Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
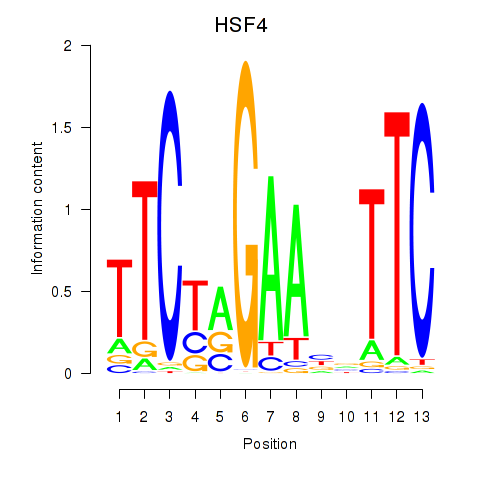
Results for HSF4
Z-value: 1.07
Transcription factors associated with HSF4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HSF4
|
ENSG00000102878.11 | HSF4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| HSF4 | hg19_v2_chr16_+_67198683_67198715 | -0.16 | 5.5e-01 | Click! |
Activity profile of HSF4 motif
Sorted Z-values of HSF4 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of HSF4
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_122933043 | 3.51 |
ENST00000534624.1 ENST00000453788.2 ENST00000527387.1 |
HSPA8 |
heat shock 70kDa protein 8 |
| chr11_-_122932730 | 3.49 |
ENST00000532182.1 ENST00000524590.1 ENST00000528292.1 ENST00000533540.1 ENST00000525463.1 |
HSPA8 |
heat shock 70kDa protein 8 |
| chr11_-_111781610 | 2.62 |
ENST00000525823.1 |
CRYAB |
crystallin, alpha B |
| chr11_-_111781454 | 2.49 |
ENST00000533280.1 |
CRYAB |
crystallin, alpha B |
| chr7_-_56119156 | 2.40 |
ENST00000421312.1 ENST00000416592.1 |
PSPH |
phosphoserine phosphatase |
| chr11_+_63953691 | 2.32 |
ENST00000543847.1 |
STIP1 |
stress-induced-phosphoprotein 1 |
| chr7_-_56118981 | 2.20 |
ENST00000419984.2 ENST00000413218.1 ENST00000424596.1 |
PSPH |
phosphoserine phosphatase |
| chr11_-_111794446 | 2.19 |
ENST00000527950.1 |
CRYAB |
crystallin, alpha B |
| chr11_-_111782696 | 2.11 |
ENST00000227251.3 ENST00000526180.1 |
CRYAB |
crystallin, alpha B |
| chr9_-_88715044 | 2.04 |
ENST00000388711.3 ENST00000466178.1 |
GOLM1 |
golgi membrane protein 1 |
| chr11_-_111781554 | 1.94 |
ENST00000526167.1 ENST00000528961.1 |
CRYAB |
crystallin, alpha B |
| chr11_+_63953587 | 1.86 |
ENST00000305218.4 ENST00000538945.1 |
STIP1 |
stress-induced-phosphoprotein 1 |
| chr15_-_58357932 | 1.85 |
ENST00000347587.3 |
ALDH1A2 |
aldehyde dehydrogenase 1 family, member A2 |
| chr6_-_41909191 | 1.85 |
ENST00000512426.1 ENST00000372987.4 |
CCND3 |
cyclin D3 |
| chr14_-_52535712 | 1.81 |
ENST00000216286.5 ENST00000541773.1 |
NID2 |
nidogen 2 (osteonidogen) |
| chr9_-_88714421 | 1.73 |
ENST00000388712.3 |
GOLM1 |
golgi membrane protein 1 |
| chr6_-_41909561 | 1.68 |
ENST00000372991.4 |
CCND3 |
cyclin D3 |
| chr14_-_23388338 | 1.67 |
ENST00000555209.1 ENST00000554256.1 ENST00000557403.1 ENST00000557549.1 ENST00000555676.1 ENST00000557571.1 ENST00000557464.1 ENST00000554618.1 ENST00000556862.1 ENST00000555722.1 ENST00000346528.5 ENST00000542016.2 ENST00000399922.2 ENST00000557227.1 ENST00000359890.3 |
RBM23 |
RNA binding motif protein 23 |
| chr6_-_41909466 | 1.61 |
ENST00000414200.2 |
CCND3 |
cyclin D3 |
| chr11_+_86013253 | 1.39 |
ENST00000533986.1 ENST00000278483.3 |
C11orf73 |
chromosome 11 open reading frame 73 |
| chr11_-_111782484 | 1.37 |
ENST00000533971.1 |
CRYAB |
crystallin, alpha B |
| chr3_+_138066539 | 1.32 |
ENST00000289104.4 |
MRAS |
muscle RAS oncogene homolog |
| chr7_+_157129660 | 1.28 |
ENST00000429029.2 ENST00000262177.4 ENST00000417758.1 ENST00000452797.2 ENST00000443280.1 |
DNAJB6 |
DnaJ (Hsp40) homolog, subfamily B, member 6 |
| chr3_+_158787041 | 1.26 |
ENST00000471575.1 ENST00000476809.1 ENST00000485419.1 |
IQCJ-SCHIP1 |
IQCJ-SCHIP1 readthrough |
| chr15_-_58357866 | 1.25 |
ENST00000537372.1 |
ALDH1A2 |
aldehyde dehydrogenase 1 family, member A2 |
| chr20_+_19867150 | 1.20 |
ENST00000255006.6 |
RIN2 |
Ras and Rab interactor 2 |
| chr7_+_75931861 | 1.14 |
ENST00000248553.6 |
HSPB1 |
heat shock 27kDa protein 1 |
| chr15_+_96869165 | 1.06 |
ENST00000421109.2 |
NR2F2 |
nuclear receptor subfamily 2, group F, member 2 |
| chr11_+_75273101 | 0.99 |
ENST00000533603.1 ENST00000358171.3 ENST00000526242.1 |
SERPINH1 |
serpin peptidase inhibitor, clade H (heat shock protein 47), member 1, (collagen binding protein 1) |
| chr11_+_20385327 | 0.98 |
ENST00000451739.2 ENST00000532505.1 |
HTATIP2 |
HIV-1 Tat interactive protein 2, 30kDa |
| chr6_-_106773291 | 0.97 |
ENST00000343245.3 |
ATG5 |
autophagy related 5 |
| chr11_+_20385231 | 0.95 |
ENST00000530266.1 ENST00000421577.2 ENST00000443524.2 ENST00000419348.2 |
HTATIP2 |
HIV-1 Tat interactive protein 2, 30kDa |
| chr11_+_111782934 | 0.93 |
ENST00000304298.3 |
HSPB2 |
Homo sapiens heat shock 27kDa protein 2 (HSPB2), mRNA. |
| chr11_+_75273246 | 0.92 |
ENST00000526397.1 ENST00000529643.1 ENST00000525492.1 ENST00000530284.1 ENST00000532356.1 ENST00000524558.1 |
SERPINH1 |
serpin peptidase inhibitor, clade H (heat shock protein 47), member 1, (collagen binding protein 1) |
| chr2_+_85766280 | 0.92 |
ENST00000306434.3 |
MAT2A |
methionine adenosyltransferase II, alpha |
| chr19_+_859425 | 0.88 |
ENST00000327726.6 |
CFD |
complement factor D (adipsin) |
| chr19_+_859654 | 0.88 |
ENST00000592860.1 |
CFD |
complement factor D (adipsin) |
| chr13_-_31736027 | 0.82 |
ENST00000380406.5 ENST00000320027.5 ENST00000380405.4 |
HSPH1 |
heat shock 105kDa/110kDa protein 1 |
| chr13_-_31736132 | 0.82 |
ENST00000429785.2 |
HSPH1 |
heat shock 105kDa/110kDa protein 1 |
| chr9_+_33025209 | 0.82 |
ENST00000330899.4 ENST00000544625.1 |
DNAJA1 |
DnaJ (Hsp40) homolog, subfamily A, member 1 |
| chr3_-_185655795 | 0.81 |
ENST00000342294.4 ENST00000382191.4 ENST00000453386.2 |
TRA2B |
transformer 2 beta homolog (Drosophila) |
| chr17_-_39928106 | 0.80 |
ENST00000540235.1 |
JUP |
junction plakoglobin |
| chr1_-_6453426 | 0.75 |
ENST00000545482.1 |
ACOT7 |
acyl-CoA thioesterase 7 |
| chr13_-_31736478 | 0.67 |
ENST00000445273.2 |
HSPH1 |
heat shock 105kDa/110kDa protein 1 |
| chr3_-_195603566 | 0.66 |
ENST00000424563.1 ENST00000411741.1 |
TNK2 |
tyrosine kinase, non-receptor, 2 |
| chr1_+_154947148 | 0.65 |
ENST00000368436.1 ENST00000308987.5 |
CKS1B |
CDC28 protein kinase regulatory subunit 1B |
| chr20_+_4666882 | 0.64 |
ENST00000379440.4 ENST00000430350.2 |
PRNP |
prion protein |
| chr1_+_154947126 | 0.63 |
ENST00000368439.1 |
CKS1B |
CDC28 protein kinase regulatory subunit 1B |
| chr15_+_79165222 | 0.62 |
ENST00000559930.1 |
MORF4L1 |
mortality factor 4 like 1 |
| chr15_+_85359911 | 0.60 |
ENST00000258888.5 |
ALPK3 |
alpha-kinase 3 |
| chr1_-_6453399 | 0.60 |
ENST00000608083.1 |
ACOT7 |
acyl-CoA thioesterase 7 |
| chr9_-_6015607 | 0.58 |
ENST00000259569.5 |
RANBP6 |
RAN binding protein 6 |
| chr3_-_183735731 | 0.58 |
ENST00000334444.6 |
ABCC5 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 5 |
| chr21_-_40720974 | 0.57 |
ENST00000380748.1 |
HMGN1 |
high mobility group nucleosome binding domain 1 |
| chr11_+_66512089 | 0.55 |
ENST00000524551.1 ENST00000525908.1 ENST00000360962.4 ENST00000346672.4 ENST00000527634.1 ENST00000540737.1 |
C11orf80 |
chromosome 11 open reading frame 80 |
| chr11_+_66512303 | 0.54 |
ENST00000532565.2 |
C11orf80 |
chromosome 11 open reading frame 80 |
| chr19_-_14629224 | 0.53 |
ENST00000254322.2 |
DNAJB1 |
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr19_-_14628645 | 0.51 |
ENST00000598235.1 |
DNAJB1 |
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr12_-_108154925 | 0.48 |
ENST00000228437.5 |
PRDM4 |
PR domain containing 4 |
| chr9_-_13279406 | 0.48 |
ENST00000546205.1 |
MPDZ |
multiple PDZ domain protein |
| chr4_-_25865159 | 0.48 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr7_+_56119323 | 0.45 |
ENST00000275603.4 ENST00000335503.3 ENST00000540286.1 |
CCT6A |
chaperonin containing TCP1, subunit 6A (zeta 1) |
| chr5_+_10250328 | 0.44 |
ENST00000515390.1 |
CCT5 |
chaperonin containing TCP1, subunit 5 (epsilon) |
| chr4_-_159644507 | 0.44 |
ENST00000307720.3 |
PPID |
peptidylprolyl isomerase D |
| chr11_+_20385666 | 0.43 |
ENST00000532081.1 ENST00000531058.1 |
HTATIP2 |
HIV-1 Tat interactive protein 2, 30kDa |
| chr17_-_62308087 | 0.42 |
ENST00000583097.1 |
TEX2 |
testis expressed 2 |
| chr21_-_40720995 | 0.42 |
ENST00000380749.5 |
HMGN1 |
high mobility group nucleosome binding domain 1 |
| chr11_-_82997013 | 0.42 |
ENST00000529073.1 ENST00000529611.1 |
CCDC90B |
coiled-coil domain containing 90B |
| chr1_-_26232522 | 0.40 |
ENST00000399728.1 |
STMN1 |
stathmin 1 |
| chr19_+_2269485 | 0.39 |
ENST00000582888.4 ENST00000602676.2 ENST00000322297.4 ENST00000583542.4 |
OAZ1 |
ornithine decarboxylase antizyme 1 |
| chr7_-_26240357 | 0.39 |
ENST00000354667.4 ENST00000356674.7 |
HNRNPA2B1 |
heterogeneous nuclear ribonucleoprotein A2/B1 |
| chr1_+_27248203 | 0.37 |
ENST00000321265.5 |
NUDC |
nudC nuclear distribution protein |
| chr11_-_82997371 | 0.37 |
ENST00000525503.1 |
CCDC90B |
coiled-coil domain containing 90B |
| chr11_-_82997420 | 0.36 |
ENST00000455220.2 ENST00000529689.1 |
CCDC90B |
coiled-coil domain containing 90B |
| chr17_+_1936687 | 0.36 |
ENST00000570477.1 |
DPH1 |
diphthamide biosynthesis 1 |
| chrX_-_102943022 | 0.36 |
ENST00000433176.2 |
MORF4L2 |
mortality factor 4 like 2 |
| chrX_-_102942961 | 0.35 |
ENST00000434230.1 ENST00000418819.1 ENST00000360458.1 |
MORF4L2 |
mortality factor 4 like 2 |
| chrX_-_152989531 | 0.33 |
ENST00000458587.2 ENST00000416815.1 |
BCAP31 |
B-cell receptor-associated protein 31 |
| chr3_+_101443476 | 0.32 |
ENST00000327230.4 ENST00000494050.1 |
CEP97 |
centrosomal protein 97kDa |
| chrX_-_152989798 | 0.32 |
ENST00000441714.1 ENST00000442093.1 ENST00000429550.1 ENST00000345046.6 |
BCAP31 |
B-cell receptor-associated protein 31 |
| chr1_-_242162375 | 0.31 |
ENST00000357246.3 |
MAP1LC3C |
microtubule-associated protein 1 light chain 3 gamma |
| chr1_-_45987526 | 0.29 |
ENST00000372079.1 ENST00000262746.1 ENST00000447184.1 ENST00000319248.8 |
PRDX1 |
peroxiredoxin 1 |
| chr21_+_44394620 | 0.29 |
ENST00000291547.5 |
PKNOX1 |
PBX/knotted 1 homeobox 1 |
| chr1_-_228613026 | 0.29 |
ENST00000366696.1 |
HIST3H3 |
histone cluster 3, H3 |
| chr11_-_47870091 | 0.28 |
ENST00000526870.1 |
NUP160 |
nucleoporin 160kDa |
| chr11_+_45825896 | 0.27 |
ENST00000314134.3 |
SLC35C1 |
solute carrier family 35 (GDP-fucose transporter), member C1 |
| chr12_-_110883346 | 0.27 |
ENST00000547365.1 |
ARPC3 |
actin related protein 2/3 complex, subunit 3, 21kDa |
| chr14_-_61124977 | 0.27 |
ENST00000554986.1 |
SIX1 |
SIX homeobox 1 |
| chr1_+_144811744 | 0.27 |
ENST00000338347.4 ENST00000440491.2 ENST00000375552.4 |
NBPF9 |
neuroblastoma breakpoint family, member 9 |
| chr10_-_62332357 | 0.27 |
ENST00000503366.1 |
ANK3 |
ankyrin 3, node of Ranvier (ankyrin G) |
| chr14_-_102553371 | 0.25 |
ENST00000553585.1 ENST00000216281.8 |
HSP90AA1 |
heat shock protein 90kDa alpha (cytosolic), class A member 1 |
| chr8_+_143915663 | 0.25 |
ENST00000522728.1 |
GML |
glycosylphosphatidylinositol anchored molecule like |
| chr15_+_44092784 | 0.25 |
ENST00000458412.1 |
HYPK |
huntingtin interacting protein K |
| chr14_-_68141535 | 0.25 |
ENST00000554659.1 |
VTI1B |
vesicle transport through interaction with t-SNAREs 1B |
| chr7_+_26240776 | 0.25 |
ENST00000337620.4 |
CBX3 |
chromobox homolog 3 |
| chr11_-_134123142 | 0.23 |
ENST00000392595.2 ENST00000341541.3 ENST00000352327.5 ENST00000392594.3 |
THYN1 |
thymocyte nuclear protein 1 |
| chr19_+_751122 | 0.23 |
ENST00000215582.6 |
MISP |
mitotic spindle positioning |
| chr4_+_41258786 | 0.22 |
ENST00000503431.1 ENST00000284440.4 ENST00000508768.1 ENST00000512788.1 |
UCHL1 |
ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) |
| chr19_-_17958832 | 0.22 |
ENST00000458235.1 |
JAK3 |
Janus kinase 3 |
| chr3_-_126327398 | 0.21 |
ENST00000383572.2 |
TXNRD3NB |
thioredoxin reductase 3 neighbor |
| chr21_+_44394742 | 0.21 |
ENST00000432907.2 |
PKNOX1 |
PBX/knotted 1 homeobox 1 |
| chr6_+_4890226 | 0.20 |
ENST00000343762.5 |
CDYL |
chromodomain protein, Y-like |
| chr11_+_65770227 | 0.20 |
ENST00000527348.1 |
BANF1 |
barrier to autointegration factor 1 |
| chr12_-_125401885 | 0.20 |
ENST00000542416.1 |
UBC |
ubiquitin C |
| chr20_-_60573188 | 0.20 |
ENST00000474089.1 |
TAF4 |
TAF4 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 135kDa |
| chr1_+_196743912 | 0.19 |
ENST00000367425.4 |
CFHR3 |
complement factor H-related 3 |
| chr5_+_446253 | 0.19 |
ENST00000315013.5 |
EXOC3 |
exocyst complex component 3 |
| chr19_+_41509851 | 0.19 |
ENST00000593831.1 ENST00000330446.5 |
CYP2B6 |
cytochrome P450, family 2, subfamily B, polypeptide 6 |
| chr16_+_28875126 | 0.18 |
ENST00000359285.5 ENST00000538342.1 |
SH2B1 |
SH2B adaptor protein 1 |
| chr2_-_62115659 | 0.18 |
ENST00000544185.1 |
CCT4 |
chaperonin containing TCP1, subunit 4 (delta) |
| chr10_-_115423792 | 0.18 |
ENST00000369360.3 ENST00000360478.3 ENST00000359988.3 ENST00000369358.4 |
NRAP |
nebulin-related anchoring protein |
| chr11_-_89956227 | 0.18 |
ENST00000457199.2 ENST00000530765.1 |
CHORDC1 |
cysteine and histidine-rich domain (CHORD) containing 1 |
| chrX_+_54947229 | 0.17 |
ENST00000442098.1 ENST00000430420.1 ENST00000453081.1 ENST00000173898.7 ENST00000319167.8 ENST00000375022.4 ENST00000399736.1 ENST00000440072.1 ENST00000420798.2 ENST00000431115.1 ENST00000440759.1 ENST00000375041.2 |
TRO |
trophinin |
| chr13_-_31191642 | 0.17 |
ENST00000405805.1 |
HMGB1 |
high mobility group box 1 |
| chr9_+_123970052 | 0.17 |
ENST00000373823.3 |
GSN |
gelsolin |
| chr7_-_139763521 | 0.17 |
ENST00000263549.3 |
PARP12 |
poly (ADP-ribose) polymerase family, member 12 |
| chr1_-_156308018 | 0.16 |
ENST00000496684.2 ENST00000368259.2 ENST00000368261.3 ENST00000472765.2 ENST00000533194.1 ENST00000478640.2 |
CCT3 |
chaperonin containing TCP1, subunit 3 (gamma) |
| chr11_-_108408895 | 0.16 |
ENST00000443411.1 ENST00000533052.1 |
EXPH5 |
exophilin 5 |
| chr4_+_5526883 | 0.16 |
ENST00000195455.2 |
C4orf6 |
chromosome 4 open reading frame 6 |
| chr11_-_89956461 | 0.16 |
ENST00000320585.6 |
CHORDC1 |
cysteine and histidine-rich domain (CHORD) containing 1 |
| chr12_+_103981044 | 0.15 |
ENST00000388887.2 |
STAB2 |
stabilin 2 |
| chr14_+_96722539 | 0.15 |
ENST00000553356.1 |
BDKRB1 |
bradykinin receptor B1 |
| chr2_-_62115725 | 0.15 |
ENST00000538252.1 ENST00000544079.1 ENST00000394440.3 |
CCT4 |
chaperonin containing TCP1, subunit 4 (delta) |
| chr12_+_2904102 | 0.15 |
ENST00000001008.4 |
FKBP4 |
FK506 binding protein 4, 59kDa |
| chr15_+_79165112 | 0.15 |
ENST00000426013.2 |
MORF4L1 |
mortality factor 4 like 1 |
| chr11_+_65769946 | 0.14 |
ENST00000533166.1 |
BANF1 |
barrier to autointegration factor 1 |
| chr20_-_23731893 | 0.14 |
ENST00000398402.1 |
CST1 |
cystatin SN |
| chr1_+_196857144 | 0.14 |
ENST00000367416.2 ENST00000251424.4 ENST00000367418.2 |
CFHR4 |
complement factor H-related 4 |
| chr8_+_140943416 | 0.14 |
ENST00000507535.3 |
C8orf17 |
chromosome 8 open reading frame 17 |
| chr18_+_11751493 | 0.13 |
ENST00000269162.5 |
GNAL |
guanine nucleotide binding protein (G protein), alpha activating activity polypeptide, olfactory type |
| chr1_+_12227035 | 0.13 |
ENST00000376259.3 ENST00000536782.1 |
TNFRSF1B |
tumor necrosis factor receptor superfamily, member 1B |
| chr1_-_146054494 | 0.12 |
ENST00000401009.2 |
NBPF11 |
neuroblastoma breakpoint family, member 11 |
| chr15_+_79165151 | 0.12 |
ENST00000331268.5 |
MORF4L1 |
mortality factor 4 like 1 |
| chrX_-_106243451 | 0.12 |
ENST00000355610.4 ENST00000535534.1 |
MORC4 |
MORC family CW-type zinc finger 4 |
| chr8_-_89339705 | 0.12 |
ENST00000286614.6 |
MMP16 |
matrix metallopeptidase 16 (membrane-inserted) |
| chr1_-_53704157 | 0.10 |
ENST00000371466.4 ENST00000371470.3 |
MAGOH |
mago-nashi homolog, proliferation-associated (Drosophila) |
| chr9_-_137809718 | 0.10 |
ENST00000371806.3 |
FCN1 |
ficolin (collagen/fibrinogen domain containing) 1 |
| chr21_-_30445886 | 0.10 |
ENST00000431234.1 ENST00000540844.1 ENST00000286788.4 |
CCT8 |
chaperonin containing TCP1, subunit 8 (theta) |
| chr2_-_100759037 | 0.10 |
ENST00000317233.4 ENST00000423966.1 ENST00000416492.1 |
AFF3 |
AF4/FMR2 family, member 3 |
| chr11_+_65769550 | 0.09 |
ENST00000312175.2 ENST00000445560.2 ENST00000530204.1 |
BANF1 |
barrier to autointegration factor 1 |
| chr15_-_86338134 | 0.09 |
ENST00000337975.5 |
KLHL25 |
kelch-like family member 25 |
| chr19_+_6135646 | 0.08 |
ENST00000588304.1 ENST00000588485.1 ENST00000588722.1 ENST00000591403.1 ENST00000586696.1 ENST00000589401.1 ENST00000252669.5 |
ACSBG2 |
acyl-CoA synthetase bubblegum family member 2 |
| chr22_-_32334403 | 0.08 |
ENST00000543051.1 |
C22orf24 |
chromosome 22 open reading frame 24 |
| chr20_-_23807358 | 0.08 |
ENST00000304725.2 |
CST2 |
cystatin SA |
| chr2_-_69664586 | 0.08 |
ENST00000303698.3 ENST00000394305.1 ENST00000410022.2 |
NFU1 |
NFU1 iron-sulfur cluster scaffold homolog (S. cerevisiae) |
| chr11_+_65154070 | 0.07 |
ENST00000317568.5 ENST00000531296.1 ENST00000533782.1 ENST00000355991.5 ENST00000416776.2 ENST00000526201.1 |
FRMD8 |
FERM domain containing 8 |
| chr2_-_208994548 | 0.07 |
ENST00000282141.3 |
CRYGC |
crystallin, gamma C |
| chr9_+_34989638 | 0.07 |
ENST00000453597.3 ENST00000335998.3 ENST00000312316.5 |
DNAJB5 |
DnaJ (Hsp40) homolog, subfamily B, member 5 |
| chr14_-_23540826 | 0.07 |
ENST00000357481.2 |
ACIN1 |
apoptotic chromatin condensation inducer 1 |
| chr20_-_43589109 | 0.07 |
ENST00000372813.3 |
TOMM34 |
translocase of outer mitochondrial membrane 34 |
| chr19_-_17958771 | 0.06 |
ENST00000534444.1 |
JAK3 |
Janus kinase 3 |
| chr5_+_140602904 | 0.06 |
ENST00000515856.2 ENST00000239449.4 |
PCDHB14 |
protocadherin beta 14 |
| chrX_-_154563889 | 0.06 |
ENST00000369449.2 ENST00000321926.4 |
CLIC2 |
chloride intracellular channel 2 |
| chr12_-_4754339 | 0.05 |
ENST00000228850.1 |
AKAP3 |
A kinase (PRKA) anchor protein 3 |
| chr4_+_5527117 | 0.05 |
ENST00000505296.1 |
C4orf6 |
chromosome 4 open reading frame 6 |
| chr9_-_86593238 | 0.05 |
ENST00000351839.3 |
HNRNPK |
heterogeneous nuclear ribonucleoprotein K |
| chr7_+_130020180 | 0.05 |
ENST00000481342.1 ENST00000011292.3 ENST00000604896.1 |
CPA1 |
carboxypeptidase A1 (pancreatic) |
| chr5_+_133706865 | 0.04 |
ENST00000265339.2 |
UBE2B |
ubiquitin-conjugating enzyme E2B |
| chr14_-_23540747 | 0.04 |
ENST00000555566.1 ENST00000338631.6 ENST00000557515.1 ENST00000397341.3 |
ACIN1 |
apoptotic chromatin condensation inducer 1 |
| chr15_-_86338100 | 0.04 |
ENST00000536947.1 |
KLHL25 |
kelch-like family member 25 |
| chr5_+_133707252 | 0.04 |
ENST00000506787.1 ENST00000507277.1 |
UBE2B |
ubiquitin-conjugating enzyme E2B |
| chr7_+_130020932 | 0.03 |
ENST00000484324.1 |
CPA1 |
carboxypeptidase A1 (pancreatic) |
| chr6_-_32339649 | 0.03 |
ENST00000527965.1 ENST00000532023.1 ENST00000447241.2 ENST00000375007.4 ENST00000534588.1 |
C6orf10 |
chromosome 6 open reading frame 10 |
| chr6_-_32339671 | 0.03 |
ENST00000442822.2 ENST00000375015.4 ENST00000533191.1 |
C6orf10 |
chromosome 6 open reading frame 10 |
| chr5_+_172571445 | 0.02 |
ENST00000231668.9 ENST00000351486.5 ENST00000352523.6 ENST00000393770.4 |
BNIP1 |
BCL2/adenovirus E1B 19kDa interacting protein 1 |
| chr15_-_89755034 | 0.02 |
ENST00000563254.1 |
RLBP1 |
retinaldehyde binding protein 1 |
| chr20_-_23731569 | 0.01 |
ENST00000304749.2 |
CST1 |
cystatin SN |
| chr2_+_210444142 | 0.01 |
ENST00000360351.4 ENST00000361559.4 |
MAP2 |
microtubule-associated protein 2 |
| chr5_-_140053152 | 0.01 |
ENST00000542735.1 |
DND1 |
DND microRNA-mediated repression inhibitor 1 |
| chr20_-_23669590 | 0.01 |
ENST00000217423.3 |
CST4 |
cystatin S |
| chr4_+_76932326 | 0.00 |
ENST00000513353.1 ENST00000341029.5 |
ART3 |
ADP-ribosyltransferase 3 |
| chrX_+_150345054 | 0.00 |
ENST00000218316.3 |
GPR50 |
G protein-coupled receptor 50 |
Gene Ontology Analysis
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 7.0 | GO:0098575 | lumenal side of lysosomal membrane(GO:0098575) |
| 0.8 | 12.7 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.3 | 0.9 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.2 | 1.0 | GO:0034274 | Atg12-Atg5-Atg16 complex(GO:0034274) |
| 0.1 | 0.8 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.1 | 0.7 | GO:0070436 | Grb2-EGFR complex(GO:0070436) |
| 0.1 | 0.7 | GO:0005784 | Sec61 translocon complex(GO:0005784) translocon complex(GO:0071256) |
| 0.1 | 6.4 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.1 | 0.8 | GO:0071664 | catenin-TCF7L2 complex(GO:0071664) |
| 0.1 | 1.0 | GO:0002199 | zona pellucida receptor complex(GO:0002199) |
| 0.1 | 2.2 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.1 | 0.5 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 1.6 | GO:0043189 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.0 | 0.6 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.0 | 0.2 | GO:0005927 | muscle tendon junction(GO:0005927) |
| 0.0 | 1.4 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.0 | 0.5 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.0 | 0.3 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.0 | 0.2 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.1 | GO:0043196 | varicosity(GO:0043196) |
| 0.0 | 0.1 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.0 | 0.3 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.0 | 0.1 | GO:0033503 | HULC complex(GO:0033503) |
| 0.0 | 0.3 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.0 | 0.2 | GO:0030478 | actin cap(GO:0030478) |
| 0.0 | 1.4 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 1.8 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 0.1 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.0 | 0.2 | GO:0033276 | transcription factor TFTC complex(GO:0033276) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 4.6 | GO:0004647 | phosphoserine phosphatase activity(GO:0004647) |
| 0.7 | 7.8 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.4 | 12.4 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.4 | 1.1 | GO:0008426 | protein kinase C inhibitor activity(GO:0008426) |
| 0.3 | 3.1 | GO:0001758 | retinal dehydrogenase activity(GO:0001758) |
| 0.2 | 2.3 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.2 | 1.3 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.2 | 1.0 | GO:0019776 | Atg8 ligase activity(GO:0019776) |
| 0.1 | 1.4 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.1 | 0.5 | GO:0071987 | WD40-repeat domain binding(GO:0071987) |
| 0.1 | 0.6 | GO:1903135 | cupric ion binding(GO:1903135) |
| 0.1 | 5.1 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.1 | 0.3 | GO:0005457 | GDP-fucose transmembrane transporter activity(GO:0005457) purine nucleotide-sugar transmembrane transporter activity(GO:0036080) |
| 0.1 | 0.8 | GO:0086083 | cell adhesive protein binding involved in bundle of His cell-Purkinje myocyte communication(GO:0086083) |
| 0.1 | 4.7 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.1 | 0.4 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.1 | 0.2 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.1 | 0.2 | GO:0004947 | bradykinin receptor activity(GO:0004947) |
| 0.0 | 0.1 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 0.0 | 0.6 | GO:0043225 | anion transmembrane-transporting ATPase activity(GO:0043225) |
| 0.0 | 0.4 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.0 | 0.4 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 1.0 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.0 | 1.3 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.0 | 0.3 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 3.3 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.2 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.0 | 0.7 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.0 | 0.2 | GO:0010858 | calcium-dependent protein kinase regulator activity(GO:0010858) |
| 0.0 | 1.3 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 1.1 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 0.8 | GO:0070717 | poly-purine tract binding(GO:0070717) |
| 0.0 | 0.7 | GO:0042288 | MHC class I protein binding(GO:0042288) |
| 0.0 | 0.6 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 0.6 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 0.2 | GO:0017162 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) aryl hydrocarbon receptor binding(GO:0017162) |
| 0.0 | 0.2 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.0 | 0.3 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.0 | 1.6 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 1.0 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.0 | 0.2 | GO:0008391 | arachidonic acid monooxygenase activity(GO:0008391) arachidonic acid epoxygenase activity(GO:0008392) |
| 0.0 | 0.6 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 7.0 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.2 | 4.6 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 6.4 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.4 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.0 | 1.5 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.4 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.0 | 3.9 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 1.6 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.8 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.9 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.0 | 1.0 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.6 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.6 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.3 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 1.1 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.0 | 1.1 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.3 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.9 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 0.2 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 7.0 | GO:1904764 | negative regulation of fibril organization(GO:1902904) chaperone-mediated autophagy translocation complex disassembly(GO:1904764) |
| 0.7 | 12.7 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.4 | 3.1 | GO:1903748 | negative regulation of establishment of protein localization to mitochondrion(GO:1903748) |
| 0.4 | 4.6 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.3 | 3.1 | GO:0035799 | ureter maturation(GO:0035799) |
| 0.3 | 1.4 | GO:1900535 | medium-chain fatty-acyl-CoA catabolic process(GO:0036114) long-chain fatty-acyl-CoA catabolic process(GO:0036116) palmitic acid metabolic process(GO:1900533) palmitic acid biosynthetic process(GO:1900535) |
| 0.3 | 1.0 | GO:0039689 | negative stranded viral RNA replication(GO:0039689) multi-organism biosynthetic process(GO:0044034) |
| 0.3 | 1.1 | GO:0099640 | axo-dendritic protein transport(GO:0099640) |
| 0.3 | 1.1 | GO:0009956 | radial pattern formation(GO:0009956) |
| 0.2 | 1.9 | GO:0003433 | chondrocyte development involved in endochondral bone morphogenesis(GO:0003433) |
| 0.2 | 0.9 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.2 | 1.3 | GO:0060715 | syncytiotrophoblast cell differentiation involved in labyrinthine layer development(GO:0060715) |
| 0.2 | 0.6 | GO:1902938 | regulation of intracellular calcium activated chloride channel activity(GO:1902938) |
| 0.2 | 0.8 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.2 | 0.8 | GO:0002159 | desmosome assembly(GO:0002159) endothelial cell-cell adhesion(GO:0071603) |
| 0.1 | 6.4 | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045737) |
| 0.1 | 1.5 | GO:1904869 | regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) |
| 0.1 | 0.7 | GO:1904154 | positive regulation of retrograde protein transport, ER to cytosol(GO:1904154) |
| 0.1 | 0.3 | GO:2000298 | regulation of Rho-dependent protein serine/threonine kinase activity(GO:2000298) |
| 0.1 | 0.4 | GO:0070141 | response to UV-A(GO:0070141) cellular response to UV-A(GO:0071492) |
| 0.1 | 0.3 | GO:0045210 | negative regulation of dendritic cell cytokine production(GO:0002731) FasL biosynthetic process(GO:0045210) |
| 0.1 | 0.3 | GO:0015783 | GDP-fucose transport(GO:0015783) purine nucleotide-sugar transport(GO:0036079) |
| 0.1 | 0.3 | GO:0071698 | olfactory placode formation(GO:0030910) myotome development(GO:0061055) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) positive regulation of mesenchymal cell proliferation involved in ureter development(GO:2000729) |
| 0.1 | 0.4 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) |
| 0.1 | 0.4 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.1 | 0.4 | GO:0044806 | G-quadruplex DNA unwinding(GO:0044806) |
| 0.1 | 0.2 | GO:0007412 | axon target recognition(GO:0007412) |
| 0.1 | 1.0 | GO:0090084 | negative regulation of inclusion body assembly(GO:0090084) |
| 0.1 | 0.5 | GO:0043985 | histone H4-R3 methylation(GO:0043985) |
| 0.1 | 0.3 | GO:1900827 | positive regulation of cell communication by electrical coupling(GO:0010650) maintenance of protein location in membrane(GO:0072658) maintenance of protein location in plasma membrane(GO:0072660) positive regulation of membrane depolarization during cardiac muscle cell action potential(GO:1900827) |
| 0.1 | 0.3 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.1 | 0.6 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.1 | 1.8 | GO:0071711 | basement membrane organization(GO:0071711) |
| 0.1 | 1.6 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.0 | 0.4 | GO:0017183 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.0 | 0.2 | GO:0032072 | plasmacytoid dendritic cell activation(GO:0002270) T cell mediated immune response to tumor cell(GO:0002424) regulation of T cell mediated immune response to tumor cell(GO:0002840) regulation of restriction endodeoxyribonuclease activity(GO:0032072) negative regulation of apoptotic cell clearance(GO:2000426) |
| 0.0 | 0.6 | GO:0006610 | ribosomal protein import into nucleus(GO:0006610) |
| 0.0 | 0.4 | GO:0015074 | DNA integration(GO:0015074) |
| 0.0 | 0.2 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 0.1 | GO:0002752 | cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 0.0 | 1.0 | GO:0032786 | positive regulation of DNA-templated transcription, elongation(GO:0032786) |
| 0.0 | 0.2 | GO:0044858 | plasma membrane raft distribution(GO:0044855) plasma membrane raft localization(GO:0044856) plasma membrane raft polarization(GO:0044858) regulation of plasma membrane raft polarization(GO:1903906) |
| 0.0 | 0.1 | GO:0010845 | positive regulation of reciprocal meiotic recombination(GO:0010845) |
| 0.0 | 0.3 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) negative regulation of cilium assembly(GO:1902018) |
| 0.0 | 0.7 | GO:2000369 | regulation of clathrin-mediated endocytosis(GO:2000369) |
| 0.0 | 0.3 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 3.5 | GO:0006997 | nucleus organization(GO:0006997) |
| 0.0 | 0.2 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.0 | 0.1 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.0 | 0.2 | GO:0042738 | exogenous drug catabolic process(GO:0042738) |
| 0.0 | 0.0 | GO:0072369 | regulation of lipid transport by positive regulation of transcription from RNA polymerase II promoter(GO:0072369) |
| 0.0 | 0.1 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.0 | 0.2 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.0 | 0.3 | GO:0051290 | protein heterotetramerization(GO:0051290) |
| 0.0 | 0.4 | GO:0006986 | response to unfolded protein(GO:0006986) |
| 0.0 | 0.1 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.0 | 0.2 | GO:0010992 | ubiquitin homeostasis(GO:0010992) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 5.1 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.1 | 8.4 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 1.8 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 2.0 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 0.4 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 1.1 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
| 0.0 | 1.1 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 0.9 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.0 | 1.1 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 1.7 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
| 0.0 | 0.6 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 0.4 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 0.4 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |