Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
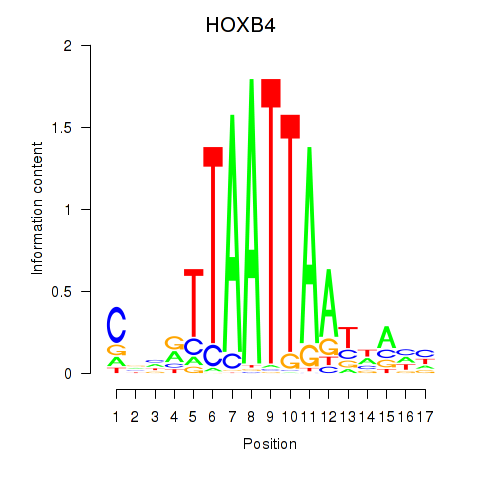
Results for HOXB4_LHX9
Z-value: 1.66
Transcription factors associated with HOXB4_LHX9
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXB4
|
ENSG00000182742.5 | HOXB4 |
|
LHX9
|
ENSG00000143355.11 | LHX9 |
Activity profile of HOXB4_LHX9 motif
Sorted Z-values of HOXB4_LHX9 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXB4_LHX9
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_169418195 | 17.92 |
ENST00000261509.6 ENST00000335742.7 |
PALLD |
palladin, cytoskeletal associated protein |
| chr17_-_39743139 | 11.85 |
ENST00000167586.6 |
KRT14 |
keratin 14 |
| chr4_+_169418255 | 8.27 |
ENST00000505667.1 ENST00000511948.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr14_-_67878917 | 6.99 |
ENST00000216446.4 |
PLEK2 |
pleckstrin 2 |
| chr12_-_8803128 | 6.80 |
ENST00000543467.1 |
MFAP5 |
microfibrillar associated protein 5 |
| chr10_-_105845674 | 5.98 |
ENST00000353479.5 ENST00000369733.3 |
COL17A1 |
collagen, type XVII, alpha 1 |
| chr11_+_5710919 | 5.55 |
ENST00000379965.3 ENST00000425490.1 |
TRIM22 |
tripartite motif containing 22 |
| chr7_+_134528635 | 5.05 |
ENST00000445569.2 |
CALD1 |
caldesmon 1 |
| chr8_-_41166953 | 4.52 |
ENST00000220772.3 |
SFRP1 |
secreted frizzled-related protein 1 |
| chr4_+_169552748 | 4.52 |
ENST00000504519.1 ENST00000512127.1 |
PALLD |
palladin, cytoskeletal associated protein |
| chr2_+_234637754 | 4.33 |
ENST00000482026.1 ENST00000609767.1 |
UGT1A3 UGT1A1 |
UDP glucuronosyltransferase 1 family, polypeptide A3 UDP glucuronosyltransferase 1 family, polypeptide A8 |
| chr7_-_87856303 | 3.26 |
ENST00000394641.3 |
SRI |
sorcin |
| chrX_+_43515467 | 3.24 |
ENST00000338702.3 ENST00000542639.1 |
MAOA |
monoamine oxidase A |
| chr3_-_149095652 | 3.14 |
ENST00000305366.3 |
TM4SF1 |
transmembrane 4 L six family member 1 |
| chr7_-_87856280 | 3.10 |
ENST00000490437.1 ENST00000431660.1 |
SRI |
sorcin |
| chr11_-_104827425 | 2.92 |
ENST00000393150.3 |
CASP4 |
caspase 4, apoptosis-related cysteine peptidase |
| chr18_+_21452804 | 2.92 |
ENST00000269217.6 |
LAMA3 |
laminin, alpha 3 |
| chr18_+_29027696 | 2.88 |
ENST00000257189.4 |
DSG3 |
desmoglein 3 |
| chr20_-_56265680 | 2.78 |
ENST00000414037.1 |
PMEPA1 |
prostate transmembrane protein, androgen induced 1 |
| chr3_-_151034734 | 2.70 |
ENST00000260843.4 |
GPR87 |
G protein-coupled receptor 87 |
| chr20_+_19867150 | 2.65 |
ENST00000255006.6 |
RIN2 |
Ras and Rab interactor 2 |
| chr8_-_49833978 | 2.60 |
ENST00000020945.1 |
SNAI2 |
snail family zinc finger 2 |
| chr8_-_49834299 | 2.59 |
ENST00000396822.1 |
SNAI2 |
snail family zinc finger 2 |
| chr12_+_26348246 | 2.56 |
ENST00000422622.2 |
SSPN |
sarcospan |
| chr18_+_21452964 | 2.54 |
ENST00000587184.1 |
LAMA3 |
laminin, alpha 3 |
| chr11_-_119991589 | 2.48 |
ENST00000526881.1 |
TRIM29 |
tripartite motif containing 29 |
| chr12_+_26348429 | 2.37 |
ENST00000242729.2 |
SSPN |
sarcospan |
| chr16_-_66584059 | 2.33 |
ENST00000417693.3 ENST00000544898.1 ENST00000569718.1 ENST00000527284.1 ENST00000299697.7 ENST00000451102.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr11_+_12766583 | 2.28 |
ENST00000361985.2 |
TEAD1 |
TEA domain family member 1 (SV40 transcriptional enhancer factor) |
| chr5_+_66300446 | 2.15 |
ENST00000261569.7 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr11_+_35201826 | 2.08 |
ENST00000531873.1 |
CD44 |
CD44 molecule (Indian blood group) |
| chr7_+_107224364 | 2.06 |
ENST00000491150.1 |
BCAP29 |
B-cell receptor-associated protein 29 |
| chr13_-_110438914 | 1.90 |
ENST00000375856.3 |
IRS2 |
insulin receptor substrate 2 |
| chr7_+_134430212 | 1.90 |
ENST00000436461.2 |
CALD1 |
caldesmon 1 |
| chr15_-_72523924 | 1.89 |
ENST00000566809.1 ENST00000567087.1 ENST00000569050.1 ENST00000568883.1 |
PKM |
pyruvate kinase, muscle |
| chr3_-_185538849 | 1.83 |
ENST00000421047.2 |
IGF2BP2 |
insulin-like growth factor 2 mRNA binding protein 2 |
| chr14_+_55595762 | 1.79 |
ENST00000254301.9 |
LGALS3 |
lectin, galactoside-binding, soluble, 3 |
| chr1_-_153518270 | 1.76 |
ENST00000354332.4 ENST00000368716.4 |
S100A4 |
S100 calcium binding protein A4 |
| chr12_-_10022735 | 1.73 |
ENST00000228438.2 |
CLEC2B |
C-type lectin domain family 2, member B |
| chr2_+_187454749 | 1.71 |
ENST00000261023.3 ENST00000374907.3 |
ITGAV |
integrin, alpha V |
| chr2_-_161056762 | 1.66 |
ENST00000428609.2 ENST00000409967.2 |
ITGB6 |
integrin, beta 6 |
| chr1_+_157963063 | 1.57 |
ENST00000360089.4 ENST00000368173.3 ENST00000392272.2 |
KIRREL |
kin of IRRE like (Drosophila) |
| chr15_+_64680003 | 1.56 |
ENST00000261884.3 |
TRIP4 |
thyroid hormone receptor interactor 4 |
| chr14_+_104182105 | 1.54 |
ENST00000311141.2 |
ZFYVE21 |
zinc finger, FYVE domain containing 21 |
| chr4_-_90757364 | 1.54 |
ENST00000508895.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr2_-_161056802 | 1.54 |
ENST00000283249.2 ENST00000409872.1 |
ITGB6 |
integrin, beta 6 |
| chr14_+_104182061 | 1.53 |
ENST00000216602.6 |
ZFYVE21 |
zinc finger, FYVE domain containing 21 |
| chr7_+_102553430 | 1.51 |
ENST00000339431.4 ENST00000249377.4 |
LRRC17 |
leucine rich repeat containing 17 |
| chr8_+_104310661 | 1.46 |
ENST00000522566.1 |
FZD6 |
frizzled family receptor 6 |
| chr7_-_107642348 | 1.45 |
ENST00000393561.1 |
LAMB1 |
laminin, beta 1 |
| chrX_-_13835147 | 1.44 |
ENST00000493677.1 ENST00000355135.2 |
GPM6B |
glycoprotein M6B |
| chr1_+_225600404 | 1.38 |
ENST00000366845.2 |
AC092811.1 |
AC092811.1 |
| chr9_+_125133315 | 1.36 |
ENST00000223423.4 ENST00000362012.2 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr12_+_81110684 | 1.34 |
ENST00000228644.3 |
MYF5 |
myogenic factor 5 |
| chr2_+_102615416 | 1.25 |
ENST00000393414.2 |
IL1R2 |
interleukin 1 receptor, type II |
| chr1_-_109584608 | 1.22 |
ENST00000400794.3 ENST00000528747.1 ENST00000369962.3 ENST00000361054.3 |
WDR47 |
WD repeat domain 47 |
| chr5_-_125930929 | 1.20 |
ENST00000553117.1 ENST00000447989.2 ENST00000409134.3 |
ALDH7A1 |
aldehyde dehydrogenase 7 family, member A1 |
| chr5_+_140593509 | 1.19 |
ENST00000341948.4 |
PCDHB13 |
protocadherin beta 13 |
| chr6_+_129204337 | 1.17 |
ENST00000421865.2 |
LAMA2 |
laminin, alpha 2 |
| chr1_-_67266939 | 1.10 |
ENST00000304526.2 |
INSL5 |
insulin-like 5 |
| chr3_+_111717600 | 1.08 |
ENST00000273368.4 |
TAGLN3 |
transgelin 3 |
| chr8_+_105235572 | 1.06 |
ENST00000523362.1 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr11_+_133938820 | 1.06 |
ENST00000299106.4 ENST00000529443.2 |
JAM3 |
junctional adhesion molecule 3 |
| chr14_+_103851712 | 1.04 |
ENST00000440884.3 ENST00000416682.2 ENST00000429436.2 ENST00000303622.9 |
MARK3 |
MAP/microtubule affinity-regulating kinase 3 |
| chr6_-_38670897 | 1.02 |
ENST00000373365.4 |
GLO1 |
glyoxalase I |
| chr6_+_114178512 | 0.94 |
ENST00000368635.4 |
MARCKS |
myristoylated alanine-rich protein kinase C substrate |
| chr3_+_111717511 | 0.93 |
ENST00000478951.1 ENST00000393917.2 |
TAGLN3 |
transgelin 3 |
| chr3_+_157154578 | 0.91 |
ENST00000295927.3 |
PTX3 |
pentraxin 3, long |
| chr6_-_52859046 | 0.88 |
ENST00000457564.1 ENST00000541324.1 ENST00000370960.1 |
GSTA4 |
glutathione S-transferase alpha 4 |
| chr16_-_28937027 | 0.87 |
ENST00000358201.4 |
RABEP2 |
rabaptin, RAB GTPase binding effector protein 2 |
| chr3_+_111718036 | 0.86 |
ENST00000455401.2 |
TAGLN3 |
transgelin 3 |
| chr12_-_10978957 | 0.85 |
ENST00000240619.2 |
TAS2R10 |
taste receptor, type 2, member 10 |
| chr5_-_16916624 | 0.85 |
ENST00000513882.1 |
MYO10 |
myosin X |
| chr10_+_64133934 | 0.84 |
ENST00000395254.3 ENST00000395255.3 ENST00000410046.3 |
ZNF365 |
zinc finger protein 365 |
| chr4_+_186990298 | 0.83 |
ENST00000296795.3 ENST00000513189.1 |
TLR3 |
toll-like receptor 3 |
| chr9_+_125132803 | 0.83 |
ENST00000540753.1 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr14_-_25078864 | 0.79 |
ENST00000216338.4 ENST00000557220.2 ENST00000382548.4 |
GZMH |
granzyme H (cathepsin G-like 2, protein h-CCPX) |
| chr10_-_75415825 | 0.75 |
ENST00000394810.2 |
SYNPO2L |
synaptopodin 2-like |
| chr2_-_74618964 | 0.73 |
ENST00000417090.1 ENST00000409868.1 |
DCTN1 |
dynactin 1 |
| chr2_+_102456277 | 0.73 |
ENST00000421882.1 |
MAP4K4 |
mitogen-activated protein kinase kinase kinase kinase 4 |
| chr19_+_58694396 | 0.73 |
ENST00000326804.4 ENST00000345813.3 ENST00000424679.2 |
ZNF274 |
zinc finger protein 274 |
| chr2_+_219110149 | 0.69 |
ENST00000456575.1 |
ARPC2 |
actin related protein 2/3 complex, subunit 2, 34kDa |
| chr2_-_74619152 | 0.67 |
ENST00000440727.1 ENST00000409240.1 |
DCTN1 |
dynactin 1 |
| chr6_-_91296602 | 0.66 |
ENST00000369325.3 ENST00000369327.3 |
MAP3K7 |
mitogen-activated protein kinase kinase kinase 7 |
| chr4_-_103749105 | 0.66 |
ENST00000394801.4 ENST00000394804.2 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chr14_-_80697396 | 0.65 |
ENST00000557010.1 |
DIO2 |
deiodinase, iodothyronine, type II |
| chrX_+_100663243 | 0.65 |
ENST00000316594.5 |
HNRNPH2 |
heterogeneous nuclear ribonucleoprotein H2 (H') |
| chr6_-_91296737 | 0.65 |
ENST00000369332.3 ENST00000369329.3 |
MAP3K7 |
mitogen-activated protein kinase kinase kinase 7 |
| chr4_-_170948361 | 0.65 |
ENST00000393702.3 |
MFAP3L |
microfibrillar-associated protein 3-like |
| chr4_-_103749313 | 0.65 |
ENST00000394803.5 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chrX_-_18690210 | 0.65 |
ENST00000379984.3 |
RS1 |
retinoschisin 1 |
| chr3_+_138340067 | 0.64 |
ENST00000479848.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr1_+_81771806 | 0.64 |
ENST00000370721.1 ENST00000370727.1 ENST00000370725.1 ENST00000370723.1 ENST00000370728.1 ENST00000370730.1 |
LPHN2 |
latrophilin 2 |
| chr3_-_122233723 | 0.63 |
ENST00000493510.1 ENST00000344337.6 ENST00000476916.1 ENST00000465882.1 |
KPNA1 |
karyopherin alpha 1 (importin alpha 5) |
| chr1_+_155278539 | 0.62 |
ENST00000447866.1 |
FDPS |
farnesyl diphosphate synthase |
| chr12_-_10151773 | 0.62 |
ENST00000298527.6 ENST00000348658.4 |
CLEC1B |
C-type lectin domain family 1, member B |
| chr4_-_57547870 | 0.61 |
ENST00000381260.3 ENST00000420433.1 ENST00000554144.1 ENST00000557328.1 |
HOPX |
HOP homeobox |
| chr16_-_66583701 | 0.60 |
ENST00000527800.1 ENST00000525974.1 ENST00000563369.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr11_-_117186946 | 0.58 |
ENST00000313005.6 ENST00000528053.1 |
BACE1 |
beta-site APP-cleaving enzyme 1 |
| chr8_+_9953214 | 0.58 |
ENST00000382490.5 |
MSRA |
methionine sulfoxide reductase A |
| chr8_-_30670384 | 0.58 |
ENST00000221138.4 ENST00000518243.1 |
PPP2CB |
protein phosphatase 2, catalytic subunit, beta isozyme |
| chr10_+_122610687 | 0.57 |
ENST00000263461.6 |
WDR11 |
WD repeat domain 11 |
| chr8_+_9953061 | 0.57 |
ENST00000522907.1 ENST00000528246.1 |
MSRA |
methionine sulfoxide reductase A |
| chr15_+_89631381 | 0.57 |
ENST00000352732.5 |
ABHD2 |
abhydrolase domain containing 2 |
| chr1_+_155278625 | 0.56 |
ENST00000368356.4 ENST00000356657.6 |
FDPS |
farnesyl diphosphate synthase |
| chr11_-_128894053 | 0.56 |
ENST00000392657.3 |
ARHGAP32 |
Rho GTPase activating protein 32 |
| chr20_+_18488137 | 0.54 |
ENST00000450074.1 ENST00000262544.2 ENST00000336714.3 ENST00000377475.3 |
SEC23B |
Sec23 homolog B (S. cerevisiae) |
| chr18_-_51751132 | 0.54 |
ENST00000256429.3 |
MBD2 |
methyl-CpG binding domain protein 2 |
| chr3_+_138340049 | 0.54 |
ENST00000464668.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr7_+_115862858 | 0.53 |
ENST00000393481.2 |
TES |
testis derived transcript (3 LIM domains) |
| chr1_+_28261492 | 0.51 |
ENST00000373894.3 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr5_-_176889381 | 0.51 |
ENST00000393563.4 ENST00000512501.1 |
DBN1 |
drebrin 1 |
| chr4_+_157997273 | 0.50 |
ENST00000541722.1 ENST00000512619.1 |
GLRB |
glycine receptor, beta |
| chrX_-_21776281 | 0.50 |
ENST00000379494.3 |
SMPX |
small muscle protein, X-linked |
| chr2_+_237994519 | 0.50 |
ENST00000392008.2 ENST00000409334.1 ENST00000409629.1 |
COPS8 |
COP9 signalosome subunit 8 |
| chr15_-_72563585 | 0.50 |
ENST00000287196.9 ENST00000260376.7 |
PARP6 |
poly (ADP-ribose) polymerase family, member 6 |
| chr20_-_1447547 | 0.49 |
ENST00000476071.1 |
NSFL1C |
NSFL1 (p97) cofactor (p47) |
| chr11_-_327537 | 0.48 |
ENST00000602735.1 |
IFITM3 |
interferon induced transmembrane protein 3 |
| chr11_-_89541743 | 0.45 |
ENST00000329758.1 |
TRIM49 |
tripartite motif containing 49 |
| chr6_+_29079668 | 0.45 |
ENST00000377169.1 |
OR2J3 |
olfactory receptor, family 2, subfamily J, member 3 |
| chrX_+_51486481 | 0.44 |
ENST00000340438.4 |
GSPT2 |
G1 to S phase transition 2 |
| chr6_+_29429217 | 0.44 |
ENST00000396792.2 |
OR2H1 |
olfactory receptor, family 2, subfamily H, member 1 |
| chr4_-_120243545 | 0.44 |
ENST00000274024.3 |
FABP2 |
fatty acid binding protein 2, intestinal |
| chr9_-_26947453 | 0.44 |
ENST00000397292.3 |
PLAA |
phospholipase A2-activating protein |
| chr20_-_1447467 | 0.43 |
ENST00000353088.2 ENST00000350991.4 |
NSFL1C |
NSFL1 (p97) cofactor (p47) |
| chr12_-_25348007 | 0.43 |
ENST00000354189.5 ENST00000545133.1 ENST00000554347.1 ENST00000395987.3 ENST00000320267.9 ENST00000395990.2 ENST00000537577.1 |
CASC1 |
cancer susceptibility candidate 1 |
| chr16_-_66764119 | 0.41 |
ENST00000569320.1 |
DYNC1LI2 |
dynein, cytoplasmic 1, light intermediate chain 2 |
| chr4_-_69536346 | 0.40 |
ENST00000338206.5 |
UGT2B15 |
UDP glucuronosyltransferase 2 family, polypeptide B15 |
| chr6_+_26402517 | 0.39 |
ENST00000414912.2 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chr4_-_70518941 | 0.38 |
ENST00000286604.4 ENST00000505512.1 ENST00000514019.1 |
UGT2A1 UGT2A1 |
UDP glucuronosyltransferase 2 family, polypeptide A1, complex locus UDP glucuronosyltransferase 2 family, polypeptide A1, complex locus |
| chrX_-_16887963 | 0.37 |
ENST00000380084.4 |
RBBP7 |
retinoblastoma binding protein 7 |
| chrX_+_55744166 | 0.36 |
ENST00000374941.4 ENST00000414239.1 |
RRAGB |
Ras-related GTP binding B |
| chr14_+_50234309 | 0.36 |
ENST00000298307.5 |
KLHDC2 |
kelch domain containing 2 |
| chrX_+_55744228 | 0.34 |
ENST00000262850.7 |
RRAGB |
Ras-related GTP binding B |
| chr15_+_67418047 | 0.33 |
ENST00000540846.2 |
SMAD3 |
SMAD family member 3 |
| chr12_+_147052 | 0.33 |
ENST00000594563.1 |
AC026369.1 |
Uncharacterized protein |
| chr1_-_211307404 | 0.32 |
ENST00000367007.4 |
KCNH1 |
potassium voltage-gated channel, subfamily H (eag-related), member 1 |
| chr3_+_149191723 | 0.32 |
ENST00000305354.4 |
TM4SF4 |
transmembrane 4 L six family member 4 |
| chr13_-_36050819 | 0.31 |
ENST00000379919.4 |
MAB21L1 |
mab-21-like 1 (C. elegans) |
| chr7_-_81399329 | 0.30 |
ENST00000453411.1 ENST00000444829.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr7_-_44122063 | 0.30 |
ENST00000335195.6 ENST00000395831.3 ENST00000414235.1 ENST00000452049.1 ENST00000242248.5 |
POLM |
polymerase (DNA directed), mu |
| chr4_-_103748880 | 0.29 |
ENST00000453744.2 ENST00000349311.8 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chr5_+_140227357 | 0.29 |
ENST00000378122.3 |
PCDHA9 |
protocadherin alpha 9 |
| chr10_-_28571015 | 0.28 |
ENST00000375719.3 ENST00000375732.1 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr3_+_148709128 | 0.28 |
ENST00000345003.4 ENST00000296048.6 ENST00000483267.1 |
GYG1 |
glycogenin 1 |
| chr11_-_18548426 | 0.28 |
ENST00000357193.3 ENST00000536719.1 |
TSG101 |
tumor susceptibility 101 |
| chr8_-_110986918 | 0.27 |
ENST00000297404.1 |
KCNV1 |
potassium channel, subfamily V, member 1 |
| chr9_-_95640218 | 0.27 |
ENST00000395506.3 ENST00000375495.3 ENST00000332591.6 |
ZNF484 |
zinc finger protein 484 |
| chr2_+_113763031 | 0.26 |
ENST00000259211.6 |
IL36A |
interleukin 36, alpha |
| chr3_+_111718173 | 0.25 |
ENST00000494932.1 |
TAGLN3 |
transgelin 3 |
| chr20_-_43133491 | 0.25 |
ENST00000411544.1 |
SERINC3 |
serine incorporator 3 |
| chr4_-_119759795 | 0.25 |
ENST00000419654.2 |
SEC24D |
SEC24 family member D |
| chr14_-_104181771 | 0.24 |
ENST00000554913.1 ENST00000554974.1 ENST00000553361.1 ENST00000555055.1 ENST00000555964.1 ENST00000556682.1 ENST00000445556.1 ENST00000553332.1 ENST00000352127.7 |
XRCC3 |
X-ray repair complementing defective repair in Chinese hamster cells 3 |
| chr5_+_140220769 | 0.23 |
ENST00000531613.1 ENST00000378123.3 |
PCDHA8 |
protocadherin alpha 8 |
| chr19_+_23945768 | 0.23 |
ENST00000486528.1 ENST00000496398.1 |
RPSAP58 |
ribosomal protein SA pseudogene 58 |
| chr10_+_5135981 | 0.23 |
ENST00000380554.3 |
AKR1C3 |
aldo-keto reductase family 1, member C3 |
| chr11_+_33061543 | 0.23 |
ENST00000432887.1 ENST00000528898.1 ENST00000531632.2 |
TCP11L1 |
t-complex 11, testis-specific-like 1 |
| chrY_-_6740649 | 0.23 |
ENST00000383036.1 ENST00000383037.4 |
AMELY |
amelogenin, Y-linked |
| chr17_-_4938712 | 0.22 |
ENST00000254853.5 ENST00000424747.1 |
SLC52A1 |
solute carrier family 52 (riboflavin transporter), member 1 |
| chr21_-_31538971 | 0.22 |
ENST00000286808.3 |
CLDN17 |
claudin 17 |
| chr6_-_52705641 | 0.21 |
ENST00000370989.2 |
GSTA5 |
glutathione S-transferase alpha 5 |
| chrX_-_138790348 | 0.20 |
ENST00000414978.1 ENST00000519895.1 |
MCF2 |
MCF.2 cell line derived transforming sequence |
| chr22_-_32766972 | 0.19 |
ENST00000382084.4 ENST00000382086.2 |
RFPL3S |
RFPL3 antisense |
| chr14_-_74551172 | 0.19 |
ENST00000553458.1 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr3_+_136649311 | 0.19 |
ENST00000469404.1 ENST00000467911.1 |
NCK1 |
NCK adaptor protein 1 |
| chr11_-_46408107 | 0.19 |
ENST00000433765.2 |
CHRM4 |
cholinergic receptor, muscarinic 4 |
| chr15_+_62853562 | 0.19 |
ENST00000561311.1 |
TLN2 |
talin 2 |
| chr13_+_48611665 | 0.18 |
ENST00000258662.2 |
NUDT15 |
nudix (nucleoside diphosphate linked moiety X)-type motif 15 |
| chrX_+_13671225 | 0.18 |
ENST00000545566.1 ENST00000544987.1 ENST00000314720.4 |
TCEANC |
transcription elongation factor A (SII) N-terminal and central domain containing |
| chrX_-_100872911 | 0.18 |
ENST00000361910.4 ENST00000539247.1 ENST00000538627.1 |
ARMCX6 |
armadillo repeat containing, X-linked 6 |
| chr1_+_196743912 | 0.18 |
ENST00000367425.4 |
CFHR3 |
complement factor H-related 3 |
| chr9_+_117904097 | 0.17 |
ENST00000374016.1 |
DEC1 |
deleted in esophageal cancer 1 |
| chr3_-_160823158 | 0.17 |
ENST00000392779.2 ENST00000392780.1 ENST00000494173.1 |
B3GALNT1 |
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) |
| chr6_-_87804815 | 0.17 |
ENST00000369582.2 |
CGA |
glycoprotein hormones, alpha polypeptide |
| chr19_+_49199209 | 0.17 |
ENST00000522966.1 ENST00000425340.2 ENST00000391876.4 |
FUT2 |
fucosyltransferase 2 (secretor status included) |
| chr15_+_89631647 | 0.17 |
ENST00000569550.1 ENST00000565066.1 ENST00000565973.1 |
ABHD2 |
abhydrolase domain containing 2 |
| chr11_+_57480046 | 0.16 |
ENST00000378312.4 ENST00000278422.4 |
TMX2 |
thioredoxin-related transmembrane protein 2 |
| chr5_+_140557371 | 0.15 |
ENST00000239444.2 |
PCDHB8 |
protocadherin beta 8 |
| chr2_-_86850949 | 0.15 |
ENST00000237455.4 |
RNF103 |
ring finger protein 103 |
| chr7_-_81399411 | 0.15 |
ENST00000423064.2 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr4_+_37455536 | 0.15 |
ENST00000381980.4 ENST00000508175.1 |
C4orf19 |
chromosome 4 open reading frame 19 |
| chr15_+_66679155 | 0.14 |
ENST00000307102.5 |
MAP2K1 |
mitogen-activated protein kinase kinase 1 |
| chr6_+_43395272 | 0.14 |
ENST00000372530.4 |
ABCC10 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 10 |
| chr4_+_88571429 | 0.14 |
ENST00000339673.6 ENST00000282479.7 |
DMP1 |
dentin matrix acidic phosphoprotein 1 |
| chr17_-_73663245 | 0.14 |
ENST00000584999.1 ENST00000317905.5 ENST00000420326.2 ENST00000340830.5 |
RECQL5 |
RecQ protein-like 5 |
| chrX_+_107334895 | 0.13 |
ENST00000372232.3 ENST00000345734.3 ENST00000372254.3 |
ATG4A |
autophagy related 4A, cysteine peptidase |
| chr1_-_233431458 | 0.12 |
ENST00000258229.9 ENST00000430153.1 |
PCNXL2 |
pecanex-like 2 (Drosophila) |
| chr4_+_156587853 | 0.12 |
ENST00000506455.1 ENST00000511108.1 |
GUCY1A3 |
guanylate cyclase 1, soluble, alpha 3 |
| chr4_-_103749205 | 0.12 |
ENST00000508249.1 |
UBE2D3 |
ubiquitin-conjugating enzyme E2D 3 |
| chr12_-_68619586 | 0.12 |
ENST00000229134.4 |
IL26 |
interleukin 26 |
| chr10_+_90424196 | 0.11 |
ENST00000394375.3 ENST00000608620.1 ENST00000238983.4 ENST00000355843.2 |
LIPF |
lipase, gastric |
| chr3_-_160823040 | 0.11 |
ENST00000484127.1 ENST00000492353.1 ENST00000473142.1 ENST00000468268.1 ENST00000460353.1 ENST00000320474.4 ENST00000392781.2 |
B3GALNT1 |
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) |
| chr11_-_64684672 | 0.11 |
ENST00000377264.3 ENST00000421419.2 |
ATG2A |
autophagy related 2A |
| chr5_-_149829244 | 0.11 |
ENST00000312037.5 |
RPS14 |
ribosomal protein S14 |
| chr4_-_74486217 | 0.11 |
ENST00000335049.5 ENST00000307439.5 |
RASSF6 |
Ras association (RalGDS/AF-6) domain family member 6 |
| chr1_-_57045228 | 0.11 |
ENST00000371250.3 |
PPAP2B |
phosphatidic acid phosphatase type 2B |
| chr1_-_77685084 | 0.11 |
ENST00000370812.3 ENST00000359130.1 ENST00000445065.1 ENST00000370813.5 |
PIGK |
phosphatidylinositol glycan anchor biosynthesis, class K |
| chr7_-_81399438 | 0.10 |
ENST00000222390.5 |
HGF |
hepatocyte growth factor (hepapoietin A; scatter factor) |
| chr12_+_49621658 | 0.10 |
ENST00000541364.1 |
TUBA1C |
tubulin, alpha 1c |
| chr19_-_58326267 | 0.10 |
ENST00000391701.1 |
ZNF552 |
zinc finger protein 552 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.2 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.2 | 4.3 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.2 | 2.9 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.1 | 2.1 | REACTOME TRAF6 MEDIATED INDUCTION OF TAK1 COMPLEX | Genes involved in TRAF6 mediated induction of TAK1 complex |
| 0.1 | 2.1 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.1 | 6.9 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 2.0 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.1 | 2.9 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.1 | 11.0 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.1 | 1.7 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.1 | 0.9 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 3.1 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.1 | 0.4 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.1 | 1.6 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.1 | 6.6 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 1.8 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
| 0.1 | 1.9 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.0 | 0.6 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 0.3 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.0 | 1.2 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 1.3 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.3 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 0.9 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.0 | 1.1 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 0.6 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 1.9 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.0 | 1.0 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 1.4 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 1.5 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.9 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.0 | 0.4 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.0 | 1.6 | REACTOME DIABETES PATHWAYS | Genes involved in Diabetes pathways |
| 0.0 | 0.6 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.0 | 0.7 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 0.3 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 0.3 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 1.2 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.4 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 11.8 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 1.7 | 5.2 | GO:0070563 | negative regulation of vitamin D receptor signaling pathway(GO:0070563) |
| 1.5 | 4.5 | GO:0090244 | Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 1.3 | 30.7 | GO:0003334 | keratinocyte development(GO:0003334) |
| 0.9 | 6.4 | GO:0006880 | intracellular sequestering of iron ion(GO:0006880) sequestering of iron ion(GO:0097577) |
| 0.9 | 4.3 | GO:0070980 | biphenyl catabolic process(GO:0070980) |
| 0.7 | 2.9 | GO:0046104 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.6 | 1.8 | GO:2000521 | negative regulation of immunological synapse formation(GO:2000521) |
| 0.5 | 4.9 | GO:0038044 | transforming growth factor-beta secretion(GO:0038044) |
| 0.4 | 11.4 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.4 | 1.5 | GO:0035880 | embryonic nail plate morphogenesis(GO:0035880) |
| 0.4 | 1.4 | GO:0021812 | neuronal-glial interaction involved in cerebral cortex radial glia guided migration(GO:0021812) |
| 0.3 | 1.9 | GO:0010746 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) |
| 0.3 | 1.9 | GO:0042866 | pyruvate biosynthetic process(GO:0042866) |
| 0.3 | 1.3 | GO:0050712 | negative regulation of interleukin-1 alpha production(GO:0032690) negative regulation of interleukin-1 alpha secretion(GO:0050712) |
| 0.3 | 1.5 | GO:0051621 | negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) regulation of peroxidase activity(GO:2000468) positive regulation of peroxidase activity(GO:2000470) |
| 0.3 | 1.5 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.3 | 1.4 | GO:0051612 | negative regulation of neurotransmitter uptake(GO:0051581) serotonin uptake(GO:0051610) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 0.3 | 1.1 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.3 | 2.9 | GO:1903265 | positive regulation of tumor necrosis factor-mediated signaling pathway(GO:1903265) |
| 0.3 | 2.1 | GO:1900625 | regulation of monocyte aggregation(GO:1900623) positive regulation of monocyte aggregation(GO:1900625) |
| 0.2 | 1.2 | GO:0033383 | geranyl diphosphate metabolic process(GO:0033383) geranyl diphosphate biosynthetic process(GO:0033384) farnesyl diphosphate biosynthetic process(GO:0045337) |
| 0.2 | 1.4 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.2 | 2.8 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.2 | 0.8 | GO:0034343 | microglial cell activation involved in immune response(GO:0002282) type III interferon production(GO:0034343) regulation of type III interferon production(GO:0034344) positive regulation of interferon-alpha biosynthetic process(GO:0045356) |
| 0.2 | 2.1 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.2 | 3.2 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.2 | 0.9 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 0.2 | 1.2 | GO:0042426 | choline catabolic process(GO:0042426) |
| 0.2 | 6.8 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.2 | 0.5 | GO:0097112 | gamma-aminobutyric acid receptor clustering(GO:0097112) |
| 0.1 | 1.0 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.1 | 0.4 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.1 | 1.0 | GO:0009438 | methylglyoxal metabolic process(GO:0009438) |
| 0.1 | 2.4 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.1 | 1.2 | GO:0030091 | protein repair(GO:0030091) |
| 0.1 | 0.4 | GO:0070370 | heat acclimation(GO:0010286) cellular heat acclimation(GO:0070370) |
| 0.1 | 1.3 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.1 | 1.1 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
| 0.1 | 1.2 | GO:0032224 | positive regulation of synaptic transmission, cholinergic(GO:0032224) |
| 0.1 | 0.7 | GO:1990253 | cellular response to leucine starvation(GO:1990253) |
| 0.1 | 2.3 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.1 | 0.5 | GO:0010643 | cell communication by chemical coupling(GO:0010643) |
| 0.1 | 5.4 | GO:0070206 | protein trimerization(GO:0070206) |
| 0.1 | 1.3 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.1 | 0.3 | GO:0070086 | ubiquitin-dependent endocytosis(GO:0070086) regulation of ubiquitin-dependent endocytosis(GO:2000395) positive regulation of ubiquitin-dependent endocytosis(GO:2000397) |
| 0.1 | 0.3 | GO:0009597 | detection of virus(GO:0009597) |
| 0.1 | 0.2 | GO:0071139 | resolution of recombination intermediates(GO:0071139) resolution of mitotic recombination intermediates(GO:0071140) |
| 0.1 | 0.6 | GO:0060665 | regulation of branching involved in salivary gland morphogenesis by mesenchymal-epithelial signaling(GO:0060665) |
| 0.1 | 2.6 | GO:1902895 | positive regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902895) |
| 0.1 | 0.7 | GO:1900112 | regulation of histone H3-K9 trimethylation(GO:1900112) |
| 0.1 | 0.6 | GO:0009642 | response to light intensity(GO:0009642) |
| 0.1 | 0.2 | GO:1903676 | regulation of cap-dependent translational initiation(GO:1903674) positive regulation of cap-dependent translational initiation(GO:1903676) |
| 0.1 | 0.7 | GO:0001514 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.1 | 0.2 | GO:0019859 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 0.1 | 3.1 | GO:0006458 | 'de novo' protein folding(GO:0006458) |
| 0.1 | 0.2 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.1 | 0.5 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.1 | 1.1 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.0 | 0.3 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.2 | GO:0006203 | dGTP catabolic process(GO:0006203) |
| 0.0 | 0.1 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 0.0 | 0.3 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.0 | 0.9 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 1.6 | GO:1901998 | toxin transport(GO:1901998) |
| 0.0 | 0.1 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.0 | 0.4 | GO:0010992 | ubiquitin homeostasis(GO:0010992) |
| 0.0 | 0.2 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.0 | 1.2 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 1.8 | GO:0035666 | TRIF-dependent toll-like receptor signaling pathway(GO:0035666) |
| 0.0 | 0.9 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.0 | 0.5 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.0 | 0.2 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 0.4 | GO:0006346 | methylation-dependent chromatin silencing(GO:0006346) |
| 0.0 | 0.6 | GO:0001829 | trophectodermal cell differentiation(GO:0001829) |
| 0.0 | 0.9 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.0 | 0.6 | GO:0030220 | platelet formation(GO:0030220) |
| 0.0 | 0.7 | GO:0048240 | sperm capacitation(GO:0048240) |
| 0.0 | 0.7 | GO:0008285 | negative regulation of cell proliferation(GO:0008285) |
| 0.0 | 0.7 | GO:0030033 | microvillus assembly(GO:0030033) |
| 0.0 | 0.9 | GO:1901685 | glutathione derivative metabolic process(GO:1901685) glutathione derivative biosynthetic process(GO:1901687) |
| 0.0 | 2.9 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 0.1 | GO:0090170 | regulation of Golgi inheritance(GO:0090170) |
| 0.0 | 0.6 | GO:0019054 | modulation by virus of host process(GO:0019054) |
| 0.0 | 2.3 | GO:1900181 | negative regulation of protein localization to nucleus(GO:1900181) |
| 0.0 | 0.1 | GO:0052565 | response to defense-related nitric oxide production by other organism involved in symbiotic interaction(GO:0052551) response to defense-related host nitric oxide production(GO:0052565) |
| 0.0 | 0.5 | GO:0046597 | negative regulation of viral entry into host cell(GO:0046597) |
| 0.0 | 0.1 | GO:1902260 | negative regulation of delayed rectifier potassium channel activity(GO:1902260) negative regulation of voltage-gated potassium channel activity(GO:1903817) |
| 0.0 | 0.1 | GO:0032053 | ciliary basal body organization(GO:0032053) |
| 0.0 | 0.3 | GO:0002566 | somatic diversification of immune receptors via somatic mutation(GO:0002566) somatic hypermutation of immunoglobulin genes(GO:0016446) |
| 0.0 | 0.9 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 0.4 | GO:0034505 | tooth mineralization(GO:0034505) |
| 0.0 | 2.0 | GO:0030838 | positive regulation of actin filament polymerization(GO:0030838) |
| 0.0 | 0.8 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.0 | 0.8 | GO:0019835 | cytolysis(GO:0019835) |
| 0.0 | 8.3 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 0.0 | GO:0030950 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) |
| 0.0 | 0.2 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.0 | 0.1 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 0.3 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.0 | 0.6 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.0 | 0.1 | GO:0048712 | negative regulation of astrocyte differentiation(GO:0048712) |
| 0.0 | 0.2 | GO:0019317 | fucose catabolic process(GO:0019317) L-fucose metabolic process(GO:0042354) L-fucose catabolic process(GO:0042355) |
| 0.0 | 0.5 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.0 | 0.4 | GO:0007608 | sensory perception of smell(GO:0007608) |
| 0.0 | 0.6 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 0.1 | GO:2001023 | regulation of response to drug(GO:2001023) |
| 0.0 | 0.2 | GO:0090110 | cargo loading into COPII-coated vesicle(GO:0090110) |
| 0.0 | 4.4 | GO:0031346 | positive regulation of cell projection organization(GO:0031346) |
| 0.0 | 0.0 | GO:0060748 | tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 8.1 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.2 | 4.9 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 11.8 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.1 | 6.0 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 4.6 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 6.1 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.1 | 2.1 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
| 0.1 | 2.8 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.1 | 1.8 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 6.5 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.0 | 1.8 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.0 | 0.8 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 1.8 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 1.9 | SIG IL4RECEPTOR IN B LYPHOCYTES | Genes related to IL4 rceptor signaling in B lymphocytes |
| 0.0 | 3.2 | PID MET PATHWAY | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) |
| 0.0 | 1.4 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.7 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.0 | 0.9 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.0 | 1.5 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.6 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.8 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.7 | PID PI3KCI AKT PATHWAY | Class I PI3K signaling events mediated by Akt |
| 0.0 | 0.7 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 0.9 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 2.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.3 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.8 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 0.7 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 0.3 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 0.5 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 11.8 | GO:1990254 | keratin filament binding(GO:1990254) |
| 1.3 | 30.0 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 1.0 | 2.9 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) thymidine kinase activity(GO:0004797) |
| 0.5 | 2.2 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.5 | 3.1 | GO:0000285 | 1-phosphatidylinositol-3-phosphate 5-kinase activity(GO:0000285) |
| 0.5 | 1.9 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.4 | 3.2 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.3 | 1.7 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.3 | 1.5 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.3 | 7.7 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.3 | 6.0 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.3 | 1.3 | GO:0004910 | interleukin-1, Type II, blocking receptor activity(GO:0004910) |
| 0.2 | 6.9 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.2 | 1.2 | GO:0004337 | dimethylallyltranstransferase activity(GO:0004161) geranyltranstransferase activity(GO:0004337) |
| 0.2 | 1.2 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.2 | 1.8 | GO:0019863 | IgE binding(GO:0019863) |
| 0.2 | 2.9 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.2 | 0.6 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.2 | 6.4 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 1.0 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.1 | 0.9 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.1 | 2.3 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 0.7 | GO:0004800 | thyroxine 5'-deiodinase activity(GO:0004800) |
| 0.1 | 2.3 | GO:0001134 | transcription factor activity, transcription factor recruiting(GO:0001134) |
| 0.1 | 1.7 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 0.5 | GO:0016934 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.1 | 2.5 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.1 | 0.3 | GO:0047273 | galactosylgalactosylglucosylceramide beta-D-acetylgalactosaminyltransferase activity(GO:0047273) |
| 0.1 | 0.8 | GO:0060002 | plus-end directed microfilament motor activity(GO:0060002) |
| 0.1 | 0.7 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.1 | 3.4 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 8.4 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 1.6 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.1 | 0.9 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.1 | 0.4 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.1 | 0.3 | GO:0008466 | glycogenin glucosyltransferase activity(GO:0008466) |
| 0.1 | 0.3 | GO:0031962 | mineralocorticoid receptor binding(GO:0031962) |
| 0.1 | 2.8 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 1.9 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.1 | 0.2 | GO:0035539 | 8-oxo-7,8-dihydroguanosine triphosphate pyrophosphatase activity(GO:0008413) NADH pyrophosphatase activity(GO:0035529) 8-oxo-7,8-dihydrodeoxyguanosine triphosphate pyrophosphatase activity(GO:0035539) |
| 0.1 | 0.2 | GO:0047718 | enone reductase activity(GO:0035671) androsterone dehydrogenase activity(GO:0047023) indanol dehydrogenase activity(GO:0047718) |
| 0.1 | 0.7 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.1 | 0.2 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.1 | 0.3 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.1 | 1.6 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.1 | 1.4 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.1 | 1.8 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.1 | 0.5 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 1.2 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.0 | 2.7 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.0 | 0.4 | GO:0003696 | satellite DNA binding(GO:0003696) |
| 0.0 | 5.6 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 0.2 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.0 | 0.2 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.1 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.0 | 0.3 | GO:0046790 | virion binding(GO:0046790) |
| 0.0 | 0.3 | GO:0015194 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.0 | 0.2 | GO:0008821 | crossover junction endodeoxyribonuclease activity(GO:0008821) |
| 0.0 | 1.3 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 2.8 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.0 | 0.5 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 1.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.4 | GO:0016004 | phospholipase activator activity(GO:0016004) |
| 0.0 | 0.1 | GO:0009378 | four-way junction helicase activity(GO:0009378) |
| 0.0 | 0.9 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 6.1 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 0.4 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.0 | 0.6 | GO:0042056 | chemoattractant activity(GO:0042056) |
| 0.0 | 1.5 | GO:0017022 | myosin binding(GO:0017022) |
| 0.0 | 1.3 | GO:0097110 | scaffold protein binding(GO:0097110) |
| 0.0 | 3.6 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.5 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.9 | GO:0004984 | olfactory receptor activity(GO:0004984) |
| 0.0 | 0.3 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 0.2 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.0 | 0.4 | GO:0004407 | histone deacetylase activity(GO:0004407) protein deacetylase activity(GO:0033558) |
| 0.0 | 0.2 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.0 | 0.3 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 4.9 | GO:0034685 | integrin alphav-beta6 complex(GO:0034685) |
| 0.9 | 6.4 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.7 | 6.9 | GO:0030478 | actin cap(GO:0030478) |
| 0.7 | 5.5 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.6 | 0.6 | GO:0019898 | extrinsic component of membrane(GO:0019898) |
| 0.5 | 30.7 | GO:0002102 | podosome(GO:0002102) |
| 0.5 | 2.9 | GO:0072557 | IPAF inflammasome complex(GO:0072557) |
| 0.5 | 1.4 | GO:0043259 | laminin-2 complex(GO:0005607) laminin-10 complex(GO:0043259) |
| 0.5 | 2.3 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) |
| 0.3 | 6.8 | GO:0043205 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.3 | 2.1 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 0.3 | 6.0 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.3 | 11.8 | GO:0045095 | keratin filament(GO:0045095) |
| 0.2 | 0.7 | GO:0036195 | muscle cell projection(GO:0036194) muscle cell projection membrane(GO:0036195) |
| 0.2 | 0.9 | GO:1990730 | VCP-NSFL1C complex(GO:1990730) |
| 0.2 | 4.3 | GO:0034663 | endoplasmic reticulum chaperone complex(GO:0034663) |
| 0.2 | 7.0 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.2 | 0.5 | GO:0016935 | glycine-gated chloride channel complex(GO:0016935) |
| 0.1 | 0.4 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.1 | 3.9 | GO:0030057 | desmosome(GO:0030057) |
| 0.1 | 3.9 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.1 | 0.7 | GO:1990131 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.1 | 1.3 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.1 | 1.4 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 0.5 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 0.1 | 5.5 | GO:0015030 | Cajal body(GO:0015030) |
| 0.1 | 0.8 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.1 | 1.5 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.1 | 0.7 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.1 | 1.5 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 0.3 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.0 | 0.8 | GO:0036020 | endolysosome membrane(GO:0036020) |
| 0.0 | 0.7 | GO:0043073 | germ cell nucleus(GO:0043073) |
| 0.0 | 0.3 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.2 | GO:0033061 | DNA recombinase mediator complex(GO:0033061) Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.0 | 0.3 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 1.2 | GO:0005605 | basal lamina(GO:0005605) |
| 0.0 | 0.1 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.0 | 0.8 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 1.6 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 1.4 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 1.1 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 1.6 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 2.5 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 2.2 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 0.5 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 4.7 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 2.8 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 5.9 | GO:0010008 | endosome membrane(GO:0010008) |
| 0.0 | 0.6 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 0.4 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.0 | 2.9 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.0 | 0.1 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.0 | 0.5 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.0 | 0.2 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 0.7 | GO:0005769 | early endosome(GO:0005769) |
| 0.0 | 0.4 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 0.9 | GO:0035580 | specific granule lumen(GO:0035580) |
| 0.0 | 0.1 | GO:1990635 | proximal dendrite(GO:1990635) |