Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
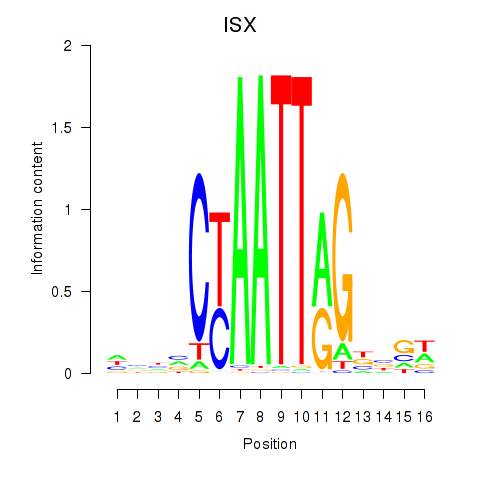
Results for ISX
Z-value: 0.41
Transcription factors associated with ISX
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ISX
|
ENSG00000175329.8 | ISX |
Activity profile of ISX motif
Sorted Z-values of ISX motif
Network of associatons between targets according to the STRING database.
First level regulatory network of ISX
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_149095652 | 1.34 |
ENST00000305366.3 |
TM4SF1 |
transmembrane 4 L six family member 1 |
| chr16_-_55866997 | 1.28 |
ENST00000360526.3 ENST00000361503.4 |
CES1 |
carboxylesterase 1 |
| chr14_-_94789663 | 0.91 |
ENST00000557225.1 ENST00000341584.3 |
SERPINA6 |
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 6 |
| chr16_-_55867146 | 0.73 |
ENST00000422046.2 |
CES1 |
carboxylesterase 1 |
| chr4_-_141348789 | 0.63 |
ENST00000414773.1 |
CLGN |
calmegin |
| chr12_-_103310987 | 0.56 |
ENST00000307000.2 |
PAH |
phenylalanine hydroxylase |
| chr4_-_141348999 | 0.50 |
ENST00000325617.5 |
CLGN |
calmegin |
| chr2_+_217524323 | 0.42 |
ENST00000456764.1 |
IGFBP2 |
insulin-like growth factor binding protein 2, 36kDa |
| chrX_+_43515467 | 0.41 |
ENST00000338702.3 ENST00000542639.1 |
MAOA |
monoamine oxidase A |
| chr10_-_99447024 | 0.40 |
ENST00000370626.3 |
AVPI1 |
arginine vasopressin-induced 1 |
| chr6_+_148663729 | 0.39 |
ENST00000367467.3 |
SASH1 |
SAM and SH3 domain containing 1 |
| chr20_-_17662705 | 0.39 |
ENST00000455029.2 |
RRBP1 |
ribosome binding protein 1 |
| chr4_-_70080449 | 0.39 |
ENST00000446444.1 |
UGT2B11 |
UDP glucuronosyltransferase 2 family, polypeptide B11 |
| chr12_-_51422017 | 0.35 |
ENST00000394904.3 |
SLC11A2 |
solute carrier family 11 (proton-coupled divalent metal ion transporter), member 2 |
| chr16_+_3070313 | 0.34 |
ENST00000326577.4 |
TNFRSF12A |
tumor necrosis factor receptor superfamily, member 12A |
| chr3_-_151034734 | 0.29 |
ENST00000260843.4 |
GPR87 |
G protein-coupled receptor 87 |
| chr16_+_3070356 | 0.28 |
ENST00000341627.5 ENST00000575124.1 ENST00000575836.1 |
TNFRSF12A |
tumor necrosis factor receptor superfamily, member 12A |
| chr1_-_68698222 | 0.27 |
ENST00000370976.3 ENST00000354777.2 ENST00000262348.4 ENST00000540432.1 |
WLS |
wntless Wnt ligand secretion mediator |
| chr1_-_209824643 | 0.26 |
ENST00000391911.1 ENST00000415782.1 |
LAMB3 |
laminin, beta 3 |
| chr1_-_68698197 | 0.24 |
ENST00000370973.2 ENST00000370971.1 |
WLS |
wntless Wnt ligand secretion mediator |
| chr20_+_44441304 | 0.24 |
ENST00000352551.5 |
UBE2C |
ubiquitin-conjugating enzyme E2C |
| chr21_+_35014783 | 0.24 |
ENST00000381291.4 ENST00000381285.4 ENST00000399367.3 ENST00000399352.1 ENST00000399355.2 ENST00000399349.1 |
ITSN1 |
intersectin 1 (SH3 domain protein) |
| chr20_+_44441215 | 0.23 |
ENST00000356455.4 ENST00000405520.1 |
UBE2C |
ubiquitin-conjugating enzyme E2C |
| chr9_+_34646624 | 0.22 |
ENST00000450095.2 ENST00000556278.1 |
GALT GALT |
galactose-1-phosphate uridylyltransferase Uncharacterized protein |
| chr20_+_44441271 | 0.21 |
ENST00000335046.3 ENST00000243893.6 |
UBE2C |
ubiquitin-conjugating enzyme E2C |
| chr5_-_95297534 | 0.21 |
ENST00000513343.1 ENST00000431061.2 |
ELL2 |
elongation factor, RNA polymerase II, 2 |
| chr8_-_82395461 | 0.20 |
ENST00000256104.4 |
FABP4 |
fatty acid binding protein 4, adipocyte |
| chr11_+_77532233 | 0.19 |
ENST00000525409.1 |
AAMDC |
adipogenesis associated, Mth938 domain containing |
| chr9_+_34646651 | 0.19 |
ENST00000378842.3 |
GALT |
galactose-1-phosphate uridylyltransferase |
| chr8_+_9413410 | 0.19 |
ENST00000520408.1 ENST00000310430.6 ENST00000522110.1 |
TNKS |
tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase |
| chr17_+_73452695 | 0.18 |
ENST00000582186.1 ENST00000582455.1 ENST00000581252.1 ENST00000579208.1 |
KIAA0195 |
KIAA0195 |
| chr9_+_99212403 | 0.17 |
ENST00000375251.3 ENST00000375249.4 |
HABP4 |
hyaluronan binding protein 4 |
| chr1_+_28261533 | 0.16 |
ENST00000411604.1 ENST00000373888.4 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr17_-_18266660 | 0.16 |
ENST00000582653.1 ENST00000352886.6 |
SHMT1 |
serine hydroxymethyltransferase 1 (soluble) |
| chr12_+_104680659 | 0.16 |
ENST00000526691.1 ENST00000531691.1 ENST00000388854.3 ENST00000354940.6 ENST00000526390.1 ENST00000531689.1 |
TXNRD1 |
thioredoxin reductase 1 |
| chr22_+_36044411 | 0.16 |
ENST00000409652.4 |
APOL6 |
apolipoprotein L, 6 |
| chr6_+_64345698 | 0.16 |
ENST00000506783.1 ENST00000481385.2 ENST00000515594.1 ENST00000494284.2 ENST00000262043.3 |
PHF3 |
PHD finger protein 3 |
| chr7_+_150929550 | 0.16 |
ENST00000482173.1 ENST00000495645.1 ENST00000035307.2 |
CHPF2 |
chondroitin polymerizing factor 2 |
| chr11_+_77532155 | 0.15 |
ENST00000532481.1 ENST00000526415.1 ENST00000393427.2 ENST00000527134.1 ENST00000304716.8 |
AAMDC |
adipogenesis associated, Mth938 domain containing |
| chr17_-_42277203 | 0.14 |
ENST00000587097.1 |
ATXN7L3 |
ataxin 7-like 3 |
| chr6_-_49681235 | 0.14 |
ENST00000339139.4 |
CRISP2 |
cysteine-rich secretory protein 2 |
| chr3_-_160823158 | 0.14 |
ENST00000392779.2 ENST00000392780.1 ENST00000494173.1 |
B3GALNT1 |
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) |
| chr4_-_186733363 | 0.13 |
ENST00000393523.2 ENST00000393528.3 ENST00000449407.2 |
SORBS2 |
sorbin and SH3 domain containing 2 |
| chr4_+_87515454 | 0.13 |
ENST00000427191.2 ENST00000436978.1 ENST00000502971.1 |
PTPN13 |
protein tyrosine phosphatase, non-receptor type 13 (APO-1/CD95 (Fas)-associated phosphatase) |
| chr17_-_18266765 | 0.13 |
ENST00000354098.3 |
SHMT1 |
serine hydroxymethyltransferase 1 (soluble) |
| chr5_+_159848807 | 0.12 |
ENST00000352433.5 |
PTTG1 |
pituitary tumor-transforming 1 |
| chr17_-_27418537 | 0.12 |
ENST00000408971.2 |
TIAF1 |
TGFB1-induced anti-apoptotic factor 1 |
| chr14_+_32798547 | 0.12 |
ENST00000557354.1 ENST00000557102.1 ENST00000557272.1 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr1_-_150669604 | 0.12 |
ENST00000427665.1 ENST00000540514.1 |
GOLPH3L |
golgi phosphoprotein 3-like |
| chr20_-_43133491 | 0.12 |
ENST00000411544.1 |
SERINC3 |
serine incorporator 3 |
| chr5_-_95297678 | 0.11 |
ENST00000237853.4 |
ELL2 |
elongation factor, RNA polymerase II, 2 |
| chr6_-_135271260 | 0.11 |
ENST00000265605.2 |
ALDH8A1 |
aldehyde dehydrogenase 8 family, member A1 |
| chr17_+_79650962 | 0.11 |
ENST00000329138.4 |
HGS |
hepatocyte growth factor-regulated tyrosine kinase substrate |
| chrX_-_24690771 | 0.11 |
ENST00000379145.1 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr17_-_48785216 | 0.11 |
ENST00000285243.6 |
ANKRD40 |
ankyrin repeat domain 40 |
| chr17_-_18266797 | 0.11 |
ENST00000316694.3 ENST00000539052.1 |
SHMT1 |
serine hydroxymethyltransferase 1 (soluble) |
| chr12_+_3000037 | 0.11 |
ENST00000544943.1 ENST00000448120.2 |
TULP3 |
tubby like protein 3 |
| chr11_-_62414070 | 0.10 |
ENST00000540933.1 ENST00000346178.4 ENST00000356638.3 ENST00000534779.1 ENST00000525994.1 |
GANAB |
glucosidase, alpha; neutral AB |
| chr3_-_160823040 | 0.10 |
ENST00000484127.1 ENST00000492353.1 ENST00000473142.1 ENST00000468268.1 ENST00000460353.1 ENST00000320474.4 ENST00000392781.2 |
B3GALNT1 |
beta-1,3-N-acetylgalactosaminyltransferase 1 (globoside blood group) |
| chr17_-_77005801 | 0.10 |
ENST00000392446.5 |
CANT1 |
calcium activated nucleotidase 1 |
| chr12_+_69201923 | 0.10 |
ENST00000462284.1 ENST00000258149.5 ENST00000356290.4 ENST00000540827.1 ENST00000428863.2 ENST00000393412.3 |
MDM2 |
MDM2 oncogene, E3 ubiquitin protein ligase |
| chr1_+_28261492 | 0.10 |
ENST00000373894.3 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr16_+_3493611 | 0.10 |
ENST00000407558.4 ENST00000572169.1 ENST00000572757.1 ENST00000573593.1 ENST00000570372.1 ENST00000424546.2 ENST00000575733.1 ENST00000573201.1 ENST00000574950.1 ENST00000573580.1 ENST00000608722.1 |
NAA60 NAA60 |
N(alpha)-acetyltransferase 60, NatF catalytic subunit N-alpha-acetyltransferase 60 |
| chr14_+_70233810 | 0.09 |
ENST00000394366.2 ENST00000553548.1 ENST00000553369.1 ENST00000557154.1 ENST00000451983.2 ENST00000553635.1 |
SRSF5 |
serine/arginine-rich splicing factor 5 |
| chr10_+_91461337 | 0.09 |
ENST00000260753.4 ENST00000416354.1 ENST00000394289.2 ENST00000371728.3 |
KIF20B |
kinesin family member 20B |
| chr5_+_159848854 | 0.09 |
ENST00000517480.1 ENST00000520452.1 ENST00000393964.1 |
PTTG1 |
pituitary tumor-transforming 1 |
| chr8_+_105235572 | 0.09 |
ENST00000523362.1 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr22_+_29138013 | 0.09 |
ENST00000216027.3 ENST00000398941.2 |
HSCB |
HscB mitochondrial iron-sulfur cluster co-chaperone |
| chr3_-_46069223 | 0.09 |
ENST00000309285.3 |
XCR1 |
chemokine (C motif) receptor 1 |
| chr2_-_183387430 | 0.09 |
ENST00000410103.1 |
PDE1A |
phosphodiesterase 1A, calmodulin-dependent |
| chr1_+_151735431 | 0.09 |
ENST00000321531.5 ENST00000315067.8 |
OAZ3 |
ornithine decarboxylase antizyme 3 |
| chr3_+_113775576 | 0.09 |
ENST00000485050.1 ENST00000281273.4 |
QTRTD1 |
queuine tRNA-ribosyltransferase domain containing 1 |
| chr12_-_10978957 | 0.09 |
ENST00000240619.2 |
TAS2R10 |
taste receptor, type 2, member 10 |
| chr2_+_87135076 | 0.09 |
ENST00000409776.2 |
RGPD1 |
RANBP2-like and GRIP domain containing 1 |
| chr1_+_28261621 | 0.08 |
ENST00000549094.1 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr6_-_135271219 | 0.08 |
ENST00000367847.2 ENST00000367845.2 |
ALDH8A1 |
aldehyde dehydrogenase 8 family, member A1 |
| chr2_+_204571375 | 0.08 |
ENST00000374478.4 |
CD28 |
CD28 molecule |
| chr16_-_31105870 | 0.07 |
ENST00000394971.3 |
VKORC1 |
vitamin K epoxide reductase complex, subunit 1 |
| chr13_+_38923959 | 0.07 |
ENST00000379649.1 ENST00000239878.4 ENST00000437952.1 ENST00000379641.1 |
UFM1 |
ubiquitin-fold modifier 1 |
| chr14_+_104182061 | 0.07 |
ENST00000216602.6 |
ZFYVE21 |
zinc finger, FYVE domain containing 21 |
| chr17_-_2966901 | 0.07 |
ENST00000575751.1 |
OR1D5 |
olfactory receptor, family 1, subfamily D, member 5 |
| chr17_+_73452545 | 0.07 |
ENST00000314256.7 |
KIAA0195 |
KIAA0195 |
| chr16_-_31106211 | 0.07 |
ENST00000532364.1 ENST00000529564.1 ENST00000319788.7 ENST00000354895.4 ENST00000394975.2 |
RP11-196G11.1 VKORC1 |
Uncharacterized protein vitamin K epoxide reductase complex, subunit 1 |
| chr2_+_86947296 | 0.07 |
ENST00000283632.4 |
RMND5A |
required for meiotic nuclear division 5 homolog A (S. cerevisiae) |
| chr6_+_160542821 | 0.06 |
ENST00000366963.4 |
SLC22A1 |
solute carrier family 22 (organic cation transporter), member 1 |
| chr16_-_31106048 | 0.06 |
ENST00000300851.6 |
VKORC1 |
vitamin K epoxide reductase complex, subunit 1 |
| chr12_+_52695617 | 0.06 |
ENST00000293525.5 |
KRT86 |
keratin 86 |
| chr6_-_39693111 | 0.06 |
ENST00000373215.3 ENST00000538893.1 ENST00000287152.7 ENST00000373216.3 |
KIF6 |
kinesin family member 6 |
| chr2_+_177015122 | 0.06 |
ENST00000468418.3 |
HOXD3 |
homeobox D3 |
| chr1_+_155278625 | 0.06 |
ENST00000368356.4 ENST00000356657.6 |
FDPS |
farnesyl diphosphate synthase |
| chr12_+_3000073 | 0.06 |
ENST00000397132.2 |
TULP3 |
tubby like protein 3 |
| chr11_-_13517565 | 0.06 |
ENST00000282091.1 ENST00000529816.1 |
PTH |
parathyroid hormone |
| chr1_+_87595433 | 0.05 |
ENST00000469312.2 ENST00000490006.2 |
RP5-1052I5.1 |
long intergenic non-protein coding RNA 1140 |
| chr7_+_144052381 | 0.05 |
ENST00000498580.1 ENST00000056217.5 |
ARHGEF5 |
Rho guanine nucleotide exchange factor (GEF) 5 |
| chr14_+_22964877 | 0.05 |
ENST00000390494.1 |
TRAJ43 |
T cell receptor alpha joining 43 |
| chr14_+_104182105 | 0.05 |
ENST00000311141.2 |
ZFYVE21 |
zinc finger, FYVE domain containing 21 |
| chr5_-_98262240 | 0.05 |
ENST00000284049.3 |
CHD1 |
chromodomain helicase DNA binding protein 1 |
| chr2_-_3521518 | 0.05 |
ENST00000382093.5 |
ADI1 |
acireductone dioxygenase 1 |
| chr8_+_9953061 | 0.05 |
ENST00000522907.1 ENST00000528246.1 |
MSRA |
methionine sulfoxide reductase A |
| chr3_+_37284668 | 0.05 |
ENST00000361924.2 ENST00000444882.1 ENST00000356847.4 ENST00000450863.2 ENST00000429018.1 |
GOLGA4 |
golgin A4 |
| chr20_+_24449821 | 0.05 |
ENST00000376862.3 |
SYNDIG1 |
synapse differentiation inducing 1 |
| chr6_+_160542870 | 0.04 |
ENST00000324965.4 ENST00000457470.2 |
SLC22A1 |
solute carrier family 22 (organic cation transporter), member 1 |
| chr7_+_115862858 | 0.04 |
ENST00000393481.2 |
TES |
testis derived transcript (3 LIM domains) |
| chr1_-_150669500 | 0.04 |
ENST00000271732.3 |
GOLPH3L |
golgi phosphoprotein 3-like |
| chr10_-_114206649 | 0.04 |
ENST00000369404.3 ENST00000369405.3 |
ZDHHC6 |
zinc finger, DHHC-type containing 6 |
| chr1_-_112281875 | 0.04 |
ENST00000527621.1 ENST00000534365.1 ENST00000357260.5 |
FAM212B |
family with sequence similarity 212, member B |
| chr14_-_20801427 | 0.04 |
ENST00000557665.1 ENST00000358932.4 ENST00000353689.4 |
CCNB1IP1 |
cyclin B1 interacting protein 1, E3 ubiquitin protein ligase |
| chr8_+_9953214 | 0.04 |
ENST00000382490.5 |
MSRA |
methionine sulfoxide reductase A |
| chr17_-_43045439 | 0.04 |
ENST00000253407.3 |
C1QL1 |
complement component 1, q subcomponent-like 1 |
| chr16_-_58585513 | 0.03 |
ENST00000245138.4 ENST00000567285.1 |
CNOT1 |
CCR4-NOT transcription complex, subunit 1 |
| chr3_+_148709128 | 0.03 |
ENST00000345003.4 ENST00000296048.6 ENST00000483267.1 |
GYG1 |
glycogenin 1 |
| chr4_+_48018781 | 0.03 |
ENST00000295461.5 |
NIPAL1 |
NIPA-like domain containing 1 |
| chr1_+_155278539 | 0.03 |
ENST00000447866.1 |
FDPS |
farnesyl diphosphate synthase |
| chr1_-_197115818 | 0.03 |
ENST00000367409.4 ENST00000294732.7 |
ASPM |
asp (abnormal spindle) homolog, microcephaly associated (Drosophila) |
| chr2_+_135809835 | 0.03 |
ENST00000264158.8 ENST00000539493.1 ENST00000442034.1 ENST00000425393.1 |
RAB3GAP1 |
RAB3 GTPase activating protein subunit 1 (catalytic) |
| chr1_-_150849174 | 0.03 |
ENST00000515192.1 |
ARNT |
aryl hydrocarbon receptor nuclear translocator |
| chr5_+_174151536 | 0.03 |
ENST00000239243.6 ENST00000507785.1 |
MSX2 |
msh homeobox 2 |
| chr1_+_220267429 | 0.03 |
ENST00000366922.1 ENST00000302637.5 |
IARS2 |
isoleucyl-tRNA synthetase 2, mitochondrial |
| chr7_-_100493482 | 0.03 |
ENST00000411582.1 ENST00000419336.2 ENST00000241069.5 ENST00000302913.4 |
ACHE |
acetylcholinesterase (Yt blood group) |
| chr12_+_15125954 | 0.03 |
ENST00000266395.2 |
PDE6H |
phosphodiesterase 6H, cGMP-specific, cone, gamma |
| chr1_+_23345930 | 0.03 |
ENST00000356634.3 |
KDM1A |
lysine (K)-specific demethylase 1A |
| chr21_+_35014706 | 0.03 |
ENST00000399353.1 ENST00000444491.1 ENST00000381318.3 |
ITSN1 |
intersectin 1 (SH3 domain protein) |
| chr3_-_15563229 | 0.02 |
ENST00000383786.5 ENST00000383787.2 ENST00000383785.2 ENST00000383788.5 ENST00000603808.1 |
COLQ |
collagen-like tail subunit (single strand of homotrimer) of asymmetric acetylcholinesterase |
| chr6_-_31125850 | 0.02 |
ENST00000507751.1 ENST00000448162.2 ENST00000502557.1 ENST00000503420.1 ENST00000507892.1 ENST00000507226.1 ENST00000513222.1 ENST00000503934.1 ENST00000396263.2 ENST00000508683.1 ENST00000428174.1 ENST00000448141.2 ENST00000507829.1 ENST00000455279.2 ENST00000376266.5 |
CCHCR1 |
coiled-coil alpha-helical rod protein 1 |
| chr12_+_122688090 | 0.02 |
ENST00000324189.4 ENST00000546192.1 |
B3GNT4 |
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 4 |
| chr6_-_32095968 | 0.02 |
ENST00000375203.3 ENST00000375201.4 |
ATF6B |
activating transcription factor 6 beta |
| chr1_-_150849047 | 0.02 |
ENST00000354396.2 ENST00000505755.1 |
ARNT |
aryl hydrocarbon receptor nuclear translocator |
| chr4_-_100242549 | 0.02 |
ENST00000305046.8 ENST00000394887.3 |
ADH1B |
alcohol dehydrogenase 1B (class I), beta polypeptide |
| chr1_-_43833628 | 0.02 |
ENST00000413844.2 ENST00000372458.3 |
ELOVL1 |
ELOVL fatty acid elongase 1 |
| chr14_-_73925225 | 0.02 |
ENST00000356296.4 ENST00000355058.3 ENST00000359560.3 ENST00000557597.1 ENST00000554394.1 ENST00000555238.1 ENST00000535282.1 ENST00000555987.1 ENST00000555394.1 ENST00000554546.1 |
NUMB |
numb homolog (Drosophila) |
| chr14_+_32798462 | 0.02 |
ENST00000280979.4 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chrX_-_6453159 | 0.02 |
ENST00000381089.3 ENST00000398729.1 |
VCX3A |
variable charge, X-linked 3A |
| chr10_-_28571015 | 0.02 |
ENST00000375719.3 ENST00000375732.1 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chrX_-_8139308 | 0.02 |
ENST00000317103.4 |
VCX2 |
variable charge, X-linked 2 |
| chr1_+_23345943 | 0.02 |
ENST00000400181.4 ENST00000542151.1 |
KDM1A |
lysine (K)-specific demethylase 1A |
| chr14_-_24711806 | 0.02 |
ENST00000540705.1 ENST00000538777.1 ENST00000558566.1 ENST00000559019.1 |
TINF2 |
TERF1 (TRF1)-interacting nuclear factor 2 |
| chr1_-_150849208 | 0.01 |
ENST00000358595.5 |
ARNT |
aryl hydrocarbon receptor nuclear translocator |
| chr6_-_33385870 | 0.01 |
ENST00000488034.1 |
CUTA |
cutA divalent cation tolerance homolog (E. coli) |
| chr7_+_130020932 | 0.01 |
ENST00000484324.1 |
CPA1 |
carboxypeptidase A1 (pancreatic) |
| chr4_-_66536057 | 0.01 |
ENST00000273854.3 |
EPHA5 |
EPH receptor A5 |
| chr6_-_33385902 | 0.01 |
ENST00000374500.5 |
CUTA |
cutA divalent cation tolerance homolog (E. coli) |
| chr10_+_124913793 | 0.01 |
ENST00000368865.4 ENST00000538238.1 ENST00000368859.2 |
BUB3 |
BUB3 mitotic checkpoint protein |
| chr22_-_29137771 | 0.01 |
ENST00000439200.1 ENST00000405598.1 ENST00000398017.2 ENST00000425190.2 ENST00000348295.3 ENST00000382578.1 ENST00000382565.1 ENST00000382566.1 ENST00000382580.2 ENST00000328354.6 |
CHEK2 |
checkpoint kinase 2 |
| chr6_-_119256311 | 0.00 |
ENST00000316316.6 |
MCM9 |
minichromosome maintenance complex component 9 |
| chr4_-_66536196 | 0.00 |
ENST00000511294.1 |
EPHA5 |
EPH receptor A5 |
| chrX_-_139587225 | 0.00 |
ENST00000370536.2 |
SOX3 |
SRY (sex determining region Y)-box 3 |
| chr6_-_33385655 | 0.00 |
ENST00000440279.3 ENST00000607266.1 |
CUTA |
cutA divalent cation tolerance homolog (E. coli) |
| chr19_-_14785622 | 0.00 |
ENST00000443157.2 |
EMR3 |
egf-like module containing, mucin-like, hormone receptor-like 3 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 0.4 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 0.3 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.2 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 0.3 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.0 | GO:0004771 | sterol esterase activity(GO:0004771) methylumbelliferyl-acetate deacetylase activity(GO:0047374) |
| 0.1 | 0.6 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 0.1 | 0.2 | GO:0047273 | galactosylgalactosylglucosylceramide beta-D-acetylgalactosaminyltransferase activity(GO:0047273) |
| 0.1 | 0.4 | GO:0070905 | serine binding(GO:0070905) |
| 0.1 | 0.3 | GO:0015094 | cadmium ion transmembrane transporter activity(GO:0015086) cobalt ion transmembrane transporter activity(GO:0015087) lead ion transmembrane transporter activity(GO:0015094) ferrous iron uptake transmembrane transporter activity(GO:0015639) |
| 0.1 | 0.2 | GO:0098626 | methylselenol reductase activity(GO:0098625) methylseleninic acid reductase activity(GO:0098626) |
| 0.1 | 0.2 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 0.1 | 0.4 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.1 | 0.2 | GO:0047057 | oxidoreductase activity, acting on the CH-OH group of donors, disulfide as acceptor(GO:0016900) vitamin-K-epoxide reductase (warfarin-sensitive) activity(GO:0047057) |
| 0.0 | 0.5 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.0 | 0.4 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.4 | GO:0070569 | uridylyltransferase activity(GO:0070569) |
| 0.0 | 0.3 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.1 | GO:0004382 | guanosine-diphosphatase activity(GO:0004382) |
| 0.0 | 1.2 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 0.1 | GO:0061663 | NEDD8 ligase activity(GO:0061663) |
| 0.0 | 0.1 | GO:0005277 | acetylcholine transmembrane transporter activity(GO:0005277) secondary active organic cation transmembrane transporter activity(GO:0008513) acetate ester transmembrane transporter activity(GO:1901375) |
| 0.0 | 0.1 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.0 | 0.4 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
| 0.0 | 0.1 | GO:0004161 | dimethylallyltranstransferase activity(GO:0004161) geranyltranstransferase activity(GO:0004337) |
| 0.0 | 0.1 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.0 | 0.1 | GO:0004874 | aryl hydrocarbon receptor activity(GO:0004874) |
| 0.0 | 0.1 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.0 | 0.3 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.0 | 0.7 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.0 | GO:0061752 | telomeric repeat-containing RNA binding(GO:0061752) |
| 0.0 | 0.1 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0070826 | paraferritin complex(GO:0070826) |
| 0.0 | 0.1 | GO:0033565 | ESCRT-0 complex(GO:0033565) |
| 0.0 | 0.3 | GO:0005610 | laminin-5 complex(GO:0005610) |
| 0.0 | 0.2 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.0 | 0.1 | GO:0034657 | GID complex(GO:0034657) |
| 0.0 | 0.6 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 0.1 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.0 | 0.3 | GO:0097431 | mitotic spindle pole(GO:0097431) |
| 0.0 | 0.5 | GO:0032839 | dendrite cytoplasm(GO:0032839) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.0 | GO:0051791 | medium-chain fatty acid metabolic process(GO:0051791) |
| 0.2 | 0.5 | GO:0061357 | positive regulation of Wnt protein secretion(GO:0061357) |
| 0.1 | 0.4 | GO:0006258 | UDP-glucose catabolic process(GO:0006258) galactose catabolic process via UDP-galactose(GO:0033499) |
| 0.1 | 0.4 | GO:1904481 | response to tetrahydrofolate(GO:1904481) cellular response to tetrahydrofolate(GO:1904482) |
| 0.1 | 0.4 | GO:1902498 | regulation of protein autoubiquitination(GO:1902498) |
| 0.1 | 0.7 | GO:0010993 | regulation of ubiquitin homeostasis(GO:0010993) free ubiquitin chain polymerization(GO:0010994) positive regulation of exit from mitosis(GO:0031536) |
| 0.1 | 0.6 | GO:0006931 | substrate-dependent cell migration, cell attachment to substrate(GO:0006931) |
| 0.1 | 0.3 | GO:0015692 | lead ion transport(GO:0015692) |
| 0.1 | 0.2 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.0 | 0.1 | GO:0032954 | regulation of cytokinetic process(GO:0032954) regulation of mitotic cytokinetic process(GO:1903436) positive regulation of mitotic cytokinetic process(GO:1903438) positive regulation of mitotic cytokinesis(GO:1903490) |
| 0.0 | 0.2 | GO:0060434 | bronchus morphogenesis(GO:0060434) |
| 0.0 | 0.6 | GO:1902222 | L-phenylalanine metabolic process(GO:0006558) L-phenylalanine catabolic process(GO:0006559) erythrose 4-phosphate/phosphoenolpyruvate family amino acid metabolic process(GO:1902221) erythrose 4-phosphate/phosphoenolpyruvate family amino acid catabolic process(GO:1902222) |
| 0.0 | 0.3 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 0.1 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) |
| 0.0 | 0.1 | GO:1904404 | cellular response to vitamin B1(GO:0071301) response to formaldehyde(GO:1904404) |
| 0.0 | 0.8 | GO:0008211 | glucocorticoid metabolic process(GO:0008211) |
| 0.0 | 1.1 | GO:0007339 | binding of sperm to zona pellucida(GO:0007339) |
| 0.0 | 0.3 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.0 | 0.6 | GO:0010226 | response to lithium ion(GO:0010226) |
| 0.0 | 0.2 | GO:0042373 | vitamin K metabolic process(GO:0042373) |
| 0.0 | 0.1 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.0 | 0.1 | GO:0008628 | hormone-mediated apoptotic signaling pathway(GO:0008628) positive regulation of osteoclast proliferation(GO:0090290) negative regulation of bone mineralization involved in bone maturation(GO:1900158) |
| 0.0 | 0.1 | GO:0033383 | geranyl diphosphate metabolic process(GO:0033383) geranyl diphosphate biosynthetic process(GO:0033384) farnesyl diphosphate biosynthetic process(GO:0045337) |
| 0.0 | 0.1 | GO:0048241 | epinephrine transport(GO:0048241) |
| 0.0 | 0.2 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.0 | 0.1 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
| 0.0 | 0.1 | GO:0009597 | detection of virus(GO:0009597) |
| 0.0 | 0.1 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.0 | 0.3 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.0 | 0.0 | GO:0051795 | positive regulation of catagen(GO:0051795) |
| 0.0 | 0.0 | GO:0046098 | guanine metabolic process(GO:0046098) |
| 0.0 | 0.1 | GO:0002408 | myeloid dendritic cell chemotaxis(GO:0002408) |
| 0.0 | 0.2 | GO:0048194 | Golgi vesicle budding(GO:0048194) |
| 0.0 | 0.1 | GO:0021615 | glossopharyngeal nerve morphogenesis(GO:0021615) |
| 0.0 | 0.4 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 0.0 | 0.0 | GO:1903371 | regulation of calcium ion-dependent exocytosis of neurotransmitter(GO:1903233) regulation of endoplasmic reticulum tubular network organization(GO:1903371) |
| 0.0 | 0.1 | GO:0042904 | 9-cis-retinoic acid biosynthetic process(GO:0042904) 9-cis-retinoic acid metabolic process(GO:0042905) |
| 0.0 | 0.1 | GO:0001507 | acetylcholine catabolic process in synaptic cleft(GO:0001507) acetylcholine catabolic process(GO:0006581) |
| 0.0 | 0.1 | GO:0098814 | spontaneous neurotransmitter secretion(GO:0061669) spontaneous synaptic transmission(GO:0098814) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.0 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.0 | 0.3 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |