Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
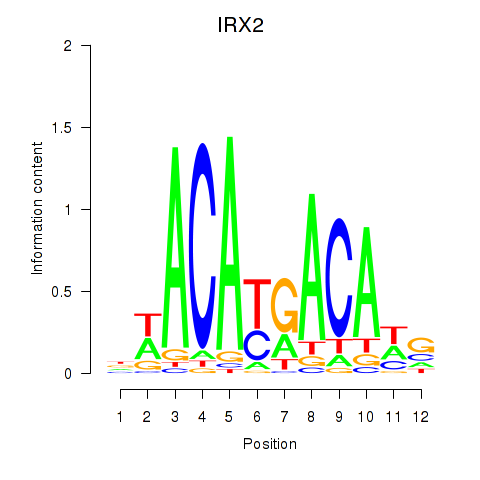
Results for IRX2
Z-value: 0.67
Transcription factors associated with IRX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
IRX2
|
ENSG00000170561.8 | IRX2 |
Activity profile of IRX2 motif
Sorted Z-values of IRX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of IRX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_88896819 | 2.26 |
ENST00000237623.7 ENST00000395080.3 ENST00000508233.1 ENST00000360804.4 |
SPP1 |
secreted phosphoprotein 1 |
| chr13_-_46679185 | 2.06 |
ENST00000439329.3 |
CPB2 |
carboxypeptidase B2 (plasma) |
| chr13_-_46679144 | 2.03 |
ENST00000181383.4 |
CPB2 |
carboxypeptidase B2 (plasma) |
| chr1_-_92371839 | 1.51 |
ENST00000370399.2 |
TGFBR3 |
transforming growth factor, beta receptor III |
| chr4_+_69681710 | 1.42 |
ENST00000265403.7 ENST00000458688.2 |
UGT2B10 |
UDP glucuronosyltransferase 2 family, polypeptide B10 |
| chr1_-_111506562 | 1.25 |
ENST00000485275.2 ENST00000369763.4 |
LRIF1 |
ligand dependent nuclear receptor interacting factor 1 |
| chr20_+_56136136 | 1.23 |
ENST00000319441.4 ENST00000543666.1 |
PCK1 |
phosphoenolpyruvate carboxykinase 1 (soluble) |
| chr7_+_150065278 | 1.12 |
ENST00000519397.1 ENST00000479668.1 ENST00000540729.1 |
REPIN1 |
replication initiator 1 |
| chr16_+_82068830 | 1.08 |
ENST00000199936.4 |
HSD17B2 |
hydroxysteroid (17-beta) dehydrogenase 2 |
| chr1_+_10292308 | 0.96 |
ENST00000377081.1 |
KIF1B |
kinesin family member 1B |
| chrX_-_70331298 | 0.94 |
ENST00000456850.2 ENST00000473378.1 ENST00000487883.1 ENST00000374202.2 |
IL2RG |
interleukin 2 receptor, gamma |
| chr3_+_193853927 | 0.86 |
ENST00000232424.3 |
HES1 |
hes family bHLH transcription factor 1 |
| chr15_-_80263506 | 0.83 |
ENST00000335661.6 |
BCL2A1 |
BCL2-related protein A1 |
| chr2_+_196313239 | 0.76 |
ENST00000413290.1 |
AC064834.1 |
AC064834.1 |
| chr8_-_124428569 | 0.76 |
ENST00000521903.1 |
ATAD2 |
ATPase family, AAA domain containing 2 |
| chr6_+_24775153 | 0.66 |
ENST00000356509.3 ENST00000230056.3 |
GMNN |
geminin, DNA replication inhibitor |
| chrX_+_9431324 | 0.65 |
ENST00000407597.2 ENST00000424279.1 ENST00000536365.1 ENST00000441088.1 ENST00000380961.1 ENST00000415293.1 |
TBL1X |
transducin (beta)-like 1X-linked |
| chr15_-_45670924 | 0.62 |
ENST00000396659.3 |
GATM |
glycine amidinotransferase (L-arginine:glycine amidinotransferase) |
| chr6_-_31620455 | 0.61 |
ENST00000437771.1 ENST00000404765.2 ENST00000375964.6 ENST00000211379.5 |
BAG6 |
BCL2-associated athanogene 6 |
| chr3_-_194072019 | 0.58 |
ENST00000429275.1 ENST00000323830.3 |
CPN2 |
carboxypeptidase N, polypeptide 2 |
| chr6_-_31620403 | 0.56 |
ENST00000451898.1 ENST00000439687.2 ENST00000362049.6 ENST00000424480.1 |
BAG6 |
BCL2-associated athanogene 6 |
| chr8_+_81397876 | 0.54 |
ENST00000430430.1 |
ZBTB10 |
zinc finger and BTB domain containing 10 |
| chr1_+_215747118 | 0.54 |
ENST00000448333.1 |
KCTD3 |
potassium channel tetramerization domain containing 3 |
| chr10_-_61122220 | 0.50 |
ENST00000422313.2 ENST00000435852.2 ENST00000442566.3 ENST00000373868.2 ENST00000277705.6 ENST00000373867.3 ENST00000419214.2 |
FAM13C |
family with sequence similarity 13, member C |
| chr5_+_156696362 | 0.49 |
ENST00000377576.3 |
CYFIP2 |
cytoplasmic FMR1 interacting protein 2 |
| chr1_-_149783914 | 0.43 |
ENST00000369167.1 ENST00000427880.2 ENST00000545683.1 |
HIST2H2BF |
histone cluster 2, H2bf |
| chr3_+_46616017 | 0.42 |
ENST00000542931.1 |
TDGF1 |
teratocarcinoma-derived growth factor 1 |
| chrX_+_30261847 | 0.41 |
ENST00000378981.3 ENST00000397550.1 |
MAGEB1 |
melanoma antigen family B, 1 |
| chr6_+_13272904 | 0.40 |
ENST00000379335.3 ENST00000379329.1 |
PHACTR1 |
phosphatase and actin regulator 1 |
| chr12_+_66218598 | 0.40 |
ENST00000541363.1 |
HMGA2 |
high mobility group AT-hook 2 |
| chr7_+_95115210 | 0.37 |
ENST00000428113.1 ENST00000325885.5 |
ASB4 |
ankyrin repeat and SOCS box containing 4 |
| chr4_-_38806404 | 0.37 |
ENST00000308979.2 ENST00000505940.1 ENST00000515861.1 |
TLR1 |
toll-like receptor 1 |
| chr15_-_20193370 | 0.35 |
ENST00000558565.2 |
IGHV3OR15-7 |
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr14_+_23790655 | 0.35 |
ENST00000397276.2 |
PABPN1 |
poly(A) binding protein, nuclear 1 |
| chr5_-_138210977 | 0.35 |
ENST00000274711.6 ENST00000521094.2 |
LRRTM2 |
leucine rich repeat transmembrane neuronal 2 |
| chr4_+_119606523 | 0.34 |
ENST00000388822.5 ENST00000506780.1 ENST00000508801.1 |
METTL14 |
methyltransferase like 14 |
| chr4_+_158142750 | 0.32 |
ENST00000505888.1 ENST00000449365.1 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr16_+_69373323 | 0.30 |
ENST00000254940.5 |
NIP7 |
NIP7, nucleolar pre-rRNA processing protein |
| chr16_-_25026641 | 0.30 |
ENST00000289968.6 ENST00000303665.5 ENST00000455311.2 ENST00000441763.2 |
ARHGAP17 |
Rho GTPase activating protein 17 |
| chr1_-_173793458 | 0.29 |
ENST00000356198.2 |
CENPL |
centromere protein L |
| chr4_+_158141899 | 0.29 |
ENST00000264426.9 ENST00000506284.1 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr19_-_40336969 | 0.28 |
ENST00000599134.1 ENST00000597634.1 ENST00000598417.1 ENST00000601274.1 ENST00000594309.1 ENST00000221801.3 |
FBL |
fibrillarin |
| chr9_+_79792410 | 0.28 |
ENST00000357409.5 |
VPS13A |
vacuolar protein sorting 13 homolog A (S. cerevisiae) |
| chr16_-_5083917 | 0.28 |
ENST00000312251.3 ENST00000381955.3 |
NAGPA |
N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase |
| chr5_+_54320078 | 0.27 |
ENST00000231009.2 |
GZMK |
granzyme K (granzyme 3; tryptase II) |
| chr2_+_171034646 | 0.26 |
ENST00000409044.3 ENST00000408978.4 |
MYO3B |
myosin IIIB |
| chr4_-_153332886 | 0.26 |
ENST00000603841.1 |
FBXW7 |
F-box and WD repeat domain containing 7, E3 ubiquitin protein ligase |
| chrX_-_118739835 | 0.26 |
ENST00000542113.1 ENST00000304449.5 |
NKRF |
NFKB repressing factor |
| chr8_-_86253888 | 0.25 |
ENST00000522389.1 ENST00000432364.2 ENST00000517618.1 |
CA1 |
carbonic anhydrase I |
| chr17_+_34842473 | 0.25 |
ENST00000490126.2 ENST00000225410.4 |
ZNHIT3 |
zinc finger, HIT-type containing 3 |
| chr17_+_34842512 | 0.25 |
ENST00000588253.1 ENST00000592616.1 ENST00000590858.1 ENST00000588357.1 |
ZNHIT3 |
zinc finger, HIT-type containing 3 |
| chr1_+_74701062 | 0.24 |
ENST00000326637.3 |
TNNI3K |
TNNI3 interacting kinase |
| chr17_+_37856299 | 0.24 |
ENST00000269571.5 |
ERBB2 |
v-erb-b2 avian erythroblastic leukemia viral oncogene homolog 2 |
| chr11_+_59824127 | 0.23 |
ENST00000278865.3 |
MS4A3 |
membrane-spanning 4-domains, subfamily A, member 3 (hematopoietic cell-specific) |
| chr2_-_70520539 | 0.23 |
ENST00000482975.2 ENST00000438261.1 |
SNRPG |
small nuclear ribonucleoprotein polypeptide G |
| chr14_+_75536280 | 0.22 |
ENST00000238686.8 |
ZC2HC1C |
zinc finger, C2HC-type containing 1C |
| chr1_+_16083154 | 0.22 |
ENST00000375771.1 |
FBLIM1 |
filamin binding LIM protein 1 |
| chr4_+_187187098 | 0.21 |
ENST00000403665.2 ENST00000264692.4 |
F11 |
coagulation factor XI |
| chrX_+_19373700 | 0.20 |
ENST00000379804.1 |
PDHA1 |
pyruvate dehydrogenase (lipoamide) alpha 1 |
| chr11_-_118305921 | 0.20 |
ENST00000532619.1 |
RP11-770J1.4 |
RP11-770J1.4 |
| chr2_+_138721850 | 0.20 |
ENST00000329366.4 ENST00000280097.3 |
HNMT |
histamine N-methyltransferase |
| chr4_-_70505358 | 0.19 |
ENST00000457664.2 ENST00000604629.1 ENST00000604021.1 |
UGT2A2 |
UDP glucuronosyltransferase 2 family, polypeptide A2 |
| chr1_-_190446759 | 0.19 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr1_+_67673297 | 0.18 |
ENST00000425614.1 ENST00000395227.1 |
IL23R |
interleukin 23 receptor |
| chr14_+_75536335 | 0.18 |
ENST00000554763.1 ENST00000439583.2 ENST00000526130.1 ENST00000525046.1 |
ZC2HC1C |
zinc finger, C2HC-type containing 1C |
| chr2_+_242577097 | 0.18 |
ENST00000419606.1 ENST00000474739.2 ENST00000396411.3 ENST00000425239.1 ENST00000400771.3 ENST00000430617.2 |
ATG4B |
autophagy related 4B, cysteine peptidase |
| chr6_-_27841289 | 0.17 |
ENST00000355981.2 |
HIST1H4L |
histone cluster 1, H4l |
| chr2_+_168675182 | 0.16 |
ENST00000305861.1 |
B3GALT1 |
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 1 |
| chr1_+_110527308 | 0.16 |
ENST00000369799.5 |
AHCYL1 |
adenosylhomocysteinase-like 1 |
| chr16_-_20367584 | 0.15 |
ENST00000570689.1 |
UMOD |
uromodulin |
| chr14_+_29236269 | 0.15 |
ENST00000313071.4 |
FOXG1 |
forkhead box G1 |
| chr4_+_114214125 | 0.15 |
ENST00000509550.1 |
ANK2 |
ankyrin 2, neuronal |
| chr5_-_142814241 | 0.14 |
ENST00000504572.1 |
NR3C1 |
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr4_-_80247162 | 0.14 |
ENST00000286794.4 |
NAA11 |
N(alpha)-acetyltransferase 11, NatA catalytic subunit |
| chr1_+_206516200 | 0.13 |
ENST00000295713.5 |
SRGAP2 |
SLIT-ROBO Rho GTPase activating protein 2 |
| chr1_-_207095324 | 0.13 |
ENST00000530505.1 ENST00000367091.3 ENST00000442471.2 |
FAIM3 |
Fas apoptotic inhibitory molecule 3 |
| chr12_+_53773944 | 0.12 |
ENST00000551969.1 ENST00000327443.4 |
SP1 |
Sp1 transcription factor |
| chr21_+_43823983 | 0.11 |
ENST00000291535.6 ENST00000450356.1 ENST00000319294.6 ENST00000398367.1 |
UBASH3A |
ubiquitin associated and SH3 domain containing A |
| chr19_-_38720294 | 0.11 |
ENST00000412732.1 ENST00000456296.1 |
DPF1 |
D4, zinc and double PHD fingers family 1 |
| chr3_+_108541545 | 0.11 |
ENST00000295756.6 |
TRAT1 |
T cell receptor associated transmembrane adaptor 1 |
| chr4_+_158141843 | 0.11 |
ENST00000509417.1 ENST00000296526.7 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr14_+_22931924 | 0.10 |
ENST00000390477.2 |
TRDC |
T cell receptor delta constant |
| chr12_+_32655048 | 0.10 |
ENST00000427716.2 ENST00000266482.3 |
FGD4 |
FYVE, RhoGEF and PH domain containing 4 |
| chr2_-_109605663 | 0.10 |
ENST00000409271.1 ENST00000258443.2 ENST00000376651.1 |
EDAR |
ectodysplasin A receptor |
| chr2_-_166060552 | 0.10 |
ENST00000283254.7 ENST00000453007.1 |
SCN3A |
sodium channel, voltage-gated, type III, alpha subunit |
| chr3_-_15140629 | 0.09 |
ENST00000507357.1 ENST00000449050.1 ENST00000253699.3 ENST00000435849.3 ENST00000476527.2 |
ZFYVE20 |
zinc finger, FYVE domain containing 20 |
| chr17_-_2614927 | 0.09 |
ENST00000435359.1 |
CLUH |
clustered mitochondria (cluA/CLU1) homolog |
| chr6_-_53013620 | 0.09 |
ENST00000259803.7 |
GCM1 |
glial cells missing homolog 1 (Drosophila) |
| chr17_-_26127525 | 0.09 |
ENST00000313735.6 |
NOS2 |
nitric oxide synthase 2, inducible |
| chr22_-_36013368 | 0.08 |
ENST00000442617.1 ENST00000397326.2 ENST00000397328.1 ENST00000451685.1 |
MB |
myoglobin |
| chr8_+_24151553 | 0.08 |
ENST00000265769.4 ENST00000540823.1 ENST00000397649.3 |
ADAM28 |
ADAM metallopeptidase domain 28 |
| chr4_-_168155417 | 0.08 |
ENST00000511269.1 ENST00000506697.1 ENST00000512042.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr16_-_4401258 | 0.07 |
ENST00000577031.1 |
PAM16 |
presequence translocase-associated motor 16 homolog (S. cerevisiae) |
| chr11_+_98891797 | 0.07 |
ENST00000527185.1 ENST00000528682.1 ENST00000524871.1 |
CNTN5 |
contactin 5 |
| chr3_-_58419537 | 0.06 |
ENST00000474765.1 ENST00000485460.1 ENST00000302746.6 ENST00000383714.4 |
PDHB |
pyruvate dehydrogenase (lipoamide) beta |
| chr21_-_31538971 | 0.06 |
ENST00000286808.3 |
CLDN17 |
claudin 17 |
| chr1_-_156698181 | 0.05 |
ENST00000313146.6 |
ISG20L2 |
interferon stimulated exonuclease gene 20kDa-like 2 |
| chrX_-_100129320 | 0.05 |
ENST00000372966.3 |
NOX1 |
NADPH oxidase 1 |
| chr1_+_113010056 | 0.05 |
ENST00000369686.5 |
WNT2B |
wingless-type MMTV integration site family, member 2B |
| chr6_-_136847610 | 0.05 |
ENST00000454590.1 ENST00000432797.2 |
MAP7 |
microtubule-associated protein 7 |
| chr17_+_44590076 | 0.05 |
ENST00000333412.3 |
LRRC37A2 |
leucine rich repeat containing 37, member A2 |
| chr19_+_13134772 | 0.04 |
ENST00000587760.1 ENST00000585575.1 |
NFIX |
nuclear factor I/X (CCAAT-binding transcription factor) |
| chr19_-_38720354 | 0.03 |
ENST00000416611.1 |
DPF1 |
D4, zinc and double PHD fingers family 1 |
| chr7_-_150020578 | 0.03 |
ENST00000478393.1 |
ACTR3C |
ARP3 actin-related protein 3 homolog C (yeast) |
| chr12_-_102513843 | 0.02 |
ENST00000551744.2 ENST00000552283.1 |
NUP37 |
nucleoporin 37kDa |
| chr7_-_107770794 | 0.02 |
ENST00000205386.4 ENST00000418464.1 ENST00000388781.3 ENST00000388780.3 ENST00000414450.2 |
LAMB4 |
laminin, beta 4 |
| chr12_+_66218212 | 0.01 |
ENST00000393578.3 ENST00000425208.2 ENST00000536545.1 ENST00000354636.3 |
HMGA2 |
high mobility group AT-hook 2 |
| chr3_+_148457585 | 0.01 |
ENST00000402260.1 |
AGTR1 |
angiotensin II receptor, type 1 |
| chr14_-_75536182 | 0.01 |
ENST00000555463.1 |
ACYP1 |
acylphosphatase 1, erythrocyte (common) type |
| chr15_+_57998923 | 0.01 |
ENST00000380557.4 |
POLR2M |
polymerase (RNA) II (DNA directed) polypeptide M |
| chr3_-_81792780 | 0.01 |
ENST00000489715.1 |
GBE1 |
glucan (1,4-alpha-), branching enzyme 1 |
| chr4_-_120243545 | 0.01 |
ENST00000274024.3 |
FABP2 |
fatty acid binding protein 2, intestinal |
| chr4_-_71705060 | 0.01 |
ENST00000514161.1 |
GRSF1 |
G-rich RNA sequence binding factor 1 |
| chr10_+_96522361 | 0.01 |
ENST00000371321.3 |
CYP2C19 |
cytochrome P450, family 2, subfamily C, polypeptide 19 |
| chr10_+_46997926 | 0.01 |
ENST00000374314.4 |
GPRIN2 |
G protein regulated inducer of neurite outgrowth 2 |
| chr3_-_113160334 | 0.01 |
ENST00000393845.2 ENST00000295868.2 |
WDR52 |
WD repeat domain 52 |
| chr1_-_157811588 | 0.01 |
ENST00000368174.4 |
CD5L |
CD5 molecule-like |
| chr14_-_106312010 | 0.00 |
ENST00000390556.2 |
IGHD |
immunoglobulin heavy constant delta |
| chr3_-_11645925 | 0.00 |
ENST00000413604.1 |
VGLL4 |
vestigial like 4 (Drosophila) |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 4.1 | GO:0003331 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.8 | 2.3 | GO:2000864 | estrogen secretion(GO:0035937) estradiol secretion(GO:0035938) regulation of estrogen secretion(GO:2000861) regulation of estradiol secretion(GO:2000864) |
| 0.5 | 1.5 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.3 | 1.0 | GO:1904647 | response to rotenone(GO:1904647) |
| 0.3 | 1.2 | GO:0006114 | glycerol biosynthetic process(GO:0006114) response to methionine(GO:1904640) |
| 0.3 | 1.2 | GO:1904327 | maintenance of unfolded protein(GO:0036506) protein localization to cytosolic proteasome complex(GO:1904327) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) protein localization to cytosolic proteasome complex involved in ERAD pathway(GO:1904379) |
| 0.3 | 0.9 | GO:0021558 | midbrain-hindbrain boundary morphogenesis(GO:0021555) trochlear nerve development(GO:0021558) regulation of timing of neuron differentiation(GO:0060164) |
| 0.1 | 0.4 | GO:0031049 | mesodermal-endodermal cell signaling(GO:0003131) programmed DNA elimination(GO:0031049) chromosome breakage(GO:0031052) histone H2A-S139 phosphorylation(GO:0035978) positive regulation of cellular response to X-ray(GO:2000685) |
| 0.1 | 0.9 | GO:0038110 | interleukin-2-mediated signaling pathway(GO:0038110) |
| 0.1 | 0.4 | GO:0042495 | detection of bacterial lipoprotein(GO:0042494) detection of triacyl bacterial lipopeptide(GO:0042495) detection of bacterial lipopeptide(GO:0070340) |
| 0.1 | 0.4 | GO:1904247 | positive regulation of polynucleotide adenylyltransferase activity(GO:1904247) |
| 0.1 | 0.7 | GO:0071163 | DNA replication preinitiation complex assembly(GO:0071163) |
| 0.1 | 1.1 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.1 | 0.8 | GO:0031936 | negative regulation of chromatin silencing(GO:0031936) |
| 0.1 | 0.3 | GO:0018364 | peptidyl-glutamine methylation(GO:0018364) |
| 0.1 | 0.3 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.1 | 0.5 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.1 | 0.2 | GO:0051697 | protein delipidation(GO:0051697) |
| 0.1 | 0.6 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.1 | 0.3 | GO:2000639 | regulation of SREBP signaling pathway(GO:2000638) negative regulation of SREBP signaling pathway(GO:2000639) |
| 0.1 | 0.2 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 0.0 | 0.1 | GO:0036371 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.0 | 0.2 | GO:0038155 | interleukin-23-mediated signaling pathway(GO:0038155) |
| 0.0 | 0.1 | GO:1900239 | phenotypic switching(GO:0036166) regulation of phenotypic switching(GO:1900239) |
| 0.0 | 0.1 | GO:0018002 | N-terminal peptidyl-serine acetylation(GO:0017198) N-terminal peptidyl-glutamic acid acetylation(GO:0018002) peptidyl-serine acetylation(GO:0030920) |
| 0.0 | 0.2 | GO:0001692 | histamine metabolic process(GO:0001692) |
| 0.0 | 0.8 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.0 | 0.2 | GO:0072233 | thick ascending limb development(GO:0072023) metanephric thick ascending limb development(GO:0072233) |
| 0.0 | 0.2 | GO:0033088 | negative regulation of immature T cell proliferation in thymus(GO:0033088) |
| 0.0 | 0.4 | GO:0002091 | negative regulation of receptor internalization(GO:0002091) |
| 0.0 | 0.3 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 0.0 | 0.1 | GO:0060018 | astrocyte fate commitment(GO:0060018) |
| 0.0 | 0.4 | GO:2001214 | positive regulation of vasculogenesis(GO:2001214) |
| 0.0 | 0.4 | GO:0008595 | tripartite regional subdivision(GO:0007351) anterior/posterior axis specification, embryo(GO:0008595) |
| 0.0 | 0.3 | GO:0033299 | secretion of lysosomal enzymes(GO:0033299) |
| 0.0 | 0.2 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.0 | 0.2 | GO:0006682 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.0 | 0.1 | GO:0043402 | glucocorticoid mediated signaling pathway(GO:0043402) |
| 0.0 | 0.1 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 0.7 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.0 | 0.2 | GO:0033623 | regulation of integrin activation(GO:0033623) |
| 0.0 | 0.1 | GO:0021815 | modulation of microtubule cytoskeleton involved in cerebral cortex radial glia guided migration(GO:0021815) |
| 0.0 | 0.1 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.0 | 0.3 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.0 | 0.1 | GO:0043353 | enucleate erythrocyte differentiation(GO:0043353) |
| 0.0 | 0.2 | GO:0086069 | bundle of His cell to Purkinje myocyte communication(GO:0086069) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.0 | 0.9 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.4 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.0 | 0.7 | REACTOME ASSOCIATION OF LICENSING FACTORS WITH THE PRE REPLICATIVE COMPLEX | Genes involved in Association of licensing factors with the pre-replicative complex |
| 0.0 | 0.7 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.0 | 1.8 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 0.4 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.2 | REACTOME PROCESSING OF CAPPED INTRONLESS PRE MRNA | Genes involved in Processing of Capped Intronless Pre-mRNA |
| 0.0 | 0.3 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 2.3 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.3 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.0 | 0.4 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 0.4 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.2 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 0.2 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.3 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.0 | 0.9 | ST INTERLEUKIN 4 PATHWAY | Interleukin 4 (IL-4) Pathway |
| 0.0 | 1.5 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.5 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.9 | PID FANCONI PATHWAY | Fanconi anemia pathway |
| 0.0 | 0.4 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.5 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.0 | 0.8 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 0.4 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 0.5 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.5 | GO:0034673 | inhibin-betaglycan-ActRII complex(GO:0034673) |
| 0.3 | 1.2 | GO:0072379 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 0.7 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) |
| 0.1 | 0.4 | GO:0035354 | Toll-like receptor 1-Toll-like receptor 2 protein complex(GO:0035354) |
| 0.1 | 0.3 | GO:0036396 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 0.1 | 0.3 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.1 | 1.1 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.1 | 0.4 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.0 | 0.2 | GO:0072536 | interleukin-23 receptor complex(GO:0072536) |
| 0.0 | 0.4 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.0 | 0.3 | GO:1990452 | Parkin-FBXW7-Cul1 ubiquitin ligase complex(GO:1990452) |
| 0.0 | 0.2 | GO:0005683 | U7 snRNP(GO:0005683) |
| 0.0 | 0.3 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 0.0 | 0.4 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.0 | 1.0 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.0 | 0.1 | GO:0031415 | NatA complex(GO:0031415) |
| 0.0 | 0.1 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 0.0 | 0.3 | GO:0031045 | dense core granule(GO:0031045) |
| 0.0 | 0.1 | GO:0044327 | dendritic spine head(GO:0044327) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) phosphoenolpyruvate carboxykinase (GTP) activity(GO:0004613) |
| 0.3 | 0.9 | GO:0004913 | interleukin-4 receptor activity(GO:0004913) interleukin-7 receptor activity(GO:0004917) |
| 0.3 | 1.1 | GO:0047006 | 17-alpha,20-alpha-dihydroxypregn-4-en-3-one dehydrogenase activity(GO:0047006) |
| 0.2 | 4.7 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.2 | 1.5 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 0.1 | 0.4 | GO:0035501 | MH1 domain binding(GO:0035501) |
| 0.1 | 0.4 | GO:0035663 | Toll-like receptor 2 binding(GO:0035663) |
| 0.1 | 0.3 | GO:0016422 | mRNA (2'-O-methyladenosine-N6-)-methyltransferase activity(GO:0016422) |
| 0.1 | 1.2 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 0.3 | GO:0004739 | pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.1 | 0.7 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.1 | 0.2 | GO:0047275 | glucosaminylgalactosylglucosylceramide beta-galactosyltransferase activity(GO:0047275) |
| 0.0 | 0.2 | GO:0042020 | interleukin-23 binding(GO:0042019) interleukin-23 receptor activity(GO:0042020) |
| 0.0 | 0.8 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 0.1 | GO:1990190 | peptide-glutamate-N-acetyltransferase activity(GO:1990190) |
| 0.0 | 0.3 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.0 | 0.3 | GO:0004064 | arylesterase activity(GO:0004064) |
| 0.0 | 0.1 | GO:0004883 | glucocorticoid receptor activity(GO:0004883) glucocorticoid-activated RNA polymerase II transcription factor binding transcription factor activity(GO:0038051) |
| 0.0 | 0.2 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.4 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 1.3 | GO:0042974 | retinoic acid receptor binding(GO:0042974) |
| 0.0 | 0.6 | GO:0016769 | transferase activity, transferring nitrogenous groups(GO:0016769) |
| 0.0 | 0.2 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.0 | 0.1 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 0.0 | 0.2 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.0 | 0.3 | GO:0008649 | rRNA methyltransferase activity(GO:0008649) |
| 0.0 | 1.0 | GO:1990939 | ATP-dependent microtubule motor activity(GO:1990939) |
| 0.0 | 0.2 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.0 | 0.4 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.5 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.0 | 0.3 | GO:0015929 | hexosaminidase activity(GO:0015929) |
| 0.0 | 1.7 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 0.9 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.4 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.2 | GO:0031005 | filamin binding(GO:0031005) |