.1 | GO:0097688 | AMPA glutamate receptor clustering(GO:0097113) glutamate receptor clustering(GO:0097688) |
| 0.0 | 0.4 | GO:0045022 | early endosome to late endosome transport(GO:0045022) |
| 0.0 | 0.2 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.1 | GO:0032468 | Golgi calcium ion homeostasis(GO:0032468) |
| 0.0 | 0.2 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 0.1 | GO:0071105 | response to interleukin-11(GO:0071105) |
| 0.0 | 0.2 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.0 | 0.1 | GO:0071550 | death-inducing signaling complex assembly(GO:0071550) |