Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
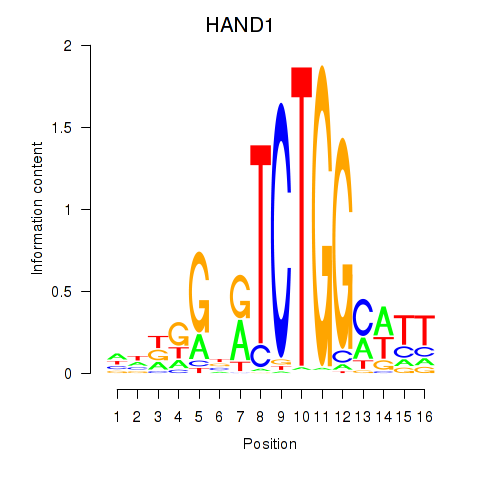
Results for HAND1
Z-value: 0.94
Transcription factors associated with HAND1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HAND1
|
ENSG00000113196.2 | HAND1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HAND1 | hg19_v2_chr5_-_153857819_153857824 | -0.60 | 1.4e-02 | Click! |
Activity profile of HAND1 motif
Sorted Z-values of HAND1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HAND1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_64216748 | 4.30 |
ENST00000585162.1 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chr2_+_228678550 | 3.53 |
ENST00000409189.3 ENST00000358813.4 |
CCL20 |
chemokine (C-C motif) ligand 20 |
| chr14_-_65409438 | 2.75 |
ENST00000557049.1 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr14_-_65409502 | 2.45 |
ENST00000389614.5 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr16_-_28621353 | 2.16 |
ENST00000567512.1 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr17_+_53342311 | 2.06 |
ENST00000226067.5 |
HLF |
hepatic leukemia factor |
| chr1_+_207262540 | 2.02 |
ENST00000452902.2 |
C4BPB |
complement component 4 binding protein, beta |
| chr16_-_28621312 | 1.98 |
ENST00000314752.7 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr1_+_207262627 | 1.91 |
ENST00000391923.1 |
C4BPB |
complement component 4 binding protein, beta |
| chr1_+_207262170 | 1.89 |
ENST00000367078.3 |
C4BPB |
complement component 4 binding protein, beta |
| chr1_+_207262578 | 1.87 |
ENST00000243611.5 ENST00000367076.3 |
C4BPB |
complement component 4 binding protein, beta |
| chr16_-_28621298 | 1.80 |
ENST00000566189.1 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr16_-_28608364 | 1.63 |
ENST00000533150.1 |
SULT1A2 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 2 |
| chr22_+_25003626 | 1.32 |
ENST00000451366.1 ENST00000406383.2 ENST00000428855.1 |
GGT1 |
gamma-glutamyltransferase 1 |
| chr16_-_28608424 | 1.25 |
ENST00000335715.4 |
SULT1A2 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 2 |
| chr3_-_114035026 | 1.24 |
ENST00000570269.1 |
RP11-553L6.5 |
RP11-553L6.5 |
| chr22_+_24999114 | 1.21 |
ENST00000412658.1 ENST00000445029.1 ENST00000419133.1 ENST00000400382.1 ENST00000438643.2 ENST00000452551.1 ENST00000400383.1 ENST00000412898.1 ENST00000400380.1 ENST00000455483.1 ENST00000430289.1 |
GGT1 |
gamma-glutamyltransferase 1 |
| chr19_-_36304201 | 0.96 |
ENST00000301175.3 |
PRODH2 |
proline dehydrogenase (oxidase) 2 |
| chr19_-_55866061 | 0.83 |
ENST00000588572.2 ENST00000593184.1 ENST00000589467.1 |
COX6B2 |
cytochrome c oxidase subunit VIb polypeptide 2 (testis) |
| chr12_+_53848505 | 0.70 |
ENST00000552819.1 ENST00000455667.3 |
PCBP2 |
poly(rC) binding protein 2 |
| chr11_+_101785727 | 0.70 |
ENST00000263468.8 |
KIAA1377 |
KIAA1377 |
| chr2_+_201997595 | 0.67 |
ENST00000470178.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr3_+_123813543 | 0.65 |
ENST00000360013.3 |
KALRN |
kalirin, RhoGEF kinase |
| chr1_+_174846570 | 0.59 |
ENST00000392064.2 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr3_-_9811595 | 0.57 |
ENST00000256460.3 |
CAMK1 |
calcium/calmodulin-dependent protein kinase I |
| chr1_-_155904187 | 0.57 |
ENST00000368321.3 ENST00000368320.3 |
KIAA0907 |
KIAA0907 |
| chr2_+_3642545 | 0.54 |
ENST00000382062.2 ENST00000236693.7 ENST00000349077.4 |
COLEC11 |
collectin sub-family member 11 |
| chr6_+_6588902 | 0.51 |
ENST00000230568.4 |
LY86 |
lymphocyte antigen 86 |
| chrX_+_49832231 | 0.49 |
ENST00000376108.3 |
CLCN5 |
chloride channel, voltage-sensitive 5 |
| chr6_+_6588316 | 0.48 |
ENST00000379953.2 |
LY86 |
lymphocyte antigen 86 |
| chr16_-_10652993 | 0.46 |
ENST00000536829.1 |
EMP2 |
epithelial membrane protein 2 |
| chr2_+_201997492 | 0.43 |
ENST00000494258.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr12_-_7261772 | 0.43 |
ENST00000545280.1 ENST00000543933.1 ENST00000545337.1 ENST00000544702.1 ENST00000266542.4 |
C1RL |
complement component 1, r subcomponent-like |
| chr1_+_52870227 | 0.42 |
ENST00000257181.9 |
PRPF38A |
pre-mRNA processing factor 38A |
| chr3_+_148415571 | 0.41 |
ENST00000497524.1 ENST00000349243.3 ENST00000542281.1 ENST00000418473.2 ENST00000404754.2 |
AGTR1 |
angiotensin II receptor, type 1 |
| chr1_+_215747118 | 0.39 |
ENST00000448333.1 |
KCTD3 |
potassium channel tetramerization domain containing 3 |
| chr17_-_43025005 | 0.39 |
ENST00000587309.1 ENST00000593135.1 ENST00000339151.4 |
KIF18B |
kinesin family member 18B |
| chr2_-_197791441 | 0.37 |
ENST00000409475.1 ENST00000354764.4 ENST00000374738.3 |
PGAP1 |
post-GPI attachment to proteins 1 |
| chr12_+_53848549 | 0.37 |
ENST00000439930.3 ENST00000548933.1 ENST00000562264.1 |
PCBP2 |
poly(rC) binding protein 2 |
| chr20_-_1165117 | 0.37 |
ENST00000381894.3 |
TMEM74B |
transmembrane protein 74B |
| chr21_+_34602377 | 0.36 |
ENST00000342101.3 ENST00000413881.1 ENST00000443073.1 |
IFNAR2 |
interferon (alpha, beta and omega) receptor 2 |
| chr12_+_121131970 | 0.36 |
ENST00000535656.1 |
MLEC |
malectin |
| chr7_-_20256965 | 0.35 |
ENST00000400331.5 ENST00000332878.4 |
MACC1 |
metastasis associated in colon cancer 1 |
| chr3_+_112051994 | 0.33 |
ENST00000473539.1 ENST00000315711.8 ENST00000383681.3 |
CD200 |
CD200 molecule |
| chr2_-_128784846 | 0.33 |
ENST00000259235.3 ENST00000357702.5 ENST00000424298.1 |
SAP130 |
Sin3A-associated protein, 130kDa |
| chr2_-_89597542 | 0.32 |
ENST00000465170.1 |
IGKV1-37 |
immunoglobulin kappa variable 1-37 (non-functional) |
| chr1_-_205601064 | 0.32 |
ENST00000357992.4 ENST00000289703.4 |
ELK4 |
ELK4, ETS-domain protein (SRF accessory protein 1) |
| chr16_+_6069586 | 0.30 |
ENST00000547372.1 |
RBFOX1 |
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr4_-_40631859 | 0.30 |
ENST00000295971.7 ENST00000319592.4 |
RBM47 |
RNA binding motif protein 47 |
| chr2_+_204732666 | 0.29 |
ENST00000295854.6 ENST00000472206.1 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
| chr11_-_72432950 | 0.29 |
ENST00000426523.1 ENST00000429686.1 |
ARAP1 |
ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 |
| chr12_+_49717019 | 0.27 |
ENST00000549275.1 ENST00000551245.1 ENST00000380327.5 ENST00000548311.1 ENST00000550346.1 ENST00000550709.1 ENST00000549534.1 ENST00000257909.3 |
TROAP |
trophinin associated protein |
| chr4_+_41614720 | 0.26 |
ENST00000509277.1 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr21_+_34602200 | 0.25 |
ENST00000382264.3 ENST00000382241.3 ENST00000404220.3 ENST00000342136.4 |
IFNAR2 |
interferon (alpha, beta and omega) receptor 2 |
| chr2_+_98262497 | 0.24 |
ENST00000258424.2 |
COX5B |
cytochrome c oxidase subunit Vb |
| chr16_+_56969284 | 0.23 |
ENST00000568358.1 |
HERPUD1 |
homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 |
| chr2_-_25475120 | 0.23 |
ENST00000380746.4 ENST00000402667.1 |
DNMT3A |
DNA (cytosine-5-)-methyltransferase 3 alpha |
| chr16_+_90089008 | 0.21 |
ENST00000268699.4 |
GAS8 |
growth arrest-specific 8 |
| chr2_-_228582709 | 0.21 |
ENST00000541617.1 ENST00000409456.2 ENST00000409287.1 ENST00000258403.3 |
SLC19A3 |
solute carrier family 19 (thiamine transporter), member 3 |
| chr5_-_150466692 | 0.21 |
ENST00000315050.7 ENST00000523338.1 ENST00000522100.1 |
TNIP1 |
TNFAIP3 interacting protein 1 |
| chr2_-_42588338 | 0.20 |
ENST00000234301.2 |
COX7A2L |
cytochrome c oxidase subunit VIIa polypeptide 2 like |
| chr1_+_28261533 | 0.20 |
ENST00000411604.1 ENST00000373888.4 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr7_+_96634850 | 0.20 |
ENST00000518156.2 |
DLX6 |
distal-less homeobox 6 |
| chr16_+_6069664 | 0.19 |
ENST00000422070.4 |
RBFOX1 |
RNA binding protein, fox-1 homolog (C. elegans) 1 |
| chr4_+_41614909 | 0.19 |
ENST00000509454.1 ENST00000396595.3 ENST00000381753.4 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chrX_+_15767971 | 0.19 |
ENST00000479740.1 ENST00000454127.2 |
CA5B |
carbonic anhydrase VB, mitochondrial |
| chr19_+_47421933 | 0.19 |
ENST00000404338.3 |
ARHGAP35 |
Rho GTPase activating protein 35 |
| chr2_+_201997676 | 0.18 |
ENST00000462763.1 ENST00000479953.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr19_+_12862486 | 0.16 |
ENST00000549706.1 |
BEST2 |
bestrophin 2 |
| chr16_+_70328680 | 0.16 |
ENST00000563206.1 ENST00000451014.3 ENST00000568625.1 |
DDX19B |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 19B |
| chr10_+_118350522 | 0.16 |
ENST00000530319.1 ENST00000527980.1 ENST00000471549.1 ENST00000534537.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr2_+_90153696 | 0.15 |
ENST00000417279.2 |
IGKV3D-15 |
immunoglobulin kappa variable 3D-15 (gene/pseudogene) |
| chr1_-_32210275 | 0.14 |
ENST00000440175.2 |
BAI2 |
brain-specific angiogenesis inhibitor 2 |
| chr10_+_118350468 | 0.14 |
ENST00000358834.4 ENST00000528052.1 ENST00000442761.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr11_-_2323290 | 0.13 |
ENST00000381153.3 |
C11orf21 |
chromosome 11 open reading frame 21 |
| chr11_-_102496063 | 0.12 |
ENST00000260228.2 |
MMP20 |
matrix metallopeptidase 20 |
| chr1_+_27719148 | 0.12 |
ENST00000374024.3 |
GPR3 |
G protein-coupled receptor 3 |
| chr10_+_72972281 | 0.10 |
ENST00000335350.6 |
UNC5B |
unc-5 homolog B (C. elegans) |
| chr11_+_2323236 | 0.10 |
ENST00000182290.4 |
TSPAN32 |
tetraspanin 32 |
| chr14_+_24422795 | 0.09 |
ENST00000313250.5 ENST00000558263.1 ENST00000543741.2 ENST00000421831.1 ENST00000397073.2 ENST00000308178.8 ENST00000382761.3 ENST00000397075.3 ENST00000397074.3 ENST00000559632.1 ENST00000558581.1 |
DHRS4 |
dehydrogenase/reductase (SDR family) member 4 |
| chr17_-_36358166 | 0.09 |
ENST00000537432.1 |
TBC1D3 |
TBC1 domain family, member 3 |
| chr19_+_36359341 | 0.09 |
ENST00000221891.4 |
APLP1 |
amyloid beta (A4) precursor-like protein 1 |
| chr17_+_66624280 | 0.09 |
ENST00000585484.1 |
RP11-118B18.1 |
RP11-118B18.1 |
| chr17_-_10372875 | 0.08 |
ENST00000255381.2 |
MYH4 |
myosin, heavy chain 4, skeletal muscle |
| chr12_+_54379569 | 0.07 |
ENST00000513209.1 |
RP11-834C11.12 |
RP11-834C11.12 |
| chr19_+_12862604 | 0.07 |
ENST00000553030.1 |
BEST2 |
bestrophin 2 |
| chr1_+_44444865 | 0.07 |
ENST00000372324.1 |
B4GALT2 |
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 2 |
| chr20_+_1115821 | 0.06 |
ENST00000435720.1 |
PSMF1 |
proteasome (prosome, macropain) inhibitor subunit 1 (PI31) |
| chr2_+_204732487 | 0.05 |
ENST00000302823.3 |
CTLA4 |
cytotoxic T-lymphocyte-associated protein 4 |
| chr17_-_40337470 | 0.05 |
ENST00000293330.1 |
HCRT |
hypocretin (orexin) neuropeptide precursor |
| chr16_-_70729496 | 0.05 |
ENST00000567648.1 |
VAC14 |
Vac14 homolog (S. cerevisiae) |
| chr16_-_89556942 | 0.04 |
ENST00000301030.4 |
ANKRD11 |
ankyrin repeat domain 11 |
| chr12_+_98909260 | 0.03 |
ENST00000556029.1 |
TMPO |
thymopoietin |
| chr7_+_76139833 | 0.02 |
ENST00000257632.5 |
UPK3B |
uroplakin 3B |
| chr1_+_11249398 | 0.02 |
ENST00000376819.3 |
ANGPTL7 |
angiopoietin-like 7 |
| chr1_+_173446405 | 0.01 |
ENST00000340385.5 |
PRDX6 |
peroxiredoxin 6 |
| chr14_+_96722539 | 0.01 |
ENST00000553356.1 |
BDKRB1 |
bradykinin receptor B1 |
| chr11_-_64527425 | 0.01 |
ENST00000377432.3 |
PYGM |
phosphorylase, glycogen, muscle |
| chr12_-_110271178 | 0.00 |
ENST00000261740.2 ENST00000392719.2 ENST00000346520.2 |
TRPV4 |
transient receptor potential cation channel, subfamily V, member 4 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 8.8 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.3 | 7.7 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.1 | 2.5 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 3.5 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 1.3 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 1.1 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.6 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.0 | 0.4 | REACTOME N GLYCAN TRIMMING IN THE ER AND CALNEXIN CALRETICULIN CYCLE | Genes involved in N-glycan trimming in the ER and Calnexin/Calreticulin cycle |
| 0.0 | 0.2 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.4 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 7.7 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 5.2 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 1.3 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 0.4 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 8.8 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) |
| 1.2 | 3.5 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 0.9 | 4.3 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.2 | 1.0 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 0.2 | 2.5 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.2 | 1.1 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.2 | 5.2 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.1 | 0.4 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.1 | 0.6 | GO:0004905 | type I interferon receptor activity(GO:0004905) |
| 0.1 | 0.2 | GO:0015403 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) |
| 0.0 | 1.3 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.2 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.0 | 0.1 | GO:0000253 | 3-keto sterol reductase activity(GO:0000253) |
| 0.0 | 0.5 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 0.5 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 0.1 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.0 | 0.4 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.1 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.0 | 0.3 | GO:0031702 | type 1 angiotensin receptor binding(GO:0031702) |
| 0.0 | 0.6 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.0 | 0.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.2 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.3 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.1 | 0.8 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.1 | 1.3 | GO:0097342 | ripoptosome(GO:0097342) |
| 0.1 | 7.1 | GO:0044215 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) |
| 0.0 | 0.4 | GO:0005818 | astral microtubule(GO:0000235) aster(GO:0005818) |
| 0.0 | 0.2 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.0 | 0.4 | GO:0071011 | precatalytic spliceosome(GO:0071011) |
| 0.0 | 0.2 | GO:0001741 | XY body(GO:0001741) |
| 0.0 | 0.3 | GO:0070822 | Sin3-type complex(GO:0070822) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.5 | GO:0045362 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 0.9 | 7.7 | GO:0045959 | regulation of complement activation, classical pathway(GO:0030450) negative regulation of complement activation, classical pathway(GO:0045959) regulation of opsonization(GO:1903027) |
| 0.4 | 4.3 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 0.4 | 2.5 | GO:0019344 | cysteine biosynthetic process(GO:0019344) |
| 0.4 | 8.8 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.2 | 1.0 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 0.1 | 0.6 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.1 | 1.3 | GO:1903944 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 1.0 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.1 | 0.4 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.1 | 0.4 | GO:0060125 | negative regulation of growth hormone secretion(GO:0060125) |
| 0.1 | 0.3 | GO:0030037 | actin filament reorganization involved in cell cycle(GO:0030037) |
| 0.1 | 0.2 | GO:0015888 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 0.1 | 0.2 | GO:0085032 | modulation of signal transduction in other organism(GO:0044501) modulation by symbiont of host signal transduction pathway(GO:0052027) modulation of signal transduction in other organism involved in symbiotic interaction(GO:0052250) modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade(GO:0085032) |
| 0.1 | 0.3 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.1 | 1.1 | GO:0075522 | IRES-dependent viral translational initiation(GO:0075522) |
| 0.1 | 0.4 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.1 | 0.3 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.0 | 0.5 | GO:0070836 | caveola assembly(GO:0070836) |
| 0.0 | 0.5 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.0 | 5.1 | GO:0098869 | cellular oxidant detoxification(GO:0098869) |
| 0.0 | 0.2 | GO:0090116 | C-5 methylation of cytosine(GO:0090116) |
| 0.0 | 0.2 | GO:1903071 | positive regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903071) |
| 0.0 | 0.2 | GO:0030950 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) |
| 0.0 | 0.2 | GO:0060294 | cilium movement involved in cell motility(GO:0060294) regulation of microtubule binding(GO:1904526) |
| 0.0 | 2.1 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.0 | 0.3 | GO:0043031 | negative regulation of macrophage activation(GO:0043031) |
| 0.0 | 0.6 | GO:0035455 | response to interferon-alpha(GO:0035455) |
| 0.0 | 0.4 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.1 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.0 | 0.1 | GO:0071874 | cellular response to norepinephrine stimulus(GO:0071874) |
| 0.0 | 0.1 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.0 | 0.0 | GO:0030886 | negative regulation of myeloid dendritic cell activation(GO:0030886) |
| 0.0 | 0.3 | GO:0070932 | histone H3 deacetylation(GO:0070932) |