Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
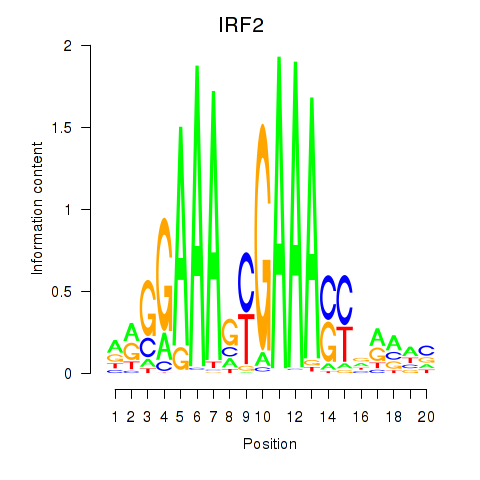
Results for IRF2_STAT2_IRF8_IRF1
Z-value: 4.88
Transcription factors associated with IRF2_STAT2_IRF8_IRF1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
IRF2
|
ENSG00000168310.6 | IRF2 |
|
STAT2
|
ENSG00000170581.9 | STAT2 |
|
IRF8
|
ENSG00000140968.6 | IRF8 |
|
IRF1
|
ENSG00000125347.9 | IRF1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| IRF8 | hg19_v2_chr16_+_85942594_85942635 | 0.85 | 3.3e-05 | Click! |
| IRF2 | hg19_v2_chr4_-_185395672_185395734 | 0.84 | 5.3e-05 | Click! |
| IRF1 | hg19_v2_chr5_-_131826457_131826514 | 0.83 | 7.0e-05 | Click! |
| STAT2 | hg19_v2_chr12_-_56753858_56753930 | -0.12 | 6.6e-01 | Click! |
Activity profile of IRF2_STAT2_IRF8_IRF1 motif
Sorted Z-values of IRF2_STAT2_IRF8_IRF1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of IRF2_STAT2_IRF8_IRF1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_79086088 | 39.26 |
ENST00000370751.5 ENST00000342282.3 |
IFI44L |
interferon-induced protein 44-like |
| chr21_+_42792442 | 24.81 |
ENST00000398600.2 |
MX1 |
myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) |
| chr6_+_32821924 | 23.87 |
ENST00000374859.2 ENST00000453265.2 |
PSMB9 |
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr10_+_91152303 | 22.66 |
ENST00000371804.3 |
IFIT1 |
interferon-induced protein with tetratricopeptide repeats 1 |
| chr1_+_948803 | 22.47 |
ENST00000379389.4 |
ISG15 |
ISG15 ubiquitin-like modifier |
| chr11_-_615942 | 20.09 |
ENST00000397562.3 ENST00000330243.5 ENST00000397570.1 ENST00000397574.2 |
IRF7 |
interferon regulatory factor 7 |
| chr4_+_89378261 | 18.72 |
ENST00000264350.3 |
HERC5 |
HECT and RLD domain containing E3 ubiquitin protein ligase 5 |
| chr16_+_12058961 | 17.15 |
ENST00000053243.1 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr1_+_79115503 | 16.52 |
ENST00000370747.4 ENST00000438486.1 ENST00000545124.1 |
IFI44 |
interferon-induced protein 44 |
| chr16_+_12059050 | 16.12 |
ENST00000396495.3 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr1_+_158979792 | 15.96 |
ENST00000359709.3 ENST00000430894.2 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr21_+_42733870 | 15.48 |
ENST00000330714.3 ENST00000436410.1 ENST00000435611.1 |
MX2 |
myxovirus (influenza virus) resistance 2 (mouse) |
| chr1_+_158979680 | 15.47 |
ENST00000368131.4 ENST00000340979.6 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr6_+_6588902 | 15.37 |
ENST00000230568.4 |
LY86 |
lymphocyte antigen 86 |
| chr1_+_158979686 | 15.27 |
ENST00000368132.3 ENST00000295809.7 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr4_-_169239921 | 15.14 |
ENST00000514995.1 ENST00000393743.3 |
DDX60 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 60 |
| chr6_+_32811885 | 14.95 |
ENST00000458296.1 ENST00000413039.1 ENST00000429600.1 ENST00000412095.1 ENST00000415067.1 ENST00000395330.1 |
TAPSAR1 PSMB9 |
TAP1 and PSMB8 antisense RNA 1 proteasome (prosome, macropain) subunit, beta type, 9 |
| chr12_+_25205446 | 14.90 |
ENST00000557489.1 ENST00000354454.3 ENST00000536173.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr2_-_163175133 | 14.04 |
ENST00000421365.2 ENST00000263642.2 |
IFIH1 |
interferon induced with helicase C domain 1 |
| chr10_-_91174215 | 13.88 |
ENST00000371837.1 |
LIPA |
lipase A, lysosomal acid, cholesterol esterase |
| chr12_+_113416191 | 13.79 |
ENST00000342315.4 ENST00000392583.2 |
OAS2 |
2'-5'-oligoadenylate synthetase 2, 69/71kDa |
| chr1_+_174843548 | 13.48 |
ENST00000478442.1 ENST00000465412.1 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr11_-_615570 | 13.37 |
ENST00000525445.1 ENST00000348655.6 ENST00000397566.1 |
IRF7 |
interferon regulatory factor 7 |
| chr22_+_18632666 | 13.12 |
ENST00000215794.7 |
USP18 |
ubiquitin specific peptidase 18 |
| chr7_+_74188309 | 12.93 |
ENST00000289473.4 ENST00000433458.1 |
NCF1 |
neutrophil cytosolic factor 1 |
| chr11_-_57335280 | 12.88 |
ENST00000287156.4 |
UBE2L6 |
ubiquitin-conjugating enzyme E2L 6 |
| chr11_+_5710919 | 12.66 |
ENST00000379965.3 ENST00000425490.1 |
TRIM22 |
tripartite motif containing 22 |
| chr6_-_32811771 | 12.02 |
ENST00000395339.3 ENST00000374882.3 |
PSMB8 |
proteasome (prosome, macropain) subunit, beta type, 8 |
| chr1_-_27998689 | 11.99 |
ENST00000339145.4 ENST00000362020.4 ENST00000361157.6 |
IFI6 |
interferon, alpha-inducible protein 6 |
| chr17_+_6659153 | 11.59 |
ENST00000441631.1 ENST00000438512.1 ENST00000346752.4 ENST00000361842.3 |
XAF1 |
XIAP associated factor 1 |
| chr6_-_32821599 | 10.95 |
ENST00000354258.4 |
TAP1 |
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr10_+_91087651 | 10.75 |
ENST00000371818.4 |
IFIT3 |
interferon-induced protein with tetratricopeptide repeats 3 |
| chr17_-_40264692 | 10.54 |
ENST00000591220.1 ENST00000251642.3 |
DHX58 |
DEXH (Asp-Glu-X-His) box polypeptide 58 |
| chr2_-_231084820 | 10.45 |
ENST00000258382.5 ENST00000338556.3 |
SP110 |
SP110 nuclear body protein |
| chr16_+_12059091 | 10.45 |
ENST00000562385.1 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr21_+_42798094 | 10.35 |
ENST00000398598.3 ENST00000455164.2 ENST00000424365.1 |
MX1 |
myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) |
| chr1_-_89531041 | 10.20 |
ENST00000370473.4 |
GBP1 |
guanylate binding protein 1, interferon-inducible |
| chr2_-_191878874 | 10.03 |
ENST00000392322.3 ENST00000392323.2 ENST00000424722.1 ENST00000361099.3 |
STAT1 |
signal transducer and activator of transcription 1, 91kDa |
| chr10_+_91174314 | 9.81 |
ENST00000371795.4 |
IFIT5 |
interferon-induced protein with tetratricopeptide repeats 5 |
| chr1_+_158801095 | 9.76 |
ENST00000368141.4 |
MNDA |
myeloid cell nuclear differentiation antigen |
| chr14_-_93214915 | 9.62 |
ENST00000553918.1 ENST00000555699.1 ENST00000553802.1 ENST00000554397.1 ENST00000554919.1 ENST00000554080.1 ENST00000553371.1 |
LGMN |
legumain |
| chr16_-_67970990 | 9.52 |
ENST00000358514.4 |
PSMB10 |
proteasome (prosome, macropain) subunit, beta type, 10 |
| chr15_+_89182178 | 9.33 |
ENST00000559876.1 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chr2_-_191878681 | 9.17 |
ENST00000409465.1 |
STAT1 |
signal transducer and activator of transcription 1, 91kDa |
| chr12_-_49319265 | 8.98 |
ENST00000552878.1 ENST00000453172.2 |
FKBP11 |
FK506 binding protein 11, 19 kDa |
| chr15_+_89181974 | 8.69 |
ENST00000306072.5 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chr14_-_93214988 | 8.55 |
ENST00000557434.1 ENST00000393218.2 ENST00000334869.4 |
LGMN |
legumain |
| chr11_+_5646213 | 8.42 |
ENST00000429814.2 |
TRIM34 |
tripartite motif containing 34 |
| chr15_+_89182156 | 8.37 |
ENST00000379224.5 |
ISG20 |
interferon stimulated exonuclease gene 20kDa |
| chr4_-_76944621 | 8.21 |
ENST00000306602.1 |
CXCL10 |
chemokine (C-X-C motif) ligand 10 |
| chr15_-_80263506 | 8.14 |
ENST00000335661.6 |
BCL2A1 |
BCL2-related protein A1 |
| chrX_+_37639264 | 7.92 |
ENST00000378588.4 |
CYBB |
cytochrome b-245, beta polypeptide |
| chr6_-_31324943 | 7.80 |
ENST00000412585.2 ENST00000434333.1 |
HLA-B |
major histocompatibility complex, class I, B |
| chr2_-_231084659 | 7.79 |
ENST00000258381.6 ENST00000358662.4 ENST00000455674.1 ENST00000392048.3 |
SP110 |
SP110 nuclear body protein |
| chr2_-_231084617 | 7.79 |
ENST00000409815.2 |
SP110 |
SP110 nuclear body protein |
| chr12_-_49318715 | 7.52 |
ENST00000444214.2 |
FKBP11 |
FK506 binding protein 11, 19 kDa |
| chr11_-_104972158 | 7.42 |
ENST00000598974.1 ENST00000593315.1 ENST00000594519.1 ENST00000415981.2 ENST00000525374.1 ENST00000375707.1 |
CASP1 CARD16 CARD17 |
caspase 1, apoptosis-related cysteine peptidase caspase recruitment domain family, member 16 caspase recruitment domain family, member 17 |
| chr12_+_25205666 | 7.37 |
ENST00000547044.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr4_+_142557717 | 7.33 |
ENST00000320650.4 ENST00000296545.7 |
IL15 |
interleukin 15 |
| chr13_+_50070077 | 7.31 |
ENST00000378319.3 ENST00000426879.1 |
PHF11 |
PHD finger protein 11 |
| chr4_+_142557771 | 7.21 |
ENST00000514653.1 |
IL15 |
interleukin 15 |
| chr11_-_60719213 | 7.16 |
ENST00000227880.3 |
SLC15A3 |
solute carrier family 15 (oligopeptide transporter), member 3 |
| chr10_+_91092241 | 7.05 |
ENST00000371811.4 |
IFIT3 |
interferon-induced protein with tetratricopeptide repeats 3 |
| chr13_+_50070491 | 6.98 |
ENST00000496612.1 ENST00000357596.3 ENST00000485919.1 ENST00000442195.1 |
PHF11 |
PHD finger protein 11 |
| chr17_+_25958174 | 6.95 |
ENST00000313648.6 ENST00000577392.1 ENST00000584661.1 ENST00000413914.2 |
LGALS9 |
lectin, galactoside-binding, soluble, 9 |
| chr2_+_231280908 | 6.77 |
ENST00000427101.2 ENST00000432979.1 |
SP100 |
SP100 nuclear antigen |
| chr12_+_113344811 | 6.75 |
ENST00000551241.1 ENST00000553185.1 ENST00000550689.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr14_+_24630465 | 6.61 |
ENST00000557894.1 ENST00000559284.1 ENST00000560275.1 |
IRF9 |
interferon regulatory factor 9 |
| chr3_+_48507210 | 6.46 |
ENST00000433541.1 ENST00000422277.2 ENST00000436480.2 ENST00000444177.1 |
TREX1 |
three prime repair exonuclease 1 |
| chr12_+_113344582 | 6.43 |
ENST00000202917.5 ENST00000445409.2 ENST00000452357.2 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr12_+_94542459 | 6.41 |
ENST00000258526.4 |
PLXNC1 |
plexin C1 |
| chr4_+_142558078 | 6.40 |
ENST00000529613.1 |
IL15 |
interleukin 15 |
| chr19_+_10196781 | 6.39 |
ENST00000253110.11 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr12_+_25205568 | 6.38 |
ENST00000548766.1 ENST00000556887.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr2_+_201981527 | 6.34 |
ENST00000441224.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr1_+_64058939 | 6.28 |
ENST00000371084.3 |
PGM1 |
phosphoglucomutase 1 |
| chr19_+_1077393 | 6.21 |
ENST00000590577.1 |
HMHA1 |
histocompatibility (minor) HA-1 |
| chr10_-_90751038 | 6.20 |
ENST00000458159.1 ENST00000415557.1 ENST00000458208.1 |
ACTA2 |
actin, alpha 2, smooth muscle, aorta |
| chr22_+_39436862 | 6.14 |
ENST00000381565.2 ENST00000452957.2 |
APOBEC3F APOBEC3G |
apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3F apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3G |
| chr12_+_113344755 | 6.06 |
ENST00000550883.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr11_+_60223312 | 6.02 |
ENST00000532491.1 ENST00000532073.1 ENST00000534668.1 ENST00000528313.1 ENST00000533306.1 |
MS4A1 |
membrane-spanning 4-domains, subfamily A, member 1 |
| chr11_+_60223225 | 5.99 |
ENST00000524807.1 ENST00000345732.4 |
MS4A1 |
membrane-spanning 4-domains, subfamily A, member 1 |
| chr6_-_29527702 | 5.95 |
ENST00000377050.4 |
UBD |
ubiquitin D |
| chr6_+_26440700 | 5.91 |
ENST00000494393.1 ENST00000482451.1 ENST00000244519.2 ENST00000339789.4 ENST00000471353.1 ENST00000361232.3 ENST00000487627.1 ENST00000496719.1 ENST00000490254.1 ENST00000487272.1 |
BTN3A3 |
butyrophilin, subfamily 3, member A3 |
| chr3_-_58196939 | 5.87 |
ENST00000394549.2 ENST00000461914.3 |
DNASE1L3 |
deoxyribonuclease I-like 3 |
| chr1_-_111746966 | 5.85 |
ENST00000369752.5 |
DENND2D |
DENN/MADD domain containing 2D |
| chr3_+_187086120 | 5.77 |
ENST00000259030.2 |
RTP4 |
receptor (chemosensory) transporter protein 4 |
| chr12_+_113376157 | 5.74 |
ENST00000228928.7 |
OAS3 |
2'-5'-oligoadenylate synthetase 3, 100kDa |
| chr3_-_49851313 | 5.73 |
ENST00000333486.3 |
UBA7 |
ubiquitin-like modifier activating enzyme 7 |
| chr11_-_104905840 | 5.68 |
ENST00000526568.1 ENST00000393136.4 ENST00000531166.1 ENST00000534497.1 ENST00000527979.1 ENST00000446369.1 ENST00000353247.5 ENST00000528974.1 ENST00000533400.1 ENST00000525825.1 ENST00000436863.3 |
CASP1 |
caspase 1, apoptosis-related cysteine peptidase |
| chr12_-_53594227 | 5.58 |
ENST00000550743.2 |
ITGB7 |
integrin, beta 7 |
| chr8_-_79717750 | 5.56 |
ENST00000263851.4 ENST00000379113.2 |
IL7 |
interleukin 7 |
| chr6_+_12958137 | 5.40 |
ENST00000457702.2 ENST00000379345.2 |
PHACTR1 |
phosphatase and actin regulator 1 |
| chr10_+_91061712 | 5.40 |
ENST00000371826.3 |
IFIT2 |
interferon-induced protein with tetratricopeptide repeats 2 |
| chr5_-_149792295 | 5.39 |
ENST00000518797.1 ENST00000524315.1 ENST00000009530.7 ENST00000377795.3 |
CD74 |
CD74 molecule, major histocompatibility complex, class II invariant chain |
| chr1_-_151319710 | 5.36 |
ENST00000290524.4 ENST00000437327.1 ENST00000452513.2 ENST00000368870.2 ENST00000452671.2 |
RFX5 |
regulatory factor X, 5 (influences HLA class II expression) |
| chr4_+_37892682 | 5.26 |
ENST00000508802.1 ENST00000261439.4 ENST00000402522.1 |
TBC1D1 |
TBC1 (tre-2/USP6, BUB2, cdc16) domain family, member 1 |
| chr11_+_1874200 | 5.20 |
ENST00000311604.3 |
LSP1 |
lymphocyte-specific protein 1 |
| chr3_+_48507621 | 5.14 |
ENST00000456089.1 |
TREX1 |
three prime repair exonuclease 1 |
| chr17_+_18380051 | 5.10 |
ENST00000581545.1 ENST00000582333.1 ENST00000328114.6 ENST00000412421.2 ENST00000583322.1 ENST00000584941.1 |
LGALS9C |
lectin, galactoside-binding, soluble, 9C |
| chr5_-_96143602 | 5.03 |
ENST00000443439.2 ENST00000503921.1 ENST00000508227.1 ENST00000507154.1 |
ERAP1 |
endoplasmic reticulum aminopeptidase 1 |
| chr5_-_95158375 | 4.75 |
ENST00000512469.2 ENST00000379979.4 ENST00000505427.1 ENST00000508780.1 |
GLRX |
glutaredoxin (thioltransferase) |
| chr3_-_58200398 | 4.72 |
ENST00000318316.3 ENST00000460422.1 ENST00000483681.1 |
DNASE1L3 |
deoxyribonuclease I-like 3 |
| chrX_+_37639302 | 4.58 |
ENST00000545017.1 ENST00000536160.1 |
CYBB |
cytochrome b-245, beta polypeptide |
| chr2_+_231280954 | 4.54 |
ENST00000409824.1 ENST00000409341.1 ENST00000409112.1 ENST00000340126.4 ENST00000341950.4 |
SP100 |
SP100 nuclear antigen |
| chr6_-_33282163 | 4.45 |
ENST00000434618.2 ENST00000456592.2 |
TAPBP |
TAP binding protein (tapasin) |
| chr10_-_101690650 | 4.43 |
ENST00000543621.1 |
DNMBP |
dynamin binding protein |
| chr4_-_156875003 | 4.36 |
ENST00000433477.3 |
CTSO |
cathepsin O |
| chr12_+_6561190 | 4.36 |
ENST00000544021.1 ENST00000266556.7 |
TAPBPL |
TAP binding protein-like |
| chr2_+_201981119 | 4.35 |
ENST00000395148.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr2_-_214014959 | 4.31 |
ENST00000442445.1 ENST00000457361.1 ENST00000342002.2 |
IKZF2 |
IKAROS family zinc finger 2 (Helios) |
| chr19_+_10196981 | 4.27 |
ENST00000591813.1 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr2_-_242089677 | 4.24 |
ENST00000405260.1 |
PASK |
PAS domain containing serine/threonine kinase |
| chr8_+_28174649 | 4.20 |
ENST00000301908.3 |
PNOC |
prepronociceptin |
| chr3_-_58196688 | 4.18 |
ENST00000486455.1 |
DNASE1L3 |
deoxyribonuclease I-like 3 |
| chr5_-_95158644 | 4.16 |
ENST00000237858.6 |
GLRX |
glutaredoxin (thioltransferase) |
| chr2_+_33701286 | 4.15 |
ENST00000403687.3 |
RASGRP3 |
RAS guanyl releasing protein 3 (calcium and DAG-regulated) |
| chr20_-_43280361 | 4.08 |
ENST00000372874.4 |
ADA |
adenosine deaminase |
| chr1_-_154580616 | 4.06 |
ENST00000368474.4 |
ADAR |
adenosine deaminase, RNA-specific |
| chr6_-_32920794 | 4.04 |
ENST00000395305.3 ENST00000395303.3 ENST00000374843.4 ENST00000429234.1 |
HLA-DMA XXbac-BPG181M17.5 |
major histocompatibility complex, class II, DM alpha Uncharacterized protein |
| chr1_-_25291475 | 3.95 |
ENST00000338888.3 ENST00000399916.1 |
RUNX3 |
runt-related transcription factor 3 |
| chr19_-_7766991 | 3.91 |
ENST00000597921.1 ENST00000346664.5 |
FCER2 |
Fc fragment of IgE, low affinity II, receptor for (CD23) |
| chr2_-_175499294 | 3.91 |
ENST00000392547.2 |
WIPF1 |
WAS/WASL interacting protein family, member 1 |
| chr1_+_241695670 | 3.90 |
ENST00000366557.4 |
KMO |
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr19_+_18284477 | 3.90 |
ENST00000407280.3 |
IFI30 |
interferon, gamma-inducible protein 30 |
| chr1_+_241695424 | 3.88 |
ENST00000366558.3 ENST00000366559.4 |
KMO |
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr20_-_43280325 | 3.64 |
ENST00000537820.1 |
ADA |
adenosine deaminase |
| chr18_-_53303123 | 3.63 |
ENST00000569357.1 ENST00000565124.1 ENST00000398339.1 |
TCF4 |
transcription factor 4 |
| chr7_+_134832808 | 3.58 |
ENST00000275767.3 |
TMEM140 |
transmembrane protein 140 |
| chr5_-_111092930 | 3.45 |
ENST00000257435.7 |
NREP |
neuronal regeneration related protein |
| chr9_+_100174232 | 3.44 |
ENST00000355295.4 |
TDRD7 |
tudor domain containing 7 |
| chr6_-_30181133 | 3.44 |
ENST00000454678.2 ENST00000434785.1 |
TRIM26 |
tripartite motif containing 26 |
| chr17_-_33759509 | 3.43 |
ENST00000304905.5 |
SLFN12 |
schlafen family member 12 |
| chr9_-_21995300 | 3.43 |
ENST00000498628.2 |
CDKN2A |
cyclin-dependent kinase inhibitor 2A |
| chr6_-_30181156 | 3.39 |
ENST00000418026.1 ENST00000416596.1 ENST00000453195.1 |
TRIM26 |
tripartite motif containing 26 |
| chr2_+_102972363 | 3.35 |
ENST00000409599.1 |
IL18R1 |
interleukin 18 receptor 1 |
| chr22_-_36556821 | 3.35 |
ENST00000531095.1 ENST00000397293.2 ENST00000349314.2 |
APOL3 |
apolipoprotein L, 3 |
| chr2_+_163175394 | 3.33 |
ENST00000446271.1 ENST00000429691.2 |
GCA |
grancalcin, EF-hand calcium binding protein |
| chr9_-_21995249 | 3.33 |
ENST00000494262.1 |
CDKN2A |
cyclin-dependent kinase inhibitor 2A |
| chr6_+_26365443 | 3.28 |
ENST00000527422.1 ENST00000356386.2 ENST00000396934.3 ENST00000377708.2 ENST00000396948.1 ENST00000508906.2 |
BTN3A2 |
butyrophilin, subfamily 3, member A2 |
| chr2_+_201994208 | 3.19 |
ENST00000440180.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr14_+_24605361 | 3.19 |
ENST00000206451.6 ENST00000559123.1 |
PSME1 |
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr9_+_100174344 | 3.16 |
ENST00000422139.2 |
TDRD7 |
tudor domain containing 7 |
| chr5_-_111093167 | 3.15 |
ENST00000446294.2 ENST00000419114.2 |
NREP |
neuronal regeneration related protein |
| chr5_+_49962772 | 3.13 |
ENST00000281631.5 ENST00000513738.1 ENST00000503665.1 ENST00000514067.2 ENST00000503046.1 |
PARP8 |
poly (ADP-ribose) polymerase family, member 8 |
| chr10_+_115439699 | 3.11 |
ENST00000369315.1 |
CASP7 |
caspase 7, apoptosis-related cysteine peptidase |
| chr16_-_11350036 | 3.10 |
ENST00000332029.2 |
SOCS1 |
suppressor of cytokine signaling 1 |
| chr10_+_97515409 | 3.04 |
ENST00000371207.3 ENST00000543964.1 |
ENTPD1 |
ectonucleoside triphosphate diphosphohydrolase 1 |
| chr9_+_33265011 | 3.04 |
ENST00000419016.2 |
CHMP5 |
charged multivesicular body protein 5 |
| chr2_+_201980827 | 3.03 |
ENST00000309955.3 ENST00000443227.1 ENST00000341222.6 ENST00000355558.4 ENST00000340870.5 ENST00000341582.6 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr6_+_29691198 | 3.03 |
ENST00000440587.2 ENST00000434407.2 |
HLA-F |
major histocompatibility complex, class I, F |
| chr2_-_152146385 | 3.01 |
ENST00000414946.1 ENST00000243346.5 |
NMI |
N-myc (and STAT) interactor |
| chr13_-_33002279 | 2.98 |
ENST00000380130.2 |
N4BP2L1 |
NEDD4 binding protein 2-like 1 |
| chr10_+_115439282 | 2.96 |
ENST00000369321.2 ENST00000345633.4 |
CASP7 |
caspase 7, apoptosis-related cysteine peptidase |
| chr4_-_153303658 | 2.94 |
ENST00000296555.5 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chr9_+_112542572 | 2.93 |
ENST00000374530.3 |
PALM2-AKAP2 |
PALM2-AKAP2 readthrough |
| chr11_+_121461097 | 2.92 |
ENST00000527934.1 |
SORL1 |
sortilin-related receptor, L(DLR class) A repeats containing |
| chr11_+_313503 | 2.92 |
ENST00000528780.1 ENST00000328221.5 |
IFITM1 |
interferon induced transmembrane protein 1 |
| chr9_+_33264861 | 2.89 |
ENST00000223500.8 |
CHMP5 |
charged multivesicular body protein 5 |
| chr1_+_12227035 | 2.87 |
ENST00000376259.3 ENST00000536782.1 |
TNFRSF1B |
tumor necrosis factor receptor superfamily, member 1B |
| chr4_-_40517984 | 2.87 |
ENST00000381795.6 |
RBM47 |
RNA binding motif protein 47 |
| chr6_-_31239846 | 2.86 |
ENST00000415537.1 ENST00000376228.5 ENST00000383329.3 |
HLA-C |
major histocompatibility complex, class I, C |
| chr4_-_185395672 | 2.85 |
ENST00000393593.3 |
IRF2 |
interferon regulatory factor 2 |
| chr2_+_85811525 | 2.84 |
ENST00000306384.4 |
VAMP5 |
vesicle-associated membrane protein 5 |
| chr1_-_159046617 | 2.84 |
ENST00000368130.4 |
AIM2 |
absent in melanoma 2 |
| chr10_+_115439630 | 2.83 |
ENST00000369318.3 |
CASP7 |
caspase 7, apoptosis-related cysteine peptidase |
| chr4_+_100737954 | 2.82 |
ENST00000296414.7 ENST00000512369.1 |
DAPP1 |
dual adaptor of phosphotyrosine and 3-phosphoinositides |
| chr3_+_119316689 | 2.81 |
ENST00000273371.4 |
PLA1A |
phospholipase A1 member A |
| chr20_-_18774614 | 2.78 |
ENST00000412553.1 |
LINC00652 |
long intergenic non-protein coding RNA 652 |
| chr2_+_201994569 | 2.77 |
ENST00000457277.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr6_+_29691056 | 2.77 |
ENST00000414333.1 ENST00000334668.4 ENST00000259951.7 |
HLA-F |
major histocompatibility complex, class I, F |
| chr3_-_119276575 | 2.76 |
ENST00000383669.3 ENST00000383668.3 |
CD80 |
CD80 molecule |
| chr20_+_44746885 | 2.73 |
ENST00000372285.3 |
CD40 |
CD40 molecule, TNF receptor superfamily member 5 |
| chr16_+_50776021 | 2.73 |
ENST00000566679.2 ENST00000564634.1 ENST00000398568.2 |
CYLD |
cylindromatosis (turban tumor syndrome) |
| chr11_-_4414880 | 2.72 |
ENST00000254436.7 ENST00000543625.1 |
TRIM21 |
tripartite motif containing 21 |
| chrX_+_80457442 | 2.71 |
ENST00000373212.5 |
SH3BGRL |
SH3 domain binding glutamic acid-rich protein like |
| chr20_+_44746939 | 2.71 |
ENST00000372276.3 |
CD40 |
CD40 molecule, TNF receptor superfamily member 5 |
| chr7_-_24797546 | 2.71 |
ENST00000414428.1 ENST00000419307.1 ENST00000342947.3 |
DFNA5 |
deafness, autosomal dominant 5 |
| chrX_+_102883620 | 2.70 |
ENST00000372626.3 |
TCEAL1 |
transcription elongation factor A (SII)-like 1 |
| chr9_+_112542591 | 2.68 |
ENST00000483909.1 ENST00000314527.4 ENST00000413420.1 ENST00000302798.7 ENST00000555236.1 ENST00000510514.5 |
PALM2 PALM2-AKAP2 AKAP2 |
paralemmin 2 PALM2-AKAP2 readthrough A kinase (PRKA) anchor protein 2 |
| chr3_+_113251143 | 2.65 |
ENST00000264852.4 ENST00000393830.3 |
SIDT1 |
SID1 transmembrane family, member 1 |
| chrX_+_102883887 | 2.62 |
ENST00000372625.3 ENST00000372624.3 |
TCEAL1 |
transcription elongation factor A (SII)-like 1 |
| chr14_+_24605389 | 2.60 |
ENST00000382708.3 ENST00000561435.1 |
PSME1 |
proteasome (prosome, macropain) activator subunit 1 (PA28 alpha) |
| chr2_+_68592305 | 2.56 |
ENST00000234313.7 |
PLEK |
pleckstrin |
| chrX_-_30595959 | 2.55 |
ENST00000378962.3 |
CXorf21 |
chromosome X open reading frame 21 |
| chr5_-_142783694 | 2.53 |
ENST00000394466.2 |
NR3C1 |
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr16_-_31214051 | 2.45 |
ENST00000350605.4 |
PYCARD |
PYD and CARD domain containing |
| chr7_-_86849025 | 2.42 |
ENST00000257637.3 |
TMEM243 |
transmembrane protein 243, mitochondrial |
| chr1_+_211500129 | 2.41 |
ENST00000427925.2 ENST00000261464.5 |
TRAF5 |
TNF receptor-associated factor 5 |
| chr17_+_41158742 | 2.40 |
ENST00000415816.2 ENST00000438323.2 |
IFI35 |
interferon-induced protein 35 |
| chr19_+_49838653 | 2.39 |
ENST00000598095.1 ENST00000426897.2 ENST00000323906.4 ENST00000535669.2 ENST00000597602.1 ENST00000595660.1 |
CD37 |
CD37 molecule |
| chr11_-_63330842 | 2.37 |
ENST00000255695.1 |
HRASLS2 |
HRAS-like suppressor 2 |
| chr1_+_111772314 | 2.37 |
ENST00000466741.1 ENST00000477185.2 |
CHI3L2 |
chitinase 3-like 2 |
| chr1_+_211499957 | 2.34 |
ENST00000336184.2 |
TRAF5 |
TNF receptor-associated factor 5 |
| chr16_+_29831715 | 2.28 |
ENST00000563915.1 ENST00000357402.5 |
MVP |
major vault protein |
| chr3_-_121379739 | 2.26 |
ENST00000428394.2 ENST00000314583.3 |
HCLS1 |
hematopoietic cell-specific Lyn substrate 1 |
| chr5_+_49963239 | 2.26 |
ENST00000505554.1 |
PARP8 |
poly (ADP-ribose) polymerase family, member 8 |
| chr12_-_10022735 | 2.24 |
ENST00000228438.2 |
CLEC2B |
C-type lectin domain family 2, member B |
| chr18_+_48405463 | 2.21 |
ENST00000382927.3 |
ME2 |
malic enzyme 2, NAD(+)-dependent, mitochondrial |
| chr6_+_26402465 | 2.21 |
ENST00000476549.2 ENST00000289361.6 ENST00000450085.2 ENST00000425234.2 ENST00000427334.1 ENST00000506698.1 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.8 | 26.4 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 7.6 | 22.7 | GO:0051097 | negative regulation of helicase activity(GO:0051097) |
| 7.5 | 59.8 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 7.0 | 20.9 | GO:0045062 | extrathymic T cell selection(GO:0045062) |
| 6.9 | 34.6 | GO:2000110 | negative regulation of macrophage apoptotic process(GO:2000110) |
| 6.4 | 25.6 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 6.1 | 18.2 | GO:0006624 | vacuolar protein processing(GO:0006624) |
| 4.8 | 19.2 | GO:0072308 | negative regulation by virus of viral protein levels in host cell(GO:0046725) negative regulation of nephron tubule epithelial cell differentiation(GO:0072183) negative regulation of metanephric nephron tubule epithelial cell differentiation(GO:0072308) negative regulation of epithelial cell differentiation involved in kidney development(GO:2000697) |
| 4.6 | 50.6 | GO:0003374 | dynamin polymerization involved in membrane fission(GO:0003373) dynamin polymerization involved in mitochondrial fission(GO:0003374) |
| 3.1 | 12.5 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 2.8 | 11.3 | GO:1902044 | regulation of Fas signaling pathway(GO:1902044) |
| 2.7 | 29.5 | GO:1901740 | negative regulation of myoblast fusion(GO:1901740) |
| 2.6 | 13.1 | GO:0002906 | mature B cell apoptotic process(GO:0002901) regulation of mature B cell apoptotic process(GO:0002905) negative regulation of mature B cell apoptotic process(GO:0002906) |
| 2.5 | 12.5 | GO:1904845 | response to L-glutamine(GO:1904844) cellular response to L-glutamine(GO:1904845) |
| 2.4 | 12.1 | GO:1900041 | negative regulation of interleukin-2 secretion(GO:1900041) |
| 2.4 | 9.4 | GO:0046967 | cytosol to ER transport(GO:0046967) |
| 2.3 | 11.6 | GO:0035549 | positive regulation of interferon-beta secretion(GO:0035549) |
| 2.3 | 6.9 | GO:1904469 | positive regulation of tumor necrosis factor secretion(GO:1904469) |
| 2.1 | 17.0 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 2.1 | 6.2 | GO:1990108 | protein linear deubiquitination(GO:1990108) |
| 2.0 | 13.8 | GO:0018377 | protein myristoylation(GO:0018377) |
| 1.9 | 23.2 | GO:0035457 | cellular response to interferon-alpha(GO:0035457) |
| 1.9 | 18.7 | GO:0002480 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-independent(GO:0002480) |
| 1.6 | 7.8 | GO:0019805 | quinolinate biosynthetic process(GO:0019805) |
| 1.5 | 13.1 | GO:0060338 | regulation of type I interferon-mediated signaling pathway(GO:0060338) |
| 1.4 | 5.6 | GO:0003366 | cell-matrix adhesion involved in ameboidal cell migration(GO:0003366) |
| 1.4 | 4.1 | GO:1900368 | regulation of RNA interference(GO:1900368) negative regulation of RNA interference(GO:1900369) |
| 1.3 | 3.8 | GO:0007290 | spermatid nucleus elongation(GO:0007290) |
| 1.2 | 5.9 | GO:0070842 | aggresome assembly(GO:0070842) |
| 1.2 | 1.2 | GO:0002731 | negative regulation of dendritic cell cytokine production(GO:0002731) |
| 1.1 | 11.4 | GO:0051902 | negative regulation of mitochondrial depolarization(GO:0051902) |
| 1.1 | 53.6 | GO:0032731 | positive regulation of interleukin-1 beta production(GO:0032731) |
| 1.1 | 8.9 | GO:0072734 | response to staurosporine(GO:0072733) cellular response to staurosporine(GO:0072734) |
| 1.1 | 2.1 | GO:0051834 | evasion or tolerance of host defenses by virus(GO:0019049) avoidance of host defenses(GO:0044413) evasion or tolerance of host defenses(GO:0044415) avoidance of defenses of other organism involved in symbiotic interaction(GO:0051832) evasion or tolerance of defenses of other organism involved in symbiotic interaction(GO:0051834) |
| 1.0 | 6.2 | GO:0072007 | mesangial cell differentiation(GO:0072007) glomerular mesangial cell differentiation(GO:0072008) mesangial cell development(GO:0072143) glomerular mesangial cell development(GO:0072144) mesenchyme migration(GO:0090131) |
| 1.0 | 17.4 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) |
| 1.0 | 4.0 | GO:0002503 | peptide antigen assembly with MHC class II protein complex(GO:0002503) |
| 1.0 | 3.0 | GO:0032687 | negative regulation of interferon-alpha production(GO:0032687) |
| 1.0 | 9.0 | GO:0016553 | base conversion or substitution editing(GO:0016553) |
| 1.0 | 14.7 | GO:0010623 | programmed cell death involved in cell development(GO:0010623) |
| 1.0 | 2.9 | GO:1905246 | regulation of choline O-acetyltransferase activity(GO:1902769) positive regulation of choline O-acetyltransferase activity(GO:1902771) negative regulation of tau-protein kinase activity(GO:1902948) positive regulation of early endosome to recycling endosome transport(GO:1902955) negative regulation of aspartic-type endopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902960) negative regulation of neurofibrillary tangle assembly(GO:1902997) negative regulation of aspartic-type peptidase activity(GO:1905246) |
| 1.0 | 1.0 | GO:1905154 | negative regulation of membrane invagination(GO:1905154) |
| 0.8 | 3.9 | GO:0002925 | positive regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002925) |
| 0.8 | 5.4 | GO:0033590 | response to cobalamin(GO:0033590) cellular response to erythropoietin(GO:0036018) |
| 0.7 | 1.5 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.7 | 1.4 | GO:0006059 | hexitol metabolic process(GO:0006059) |
| 0.7 | 4.2 | GO:0070092 | regulation of glucagon secretion(GO:0070092) |
| 0.7 | 69.0 | GO:0042590 | antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 0.7 | 41.6 | GO:0002260 | lymphocyte homeostasis(GO:0002260) |
| 0.7 | 5.4 | GO:0097039 | protein linear polyubiquitination(GO:0097039) |
| 0.7 | 7.4 | GO:2000111 | positive regulation of macrophage apoptotic process(GO:2000111) |
| 0.6 | 4.4 | GO:1902031 | regulation of NADP metabolic process(GO:1902031) |
| 0.6 | 2.5 | GO:1903984 | positive regulation of TRAIL-activated apoptotic signaling pathway(GO:1903984) |
| 0.6 | 1.9 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.6 | 4.6 | GO:0046598 | positive regulation of viral entry into host cell(GO:0046598) |
| 0.6 | 4.6 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 0.6 | 3.9 | GO:0098704 | fructose transport(GO:0015755) fructose import(GO:0032445) carbohydrate import into cell(GO:0097319) carbohydrate import across plasma membrane(GO:0098704) fructose import across plasma membrane(GO:1990539) |
| 0.6 | 10.6 | GO:0030889 | negative regulation of B cell proliferation(GO:0030889) |
| 0.6 | 3.3 | GO:0035655 | interleukin-18-mediated signaling pathway(GO:0035655) cellular response to interleukin-18(GO:0071351) |
| 0.6 | 8.9 | GO:0045838 | positive regulation of membrane potential(GO:0045838) |
| 0.5 | 12.0 | GO:0042100 | B cell proliferation(GO:0042100) |
| 0.5 | 58.7 | GO:0060333 | interferon-gamma-mediated signaling pathway(GO:0060333) |
| 0.5 | 2.1 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.5 | 6.4 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 0.5 | 5.9 | GO:1904896 | ESCRT complex disassembly(GO:1904896) ESCRT III complex disassembly(GO:1904903) |
| 0.5 | 2.7 | GO:0060337 | type I interferon signaling pathway(GO:0060337) cellular response to type I interferon(GO:0071357) |
| 0.4 | 8.1 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.4 | 4.6 | GO:0036152 | phosphatidylethanolamine acyl-chain remodeling(GO:0036152) |
| 0.4 | 1.2 | GO:0035621 | ER to Golgi ceramide transport(GO:0035621) |
| 0.4 | 1.2 | GO:1900110 | negative regulation of histone H3-K9 dimethylation(GO:1900110) |
| 0.4 | 2.8 | GO:0045425 | granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0042253) regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045423) positive regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045425) |
| 0.4 | 4.2 | GO:0006030 | chitin metabolic process(GO:0006030) chitin catabolic process(GO:0006032) |
| 0.4 | 1.1 | GO:1904924 | negative regulation of mitophagy in response to mitochondrial depolarization(GO:1904924) |
| 0.4 | 2.5 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) |
| 0.4 | 3.5 | GO:0021891 | olfactory bulb interneuron development(GO:0021891) |
| 0.3 | 1.3 | GO:0042357 | thiamine diphosphate metabolic process(GO:0042357) |
| 0.3 | 2.0 | GO:0007352 | zygotic specification of dorsal/ventral axis(GO:0007352) |
| 0.3 | 1.3 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.3 | 2.9 | GO:0051044 | positive regulation of membrane protein ectodomain proteolysis(GO:0051044) |
| 0.3 | 1.6 | GO:0016199 | axon midline choice point recognition(GO:0016199) cellular response to norepinephrine stimulus(GO:0071874) |
| 0.3 | 1.2 | GO:0090234 | regulation of kinetochore assembly(GO:0090234) |
| 0.3 | 49.7 | GO:0006906 | vesicle fusion(GO:0006906) |
| 0.3 | 0.9 | GO:1990737 | regulation of endoplasmic reticulum stress-induced eIF2 alpha phosphorylation(GO:0060734) response to manganese-induced endoplasmic reticulum stress(GO:1990737) |
| 0.3 | 1.8 | GO:0000821 | regulation of arginine metabolic process(GO:0000821) basic amino acid transmembrane transport(GO:1990822) |
| 0.3 | 14.8 | GO:0034383 | low-density lipoprotein particle clearance(GO:0034383) |
| 0.3 | 13.6 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.3 | 4.5 | GO:0031953 | negative regulation of protein autophosphorylation(GO:0031953) |
| 0.3 | 0.8 | GO:0010716 | negative regulation of extracellular matrix disassembly(GO:0010716) |
| 0.3 | 9.7 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.3 | 0.5 | GO:0045994 | positive regulation of translational initiation by iron(GO:0045994) |
| 0.2 | 4.2 | GO:0072643 | interferon-gamma secretion(GO:0072643) |
| 0.2 | 0.2 | GO:0035990 | tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) |
| 0.2 | 5.6 | GO:0036150 | phosphatidylserine acyl-chain remodeling(GO:0036150) |
| 0.2 | 2.7 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.2 | 0.9 | GO:0006780 | uroporphyrinogen III biosynthetic process(GO:0006780) uroporphyrinogen III metabolic process(GO:0046502) |
| 0.2 | 6.5 | GO:0002089 | lens morphogenesis in camera-type eye(GO:0002089) |
| 0.2 | 0.7 | GO:1990169 | detoxification of copper ion(GO:0010273) elastin metabolic process(GO:0051541) cellular response to lead ion(GO:0071284) stress response to copper ion(GO:1990169) |
| 0.2 | 1.5 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.2 | 2.4 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) |
| 0.2 | 10.8 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.2 | 0.4 | GO:0035787 | cell migration involved in kidney development(GO:0035787) cell migration involved in metanephros development(GO:0035788) metanephric mesenchymal cell migration(GO:0035789) positive regulation of metanephric mesenchymal cell migration by platelet-derived growth factor receptor-beta signaling pathway(GO:0035793) regulation of metanephric mesenchymal cell migration by platelet-derived growth factor receptor-beta signaling pathway(GO:1900238) regulation of metanephric mesenchymal cell migration(GO:2000589) positive regulation of metanephric mesenchymal cell migration(GO:2000591) |
| 0.2 | 1.0 | GO:0060266 | negative regulation of respiratory burst involved in inflammatory response(GO:0060266) |
| 0.2 | 1.9 | GO:0001783 | B cell apoptotic process(GO:0001783) |
| 0.2 | 5.0 | GO:0030890 | positive regulation of B cell proliferation(GO:0030890) |
| 0.2 | 3.0 | GO:0034656 | nucleobase-containing small molecule catabolic process(GO:0034656) |
| 0.2 | 0.5 | GO:0055099 | detection of endogenous stimulus(GO:0009726) response to high density lipoprotein particle(GO:0055099) |
| 0.2 | 2.6 | GO:0030836 | positive regulation of actin filament depolymerization(GO:0030836) |
| 0.2 | 1.2 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.2 | 0.2 | GO:0072179 | nephric duct formation(GO:0072179) |
| 0.2 | 1.3 | GO:0031053 | primary miRNA processing(GO:0031053) |
| 0.2 | 4.9 | GO:0035456 | response to interferon-beta(GO:0035456) |
| 0.2 | 1.6 | GO:0010506 | regulation of autophagy(GO:0010506) |
| 0.2 | 0.5 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.2 | 0.3 | GO:0061428 | negative regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061428) |
| 0.1 | 1.1 | GO:0060315 | negative regulation of ryanodine-sensitive calcium-release channel activity(GO:0060315) |
| 0.1 | 0.4 | GO:0045345 | positive regulation of MHC class I biosynthetic process(GO:0045345) |
| 0.1 | 1.3 | GO:0038092 | nodal signaling pathway(GO:0038092) |
| 0.1 | 0.3 | GO:0070987 | error-free translesion synthesis(GO:0070987) |
| 0.1 | 2.4 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.1 | 0.7 | GO:0046087 | cytidine catabolic process(GO:0006216) cytidine deamination(GO:0009972) cytidine metabolic process(GO:0046087) |
| 0.1 | 1.1 | GO:0045905 | translational frameshifting(GO:0006452) positive regulation of translational termination(GO:0045905) |
| 0.1 | 1.0 | GO:0015939 | pantothenate metabolic process(GO:0015939) |
| 0.1 | 0.1 | GO:1902523 | positive regulation of protein K63-linked ubiquitination(GO:1902523) |
| 0.1 | 0.8 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.1 | 1.4 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.1 | 0.2 | GO:0010700 | negative regulation of norepinephrine secretion(GO:0010700) |
| 0.1 | 1.3 | GO:0006554 | lysine catabolic process(GO:0006554) |
| 0.1 | 3.2 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.1 | 1.2 | GO:1902004 | positive regulation of beta-amyloid formation(GO:1902004) |
| 0.1 | 2.0 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.1 | 0.5 | GO:0001831 | trophectodermal cellular morphogenesis(GO:0001831) |
| 0.1 | 2.4 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 0.6 | GO:2001023 | regulation of response to drug(GO:2001023) |
| 0.1 | 0.2 | GO:0036496 | regulation of translational initiation by eIF2 alpha dephosphorylation(GO:0036496) |
| 0.1 | 0.2 | GO:1900227 | positive regulation of NLRP3 inflammasome complex assembly(GO:1900227) |
| 0.1 | 3.4 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.1 | 0.3 | GO:1903598 | receptor-mediated virion attachment to host cell(GO:0046813) positive regulation of gap junction assembly(GO:1903598) |
| 0.1 | 2.0 | GO:0032012 | regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 0.3 | GO:1901339 | activation of store-operated calcium channel activity(GO:0032237) regulation of store-operated calcium channel activity(GO:1901339) positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.1 | 1.5 | GO:0032836 | glomerular basement membrane development(GO:0032836) |
| 0.1 | 0.5 | GO:0070836 | plasma membrane raft assembly(GO:0044854) plasma membrane raft organization(GO:0044857) caveola assembly(GO:0070836) |
| 0.1 | 0.5 | GO:0032916 | transforming growth factor beta3 production(GO:0032907) regulation of transforming growth factor beta3 production(GO:0032910) positive regulation of transforming growth factor beta3 production(GO:0032916) chemotaxis to arachidonic acid(GO:0034670) response to arachidonic acid(GO:1904550) |
| 0.1 | 0.3 | GO:0060474 | positive regulation of sperm motility involved in capacitation(GO:0060474) |
| 0.1 | 1.0 | GO:1900028 | negative regulation of ruffle assembly(GO:1900028) |
| 0.1 | 3.5 | GO:0006536 | glutamate metabolic process(GO:0006536) |
| 0.1 | 25.5 | GO:0009615 | response to virus(GO:0009615) |
| 0.1 | 1.2 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.1 | 0.3 | GO:0048205 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 0.1 | 1.3 | GO:0032897 | negative regulation of viral transcription(GO:0032897) |
| 0.1 | 0.4 | GO:0071494 | cellular response to UV-C(GO:0071494) |
| 0.1 | 1.9 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.1 | 1.2 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 1.3 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.1 | 0.2 | GO:0061537 | glycine secretion(GO:0061536) glycine secretion, neurotransmission(GO:0061537) |
| 0.1 | 0.4 | GO:1900165 | negative regulation of interleukin-6 secretion(GO:1900165) |
| 0.1 | 0.4 | GO:1902035 | positive regulation of hematopoietic stem cell proliferation(GO:1902035) |
| 0.1 | 0.7 | GO:0015886 | heme transport(GO:0015886) |
| 0.1 | 1.4 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.1 | 1.1 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.1 | 2.6 | GO:0030835 | negative regulation of actin filament depolymerization(GO:0030835) |
| 0.1 | 0.8 | GO:2000659 | regulation of interleukin-1-mediated signaling pathway(GO:2000659) |
| 0.1 | 3.8 | GO:0002456 | T cell mediated immunity(GO:0002456) |
| 0.1 | 0.8 | GO:0051823 | regulation of synapse structural plasticity(GO:0051823) |
| 0.1 | 0.5 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.1 | 0.7 | GO:0043473 | pigmentation(GO:0043473) |
| 0.1 | 0.1 | GO:0016048 | detection of temperature stimulus(GO:0016048) sensory perception of temperature stimulus(GO:0050951) |
| 0.1 | 0.4 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.0 | 0.2 | GO:0021691 | cerebellar Purkinje cell layer maturation(GO:0021691) |
| 0.0 | 0.1 | GO:0021779 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) oligodendrocyte cell fate specification(GO:0021778) oligodendrocyte cell fate commitment(GO:0021779) glial cell fate specification(GO:0021780) |
| 0.0 | 0.2 | GO:0035279 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing by siRNA(GO:0090625) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.0 | 1.9 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.0 | 0.3 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.0 | 0.2 | GO:0071346 | cellular response to interferon-gamma(GO:0071346) |
| 0.0 | 1.0 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 2.7 | GO:0042490 | mechanoreceptor differentiation(GO:0042490) inner ear receptor cell differentiation(GO:0060113) |
| 0.0 | 0.3 | GO:0009804 | coumarin metabolic process(GO:0009804) |
| 0.0 | 0.1 | GO:0002589 | regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002589) negative regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002590) |
| 0.0 | 2.0 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.0 | 0.4 | GO:0051972 | regulation of telomerase activity(GO:0051972) |
| 0.0 | 0.0 | GO:0001302 | replicative cell aging(GO:0001302) |
| 0.0 | 0.2 | GO:0045137 | development of primary sexual characteristics(GO:0045137) |
| 0.0 | 0.1 | GO:0070781 | arginine biosynthetic process via ornithine(GO:0042450) response to biotin(GO:0070781) |
| 0.0 | 0.2 | GO:0006436 | tryptophanyl-tRNA aminoacylation(GO:0006436) |
| 0.0 | 0.6 | GO:2000574 | regulation of microtubule motor activity(GO:2000574) |
| 0.0 | 0.3 | GO:0035307 | positive regulation of dephosphorylation(GO:0035306) positive regulation of protein dephosphorylation(GO:0035307) |
| 0.0 | 0.2 | GO:1902904 | vacuolar transmembrane transport(GO:0034486) late endosomal microautophagy(GO:0061738) chaperone-mediated protein transport involved in chaperone-mediated autophagy(GO:0061741) negative regulation of fibril organization(GO:1902904) chaperone-mediated autophagy translocation complex disassembly(GO:1904764) |
| 0.0 | 2.4 | GO:0006968 | cellular defense response(GO:0006968) |
| 0.0 | 0.0 | GO:0042320 | regulation of circadian sleep/wake cycle, REM sleep(GO:0042320) |
| 0.0 | 0.4 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.0 | 0.8 | GO:0035338 | long-chain fatty-acyl-CoA biosynthetic process(GO:0035338) |
| 0.0 | 0.1 | GO:0005989 | lactose metabolic process(GO:0005988) lactose biosynthetic process(GO:0005989) |
| 0.0 | 0.1 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.0 | 3.3 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.0 | 0.3 | GO:0042501 | serine phosphorylation of STAT protein(GO:0042501) |
| 0.0 | 2.3 | GO:0048477 | oogenesis(GO:0048477) |
| 0.0 | 2.9 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 0.1 | GO:0044333 | Wnt signaling pathway involved in digestive tract morphogenesis(GO:0044333) |
| 0.0 | 0.2 | GO:0038161 | prolactin signaling pathway(GO:0038161) |
| 0.0 | 3.1 | GO:0051603 | proteolysis involved in cellular protein catabolic process(GO:0051603) |
| 0.0 | 0.2 | GO:0035795 | negative regulation of mitochondrial membrane permeability(GO:0035795) |
| 0.0 | 0.1 | GO:0002384 | hepatic immune response(GO:0002384) |
| 0.0 | 0.1 | GO:0010529 | regulation of transposition(GO:0010528) negative regulation of transposition(GO:0010529) |
| 0.0 | 0.5 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.0 | 0.6 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 4.4 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.0 | 0.2 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 0.0 | 1.0 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 0.1 | GO:0090394 | negative regulation of excitatory postsynaptic potential(GO:0090394) |
| 0.0 | 0.4 | GO:0042167 | porphyrin-containing compound catabolic process(GO:0006787) tetrapyrrole catabolic process(GO:0033015) heme catabolic process(GO:0042167) pigment catabolic process(GO:0046149) |
| 0.0 | 0.1 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.0 | 0.3 | GO:0051386 | regulation of neurotrophin TRK receptor signaling pathway(GO:0051386) |
| 0.0 | 0.7 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.1 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.0 | 0.2 | GO:0010886 | positive regulation of cholesterol storage(GO:0010886) |
| 0.0 | 0.1 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 0.1 | GO:0044256 | angiotensin catabolic process in blood(GO:0002005) multicellular organismal protein catabolic process(GO:0044254) protein digestion(GO:0044256) multicellular organismal macromolecule catabolic process(GO:0044266) multicellular organismal protein metabolic process(GO:0044268) |
| 0.0 | 0.1 | GO:0032480 | negative regulation of type I interferon production(GO:0032480) |
| 0.0 | 0.2 | GO:0048839 | inner ear development(GO:0048839) |
| 0.0 | 0.1 | GO:0060244 | negative regulation of cell proliferation involved in contact inhibition(GO:0060244) |
| 0.0 | 0.2 | GO:0030091 | protein repair(GO:0030091) |
| 0.0 | 0.8 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.0 | 0.3 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.0 | 0.1 | GO:1902255 | positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator(GO:1902255) |
| 0.0 | 0.1 | GO:1902775 | mitochondrial ribosome assembly(GO:0061668) mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.0 | 2.4 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.0 | 1.5 | GO:0006687 | glycosphingolipid metabolic process(GO:0006687) |
| 0.0 | 0.2 | GO:0060213 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) |
| 0.0 | 0.6 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.0 | 1.8 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 0.4 | GO:0042059 | negative regulation of epidermal growth factor receptor signaling pathway(GO:0042059) |
| 0.0 | 0.5 | GO:0045599 | negative regulation of fat cell differentiation(GO:0045599) |
| 0.0 | 0.1 | GO:1902259 | regulation of delayed rectifier potassium channel activity(GO:1902259) |
| 0.0 | 0.2 | GO:0060711 | labyrinthine layer development(GO:0060711) |
| 0.0 | 1.0 | GO:0032435 | negative regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032435) |
| 0.0 | 0.2 | GO:0006413 | translational initiation(GO:0006413) |
| 0.0 | 0.0 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.0 | 0.4 | GO:0031055 | chromatin remodeling at centromere(GO:0031055) CENP-A containing nucleosome assembly(GO:0034080) CENP-A containing chromatin organization(GO:0061641) |
| 0.0 | 0.1 | GO:2000158 | positive regulation of ubiquitin-specific protease activity(GO:2000158) |
| 0.0 | 0.2 | GO:0000725 | double-strand break repair via homologous recombination(GO:0000724) recombinational repair(GO:0000725) |
| 0.0 | 0.1 | GO:1903976 | negative regulation of glial cell migration(GO:1903976) |
| 0.0 | 0.1 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.0 | GO:0032488 | Cdc42 protein signal transduction(GO:0032488) |
| 0.0 | 1.7 | GO:0015992 | proton transport(GO:0015992) |
| 0.0 | 0.1 | GO:0032790 | ribosome disassembly(GO:0032790) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.8 | 26.4 | GO:0008859 | exoribonuclease II activity(GO:0008859) |
| 5.3 | 31.6 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 3.5 | 34.5 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 3.2 | 19.2 | GO:0046979 | TAP2 binding(GO:0046979) |
| 2.8 | 13.9 | GO:0004771 | sterol esterase activity(GO:0004771) |
| 2.4 | 7.2 | GO:0022897 | peptide:proton symporter activity(GO:0015333) proton-dependent peptide secondary active transmembrane transporter activity(GO:0022897) |
| 1.9 | 56.2 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 1.9 | 22.5 | GO:0031386 | protein tag(GO:0031386) |
| 1.8 | 12.9 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 1.6 | 8.0 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 1.6 | 9.5 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 1.4 | 11.3 | GO:0070087 | chromo shadow domain binding(GO:0070087) |
| 1.4 | 42.0 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 1.2 | 6.2 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 1.2 | 2.5 | GO:0097153 | cysteine-type endopeptidase activity involved in apoptotic process(GO:0097153) |
| 1.1 | 7.8 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 1.1 | 8.7 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 1.1 | 6.4 | GO:0004614 | phosphoglucomutase activity(GO:0004614) |
| 1.0 | 3.0 | GO:0004694 | eukaryotic translation initiation factor 2alpha kinase activity(GO:0004694) |
| 1.0 | 6.9 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 1.0 | 4.8 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 1.0 | 5.7 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.9 | 4.6 | GO:0038051 | glucocorticoid receptor activity(GO:0004883) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.9 | 8.9 | GO:0015038 | glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.9 | 3.5 | GO:0005151 | interleukin-1, Type II receptor binding(GO:0005151) |
| 0.8 | 19.7 | GO:0050664 | oxidoreductase activity, acting on NAD(P)H, oxygen as acceptor(GO:0050664) |
| 0.7 | 4.4 | GO:0008948 | malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) oxaloacetate decarboxylase activity(GO:0008948) |
| 0.7 | 5.0 | GO:0052739 | phosphatidylserine 1-acylhydrolase activity(GO:0052739) |
| 0.7 | 3.5 | GO:0047820 | D-glutamate cyclase activity(GO:0047820) |
| 0.6 | 1.9 | GO:0015526 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.6 | 6.2 | GO:1990380 | Lys63-specific deubiquitinase activity(GO:0061578) Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.6 | 4.8 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.6 | 7.6 | GO:0001011 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.6 | 3.9 | GO:0005353 | fructose transmembrane transporter activity(GO:0005353) |
| 0.5 | 15.9 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.5 | 3.5 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.5 | 3.9 | GO:0019863 | IgE binding(GO:0019863) |
| 0.5 | 8.1 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.5 | 21.3 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.5 | 13.7 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.4 | 9.9 | GO:0000339 | RNA cap binding(GO:0000339) |
| 0.4 | 2.7 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.4 | 2.1 | GO:0008174 | mRNA methyltransferase activity(GO:0008174) |
| 0.4 | 2.5 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.4 | 1.2 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.4 | 8.1 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.4 | 4.2 | GO:0008061 | chitinase activity(GO:0004568) chitin binding(GO:0008061) |
| 0.4 | 4.5 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.3 | 0.7 | GO:0032767 | copper-dependent protein binding(GO:0032767) |
| 0.3 | 1.3 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.3 | 1.6 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.3 | 34.9 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.3 | 0.9 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.3 | 81.9 | GO:0001047 | core promoter binding(GO:0001047) |
| 0.3 | 1.2 | GO:0097001 | ceramide binding(GO:0097001) |
| 0.3 | 0.9 | GO:0010997 | anaphase-promoting complex binding(GO:0010997) phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 0.3 | 5.9 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.3 | 26.9 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.3 | 8.5 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.3 | 10.1 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.3 | 4.9 | GO:0005522 | profilin binding(GO:0005522) |
| 0.3 | 16.3 | GO:0004520 | endodeoxyribonuclease activity(GO:0004520) |
| 0.3 | 4.6 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.3 | 0.5 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.3 | 10.4 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) |
| 0.2 | 1.2 | GO:0016402 | pristanoyl-CoA oxidase activity(GO:0016402) |
| 0.2 | 5.4 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.2 | 3.0 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.2 | 1.0 | GO:0017159 | pantetheine hydrolase activity(GO:0017159) |
| 0.2 | 0.6 | GO:0031735 | CCR10 chemokine receptor binding(GO:0031735) |
| 0.2 | 1.8 | GO:1901612 | cardiolipin binding(GO:1901612) |
| 0.2 | 1.2 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 0.2 | 1.1 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.2 | 5.5 | GO:0042605 | peptide antigen binding(GO:0042605) |
| 0.2 | 2.9 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.2 | 82.5 | GO:0005525 | GTP binding(GO:0005525) |
| 0.2 | 0.5 | GO:0102008 | cytosolic dipeptidase activity(GO:0102008) |
| 0.2 | 0.5 | GO:0034041 | sterol-transporting ATPase activity(GO:0034041) |
| 0.2 | 1.3 | GO:0016778 | diphosphotransferase activity(GO:0016778) |
| 0.2 | 1.2 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.2 | 1.8 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.2 | 0.5 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.2 | 0.5 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.1 | 2.2 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.1 | 2.3 | GO:0004908 | interleukin-1 receptor activity(GO:0004908) |
| 0.1 | 1.2 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 0.1 | 5.3 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 0.4 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.1 | 0.5 | GO:0031208 | POZ domain binding(GO:0031208) |
| 0.1 | 3.2 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.1 | 1.6 | GO:0004551 | nucleotide diphosphatase activity(GO:0004551) |
| 0.1 | 15.1 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.1 | 5.1 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 1.7 | GO:0044548 | S100 protein binding(GO:0044548) |
| 0.1 | 0.5 | GO:0001225 | RNA polymerase II transcription coactivator binding(GO:0001225) |
| 0.1 | 2.0 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.1 | 1.4 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.1 | 0.4 | GO:0004074 | biliverdin reductase activity(GO:0004074) |
| 0.1 | 1.0 | GO:0052658 | inositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052658) |
| 0.1 | 4.1 | GO:0016667 | oxidoreductase activity, acting on a sulfur group of donors(GO:0016667) |
| 0.1 | 0.3 | GO:0004060 | arylamine N-acetyltransferase activity(GO:0004060) |
| 0.1 | 0.6 | GO:0043140 | ATP-dependent 3'-5' DNA helicase activity(GO:0043140) |
| 0.1 | 0.3 | GO:0019826 | oxygen sensor activity(GO:0019826) |
| 0.1 | 0.3 | GO:0016160 | amylase activity(GO:0016160) |
| 0.1 | 0.7 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.1 | 1.1 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.1 | 0.3 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.1 | 3.1 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.1 | 0.2 | GO:0008267 | poly-glutamine tract binding(GO:0008267) |
| 0.1 | 0.4 | GO:0003756 | protein disulfide isomerase activity(GO:0003756) intramolecular oxidoreductase activity, transposing S-S bonds(GO:0016864) |
| 0.1 | 1.0 | GO:0017017 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) |
| 0.1 | 0.4 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.1 | 1.7 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.1 | 0.6 | GO:0098505 | single-stranded telomeric DNA binding(GO:0043047) G-rich strand telomeric DNA binding(GO:0098505) |
| 0.1 | 1.2 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.1 | 0.1 | GO:0001224 | RNA polymerase II transcription cofactor binding(GO:0001224) |
| 0.1 | 2.6 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 0.2 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 0.1 | 0.8 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 2.4 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.1 | 1.7 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.1 | 0.3 | GO:0008241 | peptidyl-dipeptidase activity(GO:0008241) |
| 0.1 | 0.6 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.1 | 1.2 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 1.4 | GO:0004623 | phospholipase A2 activity(GO:0004623) |
| 0.0 | 0.2 | GO:0001595 | angiotensin receptor activity(GO:0001595) angiotensin type II receptor activity(GO:0004945) |
| 0.0 | 2.3 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.8 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 4.4 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 1.1 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.6 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) |
| 0.0 | 0.6 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.0 | 1.1 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.0 | 0.2 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.0 | 0.2 | GO:0004830 | tryptophan-tRNA ligase activity(GO:0004830) |
| 0.0 | 1.5 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.0 | 0.5 | GO:0035325 | Toll-like receptor binding(GO:0035325) |
| 0.0 | 0.5 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.0 | 0.7 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.1 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.0 | 0.3 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.0 | 0.4 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.0 | 0.2 | GO:0004925 | prolactin receptor activity(GO:0004925) |
| 0.0 | 1.3 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 25.2 | GO:0008270 | zinc ion binding(GO:0008270) |
| 0.0 | 0.5 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.4 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.0 | 0.9 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) |
| 0.0 | 3.1 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.3 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.1 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.0 | 1.1 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.6 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.0 | 0.3 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 2.5 | GO:0047485 | protein N-terminus binding(GO:0047485) |
| 0.0 | 3.7 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 0.5 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.2 | GO:0000182 | rDNA binding(GO:0000182) |
| 0.0 | 0.4 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.0 | 1.7 | GO:0003690 | double-stranded DNA binding(GO:0003690) |
| 0.0 | 0.1 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.0 | 0.1 | GO:0019981 | interleukin-6 receptor activity(GO:0004915) interleukin-6 binding(GO:0019981) |
| 0.0 | 0.5 | GO:0048487 | beta-tubulin binding(GO:0048487) |
| 0.0 | 11.9 | GO:0044212 | transcription regulatory region DNA binding(GO:0044212) |
| 0.0 | 0.2 | GO:0008865 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.0 | 0.2 | GO:0005131 | growth hormone receptor binding(GO:0005131) |
| 0.0 | 0.2 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.0 | 0.3 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 0.4 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 1.4 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.0 | 1.2 | GO:0004871 | signal transducer activity(GO:0004871) |
| 0.0 | 0.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.0 | 0.1 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.0 | 0.9 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.0 | 0.2 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 0.8 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.2 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.1 | GO:0005168 | neurotrophin TRKA receptor binding(GO:0005168) |
| 0.0 | 1.5 | GO:0005057 | receptor signaling protein activity(GO:0005057) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.5 | 60.4 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 2.9 | 20.2 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 2.6 | 13.1 | GO:0097179 | protease inhibitor complex(GO:0097179) |
| 2.0 | 6.1 | GO:0030895 | apolipoprotein B mRNA editing enzyme complex(GO:0030895) |
| 2.0 | 11.8 | GO:0042825 | TAP complex(GO:0042825) |
| 1.7 | 11.9 | GO:0030870 | Mre11 complex(GO:0030870) |
| 1.6 | 12.9 | GO:0032010 | phagolysosome(GO:0032010) |
| 1.4 | 5.6 | GO:0034669 | integrin alpha4-beta7 complex(GO:0034669) |
| 1.3 | 7.8 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 1.2 | 8.3 | GO:0043196 | varicosity(GO:0043196) |
| 1.2 | 12.8 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 1.1 | 5.4 | GO:0071797 | LUBAC complex(GO:0071797) |
| 1.0 | 20.0 | GO:0031265 | CD95 death-inducing signaling complex(GO:0031265) |
| 1.0 | 12.5 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 1.0 | 12.4 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.8 | 3.4 | GO:0097454 | Schwann cell microvillus(GO:0097454) |
| 0.8 | 4.1 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.8 | 5.4 | GO:0035692 | macrophage migration inhibitory factor receptor complex(GO:0035692) |
| 0.6 | 4.5 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.6 | 6.8 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.6 | 6.5 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.6 | 2.8 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 0.5 | 4.9 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.5 | 37.9 | GO:0015030 | Cajal body(GO:0015030) |
| 0.5 | 1.6 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.4 | 1.6 | GO:0031905 | early endosome lumen(GO:0031905) |
| 0.4 | 29.9 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.4 | 1.1 | GO:0071748 | IgA immunoglobulin complex(GO:0071745) IgA immunoglobulin complex, circulating(GO:0071746) monomeric IgA immunoglobulin complex(GO:0071748) polymeric IgA immunoglobulin complex(GO:0071749) secretory IgA immunoglobulin complex(GO:0071751) |
| 0.3 | 4.1 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.3 | 1.7 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.3 | 1.0 | GO:0036398 | TCR signalosome(GO:0036398) |
| 0.3 | 0.9 | GO:0020016 | ciliary pocket(GO:0020016) ciliary pocket membrane(GO:0020018) |
| 0.3 | 4.6 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.3 | 0.6 | GO:0031213 | RSF complex(GO:0031213) |
| 0.3 | 18.6 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.3 | 1.3 | GO:0072559 | NLRP3 inflammasome complex(GO:0072559) |
| 0.2 | 5.0 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.2 | 4.0 | GO:0042611 | MHC protein complex(GO:0042611) MHC class II protein complex(GO:0042613) |
| 0.2 | 1.0 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.2 | 1.5 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.2 | 17.3 | GO:0018995 | host(GO:0018995) host cell(GO:0043657) |
| 0.2 | 0.5 | GO:0008282 | ATP-sensitive potassium channel complex(GO:0008282) |
| 0.2 | 5.2 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
| 0.1 | 0.4 | GO:0033565 | ESCRT-0 complex(GO:0033565) |
| 0.1 | 1.8 | GO:0070419 | nonhomologous end joining complex(GO:0070419) |
| 0.1 | 12.6 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.1 | 5.5 | GO:0097542 | ciliary tip(GO:0097542) |
| 0.1 | 5.4 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 52.2 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.1 | 38.9 | GO:0031965 | nuclear membrane(GO:0031965) |
| 0.1 | 0.2 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.1 | 12.4 | GO:0001650 | fibrillar center(GO:0001650) |
| 0.1 | 0.8 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.1 | 0.9 | GO:0035749 | myelin sheath adaxonal region(GO:0035749) |
| 0.1 | 1.3 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.1 | 12.7 | GO:0045111 | intermediate filament cytoskeleton(GO:0045111) |
| 0.1 | 2.7 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.1 | 29.5 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.1 | 3.4 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 0.6 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 0.5 | GO:0016461 | unconventional myosin complex(GO:0016461) |
| 0.1 | 22.3 | GO:0009897 | external side of plasma membrane(GO:0009897) |
| 0.1 | 8.0 | GO:0005635 | nuclear envelope(GO:0005635) |
| 0.1 | 1.9 | GO:0051233 | spindle midzone(GO:0051233) |
| 0.1 | 0.5 | GO:0032009 | early phagosome(GO:0032009) |
| 0.1 | 1.0 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.1 | 11.0 | GO:0005741 | mitochondrial outer membrane(GO:0005741) organelle outer membrane(GO:0031968) |
| 0.1 | 4.9 | GO:1904724 | tertiary granule lumen(GO:1904724) |
| 0.1 | 0.3 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.1 | 0.2 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.1 | 0.5 | GO:0005827 | polar microtubule(GO:0005827) |
| 0.1 | 0.7 | GO:0031082 | BLOC complex(GO:0031082) |
| 0.1 | 2.8 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 2.1 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 3.6 | GO:0043679 | axon terminus(GO:0043679) |
| 0.0 | 0.3 | GO:0097442 | CA3 pyramidal cell dendrite(GO:0097442) |
| 0.0 | 0.4 | GO:0071682 | endocytic vesicle lumen(GO:0071682) |
| 0.0 | 64.6 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 0.2 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 3.5 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 27.5 | GO:0005768 | endosome(GO:0005768) |
| 0.0 | 0.2 | GO:0072487 | MSL complex(GO:0072487) |
| 0.0 | 0.2 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.0 | 9.1 | GO:0000323 | lytic vacuole(GO:0000323) lysosome(GO:0005764) |
| 0.0 | 0.2 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.0 | 0.1 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 0.0 | 0.3 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 0.3 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 0.6 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 2.1 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 2.5 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 1.6 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.0 | 9.4 | GO:0016020 | membrane(GO:0016020) |
| 0.0 | 0.4 | GO:0035580 | specific granule lumen(GO:0035580) |
| 0.0 | 2.1 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 0.3 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.3 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.0 | 1.0 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 0.4 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.0 | 0.8 | GO:0000922 | spindle pole(GO:0000922) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 24.3 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.7 | 20.5 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.5 | 1.4 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.4 | 10.6 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.3 | 8.5 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.3 | 21.5 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.3 | 22.9 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.3 | 43.6 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.3 | 3.1 | SIG IL4RECEPTOR IN B LYPHOCYTES | Genes related to IL4 rceptor signaling in B lymphocytes |
| 0.2 | 8.0 | PID NFKAPPAB CANONICAL PATHWAY | Canonical NF-kappaB pathway |
| 0.2 | 9.7 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.2 | 5.6 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.2 | 6.4 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.2 | 8.0 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.2 | 12.2 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.2 | 2.6 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.2 | 6.1 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.1 | 8.1 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.1 | 7.8 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.1 | 3.5 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.1 | 4.9 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 4.1 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.1 | 5.4 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 19.3 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 0.8 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.1 | 0.6 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 4.0 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 1.5 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 0.9 | PID CD8 TCR PATHWAY | TCR signaling in naïve CD8+ T cells |
| 0.1 | 1.3 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 1.6 | PID PI3KCI PATHWAY | Class I PI3K signaling events |
| 0.1 | 1.2 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 1.8 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.1 | 6.0 | PID PDGFRB PATHWAY | PDGFR-beta signaling pathway |
| 0.1 | 1.9 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.1 | 1.1 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.1 | 10.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.1 | 2.5 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.1 | 1.5 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 3.6 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.8 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 1.5 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 1.9 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 0.3 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 0.1 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.0 | 0.4 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.0 | 0.6 | ST P38 MAPK PATHWAY | p38 MAPK Pathway |
| 0.0 | 0.5 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 0.3 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.0 | 0.5 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.0 | 0.3 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
| 0.0 | 1.0 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 3.8 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.2 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.0 | 0.2 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.0 | 0.3 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.2 | 290.3 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 1.2 | 17.3 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 1.1 | 56.8 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.8 | 16.0 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.8 | 58.2 | REACTOME VIF MEDIATED DEGRADATION OF APOBEC3G | Genes involved in Vif-mediated degradation of APOBEC3G |
| 0.7 | 2.2 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.7 | 9.1 | REACTOME ER PHAGOSOME PATHWAY | Genes involved in ER-Phagosome pathway |
| 0.6 | 16.2 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.6 | 11.6 | REACTOME ANTIGEN PROCESSING CROSS PRESENTATION | Genes involved in Antigen processing-Cross presentation |
| 0.4 | 9.5 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.3 | 18.4 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.3 | 9.9 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.3 | 10.4 | REACTOME RIG I MDA5 MEDIATED INDUCTION OF IFN ALPHA BETA PATHWAYS | Genes involved in RIG-I/MDA5 mediated induction of IFN-alpha/beta pathways |
| 0.3 | 5.9 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.3 | 8.9 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.3 | 5.6 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.3 | 1.7 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.3 | 6.5 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.2 | 2.8 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.2 | 6.3 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.2 | 1.9 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.2 | 2.4 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.2 | 5.2 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.2 | 9.9 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 3.1 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.1 | 8.5 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.1 | 2.8 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.1 | 12.3 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.1 | 1.9 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.1 | 11.3 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.1 | 6.0 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.1 | 9.2 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.1 | 0.2 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.1 | 5.1 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.1 | 1.7 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.1 | 2.5 | REACTOME PERK REGULATED GENE EXPRESSION | Genes involved in PERK regulated gene expression |
| 0.1 | 1.3 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.1 | 1.3 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 4.1 | REACTOME DOWNSTREAM SIGNALING EVENTS OF B CELL RECEPTOR BCR | Genes involved in Downstream Signaling Events Of B Cell Receptor (BCR) |
| 0.1 | 0.7 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.1 | 1.1 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.0 | 0.8 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.0 | 1.3 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 1.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.8 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 1.2 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.0 | 1.7 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 0.6 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.0 | 2.2 | REACTOME INTERFERON SIGNALING | Genes involved in Interferon Signaling |
| 0.0 | 1.1 | REACTOME PTM GAMMA CARBOXYLATION HYPUSINE FORMATION AND ARYLSULFATASE ACTIVATION | Genes involved in PTM: gamma carboxylation, hypusine formation and arylsulfatase activation |
| 0.0 | 0.3 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 0.7 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 2.1 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.5 | REACTOME INWARDLY RECTIFYING K CHANNELS | Genes involved in Inwardly rectifying K+ channels |
| 0.0 | 0.2 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.0 | 0.4 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 0.4 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 0.5 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 0.2 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.0 | 0.5 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.0 | 0.1 | REACTOME NUCLEOTIDE BINDING DOMAIN LEUCINE RICH REPEAT CONTAINING RECEPTOR NLR SIGNALING PATHWAYS | Genes involved in Nucleotide-binding domain, leucine rich repeat containing receptor (NLR) signaling pathways |
| 0.0 | 0.3 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.2 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.0 | 1.9 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.2 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.0 | 0.2 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 2.0 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |