Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
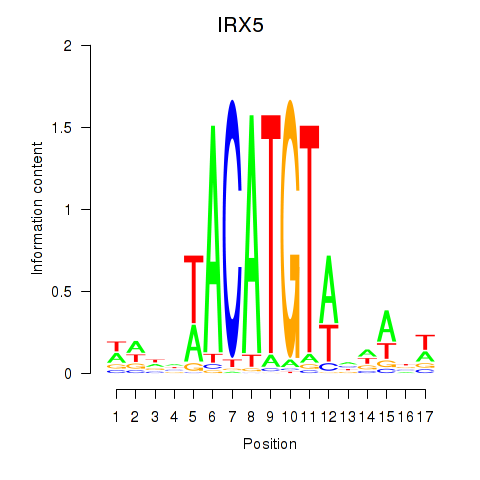
Results for IRX5
Z-value: 0.66
Transcription factors associated with IRX5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
IRX5
|
ENSG00000176842.10 | IRX5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| IRX5 | hg19_v2_chr16_+_54964740_54964789 | -0.03 | 9.2e-01 | Click! |
Activity profile of IRX5 motif
Sorted Z-values of IRX5 motif
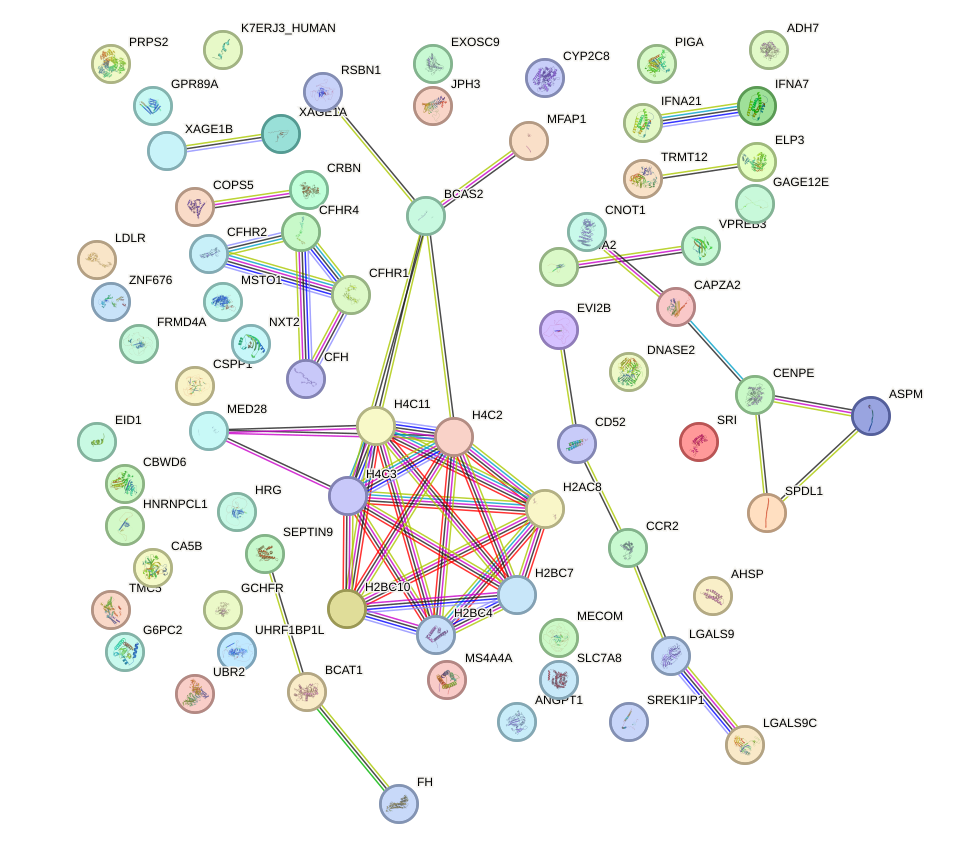
Network of associatons between targets according to the STRING database.
First level regulatory network of IRX5
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr13_-_46716969 | 4.10 |
ENST00000435666.2 |
LCP1 |
lymphocyte cytosolic protein 1 (L-plastin) |
| chrX_+_49294472 | 3.34 |
ENST00000361446.5 |
GAGE12B |
G antigen 12B |
| chrX_-_52258669 | 2.47 |
ENST00000441417.1 |
XAGE1A |
X antigen family, member 1A |
| chrX_+_52513455 | 2.36 |
ENST00000446098.1 |
XAGE1C |
X antigen family, member 1C |
| chr8_-_108510224 | 1.88 |
ENST00000517746.1 ENST00000297450.3 |
ANGPT1 |
angiopoietin 1 |
| chrX_+_52240504 | 1.57 |
ENST00000399805.2 |
XAGE1B |
X antigen family, member 1B |
| chr1_+_26644441 | 1.47 |
ENST00000374213.2 |
CD52 |
CD52 molecule |
| chr6_-_26216872 | 1.38 |
ENST00000244601.3 |
HIST1H2BG |
histone cluster 1, H2bg |
| chr7_-_87856280 | 1.20 |
ENST00000490437.1 ENST00000431660.1 |
SRI |
sorcin |
| chr7_-_87856303 | 1.14 |
ENST00000394641.3 |
SRI |
sorcin |
| chr11_+_60223225 | 0.95 |
ENST00000524807.1 ENST00000345732.4 |
MS4A1 |
membrane-spanning 4-domains, subfamily A, member 1 |
| chr16_+_31539197 | 0.92 |
ENST00000564707.1 |
AHSP |
alpha hemoglobin stabilizing protein |
| chr15_+_49170083 | 0.84 |
ENST00000530028.2 |
EID1 |
EP300 interacting inhibitor of differentiation 1 |
| chr17_-_29641104 | 0.82 |
ENST00000577894.1 ENST00000330927.4 |
EVI2B |
ecotropic viral integration site 2B |
| chrX_+_12809463 | 0.80 |
ENST00000380663.3 ENST00000380668.5 ENST00000398491.2 ENST00000489404.1 |
PRPS2 |
phosphoribosyl pyrophosphate synthetase 2 |
| chr16_+_31539183 | 0.78 |
ENST00000302312.4 |
AHSP |
alpha hemoglobin stabilizing protein |
| chr15_+_41057818 | 0.77 |
ENST00000558467.1 |
GCHFR |
GTP cyclohydrolase I feedback regulator |
| chr11_+_60048053 | 0.73 |
ENST00000337908.4 |
MS4A4A |
membrane-spanning 4-domains, subfamily A, member 4A |
| chr1_-_241683001 | 0.63 |
ENST00000366560.3 |
FH |
fumarate hydratase |
| chr17_-_29641084 | 0.62 |
ENST00000544462.1 |
EVI2B |
ecotropic viral integration site 2B |
| chr2_-_183106641 | 0.60 |
ENST00000346717.4 |
PDE1A |
phosphodiesterase 1A, calmodulin-dependent |
| chr3_-_168865522 | 0.59 |
ENST00000464456.1 |
MECOM |
MDS1 and EVI1 complex locus |
| chr12_+_32655048 | 0.56 |
ENST00000427716.2 ENST00000266482.3 |
FGD4 |
FYVE, RhoGEF and PH domain containing 4 |
| chr17_+_18380051 | 0.54 |
ENST00000581545.1 ENST00000582333.1 ENST00000328114.6 ENST00000412421.2 ENST00000583322.1 ENST00000584941.1 |
LGALS9C |
lectin, galactoside-binding, soluble, 9C |
| chr14_-_23623577 | 0.54 |
ENST00000422941.2 ENST00000453702.1 |
SLC7A8 |
solute carrier family 7 (amino acid transporter light chain, L system), member 8 |
| chr14_-_106471723 | 0.53 |
ENST00000390595.2 |
IGHV1-3 |
immunoglobulin heavy variable 1-3 |
| chr2_+_58655461 | 0.53 |
ENST00000429095.1 ENST00000429664.1 ENST00000452840.1 |
AC007092.1 |
long intergenic non-protein coding RNA 1122 |
| chr11_+_31531291 | 0.52 |
ENST00000350638.5 ENST00000379163.5 ENST00000395934.2 |
ELP4 |
elongator acetyltransferase complex subunit 4 |
| chr12_-_25102252 | 0.51 |
ENST00000261192.7 |
BCAT1 |
branched chain amino-acid transaminase 1, cytosolic |
| chr2_+_160590469 | 0.51 |
ENST00000409591.1 |
MARCH7 |
membrane-associated ring finger (C3HC4) 7, E3 ubiquitin protein ligase |
| chr8_-_101962777 | 0.50 |
ENST00000395951.3 |
YWHAZ |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta |
| chr14_-_106963409 | 0.49 |
ENST00000390621.2 |
IGHV1-45 |
immunoglobulin heavy variable 1-45 |
| chr6_+_42584847 | 0.49 |
ENST00000372883.3 |
UBR2 |
ubiquitin protein ligase E3 component n-recognin 2 |
| chr6_+_26217159 | 0.48 |
ENST00000303910.2 |
HIST1H2AE |
histone cluster 1, H2ae |
| chr8_+_27947746 | 0.46 |
ENST00000521015.1 ENST00000521570.1 |
ELP3 |
elongator acetyltransferase complex subunit 3 |
| chr8_+_125463048 | 0.45 |
ENST00000328599.3 |
TRMT12 |
tRNA methyltransferase 12 homolog (S. cerevisiae) |
| chr16_+_87636474 | 0.45 |
ENST00000284262.2 |
JPH3 |
junctophilin 3 |
| chr4_+_122722466 | 0.45 |
ENST00000243498.5 ENST00000379663.3 ENST00000509800.1 |
EXOSC9 |
exosome component 9 |
| chr14_-_106845789 | 0.44 |
ENST00000390617.2 |
IGHV3-35 |
immunoglobulin heavy variable 3-35 (non-functional) |
| chr8_-_67976509 | 0.43 |
ENST00000518747.1 |
COPS5 |
COP9 signalosome subunit 5 |
| chr3_+_173116225 | 0.40 |
ENST00000457714.1 |
NLGN1 |
neuroligin 1 |
| chr21_-_34185944 | 0.39 |
ENST00000479548.1 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr4_+_17616253 | 0.39 |
ENST00000237380.7 |
MED28 |
mediator complex subunit 28 |
| chr15_-_64665911 | 0.38 |
ENST00000606793.1 ENST00000561349.1 ENST00000560278.1 |
CTD-2116N17.1 |
Uncharacterized protein |
| chr17_-_47723943 | 0.38 |
ENST00000510476.1 ENST00000503676.1 |
SPOP |
speckle-type POZ protein |
| chr17_+_25958174 | 0.37 |
ENST00000313648.6 ENST00000577392.1 ENST00000584661.1 ENST00000413914.2 |
LGALS9 |
lectin, galactoside-binding, soluble, 9 |
| chr8_+_84824920 | 0.37 |
ENST00000523678.1 |
RP11-120I21.2 |
RP11-120I21.2 |
| chr4_-_104119528 | 0.37 |
ENST00000380026.3 ENST00000503705.1 ENST00000265148.3 |
CENPE |
centromere protein E, 312kDa |
| chr2_+_169757750 | 0.37 |
ENST00000375363.3 ENST00000429379.2 ENST00000421979.1 |
G6PC2 |
glucose-6-phosphatase, catalytic, 2 |
| chr14_-_23877474 | 0.36 |
ENST00000405093.3 |
MYH6 |
myosin, heavy chain 6, cardiac muscle, alpha |
| chr10_-_14050522 | 0.35 |
ENST00000342409.2 |
FRMD4A |
FERM domain containing 4A |
| chrX_-_15353629 | 0.35 |
ENST00000333590.4 ENST00000428964.1 ENST00000542278.1 |
PIGA |
phosphatidylinositol glycan anchor biosynthesis, class A |
| chr7_+_77469439 | 0.35 |
ENST00000450574.1 ENST00000416283.2 ENST00000248550.7 |
PHTF2 |
putative homeodomain transcription factor 2 |
| chr5_+_169010638 | 0.34 |
ENST00000265295.4 ENST00000506574.1 ENST00000515224.1 ENST00000508247.1 ENST00000513941.1 |
SPDL1 |
spindle apparatus coiled-coil protein 1 |
| chr15_-_44116873 | 0.34 |
ENST00000267812.3 |
MFAP1 |
microfibrillar-associated protein 1 |
| chr4_-_100356844 | 0.33 |
ENST00000437033.2 |
ADH7 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide |
| chr1_-_197115818 | 0.32 |
ENST00000367409.4 ENST00000294732.7 |
ASPM |
asp (abnormal spindle) homolog, microcephaly associated (Drosophila) |
| chr1_-_114355083 | 0.32 |
ENST00000261441.5 |
RSBN1 |
round spermatid basic protein 1 |
| chr3_+_52245458 | 0.32 |
ENST00000459884.1 |
ALAS1 |
aminolevulinate, delta-, synthase 1 |
| chr10_-_96829246 | 0.32 |
ENST00000371270.3 ENST00000535898.1 ENST00000539050.1 |
CYP2C8 |
cytochrome P450, family 2, subfamily C, polypeptide 8 |
| chr4_+_80584903 | 0.31 |
ENST00000506460.1 |
RP11-452C8.1 |
RP11-452C8.1 |
| chr17_+_58018269 | 0.31 |
ENST00000591035.1 |
RP11-178C3.1 |
Uncharacterized protein |
| chr5_-_64064508 | 0.31 |
ENST00000513458.4 |
SREK1IP1 |
SREK1-interacting protein 1 |
| chr19_-_12992244 | 0.31 |
ENST00000538460.1 |
DNASE2 |
deoxyribonuclease II, lysosomal |
| chr6_+_37012607 | 0.29 |
ENST00000423336.1 |
COX6A1P2 |
cytochrome c oxidase subunit VIa polypeptide 1 pseudogene 2 |
| chr3_+_186383741 | 0.27 |
ENST00000232003.4 |
HRG |
histidine-rich glycoprotein |
| chr13_-_88323218 | 0.27 |
ENST00000436290.2 ENST00000453832.2 ENST00000606590.1 |
MIR4500HG |
MIR4500 host gene (non-protein coding) |
| chr14_-_21737551 | 0.26 |
ENST00000554891.1 ENST00000555883.1 ENST00000553753.1 ENST00000555914.1 ENST00000557336.1 ENST00000555215.1 ENST00000556628.1 ENST00000555137.1 ENST00000556226.1 ENST00000555309.1 ENST00000556142.1 ENST00000554969.1 ENST00000554455.1 ENST00000556513.1 ENST00000557201.1 ENST00000420743.2 ENST00000557768.1 ENST00000553300.1 ENST00000554383.1 ENST00000554539.1 |
HNRNPC |
heterogeneous nuclear ribonucleoprotein C (C1/C2) |
| chr16_-_58585513 | 0.26 |
ENST00000245138.4 ENST00000567285.1 |
CNOT1 |
CCR4-NOT transcription complex, subunit 1 |
| chr1_-_115124257 | 0.26 |
ENST00000369541.3 |
BCAS2 |
breast carcinoma amplified sequence 2 |
| chr9_+_42704004 | 0.25 |
ENST00000457288.1 |
CBWD7 |
COBW domain containing 7 |
| chr3_-_3221358 | 0.25 |
ENST00000424814.1 ENST00000450014.1 ENST00000231948.4 ENST00000432408.2 |
CRBN |
cereblon |
| chr16_+_19467772 | 0.25 |
ENST00000219821.5 ENST00000561503.1 ENST00000564959.1 |
TMC5 |
transmembrane channel-like 5 |
| chr1_+_155579979 | 0.25 |
ENST00000452804.2 ENST00000538143.1 ENST00000245564.2 ENST00000368341.4 |
MSTO1 |
misato 1, mitochondrial distribution and morphology regulator |
| chr8_+_67976593 | 0.24 |
ENST00000262210.5 ENST00000412460.1 |
CSPP1 |
centrosome and spindle pole associated protein 1 |
| chr21_-_34186006 | 0.24 |
ENST00000490358.1 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr3_+_46395219 | 0.24 |
ENST00000445132.2 ENST00000292301.4 |
CCR2 |
chemokine (C-C motif) receptor 2 |
| chrX_+_108779004 | 0.23 |
ENST00000218004.1 |
NXT2 |
nuclear transport factor 2-like export factor 2 |
| chr9_-_21202204 | 0.23 |
ENST00000239347.3 |
IFNA7 |
interferon, alpha 7 |
| chr9_-_21166659 | 0.22 |
ENST00000380225.1 |
IFNA21 |
interferon, alpha 21 |
| chr19_-_22379753 | 0.22 |
ENST00000397121.2 |
ZNF676 |
zinc finger protein 676 |
| chrX_+_15767971 | 0.22 |
ENST00000479740.1 ENST00000454127.2 |
CA5B |
carbonic anhydrase VB, mitochondrial |
| chr17_+_75447326 | 0.22 |
ENST00000591088.1 |
SEPT9 |
septin 9 |
| chr17_-_42327236 | 0.22 |
ENST00000399246.2 |
AC003102.1 |
AC003102.1 |
| chr2_-_40657397 | 0.22 |
ENST00000408028.2 ENST00000332839.4 ENST00000406391.2 ENST00000542024.1 ENST00000542756.1 ENST00000405901.3 |
SLC8A1 |
solute carrier family 8 (sodium/calcium exchanger), member 1 |
| chr14_-_106967788 | 0.21 |
ENST00000390622.2 |
IGHV1-46 |
immunoglobulin heavy variable 1-46 |
| chr1_-_145826450 | 0.21 |
ENST00000462900.2 |
GPR89A |
G protein-coupled receptor 89A |
| chr19_-_12992274 | 0.20 |
ENST00000592506.1 ENST00000222219.3 |
DNASE2 |
deoxyribonuclease II, lysosomal |
| chr1_+_196857144 | 0.19 |
ENST00000367416.2 ENST00000251424.4 ENST00000367418.2 |
CFHR4 |
complement factor H-related 4 |
| chr1_+_196788887 | 0.18 |
ENST00000320493.5 ENST00000367424.4 ENST00000367421.3 |
CFHR1 CFHR2 |
complement factor H-related 1 complement factor H-related 2 |
| chr22_-_24096562 | 0.18 |
ENST00000398465.3 |
VPREB3 |
pre-B lymphocyte 3 |
| chr6_-_154677900 | 0.18 |
ENST00000265198.4 ENST00000520261.1 |
IPCEF1 |
interaction protein for cytohesin exchange factors 1 |
| chr12_-_100486668 | 0.18 |
ENST00000550544.1 ENST00000551980.1 ENST00000548045.1 ENST00000545232.2 ENST00000551973.1 |
UHRF1BP1L |
UHRF1 binding protein 1-like |
| chr21_-_34185989 | 0.18 |
ENST00000487113.1 ENST00000382373.4 |
C21orf62 |
chromosome 21 open reading frame 62 |
| chr7_+_116502527 | 0.18 |
ENST00000361183.3 |
CAPZA2 |
capping protein (actin filament) muscle Z-line, alpha 2 |
| chr3_-_9994021 | 0.17 |
ENST00000411976.2 ENST00000412055.1 |
PRRT3 |
proline-rich transmembrane protein 3 |
| chr6_+_26204825 | 0.17 |
ENST00000360441.4 |
HIST1H4E |
histone cluster 1, H4e |
| chr12_-_76462713 | 0.16 |
ENST00000552056.1 |
NAP1L1 |
nucleosome assembly protein 1-like 1 |
| chr14_+_51706886 | 0.16 |
ENST00000457354.2 |
TMX1 |
thioredoxin-related transmembrane protein 1 |
| chr15_+_66585879 | 0.15 |
ENST00000319212.4 |
DIS3L |
DIS3 mitotic control homolog (S. cerevisiae)-like |
| chr3_+_180586536 | 0.15 |
ENST00000465551.1 |
FXR1 |
fragile X mental retardation, autosomal homolog 1 |
| chr11_-_89653576 | 0.14 |
ENST00000420869.1 |
TRIM49D1 |
tripartite motif containing 49D1 |
| chr12_+_106751436 | 0.14 |
ENST00000228347.4 |
POLR3B |
polymerase (RNA) III (DNA directed) polypeptide B |
| chr6_+_27925019 | 0.14 |
ENST00000244623.1 |
OR2B6 |
olfactory receptor, family 2, subfamily B, member 6 |
| chr9_+_105757590 | 0.14 |
ENST00000374798.3 ENST00000487798.1 |
CYLC2 |
cylicin, basic protein of sperm head cytoskeleton 2 |
| chr1_+_196743912 | 0.14 |
ENST00000367425.4 |
CFHR3 |
complement factor H-related 3 |
| chr14_+_73563735 | 0.14 |
ENST00000532192.1 |
RBM25 |
RNA binding motif protein 25 |
| chr10_-_35379524 | 0.13 |
ENST00000374751.3 ENST00000374742.1 ENST00000602371.1 |
CUL2 |
cullin 2 |
| chr7_+_66386204 | 0.13 |
ENST00000341567.4 ENST00000607045.1 |
TMEM248 |
transmembrane protein 248 |
| chr11_-_22647350 | 0.12 |
ENST00000327470.3 |
FANCF |
Fanconi anemia, complementation group F |
| chr12_-_4553385 | 0.12 |
ENST00000543077.1 |
FGF6 |
fibroblast growth factor 6 |
| chr19_-_50266580 | 0.11 |
ENST00000246801.3 |
TSKS |
testis-specific serine kinase substrate |
| chr12_-_10282836 | 0.11 |
ENST00000304084.8 ENST00000353231.5 ENST00000525605.1 |
CLEC7A |
C-type lectin domain family 7, member A |
| chr9_-_37465396 | 0.10 |
ENST00000307750.4 |
ZBTB5 |
zinc finger and BTB domain containing 5 |
| chr21_-_43816052 | 0.10 |
ENST00000398405.1 |
TMPRSS3 |
transmembrane protease, serine 3 |
| chr4_+_76995855 | 0.09 |
ENST00000355810.4 ENST00000349321.3 |
ART3 |
ADP-ribosyltransferase 3 |
| chr8_+_12803176 | 0.09 |
ENST00000524591.2 |
KIAA1456 |
KIAA1456 |
| chr3_-_72897545 | 0.09 |
ENST00000325599.8 |
SHQ1 |
SHQ1, H/ACA ribonucleoprotein assembly factor |
| chr12_-_10282742 | 0.09 |
ENST00000298523.5 ENST00000396484.2 ENST00000310002.4 |
CLEC7A |
C-type lectin domain family 7, member A |
| chr2_+_113816215 | 0.09 |
ENST00000346807.3 |
IL36RN |
interleukin 36 receptor antagonist |
| chr1_+_196912902 | 0.09 |
ENST00000476712.2 ENST00000367415.5 |
CFHR2 |
complement factor H-related 2 |
| chr1_+_22778337 | 0.08 |
ENST00000404138.1 ENST00000400239.2 ENST00000375647.4 ENST00000374651.4 |
ZBTB40 |
zinc finger and BTB domain containing 40 |
| chr12_-_21757774 | 0.08 |
ENST00000261195.2 |
GYS2 |
glycogen synthase 2 (liver) |
| chr7_+_20686946 | 0.08 |
ENST00000443026.2 ENST00000406935.1 |
ABCB5 |
ATP-binding cassette, sub-family B (MDR/TAP), member 5 |
| chr18_-_32924372 | 0.08 |
ENST00000261332.6 ENST00000399061.3 |
ZNF24 |
zinc finger protein 24 |
| chr1_-_152086556 | 0.07 |
ENST00000368804.1 |
TCHH |
trichohyalin |
| chr17_+_3379284 | 0.07 |
ENST00000263080.2 |
ASPA |
aspartoacylase |
| chr18_+_6729725 | 0.07 |
ENST00000400091.2 ENST00000583410.1 ENST00000584387.1 |
ARHGAP28 |
Rho GTPase activating protein 28 |
| chr11_-_118305921 | 0.06 |
ENST00000532619.1 |
RP11-770J1.4 |
RP11-770J1.4 |
| chr11_+_61891445 | 0.06 |
ENST00000394818.3 ENST00000533896.1 ENST00000278849.4 |
INCENP |
inner centromere protein antigens 135/155kDa |
| chr13_+_96085847 | 0.06 |
ENST00000376873.3 |
CLDN10 |
claudin 10 |
| chr10_+_118350468 | 0.06 |
ENST00000358834.4 ENST00000528052.1 ENST00000442761.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr3_-_50378235 | 0.05 |
ENST00000357043.2 |
RASSF1 |
Ras association (RalGDS/AF-6) domain family member 1 |
| chr1_-_3566590 | 0.05 |
ENST00000424367.1 ENST00000378322.3 |
WRAP73 |
WD repeat containing, antisense to TP73 |
| chr1_+_10509971 | 0.05 |
ENST00000320498.4 |
CORT |
cortistatin |
| chr21_+_38792602 | 0.05 |
ENST00000398960.2 ENST00000398956.2 |
DYRK1A |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A |
| chr10_+_118350522 | 0.05 |
ENST00000530319.1 ENST00000527980.1 ENST00000471549.1 ENST00000534537.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr1_+_220267429 | 0.05 |
ENST00000366922.1 ENST00000302637.5 |
IARS2 |
isoleucyl-tRNA synthetase 2, mitochondrial |
| chr6_-_111804905 | 0.05 |
ENST00000358835.3 ENST00000435970.1 |
REV3L |
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr1_+_67632083 | 0.04 |
ENST00000347310.5 ENST00000371002.1 |
IL23R |
interleukin 23 receptor |
| chr6_+_146920116 | 0.04 |
ENST00000367493.3 |
ADGB |
androglobin |
| chr2_+_191792376 | 0.04 |
ENST00000409428.1 ENST00000409215.1 |
GLS |
glutaminase |
| chr3_+_53528659 | 0.04 |
ENST00000350061.5 |
CACNA1D |
calcium channel, voltage-dependent, L type, alpha 1D subunit |
| chr2_-_127963343 | 0.04 |
ENST00000335247.7 |
CYP27C1 |
cytochrome P450, family 27, subfamily C, polypeptide 1 |
| chr1_-_116383738 | 0.03 |
ENST00000320238.3 |
NHLH2 |
nescient helix loop helix 2 |
| chr6_+_29364416 | 0.03 |
ENST00000383555.2 |
OR12D2 |
olfactory receptor, family 12, subfamily D, member 2 (gene/pseudogene) |
| chr5_+_128300810 | 0.03 |
ENST00000262462.4 |
SLC27A6 |
solute carrier family 27 (fatty acid transporter), member 6 |
| chr6_-_111804393 | 0.03 |
ENST00000368802.3 ENST00000368805.1 |
REV3L |
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr14_+_50291993 | 0.03 |
ENST00000595378.1 |
AL627171.2 |
HCG1786899; PRO2610; Uncharacterized protein |
| chr2_+_173686303 | 0.03 |
ENST00000397087.3 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr1_+_196743943 | 0.03 |
ENST00000471440.2 ENST00000391985.3 |
CFHR3 |
complement factor H-related 3 |
| chr10_-_115904361 | 0.03 |
ENST00000428953.1 ENST00000543782.1 |
C10orf118 |
chromosome 10 open reading frame 118 |
| chr1_+_158969752 | 0.03 |
ENST00000566111.1 |
IFI16 |
interferon, gamma-inducible protein 16 |
| chr12_+_21679220 | 0.03 |
ENST00000256969.2 |
C12orf39 |
chromosome 12 open reading frame 39 |
| chr6_+_146920097 | 0.02 |
ENST00000397944.3 ENST00000522242.1 |
ADGB |
androglobin |
| chr1_+_113010056 | 0.02 |
ENST00000369686.5 |
WNT2B |
wingless-type MMTV integration site family, member 2B |
| chr5_+_54455946 | 0.02 |
ENST00000503787.1 ENST00000296734.6 ENST00000515370.1 |
GPX8 |
glutathione peroxidase 8 (putative) |
| chr3_+_155838337 | 0.02 |
ENST00000490337.1 ENST00000389636.5 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr3_-_46068969 | 0.02 |
ENST00000542109.1 ENST00000395946.2 |
XCR1 |
chemokine (C motif) receptor 1 |
| chrX_-_33229636 | 0.02 |
ENST00000357033.4 |
DMD |
dystrophin |
| chr14_+_22386325 | 0.02 |
ENST00000390439.2 |
TRAV13-2 |
T cell receptor alpha variable 13-2 |
| chr14_-_23876801 | 0.02 |
ENST00000356287.3 |
MYH6 |
myosin, heavy chain 6, cardiac muscle, alpha |
| chrY_+_23698778 | 0.01 |
ENST00000303902.5 |
RBMY1A1 |
RNA binding motif protein, Y-linked, family 1, member A1 |
| chr20_+_10199468 | 0.01 |
ENST00000254976.2 ENST00000304886.2 |
SNAP25 |
synaptosomal-associated protein, 25kDa |
| chr12_+_5541267 | 0.01 |
ENST00000423158.3 |
NTF3 |
neurotrophin 3 |
| chrY_-_24038660 | 0.01 |
ENST00000382677.3 |
RBMY1D |
RNA binding motif protein, Y-linked, family 1, member D |
| chr4_+_56212270 | 0.01 |
ENST00000264228.4 |
SRD5A3 |
steroid 5 alpha-reductase 3 |
| chr6_+_29429217 | 0.01 |
ENST00000396792.2 |
OR2H1 |
olfactory receptor, family 2, subfamily H, member 1 |
| chr15_+_66585555 | 0.01 |
ENST00000319194.5 ENST00000525134.2 ENST00000441424.2 |
DIS3L |
DIS3 mitotic control homolog (S. cerevisiae)-like |
| chr3_+_171561127 | 0.01 |
ENST00000334567.5 ENST00000450693.1 |
TMEM212 |
transmembrane protein 212 |
| chr7_-_80141328 | 0.01 |
ENST00000398291.3 |
GNAT3 |
guanine nucleotide binding protein, alpha transducing 3 |
| chr12_-_11036844 | 0.00 |
ENST00000428168.2 |
PRH1 |
proline-rich protein HaeIII subfamily 1 |
| chr16_+_19535133 | 0.00 |
ENST00000396212.2 ENST00000381396.5 |
CCP110 |
centriolar coiled coil protein 110kDa |
| chr4_+_159131630 | 0.00 |
ENST00000504569.1 ENST00000509278.1 ENST00000514558.1 ENST00000503200.1 |
TMEM144 |
transmembrane protein 144 |
Gene Ontology Analysis
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.3 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.2 | 4.1 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.2 | 0.8 | GO:0002189 | ribose phosphate diphosphokinase complex(GO:0002189) |
| 0.1 | 1.7 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.1 | 0.5 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.1 | 1.0 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 0.7 | GO:0000940 | condensed chromosome outer kinetochore(GO:0000940) |
| 0.0 | 0.3 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 0.1 | GO:0033011 | perinuclear theca(GO:0033011) cytoskeletal calyx(GO:0033150) |
| 0.0 | 0.6 | GO:0045239 | tricarboxylic acid cycle enzyme complex(GO:0045239) |
| 0.0 | 0.5 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 0.4 | GO:0000506 | glycosylphosphatidylinositol-N-acetylglucosaminyltransferase (GPI-GnT) complex(GO:0000506) |
| 0.0 | 1.5 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.1 | GO:0030891 | VCB complex(GO:0030891) |
| 0.0 | 0.4 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.3 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.0 | 0.2 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 0.3 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.0 | 1.9 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.4 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.0 | 0.3 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 0.2 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.0 | 0.3 | GO:0001527 | microfibril(GO:0001527) fibril(GO:0043205) |
| 0.0 | 0.4 | GO:0032982 | myosin filament(GO:0032982) |
| 0.0 | 0.1 | GO:0000801 | central element(GO:0000801) |
| 0.0 | 0.2 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.0 | 0.4 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.3 | GO:0036019 | endolysosome(GO:0036019) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.9 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.1 | 0.5 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.0 | 0.3 | REACTOME ETHANOL OXIDATION | Genes involved in Ethanol oxidation |
| 0.0 | 2.0 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.8 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 0.5 | REACTOME DESTABILIZATION OF MRNA BY KSRP | Genes involved in Destabilization of mRNA by KSRP |
| 0.0 | 0.6 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.2 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.0 | 0.5 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.3 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.0 | 0.6 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 0.2 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.5 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.0 | 0.5 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.0 | 0.3 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 0.5 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.7 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.3 | 0.8 | GO:0044549 | GTP cyclohydrolase binding(GO:0044549) |
| 0.2 | 0.7 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.2 | 0.5 | GO:0004531 | deoxyribonuclease II activity(GO:0004531) |
| 0.1 | 0.8 | GO:0004749 | ribose phosphate diphosphokinase activity(GO:0004749) |
| 0.1 | 0.3 | GO:0016749 | 5-aminolevulinate synthase activity(GO:0003870) N-succinyltransferase activity(GO:0016749) |
| 0.1 | 1.0 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.1 | 0.5 | GO:0070728 | leucine binding(GO:0070728) |
| 0.1 | 0.4 | GO:0050309 | glucose-6-phosphatase activity(GO:0004346) sugar-terminal-phosphatase activity(GO:0050309) |
| 0.1 | 0.6 | GO:0048101 | calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.1 | 0.5 | GO:0052654 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.1 | 0.3 | GO:0035276 | aldehyde oxidase activity(GO:0004031) ethanol binding(GO:0035276) |
| 0.1 | 0.2 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.1 | 0.2 | GO:0099580 | ion antiporter activity involved in regulation of postsynaptic membrane potential(GO:0099580) |
| 0.1 | 0.4 | GO:0017018 | myosin phosphatase activity(GO:0017018) |
| 0.1 | 2.4 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 0.3 | GO:0034875 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 0.1 | 0.4 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.0 | 0.4 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.0 | 0.5 | GO:0019534 | toxin transporter activity(GO:0019534) |
| 0.0 | 0.4 | GO:0017176 | phosphatidylinositol N-acetylglucosaminyltransferase activity(GO:0017176) |
| 0.0 | 0.5 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.0 | 1.0 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.0 | 0.4 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 1.5 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.0 | 1.9 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.8 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 0.3 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.0 | 0.5 | GO:0099604 | calcium-release channel activity(GO:0015278) ligand-gated calcium channel activity(GO:0099604) |
| 0.0 | 0.1 | GO:0003956 | NAD(P)+-protein-arginine ADP-ribosyltransferase activity(GO:0003956) |
| 0.0 | 0.1 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.0 | 0.1 | GO:0005152 | interleukin-1 receptor antagonist activity(GO:0005152) |
| 0.0 | 4.1 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.2 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.0 | 0.0 | GO:0004822 | isoleucine-tRNA ligase activity(GO:0004822) |
| 0.0 | 0.3 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.2 | GO:0002151 | G-quadruplex RNA binding(GO:0002151) RNA strand annealing activity(GO:0033592) |
| 0.0 | 0.1 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.0 | 0.2 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.6 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 0.5 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.0 | 0.6 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.0 | 0.3 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.9 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.5 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.0 | 0.6 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.3 | GO:0097577 | intracellular sequestering of iron ion(GO:0006880) sequestering of iron ion(GO:0097577) |
| 0.3 | 1.9 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.3 | 4.1 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.3 | 0.8 | GO:0043095 | regulation of GTP cyclohydrolase I activity(GO:0043095) negative regulation of GTP cyclohydrolase I activity(GO:0043105) |
| 0.2 | 0.6 | GO:0006106 | fumarate metabolic process(GO:0006106) |
| 0.2 | 0.8 | GO:0046391 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 0.1 | 0.4 | GO:0099543 | retrograde trans-synaptic signaling by soluble gas(GO:0098923) trans-synaptic signaling by soluble gas(GO:0099543) trans-synaptic signaling by trans-synaptic complex(GO:0099545) |
| 0.1 | 0.4 | GO:0032831 | regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032829) positive regulation of CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation(GO:0032831) positive regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001190) |
| 0.1 | 0.5 | GO:0034473 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) |
| 0.1 | 0.3 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.1 | 0.5 | GO:0009099 | branched-chain amino acid biosynthetic process(GO:0009082) leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.1 | 0.5 | GO:0071233 | cellular response to leucine(GO:0071233) |
| 0.1 | 0.2 | GO:2000439 | negative regulation of eosinophil activation(GO:1902567) positive regulation of monocyte extravasation(GO:2000439) |
| 0.1 | 1.4 | GO:0020027 | hemoglobin metabolic process(GO:0020027) |
| 0.1 | 0.3 | GO:2001027 | negative regulation of endothelial cell chemotaxis(GO:2001027) |
| 0.1 | 0.1 | GO:0006533 | aspartate catabolic process(GO:0006533) |
| 0.1 | 0.4 | GO:2000676 | positive regulation of type B pancreatic cell apoptotic process(GO:2000676) |
| 0.1 | 0.4 | GO:0007079 | mitotic chromosome movement towards spindle pole(GO:0007079) |
| 0.1 | 0.3 | GO:0051661 | maintenance of centrosome location(GO:0051661) |
| 0.0 | 0.3 | GO:0002933 | lipid hydroxylation(GO:0002933) |
| 0.0 | 1.5 | GO:0045730 | respiratory burst(GO:0045730) |
| 0.0 | 0.3 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.0 | 0.6 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.0 | 0.4 | GO:0051151 | negative regulation of smooth muscle cell differentiation(GO:0051151) |
| 0.0 | 0.4 | GO:0055009 | atrial cardiac muscle tissue development(GO:0003228) cardiac muscle fiber development(GO:0048739) atrial cardiac muscle tissue morphogenesis(GO:0055009) |
| 0.0 | 0.5 | GO:0090168 | Golgi reassembly(GO:0090168) |
| 0.0 | 0.2 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
| 0.0 | 0.3 | GO:0034501 | protein localization to kinetochore(GO:0034501) |
| 0.0 | 0.4 | GO:0000338 | protein deneddylation(GO:0000338) |
| 0.0 | 0.3 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) |
| 0.0 | 0.5 | GO:0033141 | positive regulation of peptidyl-serine phosphorylation of STAT protein(GO:0033141) |
| 0.0 | 1.5 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.0 | 0.1 | GO:2000232 | regulation of rRNA processing(GO:2000232) |
| 0.0 | 0.1 | GO:1904425 | negative regulation of GTP binding(GO:1904425) |
| 0.0 | 0.3 | GO:0090073 | positive regulation of protein homodimerization activity(GO:0090073) |
| 0.0 | 0.3 | GO:0045919 | positive regulation of cytolysis(GO:0045919) |
| 0.0 | 0.5 | GO:0002643 | regulation of tolerance induction(GO:0002643) |
| 0.0 | 0.1 | GO:0048749 | compound eye development(GO:0048749) |
| 0.0 | 0.8 | GO:0035065 | regulation of histone acetylation(GO:0035065) |
| 0.0 | 0.4 | GO:0016254 | preassembly of GPI anchor in ER membrane(GO:0016254) |
| 0.0 | 0.6 | GO:0071425 | hematopoietic stem cell proliferation(GO:0071425) |
| 0.0 | 0.5 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.0 | 0.0 | GO:0006428 | isoleucyl-tRNA aminoacylation(GO:0006428) |
| 0.0 | 0.1 | GO:1902714 | negative regulation of interferon-gamma secretion(GO:1902714) |
| 0.0 | 0.2 | GO:0016075 | rRNA catabolic process(GO:0016075) |
| 0.0 | 0.5 | GO:0035640 | exploration behavior(GO:0035640) |
| 0.0 | 0.2 | GO:0048311 | mitochondrion distribution(GO:0048311) |
| 0.0 | 0.3 | GO:0032211 | negative regulation of telomere maintenance via telomerase(GO:0032211) |
| 0.0 | 0.2 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.4 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.0 | 1.0 | GO:0043966 | histone H3 acetylation(GO:0043966) |
| 0.0 | 1.0 | GO:0042100 | B cell proliferation(GO:0042100) |
| 0.0 | 0.4 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |