Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
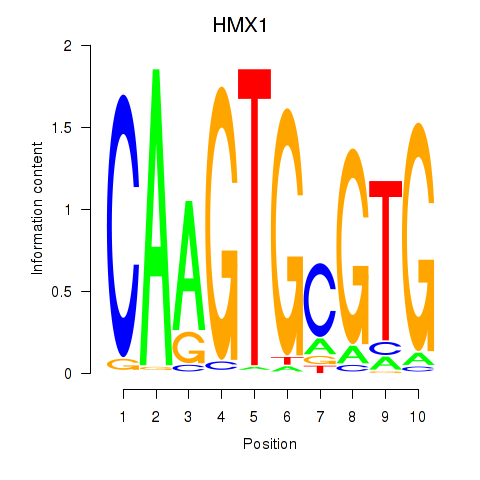
Results for HMX1
Z-value: 1.55
Transcription factors associated with HMX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HMX1
|
ENSG00000215612.5 | HMX1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HMX1 | hg19_v2_chr4_-_8873531_8873543 | -0.57 | 2.2e-02 | Click! |
Activity profile of HMX1 motif
Sorted Z-values of HMX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HMX1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_81046841 | 4.44 |
ENST00000509013.2 ENST00000505980.1 ENST00000509053.1 |
SSBP2 |
single-stranded DNA binding protein 2 |
| chr5_-_81046922 | 4.28 |
ENST00000514493.1 ENST00000320672.4 |
SSBP2 |
single-stranded DNA binding protein 2 |
| chr5_-_81046904 | 3.94 |
ENST00000515395.1 |
SSBP2 |
single-stranded DNA binding protein 2 |
| chr14_+_31343747 | 3.67 |
ENST00000216361.4 ENST00000396618.3 ENST00000475087.1 |
COCH |
cochlin |
| chr14_+_31343951 | 3.65 |
ENST00000556908.1 ENST00000555881.1 ENST00000460581.2 |
COCH |
cochlin |
| chr7_+_128470431 | 3.59 |
ENST00000325888.8 ENST00000346177.6 |
FLNC |
filamin C, gamma |
| chrX_+_37545012 | 3.13 |
ENST00000378616.3 |
XK |
X-linked Kx blood group (McLeod syndrome) |
| chr7_+_50344289 | 3.06 |
ENST00000413698.1 ENST00000359197.5 ENST00000331340.3 ENST00000357364.4 ENST00000343574.5 ENST00000349824.4 ENST00000346667.4 ENST00000440768.2 |
IKZF1 |
IKAROS family zinc finger 1 (Ikaros) |
| chr7_-_150675372 | 2.93 |
ENST00000262186.5 |
KCNH2 |
potassium voltage-gated channel, subfamily H (eag-related), member 2 |
| chr17_+_40440481 | 2.47 |
ENST00000590726.2 ENST00000452307.2 ENST00000444283.1 ENST00000588868.1 |
STAT5A |
signal transducer and activator of transcription 5A |
| chr11_-_417388 | 2.38 |
ENST00000332725.3 |
SIGIRR |
single immunoglobulin and toll-interleukin 1 receptor (TIR) domain |
| chr11_-_417308 | 2.25 |
ENST00000397632.3 ENST00000382520.2 |
SIGIRR |
single immunoglobulin and toll-interleukin 1 receptor (TIR) domain |
| chr15_-_58306295 | 2.10 |
ENST00000559517.1 |
ALDH1A2 |
aldehyde dehydrogenase 1 family, member A2 |
| chr3_+_151986709 | 2.05 |
ENST00000495875.2 ENST00000493459.1 ENST00000324210.5 ENST00000459747.1 |
MBNL1 |
muscleblind-like splicing regulator 1 |
| chr17_-_62097927 | 2.03 |
ENST00000578313.1 ENST00000584084.1 ENST00000579788.1 ENST00000579687.1 ENST00000578379.1 ENST00000578892.1 ENST00000412356.1 ENST00000418105.1 |
ICAM2 |
intercellular adhesion molecule 2 |
| chr5_-_37371163 | 1.93 |
ENST00000513532.1 |
NUP155 |
nucleoporin 155kDa |
| chr16_+_85645007 | 1.87 |
ENST00000405402.2 |
GSE1 |
Gse1 coiled-coil protein |
| chr17_+_79989500 | 1.86 |
ENST00000306897.4 |
RAC3 |
ras-related C3 botulinum toxin substrate 3 (rho family, small GTP binding protein Rac3) |
| chr17_+_79989937 | 1.85 |
ENST00000580965.1 |
RAC3 |
ras-related C3 botulinum toxin substrate 3 (rho family, small GTP binding protein Rac3) |
| chr13_+_42031679 | 1.82 |
ENST00000379359.3 |
RGCC |
regulator of cell cycle |
| chr17_-_62097904 | 1.81 |
ENST00000583366.1 |
ICAM2 |
intercellular adhesion molecule 2 |
| chr5_-_37371278 | 1.71 |
ENST00000231498.3 |
NUP155 |
nucleoporin 155kDa |
| chr1_-_28241024 | 1.66 |
ENST00000313433.7 ENST00000444045.1 |
RPA2 |
replication protein A2, 32kDa |
| chr8_+_145438870 | 1.66 |
ENST00000527931.1 |
FAM203B |
family with sequence similarity 203, member B |
| chr17_+_79990058 | 1.63 |
ENST00000584341.1 |
RAC3 |
ras-related C3 botulinum toxin substrate 3 (rho family, small GTP binding protein Rac3) |
| chr4_-_144940477 | 1.60 |
ENST00000513128.1 ENST00000429670.2 ENST00000502664.1 |
GYPB |
glycophorin B (MNS blood group) |
| chr16_+_88872176 | 1.58 |
ENST00000569140.1 |
CDT1 |
chromatin licensing and DNA replication factor 1 |
| chr1_+_15986364 | 1.58 |
ENST00000345034.1 |
RSC1A1 |
regulatory solute carrier protein, family 1, member 1 |
| chr1_-_51425902 | 1.46 |
ENST00000396153.2 |
FAF1 |
Fas (TNFRSF6) associated factor 1 |
| chr1_-_54405773 | 1.46 |
ENST00000371376.1 |
HSPB11 |
heat shock protein family B (small), member 11 |
| chr3_+_69788576 | 1.42 |
ENST00000352241.4 ENST00000448226.2 |
MITF |
microphthalmia-associated transcription factor |
| chr11_+_59522900 | 1.41 |
ENST00000529177.1 |
STX3 |
syntaxin 3 |
| chr3_+_20081515 | 1.38 |
ENST00000263754.4 |
KAT2B |
K(lysine) acetyltransferase 2B |
| chr1_-_26232951 | 1.34 |
ENST00000426559.2 ENST00000455785.2 |
STMN1 |
stathmin 1 |
| chr4_-_144826682 | 1.24 |
ENST00000358615.4 ENST00000437468.2 |
GYPE |
glycophorin E (MNS blood group) |
| chr17_-_41132010 | 1.22 |
ENST00000409103.1 ENST00000360221.4 |
PTGES3L-AARSD1 |
PTGES3L-AARSD1 readthrough |
| chr6_+_33043703 | 1.15 |
ENST00000418931.2 ENST00000535465.1 |
HLA-DPB1 |
major histocompatibility complex, class II, DP beta 1 |
| chr6_+_163835669 | 1.14 |
ENST00000453779.2 ENST00000275262.7 ENST00000392127.2 ENST00000361752.3 |
QKI |
QKI, KH domain containing, RNA binding |
| chr1_-_51425772 | 1.06 |
ENST00000371778.4 |
FAF1 |
Fas (TNFRSF6) associated factor 1 |
| chr3_-_13461807 | 1.05 |
ENST00000254508.5 |
NUP210 |
nucleoporin 210kDa |
| chr9_-_74525658 | 1.04 |
ENST00000333421.6 |
ABHD17B |
abhydrolase domain containing 17B |
| chr20_+_30639991 | 1.04 |
ENST00000534862.1 ENST00000538448.1 ENST00000375862.2 |
HCK |
hemopoietic cell kinase |
| chr11_+_1411503 | 1.04 |
ENST00000526678.1 |
BRSK2 |
BR serine/threonine kinase 2 |
| chr11_+_59522837 | 1.04 |
ENST00000437946.2 |
STX3 |
syntaxin 3 |
| chr17_-_47755436 | 1.03 |
ENST00000505581.1 ENST00000514121.1 ENST00000393328.2 ENST00000509079.1 ENST00000393331.3 ENST00000347630.2 ENST00000504102.1 |
SPOP |
speckle-type POZ protein |
| chr2_+_46769798 | 1.03 |
ENST00000238738.4 |
RHOQ |
ras homolog family member Q |
| chr11_+_105948216 | 1.02 |
ENST00000278618.4 |
AASDHPPT |
aminoadipate-semialdehyde dehydrogenase-phosphopantetheinyl transferase |
| chr6_-_32157947 | 0.99 |
ENST00000375050.4 |
PBX2 |
pre-B-cell leukemia homeobox 2 |
| chr20_+_30640004 | 0.98 |
ENST00000520553.1 ENST00000518730.1 ENST00000375852.2 |
HCK |
hemopoietic cell kinase |
| chr5_+_10353780 | 0.97 |
ENST00000449913.2 ENST00000503788.1 ENST00000274140.5 |
MARCH6 |
membrane-associated ring finger (C3HC4) 6, E3 ubiquitin protein ligase |
| chr16_+_85646763 | 0.95 |
ENST00000411612.1 ENST00000253458.7 |
GSE1 |
Gse1 coiled-coil protein |
| chr1_+_150521876 | 0.95 |
ENST00000369041.5 ENST00000271643.4 ENST00000538795.1 |
ADAMTSL4 AL356356.1 |
ADAMTS-like 4 Protein LOC100996516 |
| chr16_-_1823114 | 0.94 |
ENST00000177742.3 ENST00000397375.2 |
MRPS34 |
mitochondrial ribosomal protein S34 |
| chr20_-_41818373 | 0.93 |
ENST00000373187.1 ENST00000356100.2 ENST00000373184.1 ENST00000373190.1 |
PTPRT |
protein tyrosine phosphatase, receptor type, T |
| chr3_-_49066811 | 0.93 |
ENST00000442157.1 ENST00000326739.4 |
IMPDH2 |
IMP (inosine 5'-monophosphate) dehydrogenase 2 |
| chr1_-_25256368 | 0.92 |
ENST00000308873.6 |
RUNX3 |
runt-related transcription factor 3 |
| chr9_-_126692386 | 0.92 |
ENST00000373624.2 ENST00000394219.3 ENST00000373620.3 ENST00000394215.2 ENST00000373618.1 |
DENND1A |
DENN/MADD domain containing 1A |
| chr11_+_66624527 | 0.92 |
ENST00000393952.3 |
LRFN4 |
leucine rich repeat and fibronectin type III domain containing 4 |
| chrX_+_69509927 | 0.89 |
ENST00000374403.3 |
KIF4A |
kinesin family member 4A |
| chr1_+_110163709 | 0.84 |
ENST00000369840.2 ENST00000527846.1 |
AMPD2 |
adenosine monophosphate deaminase 2 |
| chr2_+_30454390 | 0.84 |
ENST00000395323.3 ENST00000406087.1 ENST00000404397.1 |
LBH |
limb bud and heart development |
| chr14_-_55369525 | 0.83 |
ENST00000543643.2 ENST00000536224.2 ENST00000395514.1 ENST00000491895.2 |
GCH1 |
GTP cyclohydrolase 1 |
| chr17_-_55038375 | 0.83 |
ENST00000240316.4 |
COIL |
coilin |
| chr13_+_43148281 | 0.82 |
ENST00000239849.6 ENST00000398795.2 ENST00000544862.1 |
TNFSF11 |
tumor necrosis factor (ligand) superfamily, member 11 |
| chr1_-_28241226 | 0.82 |
ENST00000373912.3 ENST00000373909.3 |
RPA2 |
replication protein A2, 32kDa |
| chr17_-_78450398 | 0.81 |
ENST00000306773.4 |
NPTX1 |
neuronal pentraxin I |
| chr22_-_39268308 | 0.80 |
ENST00000407418.3 |
CBX6 |
chromobox homolog 6 |
| chr6_-_31550192 | 0.80 |
ENST00000429299.2 ENST00000446745.2 |
LTB |
lymphotoxin beta (TNF superfamily, member 3) |
| chrX_-_153744507 | 0.80 |
ENST00000442929.1 ENST00000426266.1 ENST00000359889.5 ENST00000369641.3 ENST00000447601.2 ENST00000434658.2 |
FAM3A |
family with sequence similarity 3, member A |
| chr4_+_86396265 | 0.80 |
ENST00000395184.1 |
ARHGAP24 |
Rho GTPase activating protein 24 |
| chr17_-_41132410 | 0.80 |
ENST00000409446.3 ENST00000453594.1 ENST00000409399.1 ENST00000421990.2 |
PTGES3L PTGES3L-AARSD1 |
prostaglandin E synthase 3 (cytosolic)-like PTGES3L-AARSD1 readthrough |
| chrX_-_153744434 | 0.77 |
ENST00000369643.1 ENST00000393572.1 |
FAM3A |
family with sequence similarity 3, member A |
| chr6_+_30524663 | 0.76 |
ENST00000376560.3 |
PRR3 |
proline rich 3 |
| chr9_-_74525847 | 0.76 |
ENST00000377041.2 |
ABHD17B |
abhydrolase domain containing 17B |
| chr7_-_752577 | 0.75 |
ENST00000544935.1 ENST00000430040.1 ENST00000456696.2 ENST00000406797.1 |
PRKAR1B |
protein kinase, cAMP-dependent, regulatory, type I, beta |
| chr12_-_7077125 | 0.74 |
ENST00000545555.2 |
PHB2 |
prohibitin 2 |
| chr6_+_20403997 | 0.74 |
ENST00000535432.1 |
E2F3 |
E2F transcription factor 3 |
| chr11_-_128392085 | 0.73 |
ENST00000526145.2 ENST00000531611.1 ENST00000319397.6 ENST00000345075.4 ENST00000535549.1 |
ETS1 |
v-ets avian erythroblastosis virus E26 oncogene homolog 1 |
| chr9_-_136024721 | 0.73 |
ENST00000393160.3 |
RALGDS |
ral guanine nucleotide dissociation stimulator |
| chr1_-_24194771 | 0.71 |
ENST00000374479.3 |
FUCA1 |
fucosidase, alpha-L- 1, tissue |
| chr19_-_14316980 | 0.70 |
ENST00000361434.3 ENST00000340736.6 |
LPHN1 |
latrophilin 1 |
| chr19_-_13068012 | 0.68 |
ENST00000316939.1 |
GADD45GIP1 |
growth arrest and DNA-damage-inducible, gamma interacting protein 1 |
| chr11_-_62607036 | 0.67 |
ENST00000311713.7 ENST00000278856.4 |
WDR74 |
WD repeat domain 74 |
| chr6_-_89927151 | 0.67 |
ENST00000454853.2 |
GABRR1 |
gamma-aminobutyric acid (GABA) A receptor, rho 1 |
| chr1_-_26232522 | 0.66 |
ENST00000399728.1 |
STMN1 |
stathmin 1 |
| chr3_-_138553779 | 0.66 |
ENST00000461451.1 |
PIK3CB |
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit beta |
| chr6_-_7911042 | 0.66 |
ENST00000379757.4 |
TXNDC5 |
thioredoxin domain containing 5 (endoplasmic reticulum) |
| chr21_-_40685477 | 0.65 |
ENST00000342449.3 |
BRWD1 |
bromodomain and WD repeat domain containing 1 |
| chr1_+_110163202 | 0.65 |
ENST00000531203.1 ENST00000256578.3 |
AMPD2 |
adenosine monophosphate deaminase 2 |
| chrX_+_110924346 | 0.65 |
ENST00000371979.3 ENST00000251943.4 ENST00000486353.1 ENST00000394780.3 ENST00000495283.1 |
ALG13 |
ALG13, UDP-N-acetylglucosaminyltransferase subunit |
| chrX_+_131157290 | 0.64 |
ENST00000394334.2 |
MST4 |
Serine/threonine-protein kinase MST4 |
| chr3_-_138553594 | 0.63 |
ENST00000477593.1 ENST00000483968.1 |
PIK3CB |
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit beta |
| chr14_-_101034407 | 0.63 |
ENST00000443071.2 ENST00000557378.1 |
BEGAIN |
brain-enriched guanylate kinase-associated |
| chr19_-_1401486 | 0.62 |
ENST00000252288.2 ENST00000447102.3 |
GAMT |
guanidinoacetate N-methyltransferase |
| chr19_+_41305406 | 0.62 |
ENST00000406058.2 ENST00000593726.1 |
EGLN2 |
egl-9 family hypoxia-inducible factor 2 |
| chr11_+_6502675 | 0.61 |
ENST00000254616.6 ENST00000530751.1 |
TIMM10B |
translocase of inner mitochondrial membrane 10 homolog B (yeast) |
| chr7_-_37488834 | 0.61 |
ENST00000310758.4 |
ELMO1 |
engulfment and cell motility 1 |
| chr21_-_42879909 | 0.60 |
ENST00000458356.1 ENST00000398585.3 ENST00000424093.1 |
TMPRSS2 |
transmembrane protease, serine 2 |
| chr1_-_153949751 | 0.60 |
ENST00000428469.1 |
JTB |
jumping translocation breakpoint |
| chrX_+_131157322 | 0.60 |
ENST00000481105.1 ENST00000354719.6 ENST00000394335.2 |
MST4 |
Serine/threonine-protein kinase MST4 |
| chr6_-_8102279 | 0.59 |
ENST00000488226.2 |
EEF1E1 |
eukaryotic translation elongation factor 1 epsilon 1 |
| chr1_+_110163682 | 0.59 |
ENST00000358729.4 |
AMPD2 |
adenosine monophosphate deaminase 2 |
| chr1_+_193091080 | 0.59 |
ENST00000367435.3 |
CDC73 |
cell division cycle 73 |
| chr19_+_41313017 | 0.58 |
ENST00000595621.1 ENST00000595051.1 |
EGLN2 |
egl-9 family hypoxia-inducible factor 2 |
| chr17_+_79679299 | 0.57 |
ENST00000331531.5 |
SLC25A10 |
solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr20_-_41818536 | 0.57 |
ENST00000373193.3 ENST00000373198.4 ENST00000373201.1 |
PTPRT |
protein tyrosine phosphatase, receptor type, T |
| chr22_-_37640456 | 0.57 |
ENST00000405484.1 ENST00000441619.1 ENST00000406508.1 |
RAC2 |
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr1_-_109940550 | 0.56 |
ENST00000256637.6 |
SORT1 |
sortilin 1 |
| chr14_+_90422239 | 0.56 |
ENST00000393452.3 ENST00000554180.1 ENST00000393454.2 ENST00000553617.1 ENST00000335725.4 ENST00000357382.3 ENST00000556867.1 ENST00000553527.1 |
TDP1 |
tyrosyl-DNA phosphodiesterase 1 |
| chr12_+_53689309 | 0.56 |
ENST00000351500.3 ENST00000550846.1 ENST00000334478.4 ENST00000549759.1 |
PFDN5 |
prefoldin subunit 5 |
| chr1_+_95583479 | 0.55 |
ENST00000455656.1 ENST00000604534.1 |
TMEM56 RP11-57H12.6 |
transmembrane protein 56 TMEM56-RWDD3 readthrough |
| chr6_-_8102714 | 0.55 |
ENST00000502429.1 ENST00000429723.2 ENST00000507463.1 ENST00000379715.5 |
EEF1E1 |
eukaryotic translation elongation factor 1 epsilon 1 |
| chr4_+_83956312 | 0.55 |
ENST00000509317.1 ENST00000503682.1 ENST00000511653.1 |
COPS4 |
COP9 signalosome subunit 4 |
| chr4_+_83956237 | 0.54 |
ENST00000264389.2 |
COPS4 |
COP9 signalosome subunit 4 |
| chr1_-_115259337 | 0.53 |
ENST00000369535.4 |
NRAS |
neuroblastoma RAS viral (v-ras) oncogene homolog |
| chr14_+_72399114 | 0.52 |
ENST00000553525.1 ENST00000555571.1 |
RGS6 |
regulator of G-protein signaling 6 |
| chr22_-_37640277 | 0.52 |
ENST00000401529.3 ENST00000249071.6 |
RAC2 |
ras-related C3 botulinum toxin substrate 2 (rho family, small GTP binding protein Rac2) |
| chr15_+_81489213 | 0.51 |
ENST00000559383.1 ENST00000394660.2 |
IL16 |
interleukin 16 |
| chr3_-_195270162 | 0.51 |
ENST00000438848.1 ENST00000328432.3 |
PPP1R2 |
protein phosphatase 1, regulatory (inhibitor) subunit 2 |
| chrX_+_135579670 | 0.50 |
ENST00000218364.4 |
HTATSF1 |
HIV-1 Tat specific factor 1 |
| chr1_-_46598284 | 0.49 |
ENST00000423209.1 ENST00000262741.5 |
PIK3R3 |
phosphoinositide-3-kinase, regulatory subunit 3 (gamma) |
| chr12_+_70133152 | 0.49 |
ENST00000550536.1 ENST00000362025.5 |
RAB3IP |
RAB3A interacting protein |
| chr17_+_79679369 | 0.48 |
ENST00000350690.5 |
SLC25A10 |
solute carrier family 25 (mitochondrial carrier; dicarboxylate transporter), member 10 |
| chr21_-_42880075 | 0.48 |
ENST00000332149.5 |
TMPRSS2 |
transmembrane protease, serine 2 |
| chr15_+_89346699 | 0.48 |
ENST00000558207.1 |
ACAN |
aggrecan |
| chr11_-_19262486 | 0.48 |
ENST00000250024.4 |
E2F8 |
E2F transcription factor 8 |
| chr5_+_156693159 | 0.48 |
ENST00000347377.6 |
CYFIP2 |
cytoplasmic FMR1 interacting protein 2 |
| chr1_-_207095324 | 0.47 |
ENST00000530505.1 ENST00000367091.3 ENST00000442471.2 |
FAIM3 |
Fas apoptotic inhibitory molecule 3 |
| chr2_-_175462456 | 0.46 |
ENST00000409891.1 ENST00000410117.1 |
WIPF1 |
WAS/WASL interacting protein family, member 1 |
| chr17_-_8151353 | 0.45 |
ENST00000315684.8 |
CTC1 |
CTS telomere maintenance complex component 1 |
| chr2_-_175462934 | 0.45 |
ENST00000392546.2 ENST00000436221.1 |
WIPF1 |
WAS/WASL interacting protein family, member 1 |
| chr22_-_29196030 | 0.44 |
ENST00000405219.3 |
XBP1 |
X-box binding protein 1 |
| chr13_+_37574678 | 0.43 |
ENST00000389704.3 |
EXOSC8 |
exosome component 8 |
| chr14_-_25103388 | 0.43 |
ENST00000526004.1 ENST00000415355.3 |
GZMB |
granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) |
| chr1_-_207095212 | 0.43 |
ENST00000420007.2 |
FAIM3 |
Fas apoptotic inhibitory molecule 3 |
| chr5_+_156693091 | 0.43 |
ENST00000318218.6 ENST00000442283.2 ENST00000522463.1 ENST00000521420.1 |
CYFIP2 |
cytoplasmic FMR1 interacting protein 2 |
| chr11_-_67275542 | 0.42 |
ENST00000531506.1 |
CDK2AP2 |
cyclin-dependent kinase 2 associated protein 2 |
| chr2_-_24583168 | 0.42 |
ENST00000361999.3 |
ITSN2 |
intersectin 2 |
| chr5_+_176784837 | 0.42 |
ENST00000408923.3 |
RGS14 |
regulator of G-protein signaling 14 |
| chr12_-_49259643 | 0.41 |
ENST00000309739.5 |
RND1 |
Rho family GTPase 1 |
| chr3_+_39093481 | 0.41 |
ENST00000302313.5 ENST00000544962.1 ENST00000396258.3 ENST00000418020.1 |
WDR48 |
WD repeat domain 48 |
| chr12_+_7060432 | 0.41 |
ENST00000318974.9 ENST00000456013.1 |
PTPN6 |
protein tyrosine phosphatase, non-receptor type 6 |
| chrX_-_153599578 | 0.41 |
ENST00000360319.4 ENST00000344736.4 |
FLNA |
filamin A, alpha |
| chr17_-_79876010 | 0.41 |
ENST00000328666.6 |
SIRT7 |
sirtuin 7 |
| chr9_+_10613163 | 0.40 |
ENST00000429581.2 |
RP11-87N24.2 |
RP11-87N24.2 |
| chr1_-_46598371 | 0.40 |
ENST00000372006.1 ENST00000425892.1 ENST00000420542.1 ENST00000354242.4 ENST00000340332.6 |
PIK3R3 |
phosphoinositide-3-kinase, regulatory subunit 3 (gamma) |
| chr17_-_42298201 | 0.40 |
ENST00000527034.1 |
UBTF |
upstream binding transcription factor, RNA polymerase I |
| chr4_-_151936416 | 0.40 |
ENST00000510413.1 ENST00000507224.1 |
LRBA |
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr1_+_178995021 | 0.40 |
ENST00000263733.4 |
FAM20B |
family with sequence similarity 20, member B |
| chr1_+_235490659 | 0.39 |
ENST00000488594.1 |
GGPS1 |
geranylgeranyl diphosphate synthase 1 |
| chr14_-_25103472 | 0.39 |
ENST00000216341.4 ENST00000382542.1 ENST00000382540.1 |
GZMB |
granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) |
| chr4_-_151936865 | 0.39 |
ENST00000535741.1 |
LRBA |
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr2_-_27545431 | 0.38 |
ENST00000233545.2 |
MPV17 |
MpV17 mitochondrial inner membrane protein |
| chr12_+_70132632 | 0.38 |
ENST00000378815.6 ENST00000483530.2 ENST00000325555.9 |
RAB3IP |
RAB3A interacting protein |
| chrX_+_135579238 | 0.38 |
ENST00000535601.1 ENST00000448450.1 ENST00000425695.1 |
HTATSF1 |
HIV-1 Tat specific factor 1 |
| chr8_+_120428546 | 0.38 |
ENST00000259526.3 |
NOV |
nephroblastoma overexpressed |
| chr4_-_71705027 | 0.37 |
ENST00000545193.1 |
GRSF1 |
G-rich RNA sequence binding factor 1 |
| chr1_+_160336851 | 0.37 |
ENST00000302101.5 |
NHLH1 |
nescient helix loop helix 1 |
| chr3_-_52312636 | 0.37 |
ENST00000296490.3 |
WDR82 |
WD repeat domain 82 |
| chr4_-_109087872 | 0.37 |
ENST00000510624.1 |
LEF1 |
lymphoid enhancer-binding factor 1 |
| chr12_+_41831485 | 0.37 |
ENST00000539469.2 ENST00000298919.7 |
PDZRN4 |
PDZ domain containing ring finger 4 |
| chr16_+_19535133 | 0.36 |
ENST00000396212.2 ENST00000381396.5 |
CCP110 |
centriolar coiled coil protein 110kDa |
| chr22_-_39268192 | 0.36 |
ENST00000216083.6 |
CBX6 |
chromobox homolog 6 |
| chr4_-_140005443 | 0.36 |
ENST00000510408.1 ENST00000420916.2 ENST00000358635.3 |
ELF2 |
E74-like factor 2 (ets domain transcription factor) |
| chr17_-_8534067 | 0.36 |
ENST00000360416.3 ENST00000269243.4 |
MYH10 |
myosin, heavy chain 10, non-muscle |
| chr19_-_19006890 | 0.35 |
ENST00000247005.6 |
GDF1 |
growth differentiation factor 1 |
| chr16_+_67226019 | 0.35 |
ENST00000379378.3 |
E2F4 |
E2F transcription factor 4, p107/p130-binding |
| chr16_+_19535235 | 0.34 |
ENST00000565376.2 ENST00000396208.2 |
CCP110 |
centriolar coiled coil protein 110kDa |
| chr10_-_7708918 | 0.34 |
ENST00000256861.6 ENST00000397146.2 ENST00000446830.2 ENST00000397145.2 |
ITIH5 |
inter-alpha-trypsin inhibitor heavy chain family, member 5 |
| chr9_-_130616915 | 0.34 |
ENST00000344849.3 |
ENG |
endoglin |
| chr5_+_112043186 | 0.33 |
ENST00000509732.1 ENST00000457016.1 ENST00000507379.1 |
APC |
adenomatous polyposis coli |
| chr8_+_98881268 | 0.32 |
ENST00000254898.5 ENST00000524308.1 ENST00000522025.2 |
MATN2 |
matrilin 2 |
| chr19_-_7939319 | 0.32 |
ENST00000539422.1 |
CTD-3193O13.9 |
Protein FLJ22184 |
| chr12_-_42631529 | 0.32 |
ENST00000548917.1 |
YAF2 |
YY1 associated factor 2 |
| chr20_+_49348081 | 0.32 |
ENST00000371610.2 |
PARD6B |
par-6 family cell polarity regulator beta |
| chr4_-_140005341 | 0.32 |
ENST00000379549.2 ENST00000512627.1 |
ELF2 |
E74-like factor 2 (ets domain transcription factor) |
| chr11_-_78285804 | 0.31 |
ENST00000281038.5 ENST00000529571.1 |
NARS2 |
asparaginyl-tRNA synthetase 2, mitochondrial (putative) |
| chr1_+_39456895 | 0.31 |
ENST00000432648.3 ENST00000446189.2 ENST00000372984.4 |
AKIRIN1 |
akirin 1 |
| chr17_-_42298331 | 0.31 |
ENST00000343638.5 |
UBTF |
upstream binding transcription factor, RNA polymerase I |
| chr18_-_56940611 | 0.31 |
ENST00000256852.7 ENST00000334889.3 |
RAX |
retina and anterior neural fold homeobox |
| chr17_+_77751931 | 0.31 |
ENST00000310942.4 ENST00000269399.5 |
CBX2 |
chromobox homolog 2 |
| chr19_-_55686399 | 0.31 |
ENST00000587067.1 |
SYT5 |
synaptotagmin V |
| chr19_-_19006920 | 0.30 |
ENST00000429504.2 ENST00000427170.2 |
CERS1 |
ceramide synthase 1 |
| chr19_+_41305085 | 0.30 |
ENST00000303961.4 |
EGLN2 |
egl-9 family hypoxia-inducible factor 2 |
| chr19_-_46285646 | 0.30 |
ENST00000458663.2 |
DMPK |
dystrophia myotonica-protein kinase |
| chr7_+_155250824 | 0.30 |
ENST00000297375.4 |
EN2 |
engrailed homeobox 2 |
| chr11_-_19223523 | 0.29 |
ENST00000265968.3 |
CSRP3 |
cysteine and glycine-rich protein 3 (cardiac LIM protein) |
| chr17_-_7165662 | 0.29 |
ENST00000571881.2 ENST00000360325.7 |
CLDN7 |
claudin 7 |
| chr2_-_27545921 | 0.29 |
ENST00000402310.1 ENST00000405983.1 ENST00000403262.2 ENST00000428910.1 ENST00000402722.1 ENST00000399052.4 ENST00000380044.1 ENST00000405076.1 |
MPV17 |
MpV17 mitochondrial inner membrane protein |
| chr15_+_33010175 | 0.28 |
ENST00000300177.4 ENST00000560677.1 ENST00000560830.1 |
GREM1 |
gremlin 1, DAN family BMP antagonist |
| chr8_-_17270809 | 0.27 |
ENST00000180173.5 ENST00000521857.1 |
MTMR7 |
myotubularin related protein 7 |
| chr19_-_10341948 | 0.27 |
ENST00000590320.1 ENST00000592342.1 ENST00000588952.1 |
S1PR2 DNMT1 |
sphingosine-1-phosphate receptor 2 DNA (cytosine-5-)-methyltransferase 1 |
| chr17_+_38465441 | 0.27 |
ENST00000577646.1 ENST00000254066.5 |
RARA |
retinoic acid receptor, alpha |
| chr11_+_129245796 | 0.27 |
ENST00000281437.4 |
BARX2 |
BARX homeobox 2 |
| chr12_+_125478241 | 0.27 |
ENST00000341446.8 |
BRI3BP |
BRI3 binding protein |
| chr6_-_10882174 | 0.26 |
ENST00000379491.4 |
GCM2 |
glial cells missing homolog 2 (Drosophila) |
| chr12_+_57482665 | 0.26 |
ENST00000300131.3 |
NAB2 |
NGFI-A binding protein 2 (EGR1 binding protein 2) |
| chr19_+_35783047 | 0.26 |
ENST00000595791.1 ENST00000597035.1 ENST00000537831.2 |
MAG |
myelin associated glycoprotein |
| chr3_-_67705006 | 0.25 |
ENST00000492795.1 ENST00000493112.1 ENST00000307227.5 |
SUCLG2 |
succinate-CoA ligase, GDP-forming, beta subunit |
| chr1_+_226411319 | 0.24 |
ENST00000542034.1 ENST00000366810.5 |
MIXL1 |
Mix paired-like homeobox |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 3.1 | GO:0031133 | regulation of axon diameter(GO:0031133) |
| 1.0 | 2.9 | GO:1902303 | regulation of heart rate by hormone(GO:0003064) negative regulation of potassium ion export(GO:1902303) |
| 0.9 | 3.6 | GO:0036228 | protein targeting to nuclear inner membrane(GO:0036228) |
| 0.7 | 2.0 | GO:0002522 | leukocyte migration involved in immune response(GO:0002522) |
| 0.7 | 4.6 | GO:0045079 | negative regulation of chemokine biosynthetic process(GO:0045079) |
| 0.6 | 2.5 | GO:0000255 | allantoin metabolic process(GO:0000255) |
| 0.6 | 1.8 | GO:2000048 | negative regulation of cell-cell adhesion mediated by cadherin(GO:2000048) |
| 0.5 | 1.6 | GO:1902595 | regulation of DNA replication origin binding(GO:1902595) |
| 0.5 | 1.6 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.5 | 1.4 | GO:0035948 | positive regulation of gluconeogenesis by positive regulation of transcription from RNA polymerase II promoter(GO:0035948) |
| 0.4 | 5.3 | GO:0021894 | cerebral cortex GABAergic interneuron development(GO:0021894) |
| 0.4 | 2.0 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) |
| 0.4 | 1.1 | GO:0015709 | thiosulfate transport(GO:0015709) oxaloacetate transport(GO:0015729) malate transport(GO:0015743) malate transmembrane transport(GO:0071423) oxaloacetate(2-) transmembrane transport(GO:1902356) |
| 0.3 | 2.4 | GO:0098881 | exocytic insertion of neurotransmitter receptor to plasma membrane(GO:0098881) exocytic insertion of neurotransmitter receptor to postsynaptic membrane(GO:0098967) |
| 0.3 | 0.8 | GO:0051066 | dihydrobiopterin metabolic process(GO:0051066) |
| 0.3 | 0.8 | GO:0100009 | regulation of fever generation by regulation of prostaglandin secretion(GO:0071810) positive regulation of fever generation by positive regulation of prostaglandin secretion(GO:0071812) positive regulation of ERK1 and ERK2 cascade via TNFSF11-mediated signaling(GO:0071848) regulation of fever generation by prostaglandin secretion(GO:0100009) |
| 0.3 | 1.3 | GO:0033031 | positive regulation of neutrophil apoptotic process(GO:0033031) |
| 0.3 | 1.3 | GO:2000035 | regulation of stem cell division(GO:2000035) |
| 0.2 | 2.1 | GO:0035799 | ureter maturation(GO:0035799) |
| 0.2 | 2.0 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.2 | 0.6 | GO:0035470 | positive regulation of vascular wound healing(GO:0035470) regulation of lactation(GO:1903487) |
| 0.2 | 0.6 | GO:0021678 | fourth ventricle development(GO:0021592) third ventricle development(GO:0021678) |
| 0.2 | 0.6 | GO:0001300 | chronological cell aging(GO:0001300) |
| 0.2 | 1.5 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.2 | 0.7 | GO:0032053 | ciliary basal body organization(GO:0032053) |
| 0.2 | 0.9 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.2 | 0.9 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.1 | 2.5 | GO:0007253 | cytoplasmic sequestering of NF-kappaB(GO:0007253) |
| 0.1 | 0.7 | GO:0033600 | negative regulation of mammary gland epithelial cell proliferation(GO:0033600) |
| 0.1 | 0.7 | GO:1903862 | positive regulation of oxidative phosphorylation(GO:1903862) |
| 0.1 | 0.4 | GO:0045897 | positive regulation of transcription during mitosis(GO:0045897) |
| 0.1 | 1.1 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.1 | 0.4 | GO:0033386 | geranylgeranyl diphosphate metabolic process(GO:0033385) geranylgeranyl diphosphate biosynthetic process(GO:0033386) |
| 0.1 | 0.4 | GO:1990523 | bone regeneration(GO:1990523) |
| 0.1 | 0.1 | GO:0048073 | regulation of eye pigmentation(GO:0048073) |
| 0.1 | 0.5 | GO:2000676 | positive regulation of type B pancreatic cell apoptotic process(GO:2000676) |
| 0.1 | 2.5 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 0.8 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.1 | 1.5 | GO:0070986 | left/right axis specification(GO:0070986) |
| 0.1 | 0.4 | GO:0034473 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 0.1 | 1.1 | GO:0036066 | protein O-linked fucosylation(GO:0036066) |
| 0.1 | 0.3 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 0.1 | 1.0 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.1 | 0.4 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.1 | 0.8 | GO:0045084 | positive regulation of interleukin-12 biosynthetic process(GO:0045084) |
| 0.1 | 0.8 | GO:0021943 | formation of radial glial scaffolds(GO:0021943) |
| 0.1 | 0.5 | GO:0060718 | chorionic trophoblast cell differentiation(GO:0060718) |
| 0.1 | 1.1 | GO:0061158 | 3'-UTR-mediated mRNA destabilization(GO:0061158) |
| 0.1 | 0.3 | GO:1900158 | negative regulation of osteoclast proliferation(GO:0090291) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.1 | 0.7 | GO:1904996 | PML body organization(GO:0030578) positive regulation of leukocyte adhesion to vascular endothelial cell(GO:1904996) |
| 0.1 | 0.4 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.1 | 0.7 | GO:0016139 | glycoside catabolic process(GO:0016139) |
| 0.1 | 0.8 | GO:0060385 | axonogenesis involved in innervation(GO:0060385) |
| 0.1 | 0.2 | GO:0071409 | cellular response to cycloheximide(GO:0071409) |
| 0.1 | 0.2 | GO:0051885 | positive regulation of anagen(GO:0051885) |
| 0.1 | 0.6 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.1 | 0.5 | GO:0014722 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) |
| 0.1 | 0.3 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.1 | 0.7 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.1 | 0.3 | GO:0035995 | detection of muscle stretch(GO:0035995) |
| 0.1 | 1.1 | GO:0043517 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) |
| 0.1 | 0.4 | GO:0031914 | negative regulation of synaptic plasticity(GO:0031914) |
| 0.1 | 1.0 | GO:1904152 | regulation of retrograde protein transport, ER to cytosol(GO:1904152) |
| 0.1 | 0.3 | GO:0010216 | maintenance of DNA methylation(GO:0010216) |
| 0.1 | 1.1 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.1 | 0.3 | GO:0060010 | Sertoli cell fate commitment(GO:0060010) |
| 0.1 | 3.5 | GO:0048747 | muscle fiber development(GO:0048747) |
| 0.1 | 0.6 | GO:0038180 | nerve growth factor signaling pathway(GO:0038180) |
| 0.1 | 0.6 | GO:0000012 | single strand break repair(GO:0000012) |
| 0.1 | 0.2 | GO:0046010 | positive regulation of circadian sleep/wake cycle, non-REM sleep(GO:0046010) |
| 0.1 | 0.5 | GO:1902525 | regulation of protein monoubiquitination(GO:1902525) |
| 0.1 | 0.9 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
| 0.1 | 0.7 | GO:1902659 | regulation of glucose mediated signaling pathway(GO:1902659) |
| 0.1 | 0.2 | GO:1904383 | response to sodium phosphate(GO:1904383) |
| 0.1 | 1.0 | GO:0090005 | negative regulation of establishment of protein localization to plasma membrane(GO:0090005) |
| 0.1 | 0.2 | GO:0019482 | beta-alanine metabolic process(GO:0019482) |
| 0.0 | 0.2 | GO:0006104 | succinyl-CoA metabolic process(GO:0006104) |
| 0.0 | 0.2 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.0 | 0.3 | GO:1904885 | beta-catenin destruction complex assembly(GO:1904885) |
| 0.0 | 0.7 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.0 | 0.6 | GO:0001711 | endodermal cell fate commitment(GO:0001711) |
| 0.0 | 0.8 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.0 | 7.3 | GO:0007605 | sensory perception of sound(GO:0007605) |
| 0.0 | 0.3 | GO:0030643 | cellular phosphate ion homeostasis(GO:0030643) parathyroid gland development(GO:0060017) cellular trivalent inorganic anion homeostasis(GO:0072502) |
| 0.0 | 0.0 | GO:0055073 | cadmium ion homeostasis(GO:0055073) |
| 0.0 | 1.0 | GO:0009954 | proximal/distal pattern formation(GO:0009954) |
| 0.0 | 0.2 | GO:2000382 | positive regulation of mesoderm development(GO:2000382) |
| 0.0 | 0.6 | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process(GO:0006488) |
| 0.0 | 0.3 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.0 | 0.2 | GO:0048842 | positive regulation of axon extension involved in axon guidance(GO:0048842) |
| 0.0 | 0.3 | GO:1902949 | positive regulation of tau-protein kinase activity(GO:1902949) |
| 0.0 | 0.2 | GO:0009137 | purine nucleoside diphosphate catabolic process(GO:0009137) purine ribonucleoside diphosphate catabolic process(GO:0009181) |
| 0.0 | 0.4 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.0 | 0.9 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.0 | 0.3 | GO:1990403 | embryonic brain development(GO:1990403) |
| 0.0 | 1.4 | GO:0030318 | melanocyte differentiation(GO:0030318) |
| 0.0 | 0.4 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.0 | 0.7 | GO:0050855 | regulation of B cell receptor signaling pathway(GO:0050855) |
| 0.0 | 0.2 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 0.0 | 0.2 | GO:0060536 | growth plate cartilage chondrocyte differentiation(GO:0003418) cartilage morphogenesis(GO:0060536) |
| 0.0 | 1.1 | GO:0043304 | regulation of mast cell activation involved in immune response(GO:0033006) regulation of mast cell degranulation(GO:0043304) |
| 0.0 | 4.4 | GO:0002223 | stimulatory C-type lectin receptor signaling pathway(GO:0002223) |
| 0.0 | 0.5 | GO:2001275 | positive regulation of glucose import in response to insulin stimulus(GO:2001275) |
| 0.0 | 1.1 | GO:0006409 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.0 | 0.7 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 0.9 | GO:0032784 | regulation of DNA-templated transcription, elongation(GO:0032784) |
| 0.0 | 0.3 | GO:0048711 | positive regulation of astrocyte differentiation(GO:0048711) |
| 0.0 | 0.5 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.0 | 0.3 | GO:1904380 | endoplasmic reticulum mannose trimming(GO:1904380) |
| 0.0 | 0.0 | GO:0060830 | ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.0 | 0.1 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.0 | 2.9 | GO:0051291 | protein heterooligomerization(GO:0051291) |
| 0.0 | 0.7 | GO:0007265 | Ras protein signal transduction(GO:0007265) |
| 0.0 | 0.1 | GO:0019470 | 4-hydroxyproline catabolic process(GO:0019470) |
| 0.0 | 0.1 | GO:0045872 | positive regulation of rhodopsin gene expression(GO:0045872) |
| 0.0 | 0.2 | GO:0030220 | platelet formation(GO:0030220) |
| 0.0 | 0.1 | GO:0036289 | peptidyl-serine autophosphorylation(GO:0036289) |
| 0.0 | 2.0 | GO:0030326 | embryonic limb morphogenesis(GO:0030326) embryonic appendage morphogenesis(GO:0035113) |
| 0.0 | 0.2 | GO:0034134 | toll-like receptor 2 signaling pathway(GO:0034134) |
| 0.0 | 0.5 | GO:1903861 | regulation of dendrite extension(GO:1903859) positive regulation of dendrite extension(GO:1903861) |
| 0.0 | 0.1 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.0 | 1.2 | GO:0032729 | positive regulation of interferon-gamma production(GO:0032729) |
| 0.0 | 0.3 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.3 | GO:0070830 | bicellular tight junction assembly(GO:0070830) |
| 0.0 | 0.3 | GO:0035909 | aorta morphogenesis(GO:0035909) |
| 0.0 | 0.2 | GO:0071481 | cellular response to X-ray(GO:0071481) |
| 0.0 | 0.3 | GO:0001502 | cartilage condensation(GO:0001502) |
| 0.0 | 0.9 | GO:0070125 | mitochondrial translational elongation(GO:0070125) |
| 0.0 | 0.2 | GO:0000963 | mitochondrial RNA processing(GO:0000963) |
| 0.0 | 0.5 | GO:0043666 | regulation of phosphoprotein phosphatase activity(GO:0043666) |
| 0.0 | 0.4 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.0 | 0.7 | GO:0043277 | apoptotic cell clearance(GO:0043277) |
| 0.0 | 0.2 | GO:0031929 | TOR signaling(GO:0031929) regulation of TOR signaling(GO:0032006) |
| 0.0 | 1.8 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.3 | GO:0043303 | mast cell activation involved in immune response(GO:0002279) mast cell degranulation(GO:0043303) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.5 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.2 | 3.4 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.1 | 0.7 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.1 | 3.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 2.3 | REACTOME ASSOCIATION OF LICENSING FACTORS WITH THE PRE REPLICATIVE COMPLEX | Genes involved in Association of licensing factors with the pre-replicative complex |
| 0.1 | 4.7 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.1 | 3.3 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.1 | 1.4 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.1 | 2.4 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 2.0 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 1.0 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 2.8 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 0.9 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.1 | 1.6 | REACTOME PD1 SIGNALING | Genes involved in PD-1 signaling |
| 0.0 | 0.4 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.0 | 1.5 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.0 | 3.8 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 1.3 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.0 | 7.4 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.8 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.0 | 0.6 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.0 | 0.2 | REACTOME DESTABILIZATION OF MRNA BY BRF1 | Genes involved in Destabilization of mRNA by Butyrate Response Factor 1 (BRF1) |
| 0.0 | 4.8 | REACTOME MYD88 MAL CASCADE INITIATED ON PLASMA MEMBRANE | Genes involved in MyD88:Mal cascade initiated on plasma membrane |
| 0.0 | 1.0 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 0.7 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.0 | 0.8 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.9 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 1.1 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 0.6 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.9 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 0.7 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.0 | 0.7 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.6 | REACTOME DOUBLE STRAND BREAK REPAIR | Genes involved in Double-Strand Break Repair |
| 0.0 | 0.7 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.0 | 0.8 | REACTOME INTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Intrinsic Pathway for Apoptosis |
| 0.0 | 1.1 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 0.4 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 0.6 | REACTOME PREFOLDIN MEDIATED TRANSFER OF SUBSTRATE TO CCT TRIC | Genes involved in Prefoldin mediated transfer of substrate to CCT/TriC |
| 0.0 | 1.0 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 0.3 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.4 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.2 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.0 | 0.2 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.3 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.5 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.1 | 3.8 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.1 | 4.0 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 3.7 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 0.3 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.1 | 2.6 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.1 | 1.8 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 2.9 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 0.8 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.9 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 1.8 | PID HIF2PATHWAY | HIF-2-alpha transcription factor network |
| 0.0 | 0.7 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 7.7 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 2.0 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 2.4 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 2.5 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.0 | 1.4 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 0.6 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.0 | 0.3 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.0 | 0.3 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.0 | 0.9 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 1.1 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 1.6 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.8 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.5 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 1.0 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 0.3 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 0.5 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.2 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.0 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.5 | 2.9 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.4 | 1.1 | GO:0015131 | thiosulfate transmembrane transporter activity(GO:0015117) oxaloacetate transmembrane transporter activity(GO:0015131) |
| 0.3 | 1.0 | GO:0032427 | GBD domain binding(GO:0032427) |
| 0.3 | 1.5 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.3 | 1.4 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.3 | 2.9 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.2 | 0.9 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.2 | 2.4 | GO:0050542 | icosanoid binding(GO:0050542) arachidonic acid binding(GO:0050544) fatty acid derivative binding(GO:1901567) |
| 0.2 | 2.0 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.2 | 1.3 | GO:0035005 | 1-phosphatidylinositol-4-phosphate 3-kinase activity(GO:0035005) |
| 0.2 | 2.1 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.1 | 1.5 | GO:0019826 | oxygen sensor activity(GO:0019826) |
| 0.1 | 0.6 | GO:0070259 | tyrosyl-DNA phosphodiesterase activity(GO:0070259) |
| 0.1 | 0.8 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.1 | 0.4 | GO:0034739 | histone deacetylase activity (H4-K16 specific)(GO:0034739) |
| 0.1 | 0.2 | GO:0051880 | G-quadruplex DNA binding(GO:0051880) |
| 0.1 | 3.5 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 0.6 | GO:0010465 | nerve growth factor receptor activity(GO:0010465) |
| 0.1 | 3.2 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.1 | 12.8 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.1 | 0.4 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 0.1 | 0.7 | GO:0001165 | RNA polymerase I upstream control element sequence-specific DNA binding(GO:0001165) |
| 0.1 | 1.0 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.1 | 0.6 | GO:0005534 | galactose binding(GO:0005534) |
| 0.1 | 2.5 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.1 | 0.5 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.1 | 0.7 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.1 | 0.2 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) |
| 0.1 | 5.4 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.1 | 0.6 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.1 | 0.2 | GO:0004157 | dihydropyrimidinase activity(GO:0004157) |
| 0.1 | 0.9 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 0.3 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.0 | 1.0 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.0 | 1.0 | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups(GO:0016780) |
| 0.0 | 0.2 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.0 | 0.2 | GO:0016520 | growth hormone-releasing hormone receptor activity(GO:0016520) |
| 0.0 | 0.3 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.0 | 1.8 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.0 | 3.9 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.3 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.0 | 0.4 | GO:0004311 | farnesyltranstransferase activity(GO:0004311) |
| 0.0 | 0.2 | GO:0042608 | T cell receptor binding(GO:0042608) |
| 0.0 | 1.6 | GO:0008200 | ion channel inhibitor activity(GO:0008200) |
| 0.0 | 0.3 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.0 | 1.8 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.7 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.0 | 0.1 | GO:0016499 | orexin receptor activity(GO:0016499) |
| 0.0 | 0.2 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.0 | 1.2 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.0 | 0.3 | GO:0031433 | telethonin binding(GO:0031433) |
| 0.0 | 0.9 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 1.3 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.6 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.4 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.0 | 0.3 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.6 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 1.4 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.0 | 0.2 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.0 | 1.3 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.7 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 0.5 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.0 | 1.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.3 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 1.8 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.3 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.0 | 0.1 | GO:0005113 | patched binding(GO:0005113) |
| 0.0 | 0.1 | GO:0004703 | G-protein coupled receptor kinase activity(GO:0004703) |
| 0.0 | 0.6 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.0 | 0.1 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 0.0 | 0.7 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.4 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.0 | 0.4 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.3 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 0.9 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 2.2 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 0.1 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.0 | 0.2 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.0 | 2.1 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 1.2 | GO:0101005 | thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 0.2 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.0 | 0.1 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.0 | 0.4 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.0 | 2.1 | GO:0004713 | protein tyrosine kinase activity(GO:0004713) |
| 0.0 | 1.1 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 0.2 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 0.0 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.0 | 0.3 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.8 | GO:0005070 | SH3/SH2 adaptor activity(GO:0005070) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.6 | GO:0044611 | nuclear pore inner ring(GO:0044611) |
| 0.6 | 2.4 | GO:1990796 | photoreceptor cell terminal bouton(GO:1990796) |
| 0.6 | 2.9 | GO:1902937 | inward rectifier potassium channel complex(GO:1902937) |
| 0.3 | 0.5 | GO:0070821 | tertiary granule membrane(GO:0070821) |
| 0.2 | 3.1 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.2 | 0.6 | GO:0072563 | endothelial microparticle(GO:0072563) |
| 0.2 | 2.5 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.2 | 3.8 | GO:0031254 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.2 | 2.5 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.2 | 0.6 | GO:0043541 | UDP-N-acetylglucosamine transferase complex(GO:0043541) |
| 0.2 | 1.0 | GO:0000835 | ER ubiquitin ligase complex(GO:0000835) |
| 0.2 | 1.4 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.1 | 0.6 | GO:0097513 | myosin II filament(GO:0097513) |
| 0.1 | 0.4 | GO:0031523 | Myb complex(GO:0031523) |
| 0.1 | 5.3 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 0.3 | GO:1990015 | mesaxon(GO:0097453) ensheathing process(GO:1990015) |
| 0.1 | 0.9 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.1 | 0.3 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.1 | 2.0 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.1 | 0.6 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.1 | 0.6 | GO:0042719 | mitochondrial intermembrane space protein transporter complex(GO:0042719) |
| 0.1 | 3.6 | GO:0043034 | costamere(GO:0043034) |
| 0.1 | 0.3 | GO:1990769 | proximal neuron projection(GO:1990769) |
| 0.1 | 0.6 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.1 | 0.2 | GO:0097441 | basilar dendrite(GO:0097441) |
| 0.1 | 1.2 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.1 | 1.1 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.1 | 1.1 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 0.6 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.0 | 4.3 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 0.2 | GO:0030062 | mitochondrial tricarboxylic acid cycle enzyme complex(GO:0030062) |
| 0.0 | 0.9 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.2 | GO:0071817 | MMXD complex(GO:0071817) |
| 0.0 | 0.4 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 0.5 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.4 | GO:0072357 | PTW/PP1 phosphatase complex(GO:0072357) |
| 0.0 | 0.4 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.0 | 0.4 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.0 | 0.3 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 0.2 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 0.2 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 0.0 | 2.2 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 1.4 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 0.2 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.0 | 0.4 | GO:0000783 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) |
| 0.0 | 0.3 | GO:0005721 | pericentric heterochromatin(GO:0005721) |
| 0.0 | 1.5 | GO:0031519 | PcG protein complex(GO:0031519) |
| 0.0 | 0.7 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.7 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 0.7 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.9 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.0 | 0.6 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.0 | 1.6 | GO:0000777 | condensed chromosome kinetochore(GO:0000777) |
| 0.0 | 5.9 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 0.8 | GO:0030133 | transport vesicle(GO:0030133) |
| 0.0 | 0.7 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 1.1 | GO:0043657 | host(GO:0018995) host cell(GO:0043657) |
| 0.0 | 0.8 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 1.0 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 0.6 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.3 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |