Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
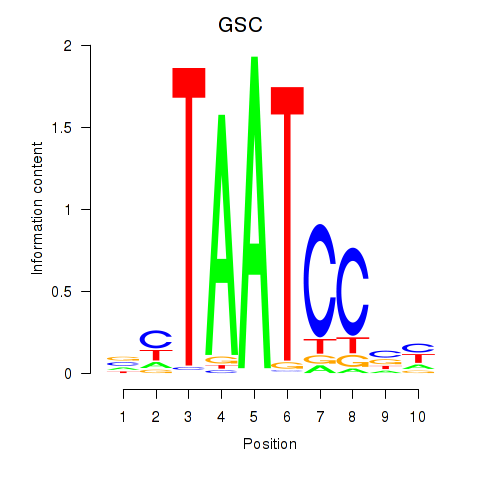
Results for GSC_GSC2
Z-value: 1.39
Transcription factors associated with GSC_GSC2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
GSC
|
ENSG00000133937.3 | GSC |
|
GSC2
|
ENSG00000063515.2 | GSC2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| GSC2 | hg19_v2_chr22_-_19137796_19137796 | 0.11 | 6.9e-01 | Click! |
Activity profile of GSC_GSC2 motif
Sorted Z-values of GSC_GSC2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of GSC_GSC2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_126716459 | 15.73 |
ENST00000309035.6 |
CTBP2 |
C-terminal binding protein 2 |
| chr2_-_175711133 | 7.58 |
ENST00000409597.1 ENST00000413882.1 |
CHN1 |
chimerin 1 |
| chr4_+_169552748 | 5.97 |
ENST00000504519.1 ENST00000512127.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr10_-_126849588 | 5.56 |
ENST00000411419.2 |
CTBP2 |
C-terminal binding protein 2 |
| chrX_+_135279179 | 5.52 |
ENST00000370676.3 |
FHL1 |
four and a half LIM domains 1 |
| chrX_-_10588459 | 4.61 |
ENST00000380782.2 |
MID1 |
midline 1 (Opitz/BBB syndrome) |
| chrX_-_10588595 | 4.27 |
ENST00000423614.1 ENST00000317552.4 |
MID1 |
midline 1 (Opitz/BBB syndrome) |
| chr9_+_116298778 | 3.32 |
ENST00000462143.1 |
RGS3 |
regulator of G-protein signaling 3 |
| chr5_+_76114758 | 3.20 |
ENST00000514165.1 ENST00000296677.4 |
F2RL1 |
coagulation factor II (thrombin) receptor-like 1 |
| chr6_+_121756809 | 2.99 |
ENST00000282561.3 |
GJA1 |
gap junction protein, alpha 1, 43kDa |
| chr8_+_70378852 | 2.96 |
ENST00000525061.1 ENST00000458141.2 ENST00000260128.4 |
SULF1 |
sulfatase 1 |
| chrX_-_48931648 | 2.94 |
ENST00000376386.3 ENST00000376390.4 |
PRAF2 |
PRA1 domain family, member 2 |
| chrX_+_100333709 | 2.94 |
ENST00000372930.4 |
TMEM35 |
transmembrane protein 35 |
| chr9_-_13279563 | 2.60 |
ENST00000541718.1 |
MPDZ |
multiple PDZ domain protein |
| chr13_+_73632897 | 2.57 |
ENST00000377687.4 |
KLF5 |
Kruppel-like factor 5 (intestinal) |
| chr14_+_96722152 | 2.45 |
ENST00000216629.6 |
BDKRB1 |
bradykinin receptor B1 |
| chr12_-_77272765 | 2.18 |
ENST00000547435.1 ENST00000552330.1 ENST00000546966.1 ENST00000311083.5 |
CSRP2 |
cysteine and glycine-rich protein 2 |
| chr2_+_192141611 | 2.11 |
ENST00000392316.1 |
MYO1B |
myosin IB |
| chrX_+_135278908 | 2.11 |
ENST00000539015.1 ENST00000370683.1 |
FHL1 |
four and a half LIM domains 1 |
| chr9_-_13279589 | 2.06 |
ENST00000319217.7 |
MPDZ |
multiple PDZ domain protein |
| chr1_+_114522049 | 1.59 |
ENST00000369551.1 ENST00000320334.4 |
OLFML3 |
olfactomedin-like 3 |
| chrX_+_99899180 | 1.49 |
ENST00000373004.3 |
SRPX2 |
sushi-repeat containing protein, X-linked 2 |
| chr15_-_52483566 | 1.39 |
ENST00000261837.7 |
GNB5 |
guanine nucleotide binding protein (G protein), beta 5 |
| chr11_-_111794446 | 1.39 |
ENST00000527950.1 |
CRYAB |
crystallin, alpha B |
| chr9_+_103204553 | 1.38 |
ENST00000502978.1 ENST00000334943.6 |
MSANTD3-TMEFF1 TMEFF1 |
MSANTD3-TMEFF1 readthrough transmembrane protein with EGF-like and two follistatin-like domains 1 |
| chr11_-_111783595 | 1.36 |
ENST00000528628.1 |
CRYAB |
crystallin, alpha B |
| chr19_-_36247910 | 1.30 |
ENST00000587965.1 ENST00000004982.3 |
HSPB6 |
heat shock protein, alpha-crystallin-related, B6 |
| chr5_-_16738451 | 1.15 |
ENST00000274203.9 ENST00000515803.1 |
MYO10 |
myosin X |
| chr5_+_140868717 | 1.04 |
ENST00000252087.1 |
PCDHGC5 |
protocadherin gamma subfamily C, 5 |
| chr1_+_169079823 | 1.02 |
ENST00000367813.3 |
ATP1B1 |
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr1_+_17248418 | 1.02 |
ENST00000375541.5 |
CROCC |
ciliary rootlet coiled-coil, rootletin |
| chr2_-_37899323 | 1.00 |
ENST00000295324.3 ENST00000457889.1 |
CDC42EP3 |
CDC42 effector protein (Rho GTPase binding) 3 |
| chr6_+_35310312 | 0.99 |
ENST00000448077.2 ENST00000360694.3 ENST00000418635.2 ENST00000444397.1 |
PPARD |
peroxisome proliferator-activated receptor delta |
| chr2_+_189156389 | 0.96 |
ENST00000409843.1 |
GULP1 |
GULP, engulfment adaptor PTB domain containing 1 |
| chr9_+_17134980 | 0.91 |
ENST00000380647.3 |
CNTLN |
centlein, centrosomal protein |
| chr1_+_244816371 | 0.89 |
ENST00000263831.7 |
DESI2 |
desumoylating isopeptidase 2 |
| chr1_+_244816237 | 0.86 |
ENST00000302550.11 |
DESI2 |
desumoylating isopeptidase 2 |
| chr15_+_96876340 | 0.86 |
ENST00000453270.2 |
NR2F2 |
nuclear receptor subfamily 2, group F, member 2 |
| chrX_+_107069063 | 0.80 |
ENST00000262843.6 |
MID2 |
midline 2 |
| chr11_+_76156045 | 0.79 |
ENST00000533988.1 ENST00000524490.1 ENST00000334736.3 ENST00000343878.3 ENST00000533972.1 |
C11orf30 |
chromosome 11 open reading frame 30 |
| chr16_-_3074231 | 0.75 |
ENST00000572355.1 ENST00000248089.3 ENST00000574980.1 ENST00000354679.3 ENST00000396916.1 ENST00000573842.1 |
HCFC1R1 |
host cell factor C1 regulator 1 (XPO1 dependent) |
| chr1_+_20396649 | 0.71 |
ENST00000375108.3 |
PLA2G5 |
phospholipase A2, group V |
| chr18_-_64271363 | 0.68 |
ENST00000262150.2 |
CDH19 |
cadherin 19, type 2 |
| chr12_+_104680659 | 0.67 |
ENST00000526691.1 ENST00000531691.1 ENST00000388854.3 ENST00000354940.6 ENST00000526390.1 ENST00000531689.1 |
TXNRD1 |
thioredoxin reductase 1 |
| chr8_-_116681221 | 0.67 |
ENST00000395715.3 |
TRPS1 |
trichorhinophalangeal syndrome I |
| chr6_+_35310391 | 0.64 |
ENST00000337400.2 ENST00000311565.4 ENST00000540939.1 |
PPARD |
peroxisome proliferator-activated receptor delta |
| chr9_+_17135016 | 0.62 |
ENST00000425824.1 ENST00000262360.5 ENST00000380641.4 |
CNTLN |
centlein, centrosomal protein |
| chr3_+_49058444 | 0.61 |
ENST00000326925.6 ENST00000395458.2 |
NDUFAF3 |
NADH dehydrogenase (ubiquinone) complex I, assembly factor 3 |
| chr17_+_44668035 | 0.60 |
ENST00000398238.4 ENST00000225282.8 |
NSF |
N-ethylmaleimide-sensitive factor |
| chr11_+_124735282 | 0.60 |
ENST00000397801.1 |
ROBO3 |
roundabout, axon guidance receptor, homolog 3 (Drosophila) |
| chrX_-_6146876 | 0.60 |
ENST00000381095.3 |
NLGN4X |
neuroligin 4, X-linked |
| chr11_-_67120974 | 0.57 |
ENST00000539074.1 ENST00000312419.3 |
POLD4 |
polymerase (DNA-directed), delta 4, accessory subunit |
| chr20_+_62496596 | 0.51 |
ENST00000369927.4 ENST00000346249.4 ENST00000348257.5 ENST00000352482.4 ENST00000351424.4 ENST00000217121.5 ENST00000358548.4 |
TPD52L2 |
tumor protein D52-like 2 |
| chr22_-_29663954 | 0.51 |
ENST00000216085.7 |
RHBDD3 |
rhomboid domain containing 3 |
| chr2_+_189156721 | 0.49 |
ENST00000409927.1 ENST00000409805.1 |
GULP1 |
GULP, engulfment adaptor PTB domain containing 1 |
| chr16_+_81272287 | 0.48 |
ENST00000425577.2 ENST00000564552.1 |
BCMO1 |
beta-carotene 15,15'-monooxygenase 1 |
| chr7_-_128415844 | 0.48 |
ENST00000249389.2 |
OPN1SW |
opsin 1 (cone pigments), short-wave-sensitive |
| chr19_+_2476116 | 0.42 |
ENST00000215631.4 ENST00000587345.1 |
GADD45B |
growth arrest and DNA-damage-inducible, beta |
| chr2_+_189156586 | 0.42 |
ENST00000409830.1 |
GULP1 |
GULP, engulfment adaptor PTB domain containing 1 |
| chrX_+_100878079 | 0.40 |
ENST00000471229.2 |
ARMCX3 |
armadillo repeat containing, X-linked 3 |
| chr13_+_114321463 | 0.40 |
ENST00000335678.6 |
GRK1 |
G protein-coupled receptor kinase 1 |
| chr12_+_81101277 | 0.39 |
ENST00000228641.3 |
MYF6 |
myogenic factor 6 (herculin) |
| chr16_-_29875057 | 0.36 |
ENST00000219789.6 |
CDIPT |
CDP-diacylglycerol--inositol 3-phosphatidyltransferase |
| chr19_+_40877583 | 0.35 |
ENST00000596470.1 |
PLD3 |
phospholipase D family, member 3 |
| chr3_+_36421826 | 0.35 |
ENST00000273183.3 |
STAC |
SH3 and cysteine rich domain |
| chr19_-_10121144 | 0.34 |
ENST00000264828.3 |
COL5A3 |
collagen, type V, alpha 3 |
| chr19_-_39881669 | 0.33 |
ENST00000221266.7 |
PAF1 |
Paf1, RNA polymerase II associated factor, homolog (S. cerevisiae) |
| chr10_-_86001210 | 0.33 |
ENST00000372105.3 |
LRIT1 |
leucine-rich repeat, immunoglobulin-like and transmembrane domains 1 |
| chr4_+_70916119 | 0.31 |
ENST00000246896.3 ENST00000511674.1 |
HTN1 |
histatin 1 |
| chr16_-_29874462 | 0.30 |
ENST00000566113.1 ENST00000569956.1 ENST00000570016.1 |
CDIPT |
CDP-diacylglycerol--inositol 3-phosphatidyltransferase |
| chr11_-_65625330 | 0.28 |
ENST00000531407.1 |
CFL1 |
cofilin 1 (non-muscle) |
| chr10_+_74451883 | 0.28 |
ENST00000373053.3 ENST00000357157.6 |
MCU |
mitochondrial calcium uniporter |
| chr3_-_170626418 | 0.27 |
ENST00000474096.1 ENST00000295822.2 |
EIF5A2 |
eukaryotic translation initiation factor 5A2 |
| chr11_+_33563821 | 0.26 |
ENST00000321505.4 ENST00000265654.5 ENST00000389726.3 |
KIAA1549L |
KIAA1549-like |
| chr11_-_62783276 | 0.26 |
ENST00000535878.1 ENST00000545207.1 |
SLC22A8 |
solute carrier family 22 (organic anion transporter), member 8 |
| chr13_-_36050819 | 0.25 |
ENST00000379919.4 |
MAB21L1 |
mab-21-like 1 (C. elegans) |
| chr8_-_87755878 | 0.24 |
ENST00000320005.5 |
CNGB3 |
cyclic nucleotide gated channel beta 3 |
| chr6_+_111580508 | 0.23 |
ENST00000368847.4 |
KIAA1919 |
KIAA1919 |
| chr10_+_52750930 | 0.23 |
ENST00000401604.2 |
PRKG1 |
protein kinase, cGMP-dependent, type I |
| chr11_-_65624415 | 0.22 |
ENST00000524553.1 ENST00000527344.1 |
CFL1 |
cofilin 1 (non-muscle) |
| chr11_+_44587141 | 0.21 |
ENST00000227155.4 ENST00000342935.3 ENST00000532544.1 |
CD82 |
CD82 molecule |
| chr6_+_31588478 | 0.21 |
ENST00000376007.4 ENST00000376033.2 |
PRRC2A |
proline-rich coiled-coil 2A |
| chr17_+_77020325 | 0.16 |
ENST00000311661.4 |
C1QTNF1 |
C1q and tumor necrosis factor related protein 1 |
| chrX_-_24665353 | 0.15 |
ENST00000379144.2 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr10_+_52751010 | 0.15 |
ENST00000373985.1 |
PRKG1 |
protein kinase, cGMP-dependent, type I |
| chr1_-_36615065 | 0.14 |
ENST00000373166.3 ENST00000373159.1 ENST00000373162.1 |
TRAPPC3 |
trafficking protein particle complex 3 |
| chr6_-_41130914 | 0.14 |
ENST00000373113.3 ENST00000338469.3 |
TREM2 |
triggering receptor expressed on myeloid cells 2 |
| chr17_+_77020224 | 0.13 |
ENST00000339142.2 |
C1QTNF1 |
C1q and tumor necrosis factor related protein 1 |
| chr7_-_97881429 | 0.13 |
ENST00000420697.1 ENST00000379795.3 ENST00000415086.1 ENST00000542604.1 ENST00000447648.2 |
TECPR1 |
tectonin beta-propeller repeat containing 1 |
| chr10_-_35104185 | 0.13 |
ENST00000374789.3 ENST00000374788.3 ENST00000346874.4 ENST00000374794.3 ENST00000350537.4 ENST00000374790.3 ENST00000374776.1 ENST00000374773.1 ENST00000545693.1 ENST00000545260.1 ENST00000340077.5 |
PARD3 |
par-3 family cell polarity regulator |
| chrX_-_24665208 | 0.12 |
ENST00000356768.4 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr6_-_41130841 | 0.12 |
ENST00000373122.4 |
TREM2 |
triggering receptor expressed on myeloid cells 2 |
| chr2_+_189156638 | 0.11 |
ENST00000410051.1 |
GULP1 |
GULP, engulfment adaptor PTB domain containing 1 |
| chr5_-_150138061 | 0.10 |
ENST00000521533.1 ENST00000424236.1 |
DCTN4 |
dynactin 4 (p62) |
| chrX_-_54384425 | 0.09 |
ENST00000375169.3 ENST00000354646.2 |
WNK3 |
WNK lysine deficient protein kinase 3 |
| chr11_-_62783303 | 0.09 |
ENST00000336232.2 ENST00000430500.2 |
SLC22A8 |
solute carrier family 22 (organic anion transporter), member 8 |
| chr22_+_29664305 | 0.09 |
ENST00000414183.2 ENST00000333395.6 ENST00000455726.1 ENST00000332035.6 |
EWSR1 |
EWS RNA-binding protein 1 |
| chr17_-_48207157 | 0.08 |
ENST00000330175.4 ENST00000503131.1 |
SAMD14 |
sterile alpha motif domain containing 14 |
| chrY_+_16634483 | 0.08 |
ENST00000382872.1 |
NLGN4Y |
neuroligin 4, Y-linked |
| chr4_+_70894130 | 0.06 |
ENST00000526767.1 ENST00000530128.1 ENST00000381057.3 |
HTN3 |
histatin 3 |
| chr1_-_47655686 | 0.06 |
ENST00000294338.2 |
PDZK1IP1 |
PDZK1 interacting protein 1 |
| chr1_-_160313025 | 0.05 |
ENST00000368069.3 ENST00000241704.7 |
COPA |
coatomer protein complex, subunit alpha |
| chr1_-_165414414 | 0.04 |
ENST00000359842.5 |
RXRG |
retinoid X receptor, gamma |
| chr1_-_43638168 | 0.04 |
ENST00000431635.2 |
EBNA1BP2 |
EBNA1 binding protein 2 |
| chr6_-_35480705 | 0.03 |
ENST00000229771.6 |
TULP1 |
tubby like protein 1 |
| chr1_+_18957500 | 0.03 |
ENST00000375375.3 |
PAX7 |
paired box 7 |
| chr4_+_170541660 | 0.03 |
ENST00000513761.1 ENST00000347613.4 |
CLCN3 |
chloride channel, voltage-sensitive 3 |
| chr2_-_30144432 | 0.03 |
ENST00000389048.3 |
ALK |
anaplastic lymphoma receptor tyrosine kinase |
| chr16_-_68269971 | 0.03 |
ENST00000565858.1 |
ESRP2 |
epithelial splicing regulatory protein 2 |
| chr12_-_113574028 | 0.01 |
ENST00000546530.1 ENST00000261729.5 |
RASAL1 |
RAS protein activator like 1 (GAP1 like) |
| chr15_+_58430567 | 0.01 |
ENST00000536493.1 |
AQP9 |
aquaporin 9 |
| chrX_-_49089771 | 0.01 |
ENST00000376251.1 ENST00000323022.5 ENST00000376265.2 |
CACNA1F |
calcium channel, voltage-dependent, L type, alpha 1F subunit |
| chr15_-_31453359 | 0.01 |
ENST00000542188.1 |
TRPM1 |
transient receptor potential cation channel, subfamily M, member 1 |
| chr15_-_31393910 | 0.01 |
ENST00000397795.2 ENST00000256552.6 ENST00000559179.1 |
TRPM1 |
transient receptor potential cation channel, subfamily M, member 1 |
| chr18_-_24443151 | 0.00 |
ENST00000440832.3 |
AQP4 |
aquaporin 4 |
Gene Ontology Analysis
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 21.3 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.2 | 4.7 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.2 | 3.0 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.2 | 2.8 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.1 | 0.6 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.1 | 8.9 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 3.2 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.1 | 0.3 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 6.0 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 1.0 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.1 | 0.3 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.1 | 1.2 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.1 | 1.0 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.0 | 0.8 | GO:0097381 | photoreceptor disc membrane(GO:0097381) |
| 0.0 | 0.3 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.0 | 2.1 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 3.6 | GO:0005795 | Golgi stack(GO:0005795) |
| 0.0 | 0.1 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.0 | 9.7 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 0.4 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.0 | 0.1 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.0 | 1.5 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 0.7 | GO:0001917 | photoreceptor inner segment(GO:0001917) |
| 0.0 | 0.1 | GO:0005869 | dynactin complex(GO:0005869) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.0 | 21.3 | GO:0016618 | hydroxypyruvate reductase activity(GO:0016618) glyoxylate reductase (NADP) activity(GO:0030267) |
| 0.8 | 2.4 | GO:0004947 | bradykinin receptor activity(GO:0004947) |
| 0.7 | 3.0 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.6 | 3.0 | GO:0086075 | gap junction channel activity involved in cardiac conduction electrical coupling(GO:0086075) |
| 0.5 | 3.2 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.3 | 6.0 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.2 | 0.7 | GO:0098626 | methylselenol reductase activity(GO:0098625) methylseleninic acid reductase activity(GO:0098626) |
| 0.2 | 0.7 | GO:0003881 | CDP-diacylglycerol-inositol 3-phosphatidyltransferase activity(GO:0003881) |
| 0.2 | 7.6 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.1 | 3.3 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.1 | 4.0 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 0.3 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.1 | 9.7 | GO:0051219 | phosphoprotein binding(GO:0051219) |
| 0.1 | 0.4 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.1 | 0.4 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.1 | 0.7 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.1 | 1.0 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.1 | 1.6 | GO:0036041 | long-chain fatty acid binding(GO:0036041) |
| 0.1 | 1.0 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.1 | 0.3 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.0 | 0.7 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 7.6 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 0.5 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.3 | GO:0070891 | lipoteichoic acid binding(GO:0070891) |
| 0.0 | 0.6 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.6 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 1.0 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 0.0 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.0 | 1.4 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 0.5 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 0.3 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.0 | 2.9 | GO:0008022 | protein C-terminus binding(GO:0008022) |
| 0.0 | 0.3 | GO:0003746 | translation elongation factor activity(GO:0003746) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.0 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 0.6 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.0 | 0.7 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.0 | 0.5 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 3.4 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 6.2 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 5.6 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 1.0 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.6 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.0 | 0.7 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 1.2 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.6 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 0.5 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.0 | 1.3 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 1.3 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.3 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.4 | 21.3 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 1.5 | 6.2 | GO:0003104 | positive regulation of glomerular filtration(GO:0003104) |
| 0.7 | 9.7 | GO:0035372 | protein localization to microtubule(GO:0035372) |
| 0.3 | 3.0 | GO:0060686 | negative regulation of prostatic bud formation(GO:0060686) |
| 0.3 | 2.6 | GO:0032534 | regulation of microvillus assembly(GO:0032534) |
| 0.3 | 6.0 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.3 | 1.0 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) positive regulation of potassium ion import(GO:1903288) |
| 0.3 | 1.0 | GO:0032053 | ciliary basal body organization(GO:0032053) positive regulation of protein localization to cilium(GO:1903566) |
| 0.2 | 7.6 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.2 | 0.9 | GO:0009956 | radial pattern formation(GO:0009956) |
| 0.2 | 0.7 | GO:0006663 | platelet activating factor biosynthetic process(GO:0006663) |
| 0.2 | 1.6 | GO:0032966 | negative regulation of collagen biosynthetic process(GO:0032966) |
| 0.2 | 2.8 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.1 | 1.5 | GO:0090050 | positive regulation of cell migration involved in sprouting angiogenesis(GO:0090050) |
| 0.1 | 7.6 | GO:0043268 | positive regulation of potassium ion transport(GO:0043268) |
| 0.1 | 0.5 | GO:0032815 | negative regulation of natural killer cell activation(GO:0032815) |
| 0.1 | 0.5 | GO:2000814 | regulation of unidimensional cell growth(GO:0051510) negative regulation of unidimensional cell growth(GO:0051511) establishment of cell polarity regulating cell shape(GO:0071964) regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000769) positive regulation of establishment or maintenance of cell polarity regulating cell shape(GO:2000771) regulation of establishment of cell polarity regulating cell shape(GO:2000782) positive regulation of establishment of cell polarity regulating cell shape(GO:2000784) regulation of barbed-end actin filament capping(GO:2000812) positive regulation of barbed-end actin filament capping(GO:2000814) |
| 0.1 | 1.0 | GO:0031274 | positive regulation of pseudopodium assembly(GO:0031274) |
| 0.1 | 1.5 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.1 | 0.3 | GO:0032499 | positive regulation of antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002582) positive regulation of antigen processing and presentation of peptide antigen(GO:0002585) positive regulation of antigen processing and presentation of peptide antigen via MHC class II(GO:0002588) detection of peptidoglycan(GO:0032499) |
| 0.1 | 1.4 | GO:1901386 | negative regulation of voltage-gated calcium channel activity(GO:1901386) |
| 0.1 | 0.6 | GO:0016199 | axon midline choice point recognition(GO:0016199) |
| 0.1 | 3.3 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.1 | 0.7 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 0.1 | 0.7 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.1 | 0.3 | GO:2000857 | positive regulation of mineralocorticoid secretion(GO:2000857) positive regulation of aldosterone secretion(GO:2000860) |
| 0.1 | 2.2 | GO:0015813 | L-glutamate transport(GO:0015813) |
| 0.1 | 2.6 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.3 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 1.3 | GO:0010667 | negative regulation of cardiac muscle cell apoptotic process(GO:0010667) |
| 0.0 | 0.3 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.0 | 0.7 | GO:0046341 | CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.0 | 0.4 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.0 | 0.3 | GO:0006452 | translational frameshifting(GO:0006452) positive regulation of translational termination(GO:0045905) |
| 0.0 | 0.1 | GO:2000686 | regulation of rubidium ion transmembrane transporter activity(GO:2000686) |
| 0.0 | 0.5 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.0 | 0.4 | GO:0060087 | relaxation of vascular smooth muscle(GO:0060087) |
| 0.0 | 0.3 | GO:0006657 | CDP-choline pathway(GO:0006657) |
| 0.0 | 0.4 | GO:1900745 | positive regulation of p38MAPK cascade(GO:1900745) |
| 0.0 | 0.6 | GO:0001921 | positive regulation of receptor recycling(GO:0001921) |
| 0.0 | 0.6 | GO:0032201 | telomere maintenance via semi-conservative replication(GO:0032201) |
| 0.0 | 0.7 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 2.0 | GO:0006911 | phagocytosis, engulfment(GO:0006911) |
| 0.0 | 2.1 | GO:0006892 | post-Golgi vesicle-mediated transport(GO:0006892) |
| 0.0 | 1.7 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.3 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 19.2 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.1 | 3.6 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.1 | 6.0 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.1 | 3.3 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.1 | 3.0 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 1.6 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 1.2 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 2.0 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 1.0 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 2.2 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 0.5 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 0.4 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 0.4 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.9 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 2.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |