Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
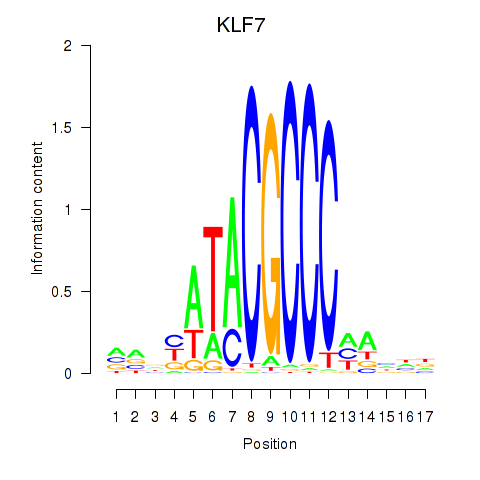
Results for KLF7
Z-value: 0.32
Transcription factors associated with KLF7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
KLF7
|
ENSG00000118263.10 | KLF7 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| KLF7 | hg19_v2_chr2_-_208030647_208030689, hg19_v2_chr2_-_208031943_208031991 | -0.09 | 7.4e-01 | Click! |
Activity profile of KLF7 motif
Sorted Z-values of KLF7 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of KLF7
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_96873921 | 0.40 |
ENST00000394166.3 |
NR2F2 |
nuclear receptor subfamily 2, group F, member 2 |
| chr19_-_39390440 | 0.31 |
ENST00000249396.7 ENST00000414941.1 ENST00000392081.2 |
SIRT2 |
sirtuin 2 |
| chr19_-_39390350 | 0.29 |
ENST00000447739.1 ENST00000358931.5 ENST00000407552.1 |
SIRT2 |
sirtuin 2 |
| chrX_-_92928557 | 0.25 |
ENST00000373079.3 ENST00000475430.2 |
NAP1L3 |
nucleosome assembly protein 1-like 3 |
| chr1_-_21948906 | 0.18 |
ENST00000374761.2 ENST00000599760.1 |
RAP1GAP |
RAP1 GTPase activating protein |
| chr7_+_27779714 | 0.17 |
ENST00000265393.6 ENST00000409980.1 ENST00000433216.2 ENST00000396319.2 |
TAX1BP1 |
Tax1 (human T-cell leukemia virus type I) binding protein 1 |
| chr17_-_19281203 | 0.17 |
ENST00000487415.2 |
B9D1 |
B9 protein domain 1 |
| chr6_+_31865552 | 0.17 |
ENST00000469372.1 ENST00000497706.1 |
C2 |
complement component 2 |
| chr17_+_41476327 | 0.17 |
ENST00000320033.4 |
ARL4D |
ADP-ribosylation factor-like 4D |
| chr1_-_19229248 | 0.16 |
ENST00000375341.3 |
ALDH4A1 |
aldehyde dehydrogenase 4 family, member A1 |
| chr19_-_14629224 | 0.16 |
ENST00000254322.2 |
DNAJB1 |
DnaJ (Hsp40) homolog, subfamily B, member 1 |
| chr2_-_188430478 | 0.16 |
ENST00000421427.1 |
TFPI |
tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) |
| chr1_-_19229014 | 0.15 |
ENST00000538839.1 ENST00000290597.5 |
ALDH4A1 |
aldehyde dehydrogenase 4 family, member A1 |
| chr6_-_33239612 | 0.15 |
ENST00000482399.1 ENST00000445902.2 |
VPS52 |
vacuolar protein sorting 52 homolog (S. cerevisiae) |
| chr5_+_141348598 | 0.15 |
ENST00000394520.2 ENST00000347642.3 |
RNF14 |
ring finger protein 14 |
| chr6_-_33239712 | 0.15 |
ENST00000436044.2 |
VPS52 |
vacuolar protein sorting 52 homolog (S. cerevisiae) |
| chrX_+_102631844 | 0.14 |
ENST00000372634.1 ENST00000299872.7 |
NGFRAP1 |
nerve growth factor receptor (TNFRSF16) associated protein 1 |
| chr19_+_39390587 | 0.14 |
ENST00000572515.1 ENST00000392079.3 ENST00000575359.1 ENST00000313582.5 |
NFKBIB |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr5_+_141348721 | 0.13 |
ENST00000507163.1 ENST00000394519.1 |
RNF14 |
ring finger protein 14 |
| chr19_+_39390320 | 0.12 |
ENST00000576510.1 |
NFKBIB |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr6_+_30585486 | 0.10 |
ENST00000259873.4 ENST00000506373.2 |
MRPS18B |
mitochondrial ribosomal protein S18B |
| chr16_+_31539183 | 0.08 |
ENST00000302312.4 |
AHSP |
alpha hemoglobin stabilizing protein |
| chr3_-_33138624 | 0.06 |
ENST00000445488.2 ENST00000307377.8 ENST00000440656.1 ENST00000436768.1 ENST00000307363.5 |
GLB1 |
galactosidase, beta 1 |
| chr16_+_31539197 | 0.05 |
ENST00000564707.1 |
AHSP |
alpha hemoglobin stabilizing protein |
| chr20_+_2821340 | 0.05 |
ENST00000380445.3 ENST00000380469.3 |
VPS16 |
vacuolar protein sorting 16 homolog (S. cerevisiae) |
| chr1_-_21503337 | 0.05 |
ENST00000400422.1 ENST00000602326.1 ENST00000411888.1 ENST00000438975.1 |
EIF4G3 |
eukaryotic translation initiation factor 4 gamma, 3 |
| chr17_+_38375574 | 0.04 |
ENST00000323571.4 ENST00000585043.1 ENST00000394103.3 ENST00000536600.1 |
WIPF2 |
WAS/WASL interacting protein family, member 2 |
| chr9_+_130565487 | 0.04 |
ENST00000373225.3 ENST00000431857.1 |
FPGS |
folylpolyglutamate synthase |
| chr17_+_73008755 | 0.03 |
ENST00000584208.1 ENST00000301585.5 |
ICT1 |
immature colon carcinoma transcript 1 |
| chr9_+_130565147 | 0.03 |
ENST00000373247.2 ENST00000373245.1 ENST00000393706.2 ENST00000373228.1 |
FPGS |
folylpolyglutamate synthase |
| chr8_-_6875778 | 0.03 |
ENST00000535841.1 ENST00000327857.2 |
DEFA1B DEFA3 |
defensin, alpha 1B defensin, alpha 3, neutrophil-specific |
| chr6_-_109703600 | 0.03 |
ENST00000512821.1 |
CD164 |
CD164 molecule, sialomucin |
| chr6_-_109703634 | 0.03 |
ENST00000324953.5 ENST00000310786.4 ENST00000275080.7 ENST00000413644.2 |
CD164 |
CD164 molecule, sialomucin |
| chr8_-_6837602 | 0.03 |
ENST00000382692.2 |
DEFA1 |
defensin, alpha 1 |
| chr17_-_18218237 | 0.02 |
ENST00000542570.1 |
TOP3A |
topoisomerase (DNA) III alpha |
| chr14_+_29234870 | 0.02 |
ENST00000382535.3 |
FOXG1 |
forkhead box G1 |
| chr6_+_33239787 | 0.02 |
ENST00000439602.2 ENST00000474973.1 |
RPS18 |
ribosomal protein S18 |
| chr19_-_7694417 | 0.02 |
ENST00000358368.4 ENST00000534844.1 |
XAB2 |
XPA binding protein 2 |
| chr3_+_186353756 | 0.01 |
ENST00000431018.1 ENST00000450521.1 ENST00000539949.1 |
FETUB |
fetuin B |
| chr13_-_108867101 | 0.01 |
ENST00000356922.4 |
LIG4 |
ligase IV, DNA, ATP-dependent |
| chr6_-_30585009 | 0.01 |
ENST00000376511.2 |
PPP1R10 |
protein phosphatase 1, regulatory subunit 10 |
| chr1_-_155243235 | 0.01 |
ENST00000355560.4 ENST00000368361.4 |
CLK2 |
CDC-like kinase 2 |
| chr1_-_157108266 | 0.00 |
ENST00000326786.4 |
ETV3 |
ets variant 3 |
| chr22_-_30987837 | 0.00 |
ENST00000335214.6 |
PES1 |
pescadillo ribosomal biogenesis factor 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:2000777 | positive regulation of oocyte maturation(GO:1900195) negative regulation of NLRP3 inflammasome complex assembly(GO:1900226) positive regulation of proteasomal ubiquitin-dependent protein catabolic process involved in cellular response to hypoxia(GO:2000777) |
| 0.1 | 0.4 | GO:0009956 | radial pattern formation(GO:0009956) |
| 0.1 | 0.3 | GO:0048611 | ectodermal digestive tract development(GO:0007439) embryonic ectodermal digestive tract development(GO:0048611) |
| 0.1 | 0.3 | GO:0019470 | 4-hydroxyproline catabolic process(GO:0019470) |
| 0.1 | 0.2 | GO:1903697 | negative regulation of microvillus assembly(GO:1903697) |
| 0.0 | 0.1 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.0 | 0.2 | GO:0007598 | blood coagulation, extrinsic pathway(GO:0007598) |
| 0.0 | 0.2 | GO:2000427 | positive regulation of apoptotic cell clearance(GO:2000427) |
| 0.0 | 0.1 | GO:1904016 | response to Thyroglobulin triiodothyronine(GO:1904016) |
| 0.0 | 0.2 | GO:0070389 | chaperone cofactor-dependent protein refolding(GO:0070389) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:1990745 | EARP complex(GO:1990745) |
| 0.1 | 0.6 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.0 | 0.2 | GO:0036038 | MKS complex(GO:0036038) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0034739 | histone deacetylase activity (H4-K16 specific)(GO:0034739) |
| 0.0 | 0.1 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.0 | 0.2 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 0.0 | 0.1 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.0 | 0.3 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |