Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
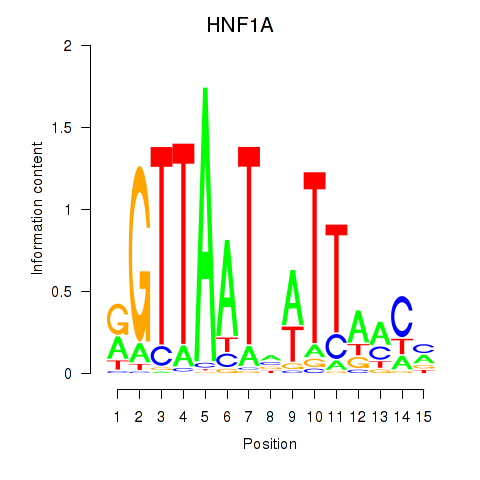
Results for HNF1A_HNF1B
Z-value: 4.44
Transcription factors associated with HNF1A_HNF1B
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HNF1A
|
ENSG00000135100.13 | HNF1A |
|
HNF1B
|
ENSG00000108753.8 | HNF1B |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HNF1A | hg19_v2_chr12_+_121416489_121416552 | 0.97 | 1.6e-10 | Click! |
Activity profile of HNF1A_HNF1B motif
Sorted Z-values of HNF1A_HNF1B motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HNF1A_HNF1B
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_74301880 | 66.80 |
ENST00000395792.2 ENST00000226359.2 |
AFP |
alpha-fetoprotein |
| chr17_-_64225508 | 41.88 |
ENST00000205948.6 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chrX_-_105282712 | 27.11 |
ENST00000372563.1 |
SERPINA7 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr9_-_123812542 | 25.50 |
ENST00000223642.1 |
C5 |
complement component 5 |
| chr4_-_110723335 | 22.70 |
ENST00000394634.2 |
CFI |
complement factor I |
| chr4_-_155511887 | 22.56 |
ENST00000302053.3 ENST00000403106.3 |
FGA |
fibrinogen alpha chain |
| chr4_-_110723194 | 22.18 |
ENST00000394635.3 |
CFI |
complement factor I |
| chr4_-_110723134 | 22.14 |
ENST00000510800.1 ENST00000512148.1 |
CFI |
complement factor I |
| chr14_-_94854926 | 18.80 |
ENST00000402629.1 ENST00000556091.1 ENST00000554720.1 |
SERPINA1 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 1 |
| chr10_+_5135981 | 17.37 |
ENST00000380554.3 |
AKR1C3 |
aldo-keto reductase family 1, member C3 |
| chr4_+_155484155 | 16.66 |
ENST00000509493.1 |
FGB |
fibrinogen beta chain |
| chr4_+_74269956 | 16.55 |
ENST00000295897.4 ENST00000415165.2 ENST00000503124.1 ENST00000509063.1 ENST00000401494.3 |
ALB |
albumin |
| chr19_-_36304201 | 16.52 |
ENST00000301175.3 |
PRODH2 |
proline dehydrogenase (oxidase) 2 |
| chr7_+_45927956 | 16.16 |
ENST00000275525.3 ENST00000457280.1 |
IGFBP1 |
insulin-like growth factor binding protein 1 |
| chr19_+_11350278 | 15.64 |
ENST00000252453.8 |
C19orf80 |
chromosome 19 open reading frame 80 |
| chr7_+_45928079 | 15.50 |
ENST00000468955.1 |
IGFBP1 |
insulin-like growth factor binding protein 1 |
| chr1_+_207277590 | 15.30 |
ENST00000367070.3 |
C4BPA |
complement component 4 binding protein, alpha |
| chr15_+_58724184 | 15.22 |
ENST00000433326.2 |
LIPC |
lipase, hepatic |
| chr10_+_101542462 | 14.70 |
ENST00000370449.4 ENST00000370434.1 |
ABCC2 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 2 |
| chr4_-_69817481 | 13.71 |
ENST00000251566.4 |
UGT2A3 |
UDP glucuronosyltransferase 2 family, polypeptide A3 |
| chr14_-_94759595 | 13.66 |
ENST00000261994.4 |
SERPINA10 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 |
| chr5_+_176513868 | 13.45 |
ENST00000292408.4 |
FGFR4 |
fibroblast growth factor receptor 4 |
| chr14_-_94759361 | 13.28 |
ENST00000393096.1 |
SERPINA10 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 |
| chr14_-_94759408 | 13.06 |
ENST00000554723.1 |
SERPINA10 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 10 |
| chr2_-_88427568 | 12.75 |
ENST00000393750.3 ENST00000295834.3 |
FABP1 |
fatty acid binding protein 1, liver |
| chr8_-_17752996 | 12.71 |
ENST00000381841.2 ENST00000427924.1 |
FGL1 |
fibrinogen-like 1 |
| chr4_+_155484103 | 12.59 |
ENST00000302068.4 |
FGB |
fibrinogen beta chain |
| chr5_+_176513895 | 12.06 |
ENST00000503708.1 ENST00000393648.2 ENST00000514472.1 ENST00000502906.1 ENST00000292410.3 ENST00000510911.1 |
FGFR4 |
fibroblast growth factor receptor 4 |
| chr4_+_100495864 | 11.63 |
ENST00000265517.5 ENST00000422897.2 |
MTTP |
microsomal triglyceride transfer protein |
| chr14_+_21156915 | 11.38 |
ENST00000397990.4 ENST00000555597.1 |
ANG RNASE4 |
angiogenin, ribonuclease, RNase A family, 5 ribonuclease, RNase A family, 4 |
| chr1_+_207262170 | 9.86 |
ENST00000367078.3 |
C4BPB |
complement component 4 binding protein, beta |
| chr6_+_7541845 | 8.95 |
ENST00000418664.2 |
DSP |
desmoplakin |
| chr7_-_50633078 | 8.80 |
ENST00000444124.2 |
DDC |
dopa decarboxylase (aromatic L-amino acid decarboxylase) |
| chr12_+_100867486 | 8.67 |
ENST00000548884.1 |
NR1H4 |
nuclear receptor subfamily 1, group H, member 4 |
| chr6_-_25930819 | 8.61 |
ENST00000360488.3 |
SLC17A2 |
solute carrier family 17, member 2 |
| chr19_+_50016610 | 8.40 |
ENST00000596975.1 |
FCGRT |
Fc fragment of IgG, receptor, transporter, alpha |
| chr6_-_25930904 | 8.02 |
ENST00000377850.3 |
SLC17A2 |
solute carrier family 17, member 2 |
| chr11_+_46740730 | 7.87 |
ENST00000311907.5 ENST00000530231.1 ENST00000442468.1 |
F2 |
coagulation factor II (thrombin) |
| chr17_+_1646130 | 7.65 |
ENST00000453066.1 ENST00000324015.3 ENST00000450523.2 ENST00000453723.1 ENST00000382061.4 |
SERPINF2 |
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 2 |
| chr1_+_207262881 | 7.58 |
ENST00000451804.2 |
C4BPB |
complement component 4 binding protein, beta |
| chr17_+_41052808 | 6.86 |
ENST00000592383.1 ENST00000253801.2 ENST00000585489.1 |
G6PC |
glucose-6-phosphatase, catalytic subunit |
| chr14_-_65409438 | 6.70 |
ENST00000557049.1 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr1_+_207262627 | 6.64 |
ENST00000391923.1 |
C4BPB |
complement component 4 binding protein, beta |
| chr22_-_30970560 | 6.63 |
ENST00000402369.1 ENST00000406361.1 |
GAL3ST1 |
galactose-3-O-sulfotransferase 1 |
| chr6_+_167704798 | 6.58 |
ENST00000230256.3 |
UNC93A |
unc-93 homolog A (C. elegans) |
| chr19_+_50016411 | 6.51 |
ENST00000426395.3 ENST00000600273.1 ENST00000599988.1 |
FCGRT |
Fc fragment of IgG, receptor, transporter, alpha |
| chr6_+_167704838 | 6.50 |
ENST00000366829.2 |
UNC93A |
unc-93 homolog A (C. elegans) |
| chr1_+_207262578 | 6.39 |
ENST00000243611.5 ENST00000367076.3 |
C4BPB |
complement component 4 binding protein, beta |
| chr17_+_79953310 | 6.34 |
ENST00000582355.2 |
ASPSCR1 |
alveolar soft part sarcoma chromosome region, candidate 1 |
| chr10_-_52645379 | 6.27 |
ENST00000395489.2 |
A1CF |
APOBEC1 complementation factor |
| chr12_+_100867694 | 6.26 |
ENST00000392986.3 ENST00000549996.1 |
NR1H4 |
nuclear receptor subfamily 1, group H, member 4 |
| chr10_-_52645416 | 6.12 |
ENST00000374001.2 ENST00000373997.3 ENST00000373995.3 ENST00000282641.2 ENST00000395495.1 ENST00000414883.1 |
A1CF |
APOBEC1 complementation factor |
| chr14_-_65409502 | 5.96 |
ENST00000389614.5 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr14_+_95027772 | 5.86 |
ENST00000555095.1 ENST00000298841.5 ENST00000554220.1 ENST00000553780.1 |
SERPINA4 SERPINA5 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 4 serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 5 |
| chr4_+_187187098 | 5.79 |
ENST00000403665.2 ENST00000264692.4 |
F11 |
coagulation factor XI |
| chr2_+_27719697 | 5.42 |
ENST00000264717.2 ENST00000424318.2 |
GCKR |
glucokinase (hexokinase 4) regulator |
| chr8_+_21911054 | 5.42 |
ENST00000519850.1 ENST00000381470.3 |
DMTN |
dematin actin binding protein |
| chr7_-_44580861 | 5.30 |
ENST00000546276.1 ENST00000289547.4 ENST00000381160.3 ENST00000423141.1 |
NPC1L1 |
NPC1-like 1 |
| chr16_-_87970122 | 5.22 |
ENST00000309893.2 |
CA5A |
carbonic anhydrase VA, mitochondrial |
| chr20_+_35973080 | 5.14 |
ENST00000445403.1 |
SRC |
v-src avian sarcoma (Schmidt-Ruppin A-2) viral oncogene homolog |
| chr5_+_175976324 | 5.05 |
ENST00000261944.5 |
CDHR2 |
cadherin-related family member 2 |
| chr14_+_39703084 | 4.96 |
ENST00000553728.1 |
RP11-407N17.3 |
cTAGE family member 5 isoform 4 |
| chr17_-_202579 | 4.86 |
ENST00000577079.1 ENST00000331302.7 ENST00000536489.2 |
RPH3AL |
rabphilin 3A-like (without C2 domains) |
| chr1_+_17906970 | 4.35 |
ENST00000375415.1 |
ARHGEF10L |
Rho guanine nucleotide exchange factor (GEF) 10-like |
| chr12_+_100897130 | 4.17 |
ENST00000551379.1 ENST00000188403.7 ENST00000551184.1 |
NR1H4 |
nuclear receptor subfamily 1, group H, member 4 |
| chr12_+_70132632 | 3.67 |
ENST00000378815.6 ENST00000483530.2 ENST00000325555.9 |
RAB3IP |
RAB3A interacting protein |
| chr16_+_20775358 | 3.58 |
ENST00000440284.2 |
ACSM3 |
acyl-CoA synthetase medium-chain family member 3 |
| chr11_-_117695449 | 3.53 |
ENST00000292079.2 |
FXYD2 |
FXYD domain containing ion transport regulator 2 |
| chr2_-_55920952 | 3.31 |
ENST00000447944.2 |
PNPT1 |
polyribonucleotide nucleotidyltransferase 1 |
| chr7_-_47579188 | 3.30 |
ENST00000398879.1 ENST00000355730.3 ENST00000442536.2 ENST00000458317.2 |
TNS3 |
tensin 3 |
| chr2_-_162931052 | 3.29 |
ENST00000360534.3 |
DPP4 |
dipeptidyl-peptidase 4 |
| chr6_+_161123270 | 3.26 |
ENST00000366924.2 ENST00000308192.9 ENST00000418964.1 |
PLG |
plasminogen |
| chr16_+_20775024 | 3.22 |
ENST00000289416.5 |
ACSM3 |
acyl-CoA synthetase medium-chain family member 3 |
| chr21_-_47575481 | 3.15 |
ENST00000291670.5 ENST00000397748.1 ENST00000359679.2 ENST00000355384.2 ENST00000397746.3 ENST00000397743.1 |
FTCD |
formimidoyltransferase cyclodeaminase |
| chr14_+_24590560 | 3.10 |
ENST00000558325.1 |
RP11-468E2.6 |
RP11-468E2.6 |
| chr4_+_8201091 | 3.06 |
ENST00000382521.3 ENST00000245105.3 ENST00000457650.2 ENST00000539824.1 |
SH3TC1 |
SH3 domain and tetratricopeptide repeats 1 |
| chr22_-_30968813 | 3.01 |
ENST00000443111.2 ENST00000443136.1 ENST00000426220.1 |
GAL3ST1 |
galactose-3-O-sulfotransferase 1 |
| chr16_+_20462783 | 2.86 |
ENST00000574251.1 ENST00000576361.1 ENST00000417235.2 ENST00000573854.1 ENST00000424070.1 ENST00000536134.1 ENST00000219054.6 ENST00000575690.1 ENST00000571894.1 |
ACSM2A |
acyl-CoA synthetase medium-chain family member 2A |
| chr2_-_169104651 | 2.84 |
ENST00000355999.4 |
STK39 |
serine threonine kinase 39 |
| chr16_-_20587599 | 2.75 |
ENST00000566384.1 ENST00000565232.1 ENST00000567001.1 ENST00000565322.1 ENST00000569344.1 ENST00000329697.6 ENST00000414188.2 ENST00000568882.1 |
ACSM2B |
acyl-CoA synthetase medium-chain family member 2B |
| chr16_-_57831914 | 2.64 |
ENST00000421376.2 |
KIFC3 |
kinesin family member C3 |
| chr6_-_52628271 | 2.54 |
ENST00000493422.1 |
GSTA2 |
glutathione S-transferase alpha 2 |
| chr12_-_24102576 | 2.40 |
ENST00000537393.1 ENST00000309359.1 ENST00000381381.2 ENST00000451604.2 |
SOX5 |
SRY (sex determining region Y)-box 5 |
| chr4_-_1166954 | 2.36 |
ENST00000514490.1 ENST00000431380.1 ENST00000503765.1 |
SPON2 |
spondin 2, extracellular matrix protein |
| chr17_+_41363854 | 2.26 |
ENST00000588693.1 ENST00000588659.1 ENST00000541594.1 ENST00000536052.1 ENST00000331615.3 |
TMEM106A |
transmembrane protein 106A |
| chrX_+_107288197 | 2.21 |
ENST00000415430.3 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr10_-_72648541 | 2.21 |
ENST00000299299.3 |
PCBD1 |
pterin-4 alpha-carbinolamine dehydratase/dimerization cofactor of hepatocyte nuclear factor 1 alpha |
| chr1_-_110284384 | 2.16 |
ENST00000540225.1 |
GSTM3 |
glutathione S-transferase mu 3 (brain) |
| chr1_+_36789335 | 2.11 |
ENST00000373137.2 |
RP11-268J15.5 |
RP11-268J15.5 |
| chr4_+_3443614 | 2.09 |
ENST00000382774.3 ENST00000511533.1 |
HGFAC |
HGF activator |
| chr16_+_89696692 | 2.08 |
ENST00000261615.4 |
DPEP1 |
dipeptidase 1 (renal) |
| chr6_-_25874440 | 2.07 |
ENST00000361703.6 ENST00000397060.4 |
SLC17A3 |
solute carrier family 17 (organic anion transporter), member 3 |
| chr16_-_15474904 | 2.02 |
ENST00000534094.1 |
NPIPA5 |
nuclear pore complex interacting protein family, member A5 |
| chr12_+_70133152 | 1.90 |
ENST00000550536.1 ENST00000362025.5 |
RAB3IP |
RAB3A interacting protein |
| chrX_+_107288239 | 1.83 |
ENST00000217957.5 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr1_+_57320437 | 1.69 |
ENST00000361249.3 |
C8A |
complement component 8, alpha polypeptide |
| chr11_-_128894053 | 1.69 |
ENST00000392657.3 |
ARHGAP32 |
Rho GTPase activating protein 32 |
| chr17_-_36904437 | 1.63 |
ENST00000585100.1 ENST00000360797.2 ENST00000578109.1 ENST00000579882.1 |
PCGF2 |
polycomb group ring finger 2 |
| chr7_+_120629653 | 1.54 |
ENST00000450913.2 ENST00000340646.5 |
CPED1 |
cadherin-like and PC-esterase domain containing 1 |
| chr4_-_166034029 | 1.46 |
ENST00000306480.6 |
TMEM192 |
transmembrane protein 192 |
| chr1_+_229406847 | 1.33 |
ENST00000366690.4 |
RAB4A |
RAB4A, member RAS oncogene family |
| chr16_-_57831676 | 1.30 |
ENST00000465878.2 ENST00000539578.1 ENST00000561524.1 |
KIFC3 |
kinesin family member C3 |
| chrX_+_120181457 | 1.24 |
ENST00000328078.1 |
GLUD2 |
glutamate dehydrogenase 2 |
| chr5_+_67586465 | 1.22 |
ENST00000336483.5 |
PIK3R1 |
phosphoinositide-3-kinase, regulatory subunit 1 (alpha) |
| chr1_-_20141763 | 1.21 |
ENST00000375121.2 |
RNF186 |
ring finger protein 186 |
| chr9_-_98079965 | 1.07 |
ENST00000289081.3 |
FANCC |
Fanconi anemia, complementation group C |
| chr3_-_49466686 | 1.00 |
ENST00000273598.3 ENST00000436744.2 |
NICN1 |
nicolin 1 |
| chr20_+_62371206 | 1.00 |
ENST00000266077.2 |
SLC2A4RG |
SLC2A4 regulator |
| chr17_+_48823896 | 0.97 |
ENST00000511974.1 |
LUC7L3 |
LUC7-like 3 (S. cerevisiae) |
| chr6_+_144904334 | 0.96 |
ENST00000367526.4 |
UTRN |
utrophin |
| chr3_-_192445289 | 0.94 |
ENST00000430714.1 ENST00000418610.1 ENST00000448795.1 ENST00000445105.2 |
FGF12 |
fibroblast growth factor 12 |
| chr4_+_39408470 | 0.83 |
ENST00000257408.4 |
KLB |
klotho beta |
| chr19_+_54466179 | 0.80 |
ENST00000270458.2 |
CACNG8 |
calcium channel, voltage-dependent, gamma subunit 8 |
| chr16_-_66952779 | 0.80 |
ENST00000570262.1 ENST00000394055.3 ENST00000299752.4 |
CDH16 |
cadherin 16, KSP-cadherin |
| chr11_+_20620946 | 0.78 |
ENST00000525748.1 |
SLC6A5 |
solute carrier family 6 (neurotransmitter transporter), member 5 |
| chr11_+_1860200 | 0.72 |
ENST00000381911.1 |
TNNI2 |
troponin I type 2 (skeletal, fast) |
| chr20_+_45947246 | 0.66 |
ENST00000599904.1 |
AL031666.2 |
HCG2018772; Uncharacterized protein; cDNA FLJ31609 fis, clone NT2RI2002852 |
| chr7_-_6098770 | 0.66 |
ENST00000536084.1 ENST00000446699.1 ENST00000199389.6 |
EIF2AK1 |
eukaryotic translation initiation factor 2-alpha kinase 1 |
| chr2_-_47572105 | 0.63 |
ENST00000419035.1 ENST00000448713.1 ENST00000450550.1 ENST00000413185.2 |
AC073283.4 |
AC073283.4 |
| chr16_-_66952742 | 0.60 |
ENST00000565235.2 ENST00000568632.1 ENST00000565796.1 |
CDH16 |
cadherin 16, KSP-cadherin |
| chr10_-_38146510 | 0.60 |
ENST00000395867.3 |
ZNF248 |
zinc finger protein 248 |
| chr15_+_48498480 | 0.54 |
ENST00000380993.3 ENST00000396577.3 |
SLC12A1 |
solute carrier family 12 (sodium/potassium/chloride transporter), member 1 |
| chr5_-_111093167 | 0.54 |
ENST00000446294.2 ENST00000419114.2 |
NREP |
neuronal regeneration related protein |
| chr7_+_130126012 | 0.54 |
ENST00000341441.5 |
MEST |
mesoderm specific transcript |
| chr20_+_57414743 | 0.52 |
ENST00000313949.7 |
GNAS |
GNAS complex locus |
| chr9_-_215744 | 0.44 |
ENST00000382387.2 |
C9orf66 |
chromosome 9 open reading frame 66 |
| chr2_-_228244013 | 0.40 |
ENST00000304568.3 |
TM4SF20 |
transmembrane 4 L six family member 20 |
| chr19_-_11639931 | 0.38 |
ENST00000592312.1 ENST00000590480.1 ENST00000585318.1 ENST00000252440.7 ENST00000417981.2 ENST00000270517.7 |
ECSIT |
ECSIT signalling integrator |
| chr19_-_11639910 | 0.37 |
ENST00000588998.1 ENST00000586149.1 |
ECSIT |
ECSIT signalling integrator |
| chr7_+_130126165 | 0.29 |
ENST00000427521.1 ENST00000416162.2 ENST00000378576.4 |
MEST |
mesoderm specific transcript |
| chr8_-_124749609 | 0.21 |
ENST00000262219.6 ENST00000419625.1 |
ANXA13 |
annexin A13 |
| chr5_+_6714718 | 0.18 |
ENST00000230859.6 ENST00000515721.1 |
PAPD7 |
PAP associated domain containing 7 |
| chr3_+_38307293 | 0.17 |
ENST00000311856.4 |
SLC22A13 |
solute carrier family 22 (organic anion/urate transporter), member 13 |
| chr4_-_70518941 | 0.16 |
ENST00000286604.4 ENST00000505512.1 ENST00000514019.1 |
UGT2A1 UGT2A1 |
UDP glucuronosyltransferase 2 family, polypeptide A1, complex locus UDP glucuronosyltransferase 2 family, polypeptide A1, complex locus |
| chr20_+_57414795 | 0.16 |
ENST00000371098.2 ENST00000371075.3 |
GNAS |
GNAS complex locus |
| chr7_+_152456904 | 0.15 |
ENST00000537264.1 |
ACTR3B |
ARP3 actin-related protein 3 homolog B (yeast) |
| chr5_-_77844974 | 0.13 |
ENST00000515007.2 |
LHFPL2 |
lipoma HMGIC fusion partner-like 2 |
| chr12_-_18243119 | 0.11 |
ENST00000538724.1 ENST00000229002.2 |
RERGL |
RERG/RAS-like |
| chr10_+_115312766 | 0.11 |
ENST00000351270.3 |
HABP2 |
hyaluronan binding protein 2 |
| chr13_+_96085847 | 0.07 |
ENST00000376873.3 |
CLDN10 |
claudin 10 |
| chr13_-_41593425 | 0.05 |
ENST00000239882.3 |
ELF1 |
E74-like factor 1 (ets domain transcription factor) |
| chr7_+_152456829 | 0.05 |
ENST00000377776.3 ENST00000256001.8 ENST00000397282.2 |
ACTR3B |
ARP3 actin-related protein 3 homolog B (yeast) |
| chr6_+_106534192 | 0.04 |
ENST00000369091.2 ENST00000369096.4 |
PRDM1 |
PR domain containing 1, with ZNF domain |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.4 | 41.9 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 6.4 | 19.1 | GO:1902122 | chenodeoxycholic acid binding(GO:1902122) |
| 4.3 | 17.4 | GO:0047718 | enone reductase activity(GO:0035671) androsterone dehydrogenase activity(GO:0047023) indanol dehydrogenase activity(GO:0047718) |
| 4.1 | 16.5 | GO:0004657 | proline dehydrogenase activity(GO:0004657) |
| 3.7 | 14.9 | GO:0019770 | IgG receptor activity(GO:0019770) |
| 2.9 | 31.7 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 2.6 | 25.5 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 2.0 | 7.9 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 1.9 | 11.4 | GO:0004522 | ribonuclease A activity(GO:0004522) |
| 1.8 | 12.8 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 1.7 | 6.9 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 1.6 | 14.1 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 1.5 | 18.2 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 1.4 | 12.4 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 1.4 | 9.6 | GO:0001733 | galactosylceramide sulfotransferase activity(GO:0001733) galactose 3-O-sulfotransferase activity(GO:0050694) |
| 1.2 | 14.7 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 1.1 | 5.4 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 1.0 | 64.8 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 1.0 | 16.5 | GO:0015643 | toxic substance binding(GO:0015643) |
| 1.0 | 9.0 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.9 | 97.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.8 | 3.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.7 | 5.1 | GO:0071253 | connexin binding(GO:0071253) |
| 0.7 | 3.3 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.6 | 2.2 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 0.5 | 24.7 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.4 | 4.9 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.4 | 12.7 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.4 | 3.3 | GO:1990405 | protein antigen binding(GO:1990405) |
| 0.3 | 3.5 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.3 | 1.2 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.3 | 5.2 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.3 | 2.1 | GO:0070573 | metallodipeptidase activity(GO:0070573) |
| 0.3 | 29.3 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.3 | 5.8 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.2 | 3.9 | GO:0008569 | ATP-dependent microtubule motor activity, minus-end-directed(GO:0008569) |
| 0.2 | 0.7 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 0.2 | 0.8 | GO:0008422 | beta-glucosidase activity(GO:0008422) |
| 0.2 | 11.6 | GO:0005548 | phospholipid transporter activity(GO:0005548) |
| 0.2 | 5.3 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.2 | 1.2 | GO:0043559 | insulin binding(GO:0043559) |
| 0.2 | 0.8 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.2 | 3.3 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.2 | 8.8 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.2 | 2.2 | GO:0043295 | glutathione binding(GO:0043295) |
| 0.1 | 12.4 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.1 | 5.4 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.1 | 5.6 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 2.4 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 1.3 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.1 | 2.5 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.1 | 20.5 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.1 | 0.7 | GO:0051430 | corticotropin-releasing hormone receptor binding(GO:0051429) corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.1 | 1.7 | GO:0001848 | complement binding(GO:0001848) |
| 0.1 | 0.7 | GO:0031014 | troponin T binding(GO:0031014) |
| 0.0 | 1.0 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.0 | 0.5 | GO:0015377 | cation:chloride symporter activity(GO:0015377) |
| 0.0 | 2.8 | GO:0004702 | receptor signaling protein serine/threonine kinase activity(GO:0004702) |
| 0.0 | 0.2 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.0 | 0.8 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.2 | GO:0015101 | organic cation transmembrane transporter activity(GO:0015101) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.6 | 72.2 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 4.1 | 12.4 | GO:0030895 | apolipoprotein B mRNA editing enzyme complex(GO:0030895) |
| 3.0 | 27.2 | GO:0005579 | membrane attack complex(GO:0005579) |
| 2.7 | 47.7 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 2.6 | 18.0 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 1.9 | 11.4 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 1.6 | 12.8 | GO:0045179 | apical cortex(GO:0045179) |
| 1.0 | 5.1 | GO:0089717 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.7 | 3.3 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.6 | 9.0 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.6 | 208.5 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.5 | 19.1 | GO:0005719 | nuclear euchromatin(GO:0005719) |
| 0.4 | 3.9 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.4 | 40.3 | GO:0044215 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) |
| 0.4 | 3.3 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.3 | 3.1 | GO:0097425 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.3 | 5.4 | GO:0031095 | platelet dense tubular network membrane(GO:0031095) |
| 0.3 | 1.2 | GO:1990578 | perinuclear endoplasmic reticulum membrane(GO:1990578) |
| 0.3 | 1.3 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 0.2 | 2.2 | GO:0035686 | sperm fibrous sheath(GO:0035686) |
| 0.2 | 3.5 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.2 | 1.6 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.2 | 6.3 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.2 | 1.1 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 1.0 | GO:0070938 | contractile ring(GO:0070938) |
| 0.1 | 7.1 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.1 | 2.1 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.1 | 5.1 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 37.4 | GO:0030133 | transport vesicle(GO:0030133) |
| 0.1 | 6.9 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 14.8 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 0.8 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.8 | GO:0071004 | U2-type prespliceosome(GO:0071004) |
| 0.0 | 50.6 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.7 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 1.5 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 13.4 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 5.6 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 10.7 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 0.1 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 3.5 | GO:0019898 | extrinsic component of membrane(GO:0019898) |
| 0.0 | 1.9 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.0 | 1.5 | GO:0005770 | late endosome(GO:0005770) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 129.6 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 1.0 | 54.2 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.7 | 49.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.4 | 19.1 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.4 | 32.7 | PID FGF PATHWAY | FGF signaling pathway |
| 0.3 | 15.3 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.3 | 69.8 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.2 | 12.7 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.1 | 9.0 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.1 | 4.4 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.1 | 10.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 2.2 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 1.3 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.0 | 2.8 | PID TCR PATHWAY | TCR signaling in naïve CD4+ T cells |
| 0.0 | 1.7 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 1.1 | PID BARD1 PATHWAY | BARD1 signaling events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.0 | 112.8 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 3.4 | 50.5 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 1.8 | 25.5 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 1.4 | 34.9 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 1.2 | 26.8 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 1.2 | 26.1 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.9 | 16.5 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.5 | 9.0 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.4 | 14.7 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.4 | 8.8 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.3 | 5.2 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.3 | 5.8 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.3 | 5.1 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.2 | 12.3 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.2 | 9.6 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.2 | 3.3 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.2 | 20.5 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.1 | 3.9 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 12.8 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.1 | 2.5 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 1.2 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.1 | 3.5 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 12.4 | REACTOME MRNA PROCESSING | Genes involved in mRNA Processing |
| 0.1 | 0.8 | REACTOME FGFR LIGAND BINDING AND ACTIVATION | Genes involved in FGFR ligand binding and activation |
| 0.0 | 1.1 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.8 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 0.7 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.0 | 0.7 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.5 | 25.5 | GO:0010760 | negative regulation of macrophage chemotaxis(GO:0010760) |
| 6.4 | 19.1 | GO:2000213 | nitrogen catabolite regulation of transcription from RNA polymerase II promoter(GO:0001079) nitrogen catabolite activation of transcription from RNA polymerase II promoter(GO:0001080) regulation of urea metabolic process(GO:0034255) intracellular bile acid receptor signaling pathway(GO:0038185) interleukin-17 secretion(GO:0072615) nitrogen catabolite regulation of transcription(GO:0090293) nitrogen catabolite activation of transcription(GO:0090294) regulation of nitrogen cycle metabolic process(GO:1903314) positive regulation of glutamate metabolic process(GO:2000213) regulation of ammonia assimilation cycle(GO:2001248) positive regulation of ammonia assimilation cycle(GO:2001250) |
| 5.8 | 17.4 | GO:0016095 | polyprenol catabolic process(GO:0016095) |
| 5.7 | 40.0 | GO:0008218 | bioluminescence(GO:0008218) |
| 5.6 | 16.8 | GO:0016999 | antibiotic metabolic process(GO:0016999) |
| 5.5 | 16.5 | GO:0019836 | hemolysis by symbiont of host erythrocytes(GO:0019836) hemolysis in other organism(GO:0044179) hemolysis in other organism involved in symbiotic interaction(GO:0052331) |
| 5.4 | 54.2 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 5.4 | 59.1 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 5.1 | 25.5 | GO:1903412 | response to bile acid(GO:1903412) |
| 5.1 | 45.8 | GO:0045959 | regulation of complement activation, classical pathway(GO:0030450) negative regulation of complement activation, classical pathway(GO:0045959) regulation of opsonization(GO:1903027) |
| 4.5 | 66.8 | GO:0001542 | ovulation from ovarian follicle(GO:0001542) |
| 2.9 | 8.8 | GO:0052314 | serotonin biosynthetic process(GO:0042427) phytoalexin metabolic process(GO:0052314) |
| 2.8 | 16.5 | GO:0010133 | proline catabolic process to glutamate(GO:0010133) |
| 2.3 | 11.4 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 2.2 | 26.4 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 2.1 | 14.9 | GO:0002414 | immunoglobulin transcytosis in epithelial cells(GO:0002414) |
| 2.1 | 12.4 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 1.9 | 5.8 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 1.7 | 5.1 | GO:0090675 | intermicrovillar adhesion(GO:0090675) |
| 1.4 | 5.4 | GO:1903300 | negative regulation of glucokinase activity(GO:0033132) negative regulation of hexokinase activity(GO:1903300) |
| 1.3 | 5.1 | GO:0071393 | cellular response to progesterone stimulus(GO:0071393) |
| 1.3 | 9.0 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 1.3 | 8.9 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 1.3 | 12.8 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 1.2 | 5.9 | GO:0061107 | seminal vesicle development(GO:0061107) |
| 1.2 | 15.2 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) |
| 1.2 | 11.6 | GO:0034197 | acylglycerol transport(GO:0034196) triglyceride transport(GO:0034197) |
| 1.1 | 3.3 | GO:0000965 | mitochondrial RNA 3'-end processing(GO:0000965) rRNA import into mitochondrion(GO:0035928) |
| 1.1 | 9.6 | GO:0019375 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 1.0 | 3.1 | GO:0019557 | histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 1.0 | 3.9 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.9 | 2.8 | GO:0023016 | signal transduction by trans-phosphorylation(GO:0023016) negative regulation of rubidium ion transport(GO:2000681) negative regulation of rubidium ion transmembrane transporter activity(GO:2000687) |
| 0.9 | 5.4 | GO:0070560 | protein secretion by platelet(GO:0070560) |
| 0.9 | 16.1 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.9 | 31.7 | GO:0036499 | PERK-mediated unfolded protein response(GO:0036499) |
| 0.8 | 3.3 | GO:0036343 | psychomotor behavior(GO:0036343) |
| 0.5 | 5.3 | GO:0098856 | intestinal cholesterol absorption(GO:0030299) intestinal lipid absorption(GO:0098856) |
| 0.5 | 68.7 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.4 | 2.2 | GO:0008065 | establishment of blood-nerve barrier(GO:0008065) |
| 0.4 | 2.4 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.4 | 0.4 | GO:0045861 | negative regulation of proteolysis(GO:0045861) |
| 0.3 | 2.9 | GO:0036112 | medium-chain fatty-acyl-CoA metabolic process(GO:0036112) |
| 0.2 | 2.2 | GO:0046146 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.2 | 3.5 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.2 | 4.0 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.2 | 18.8 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.2 | 0.7 | GO:0006447 | regulation of translational initiation by iron(GO:0006447) |
| 0.2 | 1.3 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.2 | 2.5 | GO:1901687 | glutathione derivative metabolic process(GO:1901685) glutathione derivative biosynthetic process(GO:1901687) |
| 0.1 | 13.7 | GO:0098869 | cellular oxidant detoxification(GO:0098869) |
| 0.1 | 1.2 | GO:1900103 | positive regulation of endoplasmic reticulum unfolded protein response(GO:1900103) |
| 0.1 | 6.3 | GO:0055092 | cholesterol homeostasis(GO:0042632) sterol homeostasis(GO:0055092) |
| 0.1 | 0.8 | GO:0090080 | positive regulation of MAPKKK cascade by fibroblast growth factor receptor signaling pathway(GO:0090080) |
| 0.1 | 5.2 | GO:0006730 | one-carbon metabolic process(GO:0006730) |
| 0.1 | 0.8 | GO:0015816 | glycine transport(GO:0015816) |
| 0.1 | 0.7 | GO:2000828 | post-embryonic body morphogenesis(GO:0040032) regulation of parathyroid hormone secretion(GO:2000828) |
| 0.1 | 1.0 | GO:0014894 | response to muscle inactivity involved in regulation of muscle adaptation(GO:0014877) response to denervation involved in regulation of muscle adaptation(GO:0014894) |
| 0.1 | 1.2 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.1 | 4.9 | GO:0045744 | negative regulation of G-protein coupled receptor protein signaling pathway(GO:0045744) |
| 0.1 | 3.3 | GO:0048286 | lung alveolus development(GO:0048286) |
| 0.1 | 4.5 | GO:0046324 | regulation of glucose import(GO:0046324) |
| 0.0 | 0.4 | GO:0098908 | regulation of neuronal action potential(GO:0098908) |
| 0.0 | 0.8 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.0 | 2.8 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 0.8 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 0.8 | GO:0007498 | mesoderm development(GO:0007498) |
| 0.0 | 4.9 | GO:0006612 | protein targeting to membrane(GO:0006612) |
| 0.0 | 0.2 | GO:0034356 | NAD biosynthesis via nicotinamide riboside salvage pathway(GO:0034356) |
| 0.0 | 1.6 | GO:0070301 | cellular response to hydrogen peroxide(GO:0070301) |
| 0.0 | 0.7 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.0 | 0.2 | GO:0007076 | mitotic chromosome condensation(GO:0007076) |