Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
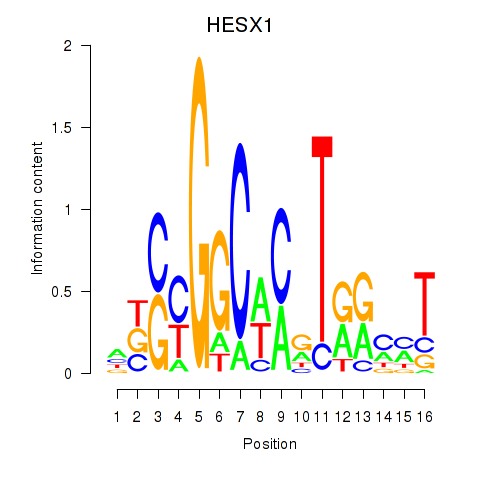
Results for HESX1
Z-value: 0.70
Transcription factors associated with HESX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HESX1
|
ENSG00000163666.4 | HESX1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HESX1 | hg19_v2_chr3_-_57233966_57234014 | -0.12 | 6.7e-01 | Click! |
Activity profile of HESX1 motif
Sorted Z-values of HESX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HESX1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_15168624 | 1.80 |
ENST00000312280.3 ENST00000494511.1 ENST00000580584.1 |
PMP22 |
peripheral myelin protein 22 |
| chr2_-_1748214 | 1.34 |
ENST00000433670.1 ENST00000425171.1 ENST00000252804.4 |
PXDN |
peroxidasin homolog (Drosophila) |
| chr7_+_130126165 | 1.20 |
ENST00000427521.1 ENST00000416162.2 ENST00000378576.4 |
MEST |
mesoderm specific transcript |
| chr6_-_167275991 | 1.15 |
ENST00000510118.1 |
RPS6KA2 |
ribosomal protein S6 kinase, 90kDa, polypeptide 2 |
| chr8_-_98290087 | 1.02 |
ENST00000322128.3 |
TSPYL5 |
TSPY-like 5 |
| chr1_+_165600083 | 0.96 |
ENST00000367889.3 |
MGST3 |
microsomal glutathione S-transferase 3 |
| chr15_+_80733570 | 0.86 |
ENST00000533983.1 ENST00000527771.1 ENST00000525103.1 |
ARNT2 |
aryl-hydrocarbon receptor nuclear translocator 2 |
| chr17_-_57184260 | 0.81 |
ENST00000376149.3 ENST00000393066.3 |
TRIM37 |
tripartite motif containing 37 |
| chr6_+_52285131 | 0.80 |
ENST00000433625.2 |
EFHC1 |
EF-hand domain (C-terminal) containing 1 |
| chr8_+_79428539 | 0.78 |
ENST00000352966.5 |
PKIA |
protein kinase (cAMP-dependent, catalytic) inhibitor alpha |
| chr5_-_179780312 | 0.73 |
ENST00000253778.8 |
GFPT2 |
glutamine-fructose-6-phosphate transaminase 2 |
| chr6_+_52285046 | 0.73 |
ENST00000371068.5 |
EFHC1 |
EF-hand domain (C-terminal) containing 1 |
| chr17_-_57184064 | 0.69 |
ENST00000262294.7 |
TRIM37 |
tripartite motif containing 37 |
| chr17_+_45286706 | 0.67 |
ENST00000393450.1 ENST00000572303.1 |
MYL4 |
myosin, light chain 4, alkali; atrial, embryonic |
| chr15_-_89089860 | 0.61 |
ENST00000558413.1 ENST00000564406.1 ENST00000268148.8 |
DET1 |
de-etiolated homolog 1 (Arabidopsis) |
| chr17_+_45286387 | 0.59 |
ENST00000572316.1 ENST00000354968.1 ENST00000576874.1 ENST00000536623.2 |
MYL4 |
myosin, light chain 4, alkali; atrial, embryonic |
| chr12_+_53491220 | 0.58 |
ENST00000548547.1 ENST00000301464.3 |
IGFBP6 |
insulin-like growth factor binding protein 6 |
| chr5_+_149340282 | 0.56 |
ENST00000286298.4 |
SLC26A2 |
solute carrier family 26 (anion exchanger), member 2 |
| chr5_+_40679584 | 0.56 |
ENST00000302472.3 |
PTGER4 |
prostaglandin E receptor 4 (subtype EP4) |
| chr8_-_101963482 | 0.56 |
ENST00000419477.2 |
YWHAZ |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta |
| chr2_-_128399706 | 0.55 |
ENST00000426981.1 |
LIMS2 |
LIM and senescent cell antigen-like domains 2 |
| chr14_+_61654271 | 0.53 |
ENST00000555185.1 ENST00000557294.1 ENST00000556778.1 |
PRKCH |
protein kinase C, eta |
| chr4_+_83956237 | 0.49 |
ENST00000264389.2 |
COPS4 |
COP9 signalosome subunit 4 |
| chr12_-_49582593 | 0.48 |
ENST00000295766.5 |
TUBA1A |
tubulin, alpha 1a |
| chr3_+_158519654 | 0.46 |
ENST00000415822.2 ENST00000392813.4 ENST00000264266.8 |
MFSD1 |
major facilitator superfamily domain containing 1 |
| chr16_+_57279248 | 0.45 |
ENST00000562023.1 ENST00000563234.1 |
ARL2BP |
ADP-ribosylation factor-like 2 binding protein |
| chr4_+_83956312 | 0.44 |
ENST00000509317.1 ENST00000503682.1 ENST00000511653.1 |
COPS4 |
COP9 signalosome subunit 4 |
| chr8_-_93029865 | 0.44 |
ENST00000422361.2 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr18_+_54318616 | 0.41 |
ENST00000254442.3 |
WDR7 |
WD repeat domain 7 |
| chr17_+_47572647 | 0.40 |
ENST00000172229.3 |
NGFR |
nerve growth factor receptor |
| chr15_-_83316711 | 0.40 |
ENST00000568128.1 |
CPEB1 |
cytoplasmic polyadenylation element binding protein 1 |
| chr2_-_197036289 | 0.39 |
ENST00000263955.4 |
STK17B |
serine/threonine kinase 17b |
| chr1_-_169337176 | 0.36 |
ENST00000472647.1 ENST00000367811.3 |
NME7 |
NME/NM23 family member 7 |
| chr18_+_54318566 | 0.34 |
ENST00000589935.1 ENST00000357574.3 |
WDR7 |
WD repeat domain 7 |
| chr21_-_34852304 | 0.33 |
ENST00000542230.2 |
TMEM50B |
transmembrane protein 50B |
| chr4_-_2264015 | 0.28 |
ENST00000337190.2 |
MXD4 |
MAX dimerization protein 4 |
| chr7_+_12727250 | 0.28 |
ENST00000404894.1 |
ARL4A |
ADP-ribosylation factor-like 4A |
| chr1_-_17216143 | 0.26 |
ENST00000457075.1 |
RP11-108M9.4 |
RP11-108M9.4 |
| chr11_-_5276008 | 0.24 |
ENST00000336906.4 |
HBG2 |
hemoglobin, gamma G |
| chr14_-_70883708 | 0.22 |
ENST00000256366.4 |
SYNJ2BP |
synaptojanin 2 binding protein |
| chr2_-_160654745 | 0.22 |
ENST00000259053.4 ENST00000429078.2 |
CD302 |
CD302 molecule |
| chr12_-_123717643 | 0.22 |
ENST00000541437.1 ENST00000606320.1 |
MPHOSPH9 |
M-phase phosphoprotein 9 |
| chr17_+_73089382 | 0.22 |
ENST00000538213.2 ENST00000584118.1 |
SLC16A5 |
solute carrier family 16 (monocarboxylate transporter), member 5 |
| chr15_+_22892663 | 0.21 |
ENST00000313077.7 ENST00000561274.1 ENST00000560848.1 |
CYFIP1 |
cytoplasmic FMR1 interacting protein 1 |
| chr6_+_31939608 | 0.21 |
ENST00000375331.2 ENST00000375333.2 |
STK19 |
serine/threonine kinase 19 |
| chr4_+_156588350 | 0.20 |
ENST00000296518.7 |
GUCY1A3 |
guanylate cyclase 1, soluble, alpha 3 |
| chr19_+_2249308 | 0.20 |
ENST00000592877.1 ENST00000221496.4 |
AMH |
anti-Mullerian hormone |
| chr11_+_73019282 | 0.18 |
ENST00000263674.3 |
ARHGEF17 |
Rho guanine nucleotide exchange factor (GEF) 17 |
| chr16_-_57219966 | 0.17 |
ENST00000565760.1 ENST00000309137.8 ENST00000570184.1 ENST00000562324.1 |
FAM192A |
family with sequence similarity 192, member A |
| chr11_+_64323098 | 0.17 |
ENST00000301891.4 |
SLC22A11 |
solute carrier family 22 (organic anion/urate transporter), member 11 |
| chr4_+_6784401 | 0.17 |
ENST00000425103.1 ENST00000307659.5 |
KIAA0232 |
KIAA0232 |
| chr2_-_89310012 | 0.17 |
ENST00000493819.1 |
IGKV1-9 |
immunoglobulin kappa variable 1-9 |
| chr15_+_80351910 | 0.17 |
ENST00000261749.6 ENST00000561060.1 |
ZFAND6 |
zinc finger, AN1-type domain 6 |
| chr21_-_38445443 | 0.16 |
ENST00000360525.4 |
PIGP |
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr1_-_165738085 | 0.16 |
ENST00000464650.1 ENST00000392129.6 |
TMCO1 |
transmembrane and coiled-coil domains 1 |
| chr19_-_50370509 | 0.16 |
ENST00000596014.1 |
PNKP |
polynucleotide kinase 3'-phosphatase |
| chr14_-_24664540 | 0.16 |
ENST00000530563.1 ENST00000528895.1 ENST00000528669.1 ENST00000532632.1 |
TM9SF1 |
transmembrane 9 superfamily member 1 |
| chr11_-_126138808 | 0.16 |
ENST00000332118.6 ENST00000532259.1 |
SRPR |
signal recognition particle receptor (docking protein) |
| chr1_-_165738072 | 0.16 |
ENST00000481278.1 |
TMCO1 |
transmembrane and coiled-coil domains 1 |
| chr7_-_74267836 | 0.16 |
ENST00000361071.5 ENST00000453619.2 ENST00000417115.2 ENST00000405086.2 |
GTF2IRD2 |
GTF2I repeat domain containing 2 |
| chr19_+_47634039 | 0.16 |
ENST00000597808.1 ENST00000413379.3 ENST00000600706.1 ENST00000540850.1 ENST00000598840.1 ENST00000600753.1 ENST00000270225.7 ENST00000392776.3 |
SAE1 |
SUMO1 activating enzyme subunit 1 |
| chr7_+_127228399 | 0.16 |
ENST00000000233.5 ENST00000415666.1 |
ARF5 |
ADP-ribosylation factor 5 |
| chr19_-_50370799 | 0.15 |
ENST00000600910.1 ENST00000322344.3 ENST00000600573.1 |
PNKP |
polynucleotide kinase 3'-phosphatase |
| chr22_+_20105259 | 0.15 |
ENST00000416427.1 ENST00000421656.1 ENST00000423859.1 ENST00000418705.2 |
RANBP1 |
RAN binding protein 1 |
| chr16_+_56659687 | 0.15 |
ENST00000568293.1 ENST00000330439.6 |
MT1E |
metallothionein 1E |
| chr8_+_120885949 | 0.15 |
ENST00000523492.1 ENST00000286234.5 |
DEPTOR |
DEP domain containing MTOR-interacting protein |
| chr10_-_75634326 | 0.14 |
ENST00000322635.3 ENST00000444854.2 ENST00000423381.1 ENST00000322680.3 ENST00000394762.2 |
CAMK2G |
calcium/calmodulin-dependent protein kinase II gamma |
| chr7_+_74508372 | 0.14 |
ENST00000356115.5 ENST00000430511.2 ENST00000312575.7 |
GTF2IRD2B |
GTF2I repeat domain containing 2B |
| chr6_-_41888814 | 0.14 |
ENST00000409060.1 ENST00000265350.4 |
MED20 |
mediator complex subunit 20 |
| chr1_+_151372010 | 0.14 |
ENST00000290541.6 |
PSMB4 |
proteasome (prosome, macropain) subunit, beta type, 4 |
| chr3_+_46921732 | 0.14 |
ENST00000418619.1 |
PTH1R |
parathyroid hormone 1 receptor |
| chr1_-_54637997 | 0.13 |
ENST00000536061.1 |
AL357673.1 |
CDNA: FLJ21031 fis, clone CAE07336; HCG1780521; Uncharacterized protein |
| chr1_-_203320617 | 0.13 |
ENST00000354955.4 |
FMOD |
fibromodulin |
| chr8_+_97274119 | 0.13 |
ENST00000455950.2 |
PTDSS1 |
phosphatidylserine synthase 1 |
| chr12_+_13197218 | 0.13 |
ENST00000197268.8 |
KIAA1467 |
KIAA1467 |
| chr14_-_106068065 | 0.12 |
ENST00000390541.2 |
IGHE |
immunoglobulin heavy constant epsilon |
| chr1_-_43638168 | 0.12 |
ENST00000431635.2 |
EBNA1BP2 |
EBNA1 binding protein 2 |
| chr2_-_96931679 | 0.12 |
ENST00000258439.3 ENST00000432959.1 |
TMEM127 |
transmembrane protein 127 |
| chr1_-_109968973 | 0.12 |
ENST00000271308.4 ENST00000538610.1 |
PSMA5 |
proteasome (prosome, macropain) subunit, alpha type, 5 |
| chr19_+_42082506 | 0.12 |
ENST00000187608.9 ENST00000401445.2 |
CEACAM21 |
carcinoembryonic antigen-related cell adhesion molecule 21 |
| chr1_+_169337172 | 0.12 |
ENST00000367807.3 ENST00000367808.3 ENST00000329281.2 ENST00000420531.1 |
BLZF1 |
basic leucine zipper nuclear factor 1 |
| chr11_+_64323156 | 0.12 |
ENST00000377585.3 |
SLC22A11 |
solute carrier family 22 (organic anion/urate transporter), member 11 |
| chr12_-_121019165 | 0.12 |
ENST00000341039.2 ENST00000357500.4 |
POP5 |
processing of precursor 5, ribonuclease P/MRP subunit (S. cerevisiae) |
| chr2_+_90248739 | 0.11 |
ENST00000468879.1 |
IGKV1D-43 |
immunoglobulin kappa variable 1D-43 |
| chr6_-_41888843 | 0.11 |
ENST00000434077.1 ENST00000409312.1 |
MED20 |
mediator complex subunit 20 |
| chrX_+_153409678 | 0.11 |
ENST00000369951.4 |
OPN1LW |
opsin 1 (cone pigments), long-wave-sensitive |
| chr1_+_152658599 | 0.10 |
ENST00000368780.3 |
LCE2B |
late cornified envelope 2B |
| chr18_-_54318353 | 0.10 |
ENST00000590954.1 ENST00000540155.1 |
TXNL1 |
thioredoxin-like 1 |
| chr8_+_146277764 | 0.09 |
ENST00000331434.6 |
C8orf33 |
chromosome 8 open reading frame 33 |
| chr21_+_38445539 | 0.09 |
ENST00000418766.1 ENST00000450533.1 ENST00000438055.1 ENST00000355666.1 ENST00000540756.1 ENST00000399010.1 |
TTC3 |
tetratricopeptide repeat domain 3 |
| chr7_+_72742162 | 0.09 |
ENST00000431982.2 |
FKBP6 |
FK506 binding protein 6, 36kDa |
| chr22_+_44351301 | 0.09 |
ENST00000350028.4 |
SAMM50 |
SAMM50 sorting and assembly machinery component |
| chr9_-_123239632 | 0.09 |
ENST00000416449.1 |
CDK5RAP2 |
CDK5 regulatory subunit associated protein 2 |
| chr11_-_1629693 | 0.09 |
ENST00000399685.1 |
KRTAP5-3 |
keratin associated protein 5-3 |
| chr12_-_54070098 | 0.09 |
ENST00000394349.3 ENST00000549164.1 |
ATP5G2 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C2 (subunit 9) |
| chr22_+_44351419 | 0.09 |
ENST00000396202.3 |
SAMM50 |
SAMM50 sorting and assembly machinery component |
| chr8_-_101734308 | 0.08 |
ENST00000519004.1 ENST00000519363.1 ENST00000520142.1 |
PABPC1 |
poly(A) binding protein, cytoplasmic 1 |
| chr16_-_66968055 | 0.08 |
ENST00000568572.1 |
FAM96B |
family with sequence similarity 96, member B |
| chr21_-_38445470 | 0.08 |
ENST00000399098.1 |
PIGP |
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr4_-_144826682 | 0.08 |
ENST00000358615.4 ENST00000437468.2 |
GYPE |
glycophorin E (MNS blood group) |
| chr1_-_40237020 | 0.07 |
ENST00000327582.5 |
OXCT2 |
3-oxoacid CoA transferase 2 |
| chr7_+_72742178 | 0.07 |
ENST00000442793.1 ENST00000413573.2 ENST00000252037.4 |
FKBP6 |
FK506 binding protein 6, 36kDa |
| chr11_-_71791518 | 0.07 |
ENST00000537217.1 ENST00000366394.3 ENST00000358965.6 ENST00000546131.1 ENST00000543937.1 ENST00000368959.5 ENST00000541641.1 |
NUMA1 |
nuclear mitotic apparatus protein 1 |
| chr14_-_25103388 | 0.07 |
ENST00000526004.1 ENST00000415355.3 |
GZMB |
granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) |
| chr2_+_90458201 | 0.06 |
ENST00000603238.1 |
CH17-132F21.1 |
Uncharacterized protein |
| chr13_-_108870623 | 0.06 |
ENST00000405925.1 |
LIG4 |
ligase IV, DNA, ATP-dependent |
| chr8_+_71581565 | 0.06 |
ENST00000408926.3 ENST00000520030.1 |
XKR9 |
XK, Kell blood group complex subunit-related family, member 9 |
| chr2_-_89247338 | 0.06 |
ENST00000496168.1 |
IGKV1-5 |
immunoglobulin kappa variable 1-5 |
| chr2_+_234216454 | 0.06 |
ENST00000447536.1 ENST00000409110.1 |
SAG |
S-antigen; retina and pineal gland (arrestin) |
| chr3_+_148415571 | 0.06 |
ENST00000497524.1 ENST00000349243.3 ENST00000542281.1 ENST00000418473.2 ENST00000404754.2 |
AGTR1 |
angiotensin II receptor, type 1 |
| chr6_-_29600832 | 0.05 |
ENST00000377016.4 ENST00000376977.3 ENST00000377034.4 |
GABBR1 |
gamma-aminobutyric acid (GABA) B receptor, 1 |
| chr15_+_80364901 | 0.05 |
ENST00000560228.1 ENST00000559835.1 ENST00000559775.1 ENST00000558688.1 ENST00000560392.1 ENST00000560976.1 ENST00000558272.1 ENST00000558390.1 |
ZFAND6 |
zinc finger, AN1-type domain 6 |
| chr1_-_204165610 | 0.05 |
ENST00000367194.4 |
KISS1 |
KiSS-1 metastasis-suppressor |
| chr1_+_46016703 | 0.05 |
ENST00000481885.1 ENST00000351829.4 ENST00000471651.1 |
AKR1A1 |
aldo-keto reductase family 1, member A1 (aldehyde reductase) |
| chr7_+_4815238 | 0.05 |
ENST00000348624.4 ENST00000401897.1 |
AP5Z1 |
adaptor-related protein complex 5, zeta 1 subunit |
| chr12_-_121476750 | 0.05 |
ENST00000543677.1 |
OASL |
2'-5'-oligoadenylate synthetase-like |
| chr12_+_54379569 | 0.05 |
ENST00000513209.1 |
RP11-834C11.12 |
RP11-834C11.12 |
| chrX_+_37850026 | 0.05 |
ENST00000341016.3 |
CXorf27 |
chromosome X open reading frame 27 |
| chr4_-_48782259 | 0.04 |
ENST00000507711.1 ENST00000358350.4 ENST00000537810.1 ENST00000264319.7 |
FRYL |
FRY-like |
| chr10_+_121652204 | 0.04 |
ENST00000369075.3 ENST00000543134.1 |
SEC23IP |
SEC23 interacting protein |
| chr2_+_176957619 | 0.04 |
ENST00000392539.3 |
HOXD13 |
homeobox D13 |
| chr3_+_155860751 | 0.04 |
ENST00000471742.1 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr20_-_590944 | 0.03 |
ENST00000246080.3 |
TCF15 |
transcription factor 15 (basic helix-loop-helix) |
| chr1_+_3614591 | 0.03 |
ENST00000378290.4 |
TP73 |
tumor protein p73 |
| chr10_+_26505594 | 0.03 |
ENST00000259271.3 |
GAD2 |
glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) |
| chr6_-_42981651 | 0.02 |
ENST00000244711.3 |
MEA1 |
male-enhanced antigen 1 |
| chr7_-_99679324 | 0.02 |
ENST00000292393.5 ENST00000413658.2 ENST00000412947.1 ENST00000441298.1 ENST00000449785.1 ENST00000299667.4 ENST00000424697.1 |
ZNF3 |
zinc finger protein 3 |
| chr1_-_31661000 | 0.02 |
ENST00000263693.1 ENST00000398657.2 ENST00000526106.1 |
NKAIN1 |
Na+/K+ transporting ATPase interacting 1 |
| chr11_-_71791435 | 0.02 |
ENST00000351960.6 ENST00000541719.1 ENST00000535111.1 |
NUMA1 |
nuclear mitotic apparatus protein 1 |
| chr18_-_18691739 | 0.02 |
ENST00000399799.2 |
ROCK1 |
Rho-associated, coiled-coil containing protein kinase 1 |
| chr3_+_39851094 | 0.02 |
ENST00000302541.6 |
MYRIP |
myosin VIIA and Rab interacting protein |
| chr20_+_35807449 | 0.02 |
ENST00000237530.6 |
RPN2 |
ribophorin II |
| chr1_+_45265897 | 0.02 |
ENST00000372201.4 |
PLK3 |
polo-like kinase 3 |
| chr13_-_52378231 | 0.02 |
ENST00000280056.2 ENST00000444610.2 |
DHRS12 |
dehydrogenase/reductase (SDR family) member 12 |
| chr14_-_25103472 | 0.02 |
ENST00000216341.4 ENST00000382542.1 ENST00000382540.1 |
GZMB |
granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) |
| chr1_+_47799446 | 0.02 |
ENST00000371873.5 |
CMPK1 |
cytidine monophosphate (UMP-CMP) kinase 1, cytosolic |
| chr10_-_101380121 | 0.02 |
ENST00000370495.4 |
SLC25A28 |
solute carrier family 25 (mitochondrial iron transporter), member 28 |
| chr1_-_20446020 | 0.01 |
ENST00000375105.3 |
PLA2G2D |
phospholipase A2, group IID |
| chr2_-_89568263 | 0.01 |
ENST00000473726.1 |
IGKV1-33 |
immunoglobulin kappa variable 1-33 |
| chr22_+_18632666 | 0.01 |
ENST00000215794.7 |
USP18 |
ubiquitin specific peptidase 18 |
| chr22_+_19419425 | 0.01 |
ENST00000333130.3 |
MRPL40 |
mitochondrial ribosomal protein L40 |
| chr3_-_42846021 | 0.01 |
ENST00000321331.7 |
HIGD1A |
HIG1 hypoxia inducible domain family, member 1A |
| chr2_+_233320827 | 0.00 |
ENST00000295463.3 |
ALPI |
alkaline phosphatase, intestinal |
| chr11_+_125774258 | 0.00 |
ENST00000263576.6 |
DDX25 |
DEAD (Asp-Glu-Ala-Asp) box helicase 25 |
| chr16_+_4784273 | 0.00 |
ENST00000299320.5 ENST00000586724.1 |
C16orf71 |
chromosome 16 open reading frame 71 |
| chr21_-_38445011 | 0.00 |
ENST00000464265.1 ENST00000399102.1 |
PIGP |
phosphatidylinositol glycan anchor biosynthesis, class P |
| chr8_+_109455845 | 0.00 |
ENST00000220853.3 |
EMC2 |
ER membrane protein complex subunit 2 |
| chr21_+_34398153 | 0.00 |
ENST00000382357.3 ENST00000430860.1 ENST00000333337.3 |
OLIG2 |
oligodendrocyte lineage transcription factor 2 |
| chrX_+_118602363 | 0.00 |
ENST00000317881.8 |
SLC25A5 |
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 |
| chr20_+_35807512 | 0.00 |
ENST00000373622.5 |
RPN2 |
ribophorin II |
| chr19_+_19496624 | 0.00 |
ENST00000494516.2 ENST00000360315.3 ENST00000252577.5 |
GATAD2A |
GATA zinc finger domain containing 2A |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0036353 | histone H2A-K119 monoubiquitination(GO:0036353) |
| 0.2 | 0.6 | GO:2000417 | negative regulation of eosinophil migration(GO:2000417) |
| 0.1 | 0.6 | GO:0050428 | purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.1 | 1.8 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.1 | 0.4 | GO:1903588 | negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903588) |
| 0.1 | 0.5 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.1 | 0.3 | GO:0098502 | DNA dephosphorylation(GO:0098502) |
| 0.1 | 0.9 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.1 | 0.7 | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process(GO:0006048) |
| 0.1 | 0.2 | GO:1903422 | negative regulation of synaptic vesicle recycling(GO:1903422) |
| 0.1 | 0.6 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.1 | 0.6 | GO:2000346 | negative regulation of hepatocyte proliferation(GO:2000346) |
| 0.1 | 0.2 | GO:2000354 | regulation of ovarian follicle development(GO:2000354) |
| 0.0 | 0.4 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.0 | 0.8 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.0 | 0.1 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.0 | 1.3 | GO:0042744 | hydrogen peroxide catabolic process(GO:0042744) |
| 0.0 | 0.2 | GO:0052565 | response to defense-related nitric oxide production by other organism involved in symbiotic interaction(GO:0052551) response to defense-related host nitric oxide production(GO:0052565) |
| 0.0 | 1.3 | GO:0002026 | regulation of the force of heart contraction(GO:0002026) |
| 0.0 | 0.3 | GO:0015747 | urate transport(GO:0015747) |
| 0.0 | 0.1 | GO:0055048 | regulation of spindle elongation(GO:0032887) regulation of mitotic spindle elongation(GO:0032888) anastral spindle assembly(GO:0055048) protein localization to spindle pole body(GO:0071988) regulation of protein localization to spindle pole body(GO:1902363) positive regulation of protein localization to spindle pole body(GO:1902365) positive regulation of mitotic spindle elongation(GO:1902846) |
| 0.0 | 0.4 | GO:2000271 | positive regulation of fibroblast apoptotic process(GO:2000271) |
| 0.0 | 0.4 | GO:0046051 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 0.0 | 1.0 | GO:1901685 | glutathione derivative metabolic process(GO:1901685) glutathione derivative biosynthetic process(GO:1901687) |
| 0.0 | 0.3 | GO:0006983 | ER overload response(GO:0006983) |
| 0.0 | 0.2 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 1.5 | GO:0021795 | cerebral cortex cell migration(GO:0021795) |
| 0.0 | 0.1 | GO:0046604 | positive regulation of mitotic centrosome separation(GO:0046604) |
| 0.0 | 0.1 | GO:0051102 | DNA ligation involved in DNA recombination(GO:0051102) |
| 0.0 | 0.2 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.0 | 1.0 | GO:0071480 | cellular response to gamma radiation(GO:0071480) |
| 0.0 | 0.2 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.0 | GO:2000777 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process involved in cellular response to hypoxia(GO:2000777) |
| 0.0 | 0.4 | GO:0051457 | maintenance of protein location in nucleus(GO:0051457) |
| 0.0 | 0.2 | GO:0045040 | protein import into mitochondrial outer membrane(GO:0045040) |
| 0.0 | 0.1 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 0.0 | 0.6 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.0 | 0.1 | GO:0007032 | endosome organization(GO:0007032) |
| 0.0 | 0.1 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.0 | 0.1 | GO:0086097 | phospholipase C-activating angiotensin-activated signaling pathway(GO:0086097) |
| 0.0 | 0.2 | GO:0006909 | phagocytosis(GO:0006909) |
| 0.0 | 0.1 | GO:0002862 | negative regulation of inflammatory response to antigenic stimulus(GO:0002862) |
| 0.0 | 0.2 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.0 | 0.1 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.0 | 0.1 | GO:0009249 | protein lipoylation(GO:0009249) |
| 0.0 | 0.1 | GO:0033686 | positive regulation of luteinizing hormone secretion(GO:0033686) |
| 0.0 | 0.1 | GO:0045792 | negative regulation of cell size(GO:0045792) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | GO:0043218 | compact myelin(GO:0043218) |
| 0.1 | 1.5 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 0.6 | GO:0016942 | insulin-like growth factor binding protein complex(GO:0016942) growth factor complex(GO:0036454) |
| 0.1 | 0.6 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 0.1 | GO:0061673 | cortical microtubule(GO:0055028) mitotic spindle astral microtubule(GO:0061673) |
| 0.0 | 0.2 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.0 | 0.5 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 0.2 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.0 | 0.1 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.0 | 0.9 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 1.6 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 0.2 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.0 | 1.3 | GO:0031672 | A band(GO:0031672) |
| 0.0 | 0.1 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 0.2 | GO:0098799 | outer mitochondrial membrane protein complex(GO:0098799) |
| 0.0 | 0.2 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.0 | 0.1 | GO:0032807 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) DNA ligase IV complex(GO:0032807) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 1.1 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.0 | 1.0 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 0.4 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.0 | 0.6 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 1.3 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.5 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.0 | 0.3 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.0 | 0.3 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.0 | 0.5 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.2 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.3 | GO:0032038 | myosin II heavy chain binding(GO:0032038) |
| 0.3 | 1.3 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 0.1 | 0.5 | GO:0004699 | calcium-independent protein kinase C activity(GO:0004699) |
| 0.1 | 0.6 | GO:0019531 | oxalate transmembrane transporter activity(GO:0019531) |
| 0.1 | 0.9 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.1 | 0.3 | GO:0051734 | ATP-dependent polydeoxyribonucleotide 5'-hydroxyl-kinase activity(GO:0046404) polydeoxyribonucleotide kinase activity(GO:0051733) ATP-dependent polynucleotide kinase activity(GO:0051734) polynucleotide phosphatase activity(GO:0098518) |
| 0.1 | 0.6 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.1 | 0.9 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.1 | 0.6 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.4 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.0 | 0.8 | GO:0004862 | cAMP-dependent protein kinase inhibitor activity(GO:0004862) |
| 0.0 | 0.7 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.0 | 1.0 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.0 | 0.3 | GO:0015143 | urate transmembrane transporter activity(GO:0015143) salt transmembrane transporter activity(GO:1901702) |
| 0.0 | 0.1 | GO:0004991 | parathyroid hormone receptor activity(GO:0004991) |
| 0.0 | 0.2 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.0 | 0.2 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 0.4 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 0.2 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 1.0 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.1 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.0 | 0.2 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.0 | 0.1 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.0 | 0.1 | GO:0008410 | CoA-transferase activity(GO:0008410) |
| 0.0 | 0.4 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 1.1 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.0 | 0.1 | GO:0047134 | protein-disulfide reductase activity(GO:0047134) |
| 0.0 | 0.1 | GO:0002046 | opsin binding(GO:0002046) |
| 0.0 | 0.1 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.0 | 1.8 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 1.7 | ST PHOSPHOINOSITIDE 3 KINASE PATHWAY | PI3K Pathway |
| 0.0 | 0.4 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.6 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.0 | 0.2 | PID ALK2 PATHWAY | ALK2 signaling events |