Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
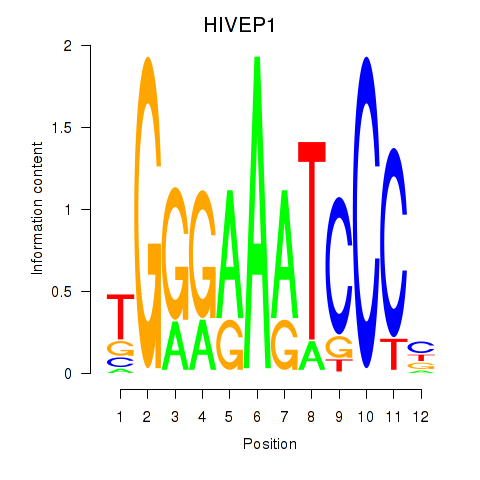
Results for HIVEP1
Z-value: 1.78
Transcription factors associated with HIVEP1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HIVEP1
|
ENSG00000095951.12 | HIVEP1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HIVEP1 | hg19_v2_chr6_+_12012536_12012571 | 0.46 | 7.2e-02 | Click! |
Activity profile of HIVEP1 motif
Sorted Z-values of HIVEP1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HIVEP1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr22_+_22930626 | 6.62 |
ENST00000390302.2 |
IGLV2-33 |
immunoglobulin lambda variable 2-33 (non-functional) |
| chr6_+_32605195 | 5.41 |
ENST00000374949.2 |
HLA-DQA1 |
major histocompatibility complex, class II, DQ alpha 1 |
| chr14_+_75988851 | 4.76 |
ENST00000555504.1 |
BATF |
basic leucine zipper transcription factor, ATF-like |
| chr12_-_9913489 | 4.70 |
ENST00000228434.3 ENST00000536709.1 |
CD69 |
CD69 molecule |
| chr6_+_32605134 | 4.46 |
ENST00000343139.5 ENST00000395363.1 ENST00000496318.1 |
HLA-DQA1 |
major histocompatibility complex, class II, DQ alpha 1 |
| chr6_+_32709119 | 4.23 |
ENST00000374940.3 |
HLA-DQA2 |
major histocompatibility complex, class II, DQ alpha 2 |
| chr6_-_32908792 | 4.20 |
ENST00000418107.2 |
HLA-DMB |
major histocompatibility complex, class II, DM beta |
| chr14_-_106322288 | 3.87 |
ENST00000390559.2 |
IGHM |
immunoglobulin heavy constant mu |
| chr6_+_32407619 | 3.76 |
ENST00000395388.2 ENST00000374982.5 |
HLA-DRA |
major histocompatibility complex, class II, DR alpha |
| chr13_-_46756351 | 3.51 |
ENST00000323076.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chr19_-_6591113 | 3.20 |
ENST00000423145.3 ENST00000245903.3 |
CD70 |
CD70 molecule |
| chr16_+_57392684 | 3.19 |
ENST00000219235.4 |
CCL22 |
chemokine (C-C motif) ligand 22 |
| chr2_+_61108650 | 3.18 |
ENST00000295025.8 |
REL |
v-rel avian reticuloendotheliosis viral oncogene homolog |
| chr19_+_4229495 | 3.09 |
ENST00000221847.5 |
EBI3 |
Epstein-Barr virus induced 3 |
| chr16_+_32077386 | 3.07 |
ENST00000354689.6 |
IGHV3OR16-9 |
immunoglobulin heavy variable 3/OR16-9 (non-functional) |
| chr3_-_124839648 | 3.04 |
ENST00000430155.2 |
SLC12A8 |
solute carrier family 12, member 8 |
| chr2_+_61108771 | 2.90 |
ENST00000394479.3 |
REL |
v-rel avian reticuloendotheliosis viral oncogene homolog |
| chr14_-_106573756 | 2.88 |
ENST00000390601.2 |
IGHV3-11 |
immunoglobulin heavy variable 3-11 (gene/pseudogene) |
| chr1_+_111770278 | 2.80 |
ENST00000369748.4 |
CHI3L2 |
chitinase 3-like 2 |
| chr1_+_111770232 | 2.77 |
ENST00000369744.2 |
CHI3L2 |
chitinase 3-like 2 |
| chr12_+_7055631 | 2.66 |
ENST00000543115.1 ENST00000399448.1 |
PTPN6 |
protein tyrosine phosphatase, non-receptor type 6 |
| chr11_+_121447469 | 2.59 |
ENST00000532694.1 ENST00000534286.1 |
SORL1 |
sortilin-related receptor, L(DLR class) A repeats containing |
| chr5_-_149792295 | 2.57 |
ENST00000518797.1 ENST00000524315.1 ENST00000009530.7 ENST00000377795.3 |
CD74 |
CD74 molecule, major histocompatibility complex, class II invariant chain |
| chr14_-_106518922 | 2.57 |
ENST00000390598.2 |
IGHV3-7 |
immunoglobulin heavy variable 3-7 |
| chr16_+_33605231 | 2.49 |
ENST00000570121.2 |
IGHV3OR16-12 |
immunoglobulin heavy variable 3/OR16-12 (non-functional) |
| chr3_-_58196939 | 2.48 |
ENST00000394549.2 ENST00000461914.3 |
DNASE1L3 |
deoxyribonuclease I-like 3 |
| chr8_-_101321584 | 2.46 |
ENST00000523167.1 |
RNF19A |
ring finger protein 19A, RBR E3 ubiquitin protein ligase |
| chr7_+_69064300 | 2.39 |
ENST00000342771.4 |
AUTS2 |
autism susceptibility candidate 2 |
| chr14_-_107049312 | 2.34 |
ENST00000390627.2 |
IGHV3-53 |
immunoglobulin heavy variable 3-53 |
| chr22_-_37640456 | 2.34 |
ENST00000405484.1 ENST00000441619.1 ENST00000406508.1 |
RAC2 |
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr6_+_138188551 | 2.29 |
ENST00000237289.4 ENST00000433680.1 |
TNFAIP3 |
tumor necrosis factor, alpha-induced protein 3 |
| chr22_-_37640277 | 2.27 |
ENST00000401529.3 ENST00000249071.6 |
RAC2 |
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr14_+_75988768 | 2.23 |
ENST00000286639.6 |
BATF |
basic leucine zipper transcription factor, ATF-like |
| chr14_-_106926724 | 2.20 |
ENST00000434710.1 |
IGHV3-43 |
immunoglobulin heavy variable 3-43 |
| chr3_-_119278446 | 2.14 |
ENST00000264246.3 |
CD80 |
CD80 molecule |
| chr22_+_37257015 | 2.14 |
ENST00000447071.1 ENST00000248899.6 ENST00000397147.4 |
NCF4 |
neutrophil cytosolic factor 4, 40kDa |
| chr14_-_35873856 | 2.13 |
ENST00000553342.1 ENST00000216797.5 ENST00000557140.1 |
NFKBIA |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha |
| chr3_-_58196688 | 2.06 |
ENST00000486455.1 |
DNASE1L3 |
deoxyribonuclease I-like 3 |
| chr13_-_46964177 | 2.02 |
ENST00000389908.3 |
KIAA0226L |
KIAA0226-like |
| chr9_-_123691047 | 2.02 |
ENST00000373887.3 |
TRAF1 |
TNF receptor-associated factor 1 |
| chrX_-_73072534 | 1.97 |
ENST00000429829.1 |
XIST |
X inactive specific transcript (non-protein coding) |
| chr14_-_106845789 | 1.96 |
ENST00000390617.2 |
IGHV3-35 |
immunoglobulin heavy variable 3-35 (non-functional) |
| chr3_-_119278376 | 1.88 |
ENST00000478182.1 |
CD80 |
CD80 molecule |
| chr14_-_106866934 | 1.79 |
ENST00000390618.2 |
IGHV3-38 |
immunoglobulin heavy variable 3-38 (non-functional) |
| chr1_-_209824643 | 1.79 |
ENST00000391911.1 ENST00000415782.1 |
LAMB3 |
laminin, beta 3 |
| chr6_+_29691198 | 1.78 |
ENST00000440587.2 ENST00000434407.2 |
HLA-F |
major histocompatibility complex, class I, F |
| chr2_-_163175133 | 1.76 |
ENST00000421365.2 ENST00000263642.2 |
IFIH1 |
interferon induced with helicase C domain 1 |
| chr6_+_29691056 | 1.73 |
ENST00000414333.1 ENST00000334668.4 ENST00000259951.7 |
HLA-F |
major histocompatibility complex, class I, F |
| chr19_-_10445399 | 1.66 |
ENST00000592945.1 |
ICAM3 |
intercellular adhesion molecule 3 |
| chr16_+_50730910 | 1.53 |
ENST00000300589.2 |
NOD2 |
nucleotide-binding oligomerization domain containing 2 |
| chr1_+_156123359 | 1.47 |
ENST00000368284.1 ENST00000368286.2 ENST00000438830.1 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr7_+_7606497 | 1.44 |
ENST00000340080.4 ENST00000405785.1 ENST00000433635.1 |
MIOS |
missing oocyte, meiosis regulator, homolog (Drosophila) |
| chr1_+_156123318 | 1.41 |
ENST00000368285.3 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr12_+_10658201 | 1.41 |
ENST00000322446.3 |
EIF2S3L |
Putative eukaryotic translation initiation factor 2 subunit 3-like protein |
| chr17_-_53499310 | 1.38 |
ENST00000262065.3 |
MMD |
monocyte to macrophage differentiation-associated |
| chr20_+_57430162 | 1.38 |
ENST00000450130.1 ENST00000349036.3 ENST00000423897.1 |
GNAS |
GNAS complex locus |
| chr5_+_125935960 | 1.33 |
ENST00000297540.4 |
PHAX |
phosphorylated adaptor for RNA export |
| chr12_-_12849073 | 1.33 |
ENST00000332427.2 ENST00000540796.1 |
GPR19 |
G protein-coupled receptor 19 |
| chr16_+_33020496 | 1.31 |
ENST00000565407.2 |
IGHV3OR16-8 |
immunoglobulin heavy variable 3/OR16-8 (non-functional) |
| chr3_+_119316689 | 1.31 |
ENST00000273371.4 |
PLA1A |
phospholipase A1 member A |
| chr10_+_114133773 | 1.30 |
ENST00000354655.4 |
ACSL5 |
acyl-CoA synthetase long-chain family member 5 |
| chr20_+_9494987 | 1.24 |
ENST00000427562.2 ENST00000246070.2 |
LAMP5 |
lysosomal-associated membrane protein family, member 5 |
| chr3_-_119396193 | 1.23 |
ENST00000484810.1 ENST00000497116.1 ENST00000261070.2 |
COX17 |
COX17 cytochrome c oxidase copper chaperone |
| chr15_+_75315896 | 1.22 |
ENST00000342932.3 ENST00000564923.1 ENST00000569562.1 ENST00000568649.1 |
PPCDC |
phosphopantothenoylcysteine decarboxylase |
| chr5_+_150591678 | 1.21 |
ENST00000523466.1 |
GM2A |
GM2 ganglioside activator |
| chr3_-_119379719 | 1.20 |
ENST00000493094.1 |
POPDC2 |
popeye domain containing 2 |
| chr2_-_27558270 | 1.18 |
ENST00000454704.1 |
GTF3C2 |
general transcription factor IIIC, polypeptide 2, beta 110kDa |
| chr16_-_33647696 | 1.18 |
ENST00000558425.1 ENST00000569103.2 |
RP11-812E19.9 |
Uncharacterized protein |
| chr4_-_76944621 | 1.18 |
ENST00000306602.1 |
CXCL10 |
chemokine (C-X-C motif) ligand 10 |
| chr16_-_11681316 | 1.17 |
ENST00000571688.1 |
LITAF |
lipopolysaccharide-induced TNF factor |
| chr14_+_103243813 | 1.17 |
ENST00000560371.1 ENST00000347662.4 ENST00000392745.2 ENST00000539721.1 ENST00000560463.1 |
TRAF3 |
TNF receptor-associated factor 3 |
| chr20_+_44746939 | 1.14 |
ENST00000372276.3 |
CD40 |
CD40 molecule, TNF receptor superfamily member 5 |
| chr20_+_44746885 | 1.13 |
ENST00000372285.3 |
CD40 |
CD40 molecule, TNF receptor superfamily member 5 |
| chr6_-_31324943 | 1.13 |
ENST00000412585.2 ENST00000434333.1 |
HLA-B |
major histocompatibility complex, class I, B |
| chr17_+_40440481 | 1.12 |
ENST00000590726.2 ENST00000452307.2 ENST00000444283.1 ENST00000588868.1 |
STAT5A |
signal transducer and activator of transcription 5A |
| chr15_+_74908147 | 1.11 |
ENST00000568139.1 ENST00000563297.1 ENST00000568488.1 ENST00000352989.5 ENST00000348245.3 |
CLK3 |
CDC-like kinase 3 |
| chr19_-_2042065 | 1.11 |
ENST00000591588.1 ENST00000591142.1 |
MKNK2 |
MAP kinase interacting serine/threonine kinase 2 |
| chr17_-_62097927 | 1.11 |
ENST00000578313.1 ENST00000584084.1 ENST00000579788.1 ENST00000579687.1 ENST00000578379.1 ENST00000578892.1 ENST00000412356.1 ENST00000418105.1 |
ICAM2 |
intercellular adhesion molecule 2 |
| chr14_-_106725723 | 1.09 |
ENST00000390609.2 |
IGHV3-23 |
immunoglobulin heavy variable 3-23 |
| chr5_-_150460914 | 1.07 |
ENST00000389378.2 |
TNIP1 |
TNFAIP3 interacting protein 1 |
| chr6_+_31554962 | 1.06 |
ENST00000376092.3 ENST00000376086.3 ENST00000303757.8 ENST00000376093.2 ENST00000376102.3 |
LST1 |
leukocyte specific transcript 1 |
| chr3_+_119316721 | 1.06 |
ENST00000488919.1 ENST00000495992.1 |
PLA1A |
phospholipase A1 member A |
| chr6_-_31239846 | 1.04 |
ENST00000415537.1 ENST00000376228.5 ENST00000383329.3 |
HLA-C |
major histocompatibility complex, class I, C |
| chr17_-_74236382 | 1.03 |
ENST00000592271.1 ENST00000319945.6 ENST00000269391.6 |
RNF157 |
ring finger protein 157 |
| chr5_-_150460539 | 1.03 |
ENST00000520931.1 ENST00000520695.1 ENST00000521591.1 ENST00000518977.1 |
TNIP1 |
TNFAIP3 interacting protein 1 |
| chr2_-_73511559 | 1.03 |
ENST00000521871.1 |
FBXO41 |
F-box protein 41 |
| chr6_+_31555045 | 1.01 |
ENST00000396101.3 ENST00000490742.1 |
LST1 |
leukocyte specific transcript 1 |
| chr11_-_57194218 | 1.00 |
ENST00000529554.1 |
SLC43A3 |
solute carrier family 43, member 3 |
| chr17_-_34207295 | 0.97 |
ENST00000463941.1 ENST00000293272.3 |
CCL5 |
chemokine (C-C motif) ligand 5 |
| chr10_+_90750378 | 0.92 |
ENST00000355740.2 ENST00000352159.4 |
FAS |
Fas cell surface death receptor |
| chr8_+_27168988 | 0.91 |
ENST00000397501.1 ENST00000338238.4 ENST00000544172.1 |
PTK2B |
protein tyrosine kinase 2 beta |
| chr9_+_139921916 | 0.91 |
ENST00000314330.2 |
C9orf139 |
chromosome 9 open reading frame 139 |
| chr17_-_62097904 | 0.89 |
ENST00000583366.1 |
ICAM2 |
intercellular adhesion molecule 2 |
| chr7_-_99698338 | 0.87 |
ENST00000354230.3 ENST00000425308.1 |
MCM7 |
minichromosome maintenance complex component 7 |
| chr1_+_155051305 | 0.85 |
ENST00000368408.3 |
EFNA3 |
ephrin-A3 |
| chr8_+_86121448 | 0.84 |
ENST00000520225.1 |
E2F5 |
E2F transcription factor 5, p130-binding |
| chrX_-_30595959 | 0.84 |
ENST00000378962.3 |
CXorf21 |
chromosome X open reading frame 21 |
| chr12_+_10658489 | 0.83 |
ENST00000538173.1 |
EIF2S3L |
Putative eukaryotic translation initiation factor 2 subunit 3-like protein |
| chr6_-_27440460 | 0.79 |
ENST00000377419.1 |
ZNF184 |
zinc finger protein 184 |
| chr17_-_62084241 | 0.79 |
ENST00000449662.2 |
ICAM2 |
intercellular adhesion molecule 2 |
| chr6_+_30850697 | 0.77 |
ENST00000509639.1 ENST00000412274.2 ENST00000507901.1 ENST00000507046.1 ENST00000437124.2 ENST00000454612.2 ENST00000396342.2 |
DDR1 |
discoidin domain receptor tyrosine kinase 1 |
| chr1_-_202129105 | 0.77 |
ENST00000367279.4 |
PTPN7 |
protein tyrosine phosphatase, non-receptor type 7 |
| chr15_+_85923856 | 0.75 |
ENST00000560302.1 ENST00000394518.2 ENST00000361243.2 ENST00000560256.1 |
AKAP13 |
A kinase (PRKA) anchor protein 13 |
| chr7_+_47694842 | 0.74 |
ENST00000408988.2 |
C7orf65 |
chromosome 7 open reading frame 65 |
| chr11_+_75526212 | 0.73 |
ENST00000356136.3 |
UVRAG |
UV radiation resistance associated |
| chr19_-_40336969 | 0.73 |
ENST00000599134.1 ENST00000597634.1 ENST00000598417.1 ENST00000601274.1 ENST00000594309.1 ENST00000221801.3 |
FBL |
fibrillarin |
| chr8_-_70747205 | 0.72 |
ENST00000260126.4 |
SLCO5A1 |
solute carrier organic anion transporter family, member 5A1 |
| chr9_-_35103105 | 0.71 |
ENST00000452248.2 ENST00000356493.5 |
STOML2 |
stomatin (EPB72)-like 2 |
| chr19_-_51522955 | 0.67 |
ENST00000358789.3 |
KLK10 |
kallikrein-related peptidase 10 |
| chr16_+_33006369 | 0.64 |
ENST00000425181.3 |
IGHV3OR16-10 |
immunoglobulin heavy variable 3/OR16-10 (non-functional) |
| chr17_-_26876350 | 0.64 |
ENST00000470125.1 |
UNC119 |
unc-119 homolog (C. elegans) |
| chr7_-_120498357 | 0.63 |
ENST00000415871.1 ENST00000222747.3 ENST00000430985.1 |
TSPAN12 |
tetraspanin 12 |
| chr10_+_124914285 | 0.62 |
ENST00000407911.2 |
BUB3 |
BUB3 mitotic checkpoint protein |
| chr8_+_54793454 | 0.62 |
ENST00000276500.4 |
RGS20 |
regulator of G-protein signaling 20 |
| chr16_+_22825475 | 0.62 |
ENST00000261374.3 |
HS3ST2 |
heparan sulfate (glucosamine) 3-O-sulfotransferase 2 |
| chr6_+_29910301 | 0.61 |
ENST00000376809.5 ENST00000376802.2 |
HLA-A |
major histocompatibility complex, class I, A |
| chr2_+_163175394 | 0.60 |
ENST00000446271.1 ENST00000429691.2 |
GCA |
grancalcin, EF-hand calcium binding protein |
| chr16_+_32063311 | 0.60 |
ENST00000426099.1 |
AC142381.1 |
AC142381.1 |
| chr1_+_37940153 | 0.59 |
ENST00000373087.6 |
ZC3H12A |
zinc finger CCCH-type containing 12A |
| chr8_+_54793425 | 0.58 |
ENST00000522225.1 |
RGS20 |
regulator of G-protein signaling 20 |
| chr6_+_87865262 | 0.57 |
ENST00000369577.3 ENST00000518845.1 ENST00000339907.4 ENST00000496806.2 |
ZNF292 |
zinc finger protein 292 |
| chr2_+_162101247 | 0.57 |
ENST00000439050.1 ENST00000436506.1 |
AC009299.3 |
AC009299.3 |
| chr7_-_22259845 | 0.57 |
ENST00000420196.1 |
RAPGEF5 |
Rap guanine nucleotide exchange factor (GEF) 5 |
| chr6_-_112115074 | 0.56 |
ENST00000368667.2 |
FYN |
FYN oncogene related to SRC, FGR, YES |
| chr19_+_56717283 | 0.56 |
ENST00000376267.1 |
ZSCAN5C |
zinc finger and SCAN domain containing 5C |
| chr1_+_46640750 | 0.56 |
ENST00000372003.1 |
TSPAN1 |
tetraspanin 1 |
| chr7_+_107110488 | 0.55 |
ENST00000304402.4 |
GPR22 |
G protein-coupled receptor 22 |
| chr19_+_45504688 | 0.54 |
ENST00000221452.8 ENST00000540120.1 ENST00000505236.1 |
RELB |
v-rel avian reticuloendotheliosis viral oncogene homolog B |
| chr7_-_99097863 | 0.54 |
ENST00000426306.2 ENST00000337673.6 |
ZNF394 |
zinc finger protein 394 |
| chr14_+_103589789 | 0.54 |
ENST00000558056.1 ENST00000560869.1 |
TNFAIP2 |
tumor necrosis factor, alpha-induced protein 2 |
| chr11_+_46316677 | 0.53 |
ENST00000534787.1 |
CREB3L1 |
cAMP responsive element binding protein 3-like 1 |
| chr5_+_49962495 | 0.52 |
ENST00000515175.1 |
PARP8 |
poly (ADP-ribose) polymerase family, member 8 |
| chr2_+_64681641 | 0.52 |
ENST00000409537.2 |
LGALSL |
lectin, galactoside-binding-like |
| chr8_-_65711310 | 0.52 |
ENST00000310193.3 |
CYP7B1 |
cytochrome P450, family 7, subfamily B, polypeptide 1 |
| chr1_-_40042073 | 0.52 |
ENST00000372858.3 |
PABPC4 |
poly(A) binding protein, cytoplasmic 4 (inducible form) |
| chr19_+_17416609 | 0.52 |
ENST00000602206.1 |
MRPL34 |
mitochondrial ribosomal protein L34 |
| chr1_-_40041925 | 0.52 |
ENST00000372862.3 |
PABPC4 |
poly(A) binding protein, cytoplasmic 4 (inducible form) |
| chr17_+_43209967 | 0.52 |
ENST00000431281.1 ENST00000591859.1 |
ACBD4 |
acyl-CoA binding domain containing 4 |
| chr19_+_48325097 | 0.52 |
ENST00000221996.7 ENST00000539067.1 |
CRX |
cone-rod homeobox |
| chr12_+_57998595 | 0.51 |
ENST00000337737.3 ENST00000548198.1 ENST00000551632.1 |
DTX3 |
deltex homolog 3 (Drosophila) |
| chr11_-_45940343 | 0.51 |
ENST00000532681.1 |
PEX16 |
peroxisomal biogenesis factor 16 |
| chr11_+_58910201 | 0.50 |
ENST00000528737.1 |
FAM111A |
family with sequence similarity 111, member A |
| chr5_+_149546334 | 0.49 |
ENST00000231656.8 |
CDX1 |
caudal type homeobox 1 |
| chr19_-_55770311 | 0.49 |
ENST00000412770.2 |
PPP6R1 |
protein phosphatase 6, regulatory subunit 1 |
| chr15_+_91643442 | 0.48 |
ENST00000394232.1 |
SV2B |
synaptic vesicle glycoprotein 2B |
| chr14_+_65878565 | 0.47 |
ENST00000556518.1 ENST00000557164.1 |
FUT8 |
fucosyltransferase 8 (alpha (1,6) fucosyltransferase) |
| chr17_+_64961026 | 0.47 |
ENST00000262138.3 |
CACNG4 |
calcium channel, voltage-dependent, gamma subunit 4 |
| chr6_+_31554779 | 0.47 |
ENST00000376090.2 |
LST1 |
leukocyte specific transcript 1 |
| chr3_-_11623804 | 0.45 |
ENST00000451674.2 |
VGLL4 |
vestigial like 4 (Drosophila) |
| chr1_+_46713404 | 0.45 |
ENST00000371975.4 ENST00000469835.1 |
RAD54L |
RAD54-like (S. cerevisiae) |
| chr1_-_31538517 | 0.45 |
ENST00000440538.2 ENST00000423018.2 ENST00000424085.2 ENST00000426105.2 ENST00000257075.5 ENST00000373747.3 ENST00000525843.1 ENST00000373742.2 |
PUM1 |
pumilio RNA-binding family member 1 |
| chr1_+_46713357 | 0.45 |
ENST00000442598.1 |
RAD54L |
RAD54-like (S. cerevisiae) |
| chr7_-_150497406 | 0.44 |
ENST00000492607.1 ENST00000326442.5 ENST00000450753.2 |
TMEM176B |
transmembrane protein 176B |
| chr19_-_52133588 | 0.44 |
ENST00000570106.2 |
SIGLEC5 |
sialic acid binding Ig-like lectin 5 |
| chr1_-_160001737 | 0.43 |
ENST00000368090.2 |
PIGM |
phosphatidylinositol glycan anchor biosynthesis, class M |
| chr2_-_100721178 | 0.43 |
ENST00000409236.2 |
AFF3 |
AF4/FMR2 family, member 3 |
| chr11_-_104480019 | 0.43 |
ENST00000536529.1 ENST00000545630.1 ENST00000538641.1 |
RP11-886D15.1 |
RP11-886D15.1 |
| chr17_-_39093672 | 0.43 |
ENST00000209718.3 ENST00000436344.3 ENST00000485751.1 |
KRT23 |
keratin 23 (histone deacetylase inducible) |
| chr3_-_121264848 | 0.42 |
ENST00000264233.5 |
POLQ |
polymerase (DNA directed), theta |
| chr11_-_46142615 | 0.42 |
ENST00000529734.1 ENST00000323180.6 |
PHF21A |
PHD finger protein 21A |
| chr3_-_119813264 | 0.42 |
ENST00000264235.8 |
GSK3B |
glycogen synthase kinase 3 beta |
| chr1_-_101491319 | 0.42 |
ENST00000342173.7 ENST00000488176.1 ENST00000370109.3 |
DPH5 |
diphthamide biosynthesis 5 |
| chr11_+_63997750 | 0.41 |
ENST00000321685.3 |
DNAJC4 |
DnaJ (Hsp40) homolog, subfamily C, member 4 |
| chr6_-_111136513 | 0.41 |
ENST00000368911.3 |
CDK19 |
cyclin-dependent kinase 19 |
| chr11_-_119999611 | 0.41 |
ENST00000529044.1 |
TRIM29 |
tripartite motif containing 29 |
| chr14_-_106586656 | 0.40 |
ENST00000390602.2 |
IGHV3-13 |
immunoglobulin heavy variable 3-13 |
| chr11_-_119999539 | 0.40 |
ENST00000541857.1 |
TRIM29 |
tripartite motif containing 29 |
| chr12_+_53662073 | 0.40 |
ENST00000553219.1 ENST00000257934.4 |
ESPL1 |
extra spindle pole bodies homolog 1 (S. cerevisiae) |
| chr1_+_156611704 | 0.40 |
ENST00000329117.5 |
BCAN |
brevican |
| chr6_-_44281043 | 0.39 |
ENST00000244571.4 |
AARS2 |
alanyl-tRNA synthetase 2, mitochondrial |
| chr6_+_43484760 | 0.39 |
ENST00000372389.3 ENST00000372344.2 ENST00000304004.3 ENST00000423780.1 |
POLR1C |
polymerase (RNA) I polypeptide C, 30kDa |
| chr5_+_142149955 | 0.39 |
ENST00000378004.3 |
ARHGAP26 |
Rho GTPase activating protein 26 |
| chr2_-_200820459 | 0.38 |
ENST00000354611.4 |
TYW5 |
tRNA-yW synthesizing protein 5 |
| chr2_+_97481974 | 0.38 |
ENST00000377060.3 ENST00000305510.3 |
CNNM3 |
cyclin M3 |
| chr3_+_141205852 | 0.38 |
ENST00000286364.3 ENST00000452898.1 |
RASA2 |
RAS p21 protein activator 2 |
| chr7_-_8301768 | 0.38 |
ENST00000265577.7 |
ICA1 |
islet cell autoantigen 1, 69kDa |
| chr8_-_9008206 | 0.38 |
ENST00000310455.3 |
PPP1R3B |
protein phosphatase 1, regulatory subunit 3B |
| chr5_+_71014990 | 0.38 |
ENST00000296777.4 |
CARTPT |
CART prepropeptide |
| chr11_+_65190245 | 0.37 |
ENST00000499732.1 ENST00000501122.2 ENST00000601801.1 |
NEAT1 |
nuclear paraspeckle assembly transcript 1 (non-protein coding) |
| chr7_-_150497621 | 0.37 |
ENST00000434545.1 |
TMEM176B |
transmembrane protein 176B |
| chr3_-_101232019 | 0.37 |
ENST00000394095.2 ENST00000394091.1 ENST00000394094.2 ENST00000358203.3 ENST00000348610.3 ENST00000314261.7 |
SENP7 |
SUMO1/sentrin specific peptidase 7 |
| chr4_+_79567314 | 0.36 |
ENST00000503539.1 ENST00000504675.1 |
RP11-792D21.2 |
long intergenic non-protein coding RNA 1094 |
| chr12_-_111021110 | 0.35 |
ENST00000354300.3 |
PPTC7 |
PTC7 protein phosphatase homolog (S. cerevisiae) |
| chr12_-_14133053 | 0.35 |
ENST00000609686.1 |
GRIN2B |
glutamate receptor, ionotropic, N-methyl D-aspartate 2B |
| chr2_-_169887827 | 0.34 |
ENST00000263817.6 |
ABCB11 |
ATP-binding cassette, sub-family B (MDR/TAP), member 11 |
| chr15_+_73976715 | 0.34 |
ENST00000558689.1 ENST00000560786.2 ENST00000561213.1 ENST00000563584.1 ENST00000561416.1 |
CD276 |
CD276 molecule |
| chr1_+_236686717 | 0.34 |
ENST00000341872.6 ENST00000450372.2 |
LGALS8 |
lectin, galactoside-binding, soluble, 8 |
| chr1_+_212738676 | 0.34 |
ENST00000366981.4 ENST00000366987.2 |
ATF3 |
activating transcription factor 3 |
| chr2_+_89999259 | 0.33 |
ENST00000558026.1 |
IGKV2D-28 |
immunoglobulin kappa variable 2D-28 |
| chr15_-_23034322 | 0.33 |
ENST00000539711.2 ENST00000560039.1 ENST00000398013.3 ENST00000337451.3 ENST00000359727.4 ENST00000398014.2 |
NIPA2 |
non imprinted in Prader-Willi/Angelman syndrome 2 |
| chr14_-_106622419 | 0.33 |
ENST00000390604.2 |
IGHV3-16 |
immunoglobulin heavy variable 3-16 (non-functional) |
| chr1_-_153066998 | 0.32 |
ENST00000368750.3 |
SPRR2E |
small proline-rich protein 2E |
| chr8_+_94752349 | 0.32 |
ENST00000391680.1 |
RBM12B-AS1 |
RBM12B antisense RNA 1 |
| chr17_-_9862772 | 0.32 |
ENST00000580865.1 ENST00000583882.1 |
GAS7 |
growth arrest-specific 7 |
| chr6_-_41863098 | 0.32 |
ENST00000373006.1 |
USP49 |
ubiquitin specific peptidase 49 |
| chr7_+_150497491 | 0.32 |
ENST00000484928.1 |
TMEM176A |
transmembrane protein 176A |
| chr12_+_53662110 | 0.32 |
ENST00000552462.1 |
ESPL1 |
extra spindle pole bodies homolog 1 (S. cerevisiae) |
| chr3_+_57261743 | 0.32 |
ENST00000288266.3 |
APPL1 |
adaptor protein, phosphotyrosine interaction, PH domain and leucine zipper containing 1 |
| chr1_-_169599314 | 0.32 |
ENST00000367786.2 ENST00000458599.2 ENST00000367795.2 ENST00000263686.6 |
SELP |
selectin P (granule membrane protein 140kDa, antigen CD62) |
| chr5_+_150040403 | 0.32 |
ENST00000517768.1 ENST00000297130.4 |
MYOZ3 |
myozenin 3 |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 17.9 | GO:0032395 | MHC class II receptor activity(GO:0032395) |
| 0.6 | 2.6 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.6 | 3.5 | GO:0046979 | TAP2 binding(GO:0046979) |
| 0.5 | 5.4 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.5 | 5.6 | GO:0004568 | chitinase activity(GO:0004568) chitin binding(GO:0008061) |
| 0.3 | 1.0 | GO:0017130 | poly(C) RNA binding(GO:0017130) |
| 0.3 | 2.4 | GO:0052739 | phosphatidylserine 1-acylhydrolase activity(GO:0052739) |
| 0.3 | 2.1 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.3 | 0.9 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 0.3 | 3.0 | GO:0022820 | potassium:chloride symporter activity(GO:0015379) potassium ion symporter activity(GO:0022820) |
| 0.3 | 14.5 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.2 | 1.2 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.2 | 2.3 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.2 | 0.6 | GO:0042806 | fucose binding(GO:0042806) |
| 0.2 | 1.2 | GO:0016531 | copper chaperone activity(GO:0016531) |
| 0.2 | 3.2 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.2 | 0.9 | GO:0043120 | tumor necrosis factor binding(GO:0043120) |
| 0.2 | 2.9 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.2 | 2.6 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.2 | 0.5 | GO:0008424 | glycoprotein 6-alpha-L-fucosyltransferase activity(GO:0008424) alpha-(1->6)-fucosyltransferase activity(GO:0046921) |
| 0.2 | 4.2 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.2 | 0.8 | GO:0038062 | protein tyrosine kinase collagen receptor activity(GO:0038062) |
| 0.2 | 1.2 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.1 | 0.4 | GO:1990404 | protein ADP-ribosylase activity(GO:1990404) |
| 0.1 | 1.0 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.1 | 6.6 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.1 | 0.4 | GO:0004813 | alanine-tRNA ligase activity(GO:0004813) |
| 0.1 | 1.4 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) corticotropin-releasing hormone receptor binding(GO:0051429) corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.1 | 2.8 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.1 | 0.5 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.1 | 0.6 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.1 | 0.7 | GO:0004372 | glycine hydroxymethyltransferase activity(GO:0004372) threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.1 | 0.6 | GO:0042610 | CD8 receptor binding(GO:0042610) |
| 0.1 | 3.2 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 2.5 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 3.5 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 0.8 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 0.7 | GO:0001094 | TFIID-class transcription factor binding(GO:0001094) |
| 0.1 | 9.4 | GO:0003823 | antigen binding(GO:0003823) |
| 0.1 | 4.0 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.1 | 0.7 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.1 | 1.2 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 0.4 | GO:0004906 | interferon-gamma receptor activity(GO:0004906) |
| 0.1 | 0.3 | GO:0003990 | acetylcholinesterase activity(GO:0003990) |
| 0.1 | 2.5 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 1.3 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.1 | 4.4 | GO:0004520 | endodeoxyribonuclease activity(GO:0004520) |
| 0.1 | 2.6 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.1 | 0.7 | GO:0036310 | annealing helicase activity(GO:0036310) |
| 0.1 | 0.9 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.1 | 0.8 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.1 | 0.4 | GO:0043142 | single-stranded DNA-dependent ATPase activity(GO:0043142) 5'-deoxyribose-5-phosphate lyase activity(GO:0051575) |
| 0.1 | 0.3 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.1 | 0.3 | GO:0086089 | voltage-gated potassium channel activity involved in atrial cardiac muscle cell action potential repolarization(GO:0086089) |
| 0.1 | 0.7 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.1 | 0.3 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.1 | 0.2 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 2.1 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.0 | 0.8 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.0 | 0.8 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.0 | 0.2 | GO:0004980 | melanocyte-stimulating hormone receptor activity(GO:0004980) |
| 0.0 | 0.3 | GO:0015125 | bile acid transmembrane transporter activity(GO:0015125) |
| 0.0 | 0.7 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 0.6 | GO:0000981 | RNA polymerase II transcription factor activity, sequence-specific DNA binding(GO:0000981) |
| 0.0 | 0.3 | GO:0043426 | MRF binding(GO:0043426) |
| 0.0 | 0.6 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.1 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.0 | 0.1 | GO:0008386 | cholesterol monooxygenase (side-chain-cleaving) activity(GO:0008386) |
| 0.0 | 0.9 | GO:0004003 | ATP-dependent DNA helicase activity(GO:0004003) |
| 0.0 | 0.1 | GO:0042019 | interleukin-23 binding(GO:0042019) interleukin-23 receptor activity(GO:0042020) |
| 0.0 | 1.2 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.7 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.0 | 0.2 | GO:0016416 | O-palmitoyltransferase activity(GO:0016416) |
| 0.0 | 0.2 | GO:0097322 | 7SK snRNA binding(GO:0097322) |
| 0.0 | 0.2 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.0 | 1.1 | GO:0001071 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
| 0.0 | 0.3 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.2 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.0 | 0.0 | GO:0001224 | RNA polymerase II transcription cofactor binding(GO:0001224) |
| 0.0 | 0.3 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 6.1 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 0.1 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.0 | 0.5 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.0 | 0.1 | GO:0004489 | methylenetetrahydrofolate reductase (NAD(P)H) activity(GO:0004489) |
| 0.0 | 0.3 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.0 | 0.7 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.1 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.0 | 1.4 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.2 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.4 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.5 | GO:0008395 | steroid hydroxylase activity(GO:0008395) |
| 0.0 | 0.1 | GO:0004996 | thyroid-stimulating hormone receptor activity(GO:0004996) |
| 0.0 | 0.3 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.3 | GO:0045295 | estrogen receptor activity(GO:0030284) gamma-catenin binding(GO:0045295) |
| 0.0 | 0.5 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.2 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) |
| 0.0 | 0.3 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 1.3 | GO:0043021 | ribonucleoprotein complex binding(GO:0043021) |
| 0.0 | 0.1 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.0 | 0.5 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.1 | GO:0022889 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.0 | 0.2 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 0.1 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 0.2 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.1 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.0 | 0.3 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.0 | 0.3 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 8.0 | GO:0002503 | peptide antigen assembly with MHC class II protein complex(GO:0002503) |
| 0.9 | 2.6 | GO:1905246 | regulation of choline O-acetyltransferase activity(GO:1902769) positive regulation of choline O-acetyltransferase activity(GO:1902771) negative regulation of tau-protein kinase activity(GO:1902948) positive regulation of early endosome to recycling endosome transport(GO:1902955) negative regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902960) negative regulation of neurofibrillary tangle assembly(GO:1902997) negative regulation of aspartic-type peptidase activity(GO:1905246) |
| 0.8 | 2.3 | GO:0034146 | B-1 B cell homeostasis(GO:0001922) toll-like receptor 5 signaling pathway(GO:0034146) regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070428) |
| 0.7 | 2.1 | GO:0070427 | nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070427) |
| 0.7 | 2.1 | GO:0052250 | modulation of signal transduction in other organism(GO:0044501) modulation by symbiont of host signal transduction pathway(GO:0052027) modulation of signal transduction in other organism involved in symbiotic interaction(GO:0052250) modulation by symbiont of host I-kappaB kinase/NF-kappaB cascade(GO:0085032) |
| 0.7 | 2.7 | GO:2000620 | positive regulation of histone H4-K16 acetylation(GO:2000620) |
| 0.7 | 2.7 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.6 | 5.9 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 0.6 | 4.0 | GO:0045425 | regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045423) positive regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045425) |
| 0.5 | 2.6 | GO:0002901 | mature B cell apoptotic process(GO:0002901) regulation of mature B cell apoptotic process(GO:0002905) negative regulation of mature B cell apoptotic process(GO:0002906) |
| 0.5 | 1.5 | GO:0060585 | detection of peptidoglycan(GO:0032499) activation of MAPK activity involved in innate immune response(GO:0035419) regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 0.5 | 5.6 | GO:0006032 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.4 | 4.6 | GO:0060753 | regulation of mast cell chemotaxis(GO:0060753) |
| 0.4 | 7.1 | GO:0072540 | T-helper 17 cell lineage commitment(GO:0072540) |
| 0.4 | 6.6 | GO:0032688 | negative regulation of interferon-beta production(GO:0032688) |
| 0.4 | 3.4 | GO:0045078 | positive regulation of interferon-gamma biosynthetic process(GO:0045078) |
| 0.3 | 1.3 | GO:0006408 | snRNA export from nucleus(GO:0006408) |
| 0.3 | 2.3 | GO:0033590 | response to cobalamin(GO:0033590) |
| 0.3 | 1.2 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.3 | 1.1 | GO:0000255 | allantoin metabolic process(GO:0000255) |
| 0.3 | 18.7 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.3 | 0.8 | GO:1900169 | regulation of glucocorticoid mediated signaling pathway(GO:1900169) |
| 0.2 | 0.7 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.2 | 1.0 | GO:2000110 | negative regulation of macrophage apoptotic process(GO:2000110) |
| 0.2 | 1.2 | GO:1901509 | regulation of endothelial tube morphogenesis(GO:1901509) |
| 0.2 | 0.9 | GO:2000537 | regulation of cGMP-mediated signaling(GO:0010752) regulation of B cell chemotaxis(GO:2000537) positive regulation of B cell chemotaxis(GO:2000538) |
| 0.2 | 4.0 | GO:0010623 | programmed cell death involved in cell development(GO:0010623) |
| 0.2 | 1.4 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
| 0.2 | 0.6 | GO:0044828 | nuclear-transcribed mRNA catabolic process, endonucleolytic cleavage-dependent decay(GO:0000294) negative regulation by host of viral genome replication(GO:0044828) |
| 0.2 | 0.9 | GO:0097527 | necroptotic signaling pathway(GO:0097527) |
| 0.2 | 0.7 | GO:0018364 | peptidyl-glutamine methylation(GO:0018364) |
| 0.2 | 0.5 | GO:0070093 | negative regulation of glucagon secretion(GO:0070093) |
| 0.2 | 0.7 | GO:0045875 | negative regulation of sister chromatid cohesion(GO:0045875) |
| 0.2 | 0.7 | GO:0090296 | regulation of mitochondrial DNA replication(GO:0090296) |
| 0.2 | 3.5 | GO:0071803 | positive regulation of podosome assembly(GO:0071803) |
| 0.2 | 1.2 | GO:0042791 | 5S class rRNA transcription from RNA polymerase III type 1 promoter(GO:0042791) tRNA transcription from RNA polymerase III promoter(GO:0042797) |
| 0.2 | 0.5 | GO:0032581 | ER-dependent peroxisome organization(GO:0032581) |
| 0.2 | 0.5 | GO:0014807 | regulation of somitogenesis(GO:0014807) |
| 0.2 | 0.6 | GO:1900186 | negative regulation of clathrin-mediated endocytosis(GO:1900186) |
| 0.2 | 1.8 | GO:0039530 | MDA-5 signaling pathway(GO:0039530) |
| 0.2 | 0.5 | GO:0036071 | N-glycan fucosylation(GO:0036071) |
| 0.2 | 1.4 | GO:2001199 | negative regulation of dendritic cell differentiation(GO:2001199) |
| 0.2 | 1.2 | GO:0006689 | ganglioside catabolic process(GO:0006689) |
| 0.2 | 0.5 | GO:1900246 | positive regulation of RIG-I signaling pathway(GO:1900246) |
| 0.1 | 0.4 | GO:0018312 | peptidyl-serine ADP-ribosylation(GO:0018312) |
| 0.1 | 0.4 | GO:0071109 | superior temporal gyrus development(GO:0071109) |
| 0.1 | 0.4 | GO:0006419 | alanyl-tRNA aminoacylation(GO:0006419) |
| 0.1 | 0.5 | GO:0046532 | regulation of photoreceptor cell differentiation(GO:0046532) |
| 0.1 | 14.7 | GO:0031295 | T cell costimulation(GO:0031295) |
| 0.1 | 0.3 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.1 | 0.5 | GO:0035754 | B cell chemotaxis(GO:0035754) |
| 0.1 | 2.4 | GO:0036150 | phosphatidylserine acyl-chain remodeling(GO:0036150) |
| 0.1 | 0.7 | GO:0019264 | glycine biosynthetic process from serine(GO:0019264) |
| 0.1 | 0.3 | GO:0045212 | negative regulation of synaptic transmission, cholinergic(GO:0032223) neurotransmitter receptor biosynthetic process(GO:0045212) |
| 0.1 | 0.3 | GO:1903625 | negative regulation of DNA catabolic process(GO:1903625) |
| 0.1 | 1.2 | GO:0097034 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.1 | 0.8 | GO:1902959 | regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902959) regulation of aspartic-type peptidase activity(GO:1905245) |
| 0.1 | 0.9 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.1 | 0.8 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 | 8.3 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.1 | 2.9 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.1 | 0.3 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.1 | 0.2 | GO:0032290 | peripheral nervous system myelin formation(GO:0032290) |
| 0.1 | 2.1 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.1 | 0.3 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.1 | 0.3 | GO:1904977 | lymphatic endothelial cell migration(GO:1904977) |
| 0.1 | 0.3 | GO:0061743 | motor learning(GO:0061743) |
| 0.1 | 0.3 | GO:0030242 | pexophagy(GO:0030242) |
| 0.1 | 0.7 | GO:0000733 | DNA strand renaturation(GO:0000733) |
| 0.1 | 0.3 | GO:0071895 | odontoblast differentiation(GO:0071895) |
| 0.1 | 0.2 | GO:0043163 | cell envelope organization(GO:0043163) external encapsulating structure organization(GO:0045229) |
| 0.1 | 0.1 | GO:2000296 | negative regulation of hydrogen peroxide catabolic process(GO:2000296) |
| 0.1 | 1.1 | GO:0071243 | cellular response to arsenic-containing substance(GO:0071243) |
| 0.1 | 1.8 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.1 | 0.3 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.1 | 2.0 | GO:0034629 | cellular protein complex localization(GO:0034629) |
| 0.1 | 0.5 | GO:1990440 | extracellular matrix constituent secretion(GO:0070278) positive regulation of transcription from RNA polymerase II promoter in response to endoplasmic reticulum stress(GO:1990440) |
| 0.1 | 1.2 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 2.6 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 0.6 | GO:0010572 | positive regulation of platelet activation(GO:0010572) |
| 0.1 | 0.3 | GO:0098914 | membrane repolarization during atrial cardiac muscle cell action potential(GO:0098914) |
| 0.1 | 0.4 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.1 | 0.4 | GO:0017183 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.0 | 4.4 | GO:0035690 | cellular response to drug(GO:0035690) |
| 0.0 | 0.2 | GO:1902572 | anagen(GO:0042640) negative regulation of serine-type endopeptidase activity(GO:1900004) negative regulation of serine-type peptidase activity(GO:1902572) |
| 0.0 | 0.2 | GO:0070980 | biphenyl catabolic process(GO:0070980) |
| 0.0 | 1.1 | GO:0035338 | long-chain fatty-acyl-CoA biosynthetic process(GO:0035338) |
| 0.0 | 6.4 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.3 | GO:0070235 | regulation of activation-induced cell death of T cells(GO:0070235) negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.3 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.0 | 0.3 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.0 | 0.1 | GO:2000612 | thyroid-stimulating hormone secretion(GO:0070460) regulation of thyroid-stimulating hormone secretion(GO:2000612) |
| 0.0 | 0.4 | GO:0046503 | glycerolipid catabolic process(GO:0046503) |
| 0.0 | 1.2 | GO:0042347 | negative regulation of NF-kappaB import into nucleus(GO:0042347) |
| 0.0 | 0.2 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.2 | GO:0032849 | regulation of cellular pH reduction(GO:0032847) positive regulation of cellular pH reduction(GO:0032849) |
| 0.0 | 0.2 | GO:0030070 | insulin processing(GO:0030070) |
| 0.0 | 0.1 | GO:0032328 | alanine transport(GO:0032328) |
| 0.0 | 0.1 | GO:0048631 | regulation of skeletal muscle tissue growth(GO:0048631) |
| 0.0 | 0.3 | GO:0061469 | regulation of type B pancreatic cell proliferation(GO:0061469) |
| 0.0 | 0.1 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.0 | 0.2 | GO:0021937 | cerebellar Purkinje cell-granule cell precursor cell signaling involved in regulation of granule cell precursor cell proliferation(GO:0021937) |
| 0.0 | 0.2 | GO:1904550 | chemotaxis to arachidonic acid(GO:0034670) response to arachidonic acid(GO:1904550) |
| 0.0 | 0.1 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.0 | 0.4 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.0 | 0.6 | GO:0046475 | glycerophospholipid catabolic process(GO:0046475) |
| 0.0 | 0.1 | GO:0045807 | positive regulation of endocytosis(GO:0045807) |
| 0.0 | 1.1 | GO:0007094 | mitotic spindle assembly checkpoint(GO:0007094) spindle assembly checkpoint(GO:0071173) |
| 0.0 | 0.5 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.0 | 0.2 | GO:1904627 | response to phorbol 13-acetate 12-myristate(GO:1904627) cellular response to phorbol 13-acetate 12-myristate(GO:1904628) |
| 0.0 | 1.0 | GO:0019835 | cytolysis(GO:0019835) |
| 0.0 | 0.1 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.0 | 3.8 | GO:0006821 | chloride transport(GO:0006821) |
| 0.0 | 0.5 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.0 | 3.5 | GO:0002223 | stimulatory C-type lectin receptor signaling pathway(GO:0002223) |
| 0.0 | 0.6 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 0.2 | GO:1901409 | positive regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901409) |
| 0.0 | 0.2 | GO:0045780 | positive regulation of bone resorption(GO:0045780) positive regulation of bone remodeling(GO:0046852) |
| 0.0 | 0.5 | GO:0030208 | dermatan sulfate biosynthetic process(GO:0030208) |
| 0.0 | 1.4 | GO:0010803 | regulation of tumor necrosis factor-mediated signaling pathway(GO:0010803) |
| 0.0 | 0.1 | GO:0032900 | negative regulation of low-density lipoprotein particle receptor catabolic process(GO:0032804) negative regulation of neurotrophin production(GO:0032900) negative regulation of transforming growth factor beta1 production(GO:0032911) |
| 0.0 | 1.5 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.0 | 0.1 | GO:0070829 | response to vitamin B2(GO:0033274) heterochromatin maintenance(GO:0070829) |
| 0.0 | 0.1 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.0 | 0.1 | GO:0060010 | Sertoli cell fate commitment(GO:0060010) |
| 0.0 | 0.1 | GO:1904274 | tricellular tight junction assembly(GO:1904274) |
| 0.0 | 0.7 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 1.1 | GO:0032945 | negative regulation of mononuclear cell proliferation(GO:0032945) negative regulation of lymphocyte proliferation(GO:0050672) |
| 0.0 | 0.1 | GO:1903071 | positive regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903071) |
| 0.0 | 0.4 | GO:0060334 | regulation of interferon-gamma-mediated signaling pathway(GO:0060334) |
| 0.0 | 0.3 | GO:0061157 | mRNA destabilization(GO:0061157) |
| 0.0 | 0.3 | GO:0030239 | myofibril assembly(GO:0030239) |
| 0.0 | 0.2 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 0.0 | 0.5 | GO:0043666 | regulation of phosphoprotein phosphatase activity(GO:0043666) |
| 0.0 | 0.2 | GO:0007617 | mating behavior(GO:0007617) multi-organism reproductive behavior(GO:0044705) |
| 0.0 | 0.7 | GO:1900181 | negative regulation of protein localization to nucleus(GO:1900181) |
| 0.0 | 0.1 | GO:0042754 | histone H3-K9 dimethylation(GO:0036123) negative regulation of circadian rhythm(GO:0042754) |
| 0.0 | 1.2 | GO:0007631 | feeding behavior(GO:0007631) |
| 0.0 | 0.5 | GO:0001510 | RNA methylation(GO:0001510) |
| 0.0 | 0.1 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.0 | 0.3 | GO:0048169 | regulation of long-term neuronal synaptic plasticity(GO:0048169) |
| 0.0 | 0.1 | GO:0042711 | maternal behavior(GO:0042711) parental behavior(GO:0060746) |
| 0.0 | 0.3 | GO:1900027 | regulation of ruffle assembly(GO:1900027) |
| 0.0 | 0.1 | GO:0006540 | glutamate decarboxylation to succinate(GO:0006540) |
| 0.0 | 0.4 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.0 | 0.3 | GO:0046710 | GDP metabolic process(GO:0046710) |
| 0.0 | 0.2 | GO:0043568 | positive regulation of insulin-like growth factor receptor signaling pathway(GO:0043568) |
| 0.0 | 0.2 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.0 | 0.3 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 24.6 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.7 | 8.7 | GO:0033256 | I-kappaB/NF-kappaB complex(GO:0033256) |
| 0.5 | 5.9 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.4 | 18.7 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.4 | 1.1 | GO:0033597 | mitotic checkpoint complex(GO:0033597) bub1-bub3 complex(GO:1990298) |
| 0.3 | 2.3 | GO:0043196 | varicosity(GO:0043196) |
| 0.3 | 1.2 | GO:0005889 | hydrogen:potassium-exchanging ATPase complex(GO:0005889) |
| 0.3 | 2.1 | GO:0032010 | phagolysosome(GO:0032010) |
| 0.2 | 0.7 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.2 | 2.7 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.2 | 1.8 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.2 | 1.4 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.2 | 1.2 | GO:0098560 | cytoplasmic side of late endosome membrane(GO:0098560) |
| 0.2 | 1.2 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.2 | 3.5 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.2 | 2.6 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.2 | 1.2 | GO:0000127 | transcription factor TFIIIC complex(GO:0000127) |
| 0.1 | 2.8 | GO:0031254 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.1 | 0.3 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.1 | 0.6 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.1 | 2.4 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.1 | 0.3 | GO:0031251 | PAN complex(GO:0031251) |
| 0.1 | 0.7 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.1 | 0.3 | GO:0034272 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.1 | 1.3 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.1 | 0.3 | GO:0031436 | BRCA1-BARD1 complex(GO:0031436) |
| 0.1 | 0.6 | GO:0034715 | pICln-Sm protein complex(GO:0034715) |
| 0.1 | 0.9 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.1 | 0.9 | GO:0042555 | MCM complex(GO:0042555) |
| 0.1 | 4.4 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
| 0.1 | 2.9 | GO:0098636 | protein complex involved in cell adhesion(GO:0098636) |
| 0.1 | 0.7 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.0 | 0.3 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 2.2 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 0.3 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.0 | 0.6 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.0 | 0.4 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.0 | 0.4 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.0 | 0.2 | GO:0097209 | epidermal lamellar body(GO:0097209) |
| 0.0 | 0.3 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.0 | 0.1 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.0 | 0.1 | GO:0072536 | interleukin-23 receptor complex(GO:0072536) |
| 0.0 | 0.3 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.0 | 0.5 | GO:0000145 | exocyst(GO:0000145) |
| 0.0 | 0.2 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.0 | 0.5 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 1.6 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.6 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.4 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.0 | 0.6 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.0 | 0.2 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) |
| 0.0 | 1.1 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 1.2 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.0 | 0.1 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.0 | 0.5 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 0.1 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.0 | 0.2 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 4.1 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.0 | 0.1 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.0 | 1.2 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) |
| 0.0 | 1.3 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 0.5 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.8 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 0.4 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.1 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 8.0 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.2 | 7.9 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.2 | 7.8 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 3.1 | PID IL27 PATHWAY | IL27-mediated signaling events |
| 0.1 | 4.4 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 4.3 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 0.4 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 2.5 | PID EPO PATHWAY | EPO signaling pathway |
| 0.1 | 0.9 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.1 | 0.8 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 1.4 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 1.3 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 0.9 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 0.8 | PID THROMBIN PAR1 PATHWAY | PAR1-mediated thrombin signaling events |
| 0.0 | 0.8 | ST G ALPHA I PATHWAY | G alpha i Pathway |
| 0.0 | 0.6 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.6 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.0 | 0.9 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
| 0.0 | 0.4 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.0 | 0.9 | PID ATR PATHWAY | ATR signaling pathway |
| 0.0 | 0.2 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.0 | 1.1 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 1.1 | PID E2F PATHWAY | E2F transcription factor network |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 14.1 | REACTOME TRANSLOCATION OF ZAP 70 TO IMMUNOLOGICAL SYNAPSE | Genes involved in Translocation of ZAP-70 to Immunological synapse |
| 0.6 | 5.9 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.3 | 2.7 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.2 | 4.6 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.1 | 2.0 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.1 | 2.7 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 4.9 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.1 | 1.2 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 1.1 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.1 | 6.2 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.1 | 3.5 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.1 | 4.8 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 1.2 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.1 | 2.1 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 0.1 | 0.9 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.1 | 1.1 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.1 | 1.7 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.1 | 4.7 | REACTOME ACTIVATION OF NF KAPPAB IN B CELLS | Genes involved in Activation of NF-kappaB in B Cells |
| 0.1 | 6.7 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 1.1 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.0 | 1.0 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.0 | 0.9 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS | Genes involved in Synthesis of bile acids and bile salts |
| 0.0 | 1.9 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.0 | 0.8 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 0.8 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.0 | 1.8 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 1.2 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.3 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 0.5 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.5 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.0 | 1.2 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.4 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.0 | 0.2 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 0.6 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.5 | REACTOME N GLYCAN ANTENNAE ELONGATION IN THE MEDIAL TRANS GOLGI | Genes involved in N-glycan antennae elongation in the medial/trans-Golgi |
| 0.0 | 0.3 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 0.3 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.4 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.4 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 0.3 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.5 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.0 | 0.2 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.3 | REACTOME PIP3 ACTIVATES AKT SIGNALING | Genes involved in PIP3 activates AKT signaling |
| 0.0 | 0.2 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |