Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
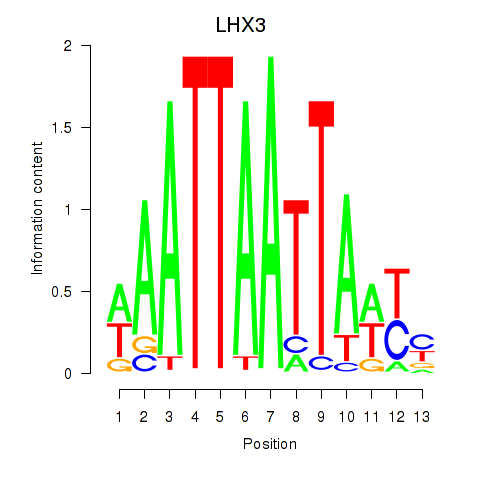
Results for LHX3
Z-value: 1.10
Transcription factors associated with LHX3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
LHX3
|
ENSG00000107187.11 | LHX3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| LHX3 | hg19_v2_chr9_-_139096955_139096979, hg19_v2_chr9_-_139094988_139095012 | -0.20 | 4.6e-01 | Click! |
Activity profile of LHX3 motif
Sorted Z-values of LHX3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of LHX3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_74301880 | 2.47 |
ENST00000395792.2 ENST00000226359.2 |
AFP |
alpha-fetoprotein |
| chr17_-_39093672 | 2.33 |
ENST00000209718.3 ENST00000436344.3 ENST00000485751.1 |
KRT23 |
keratin 23 (histone deacetylase inducible) |
| chr17_-_64225508 | 1.98 |
ENST00000205948.6 |
APOH |
apolipoprotein H (beta-2-glycoprotein I) |
| chr4_-_41884620 | 1.86 |
ENST00000504870.1 |
LINC00682 |
long intergenic non-protein coding RNA 682 |
| chr16_-_28634874 | 1.83 |
ENST00000395609.1 ENST00000350842.4 |
SULT1A1 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 1 |
| chr1_-_150208320 | 1.78 |
ENST00000534220.1 |
ANP32E |
acidic (leucine-rich) nuclear phosphoprotein 32 family, member E |
| chr4_+_155484155 | 1.76 |
ENST00000509493.1 |
FGB |
fibrinogen beta chain |
| chr5_-_78809950 | 1.71 |
ENST00000334082.6 |
HOMER1 |
homer homolog 1 (Drosophila) |
| chr7_-_87342564 | 1.69 |
ENST00000265724.3 ENST00000416177.1 |
ABCB1 |
ATP-binding cassette, sub-family B (MDR/TAP), member 1 |
| chr16_+_15489603 | 1.46 |
ENST00000568766.1 ENST00000287594.7 |
RP11-1021N1.1 MPV17L |
Uncharacterized protein MPV17 mitochondrial membrane protein-like |
| chrX_-_55057403 | 1.45 |
ENST00000396198.3 ENST00000335854.4 ENST00000455688.1 ENST00000330807.5 |
ALAS2 |
aminolevulinate, delta-, synthase 2 |
| chr11_+_59824060 | 1.44 |
ENST00000395032.2 ENST00000358152.2 |
MS4A3 |
membrane-spanning 4-domains, subfamily A, member 3 (hematopoietic cell-specific) |
| chr17_+_1674982 | 1.39 |
ENST00000572048.1 ENST00000573763.1 |
SERPINF1 |
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr11_+_59824127 | 1.37 |
ENST00000278865.3 |
MS4A3 |
membrane-spanning 4-domains, subfamily A, member 3 (hematopoietic cell-specific) |
| chr5_-_1882858 | 1.32 |
ENST00000511126.1 ENST00000231357.2 |
IRX4 |
iroquois homeobox 4 |
| chr4_+_155484103 | 1.26 |
ENST00000302068.4 |
FGB |
fibrinogen beta chain |
| chr1_-_24126023 | 1.15 |
ENST00000429356.1 |
GALE |
UDP-galactose-4-epimerase |
| chr6_+_26104104 | 1.15 |
ENST00000377803.2 |
HIST1H4C |
histone cluster 1, H4c |
| chr1_-_150208291 | 1.14 |
ENST00000533654.1 |
ANP32E |
acidic (leucine-rich) nuclear phosphoprotein 32 family, member E |
| chr14_-_21566731 | 1.14 |
ENST00000360947.3 |
ZNF219 |
zinc finger protein 219 |
| chr4_-_23891693 | 1.10 |
ENST00000264867.2 |
PPARGC1A |
peroxisome proliferator-activated receptor gamma, coactivator 1 alpha |
| chr2_-_166930131 | 1.09 |
ENST00000303395.4 ENST00000409050.1 ENST00000423058.2 ENST00000375405.3 |
SCN1A |
sodium channel, voltage-gated, type I, alpha subunit |
| chr14_+_39583427 | 0.93 |
ENST00000308317.6 ENST00000396249.2 ENST00000250379.8 ENST00000534684.2 ENST00000527381.1 |
GEMIN2 |
gem (nuclear organelle) associated protein 2 |
| chr12_+_49740700 | 0.86 |
ENST00000549441.2 ENST00000395069.3 |
DNAJC22 |
DnaJ (Hsp40) homolog, subfamily C, member 22 |
| chr1_-_150208363 | 0.85 |
ENST00000436748.2 |
ANP32E |
acidic (leucine-rich) nuclear phosphoprotein 32 family, member E |
| chr12_-_10282742 | 0.85 |
ENST00000298523.5 ENST00000396484.2 ENST00000310002.4 |
CLEC7A |
C-type lectin domain family 7, member A |
| chr3_+_157827841 | 0.84 |
ENST00000295930.3 ENST00000471994.1 ENST00000464171.1 ENST00000312179.6 ENST00000475278.2 |
RSRC1 |
arginine/serine-rich coiled-coil 1 |
| chr4_-_185275104 | 0.83 |
ENST00000317596.3 |
RP11-290F5.2 |
RP11-290F5.2 |
| chr3_+_63953415 | 0.80 |
ENST00000484332.1 |
ATXN7 |
ataxin 7 |
| chr15_+_65843130 | 0.79 |
ENST00000569894.1 |
PTPLAD1 |
protein tyrosine phosphatase-like A domain containing 1 |
| chrX_-_110655306 | 0.78 |
ENST00000371993.2 |
DCX |
doublecortin |
| chr17_-_27418537 | 0.76 |
ENST00000408971.2 |
TIAF1 |
TGFB1-induced anti-apoptotic factor 1 |
| chr3_-_141747950 | 0.75 |
ENST00000497579.1 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chrX_+_9431324 | 0.71 |
ENST00000407597.2 ENST00000424279.1 ENST00000536365.1 ENST00000441088.1 ENST00000380961.1 ENST00000415293.1 |
TBL1X |
transducin (beta)-like 1X-linked |
| chr16_+_28565230 | 0.71 |
ENST00000317058.3 |
CCDC101 |
coiled-coil domain containing 101 |
| chr17_-_60142609 | 0.70 |
ENST00000397786.2 |
MED13 |
mediator complex subunit 13 |
| chr3_-_47517302 | 0.68 |
ENST00000441517.2 ENST00000545718.1 |
SCAP |
SREBF chaperone |
| chr12_+_64798095 | 0.67 |
ENST00000332707.5 |
XPOT |
exportin, tRNA |
| chr4_+_41937131 | 0.64 |
ENST00000504986.1 ENST00000508448.1 ENST00000513702.1 ENST00000325094.5 |
TMEM33 |
transmembrane protein 33 |
| chr14_-_21567009 | 0.64 |
ENST00000556174.1 ENST00000554478.1 ENST00000553980.1 ENST00000421093.2 |
ZNF219 |
zinc finger protein 219 |
| chr7_-_140482926 | 0.62 |
ENST00000496384.2 |
BRAF |
v-raf murine sarcoma viral oncogene homolog B |
| chr5_+_59783540 | 0.61 |
ENST00000515734.2 |
PART1 |
prostate androgen-regulated transcript 1 (non-protein coding) |
| chr3_-_141719195 | 0.60 |
ENST00000397991.4 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr4_+_158142750 | 0.57 |
ENST00000505888.1 ENST00000449365.1 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr2_-_99279928 | 0.57 |
ENST00000414521.2 |
MGAT4A |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme A |
| chr17_-_72772462 | 0.57 |
ENST00000582870.1 ENST00000581136.1 ENST00000357814.3 ENST00000579218.1 ENST00000583476.1 ENST00000580301.1 ENST00000583757.1 ENST00000582524.1 |
NAT9 |
N-acetyltransferase 9 (GCN5-related, putative) |
| chr1_-_92952433 | 0.55 |
ENST00000294702.5 |
GFI1 |
growth factor independent 1 transcription repressor |
| chr2_+_234826016 | 0.55 |
ENST00000324695.4 ENST00000433712.2 |
TRPM8 |
transient receptor potential cation channel, subfamily M, member 8 |
| chr19_+_48949030 | 0.53 |
ENST00000253237.5 |
GRWD1 |
glutamate-rich WD repeat containing 1 |
| chr11_-_128894053 | 0.53 |
ENST00000392657.3 |
ARHGAP32 |
Rho GTPase activating protein 32 |
| chr3_+_69812877 | 0.51 |
ENST00000457080.1 ENST00000328528.6 |
MITF |
microphthalmia-associated transcription factor |
| chrX_+_107288197 | 0.50 |
ENST00000415430.3 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr1_-_150208412 | 0.49 |
ENST00000532744.1 ENST00000369114.5 ENST00000369115.2 ENST00000369116.4 |
ANP32E |
acidic (leucine-rich) nuclear phosphoprotein 32 family, member E |
| chr4_+_158141899 | 0.49 |
ENST00000264426.9 ENST00000506284.1 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr16_+_89334512 | 0.49 |
ENST00000602042.1 |
AC137932.1 |
AC137932.1 |
| chr18_+_72201829 | 0.49 |
ENST00000582365.1 |
CNDP1 |
carnosine dipeptidase 1 (metallopeptidase M20 family) |
| chr12_-_53994805 | 0.48 |
ENST00000328463.7 |
ATF7 |
activating transcription factor 7 |
| chr17_-_10372875 | 0.47 |
ENST00000255381.2 |
MYH4 |
myosin, heavy chain 4, skeletal muscle |
| chr8_-_57123815 | 0.44 |
ENST00000316981.3 ENST00000423799.2 ENST00000429357.2 |
PLAG1 |
pleiomorphic adenoma gene 1 |
| chr9_-_100000957 | 0.42 |
ENST00000366109.2 ENST00000607322.1 |
RP11-498P14.5 |
RP11-498P14.5 |
| chr19_-_43969796 | 0.42 |
ENST00000244333.3 |
LYPD3 |
LY6/PLAUR domain containing 3 |
| chr8_+_105352050 | 0.40 |
ENST00000297581.2 |
DCSTAMP |
dendrocyte expressed seven transmembrane protein |
| chr12_-_10282836 | 0.38 |
ENST00000304084.8 ENST00000353231.5 ENST00000525605.1 |
CLEC7A |
C-type lectin domain family 7, member A |
| chr5_+_59783941 | 0.38 |
ENST00000506884.1 ENST00000504876.2 |
PART1 |
prostate androgen-regulated transcript 1 (non-protein coding) |
| chr12_+_4385230 | 0.37 |
ENST00000536537.1 |
CCND2 |
cyclin D2 |
| chr10_-_115904361 | 0.35 |
ENST00000428953.1 ENST00000543782.1 |
C10orf118 |
chromosome 10 open reading frame 118 |
| chr19_-_49149553 | 0.34 |
ENST00000084798.4 |
CA11 |
carbonic anhydrase XI |
| chr5_-_39274617 | 0.33 |
ENST00000510188.1 |
FYB |
FYN binding protein |
| chr7_+_39017504 | 0.32 |
ENST00000403058.1 |
POU6F2 |
POU class 6 homeobox 2 |
| chr4_+_113568207 | 0.32 |
ENST00000511529.1 |
LARP7 |
La ribonucleoprotein domain family, member 7 |
| chr1_+_168250194 | 0.32 |
ENST00000367821.3 |
TBX19 |
T-box 19 |
| chrX_+_24167746 | 0.31 |
ENST00000428571.1 ENST00000539115.1 |
ZFX |
zinc finger protein, X-linked |
| chr4_+_158141843 | 0.30 |
ENST00000509417.1 ENST00000296526.7 |
GRIA2 |
glutamate receptor, ionotropic, AMPA 2 |
| chr12_-_86650045 | 0.29 |
ENST00000604798.1 |
MGAT4C |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme C (putative) |
| chr19_-_51920952 | 0.28 |
ENST00000356298.5 ENST00000339313.5 ENST00000529627.1 ENST00000439889.2 ENST00000353836.5 ENST00000432469.2 |
SIGLEC10 |
sialic acid binding Ig-like lectin 10 |
| chr12_-_86650077 | 0.28 |
ENST00000552808.2 ENST00000547225.1 |
MGAT4C |
mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme C (putative) |
| chr5_+_67586465 | 0.27 |
ENST00000336483.5 |
PIK3R1 |
phosphoinositide-3-kinase, regulatory subunit 1 (alpha) |
| chrX_+_107288239 | 0.27 |
ENST00000217957.5 |
VSIG1 |
V-set and immunoglobulin domain containing 1 |
| chr14_-_36990354 | 0.26 |
ENST00000518149.1 |
NKX2-1 |
NK2 homeobox 1 |
| chrX_+_108779004 | 0.26 |
ENST00000218004.1 |
NXT2 |
nuclear transport factor 2-like export factor 2 |
| chr17_-_39623681 | 0.26 |
ENST00000225899.3 |
KRT32 |
keratin 32 |
| chr12_-_52761262 | 0.24 |
ENST00000257901.3 |
KRT85 |
keratin 85 |
| chrX_+_84258832 | 0.24 |
ENST00000373173.2 |
APOOL |
apolipoprotein O-like |
| chr7_-_25268104 | 0.24 |
ENST00000222674.2 |
NPVF |
neuropeptide VF precursor |
| chr7_+_97361388 | 0.23 |
ENST00000350485.4 ENST00000346867.4 |
TAC1 |
tachykinin, precursor 1 |
| chr17_-_8263538 | 0.23 |
ENST00000535173.1 |
AC135178.1 |
HCG1985372; Uncharacterized protein; cDNA FLJ37541 fis, clone BRCAN2026340 |
| chr20_+_30063067 | 0.22 |
ENST00000201979.2 |
REM1 |
RAS (RAD and GEM)-like GTP-binding 1 |
| chr14_-_101295407 | 0.21 |
ENST00000596284.1 |
AL117190.2 |
AL117190.2 |
| chr3_+_138340049 | 0.20 |
ENST00000464668.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr1_-_160549235 | 0.18 |
ENST00000368054.3 ENST00000368048.3 ENST00000311224.4 ENST00000368051.3 ENST00000534968.1 |
CD84 |
CD84 molecule |
| chr8_-_25281747 | 0.18 |
ENST00000421054.2 |
GNRH1 |
gonadotropin-releasing hormone 1 (luteinizing-releasing hormone) |
| chr9_-_128246769 | 0.17 |
ENST00000444226.1 |
MAPKAP1 |
mitogen-activated protein kinase associated protein 1 |
| chr2_-_228582709 | 0.16 |
ENST00000541617.1 ENST00000409456.2 ENST00000409287.1 ENST00000258403.3 |
SLC19A3 |
solute carrier family 19 (thiamine transporter), member 3 |
| chr19_-_6279932 | 0.16 |
ENST00000252674.7 |
MLLT1 |
myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 1 |
| chr1_-_45988542 | 0.15 |
ENST00000424390.1 |
PRDX1 |
peroxiredoxin 1 |
| chr19_+_49199209 | 0.15 |
ENST00000522966.1 ENST00000425340.2 ENST00000391876.4 |
FUT2 |
fucosyltransferase 2 (secretor status included) |
| chr8_+_119294456 | 0.14 |
ENST00000366457.2 |
AC023590.1 |
Uncharacterized protein |
| chr17_-_4938712 | 0.13 |
ENST00000254853.5 ENST00000424747.1 |
SLC52A1 |
solute carrier family 52 (riboflavin transporter), member 1 |
| chr1_+_197237352 | 0.13 |
ENST00000538660.1 ENST00000367400.3 ENST00000367399.2 |
CRB1 |
crumbs homolog 1 (Drosophila) |
| chr12_-_42631529 | 0.12 |
ENST00000548917.1 |
YAF2 |
YY1 associated factor 2 |
| chrX_+_24167828 | 0.11 |
ENST00000379188.3 ENST00000419690.1 ENST00000379177.1 ENST00000304543.5 |
ZFX |
zinc finger protein, X-linked |
| chr1_+_115572415 | 0.11 |
ENST00000256592.1 |
TSHB |
thyroid stimulating hormone, beta |
| chr5_-_88119580 | 0.10 |
ENST00000539796.1 |
MEF2C |
myocyte enhancer factor 2C |
| chr4_-_120243545 | 0.10 |
ENST00000274024.3 |
FABP2 |
fatty acid binding protein 2, intestinal |
| chr6_+_72926145 | 0.09 |
ENST00000425662.2 ENST00000453976.2 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
| chr3_-_167191814 | 0.09 |
ENST00000466903.1 ENST00000264677.4 |
SERPINI2 |
serpin peptidase inhibitor, clade I (pancpin), member 2 |
| chr14_+_29241910 | 0.08 |
ENST00000399387.4 ENST00000552957.1 ENST00000548213.1 |
C14orf23 |
chromosome 14 open reading frame 23 |
| chr15_+_75970672 | 0.07 |
ENST00000435356.1 |
AC105020.1 |
Uncharacterized protein; cDNA FLJ12988 fis, clone NT2RP3000080 |
| chr13_-_70682590 | 0.06 |
ENST00000377844.4 |
KLHL1 |
kelch-like family member 1 |
| chr18_+_616672 | 0.06 |
ENST00000338387.7 |
CLUL1 |
clusterin-like 1 (retinal) |
| chr5_+_140557371 | 0.05 |
ENST00000239444.2 |
PCDHB8 |
protocadherin beta 8 |
| chr22_+_23487513 | 0.05 |
ENST00000263116.2 ENST00000341989.4 |
RAB36 |
RAB36, member RAS oncogene family |
| chr3_+_138340067 | 0.04 |
ENST00000479848.1 |
FAIM |
Fas apoptotic inhibitory molecule |
| chr5_-_59783882 | 0.03 |
ENST00000505507.2 ENST00000502484.2 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr4_+_3344141 | 0.02 |
ENST00000306648.7 |
RGS12 |
regulator of G-protein signaling 12 |
| chr11_+_5009424 | 0.01 |
ENST00000300762.1 |
MMP26 |
matrix metallopeptidase 26 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.7 | GO:0007206 | phospholipase C-activating G-protein coupled glutamate receptor signaling pathway(GO:0007206) |
| 0.4 | 1.1 | GO:0014878 | response to electrical stimulus involved in regulation of muscle adaptation(GO:0014878) regulation of progesterone biosynthetic process(GO:2000182) |
| 0.4 | 1.5 | GO:0010730 | negative regulation of hydrogen peroxide biosynthetic process(GO:0010730) |
| 0.3 | 3.0 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.3 | 1.4 | GO:0071279 | cellular response to cobalt ion(GO:0071279) |
| 0.2 | 1.7 | GO:1901529 | positive regulation of anion channel activity(GO:1901529) |
| 0.2 | 0.6 | GO:1903371 | regulation of endoplasmic reticulum tubular network organization(GO:1903371) |
| 0.2 | 0.6 | GO:0070105 | positive regulation of interleukin-6-mediated signaling pathway(GO:0070105) |
| 0.2 | 2.0 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 0.2 | 2.5 | GO:0001542 | ovulation from ovarian follicle(GO:0001542) |
| 0.1 | 0.7 | GO:0045541 | negative regulation of cholesterol biosynthetic process(GO:0045541) negative regulation of cholesterol metabolic process(GO:0090206) |
| 0.1 | 0.6 | GO:2000301 | negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.1 | 1.5 | GO:0032364 | oxygen homeostasis(GO:0032364) |
| 0.1 | 0.6 | GO:0050955 | thermoception(GO:0050955) |
| 0.1 | 0.3 | GO:0048867 | stem cell fate determination(GO:0048867) |
| 0.1 | 0.4 | GO:0072675 | multinuclear osteoclast differentiation(GO:0072674) osteoclast fusion(GO:0072675) |
| 0.1 | 1.1 | GO:0019227 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.1 | 1.2 | GO:0019388 | galactose catabolic process(GO:0019388) |
| 0.1 | 4.3 | GO:0043486 | histone exchange(GO:0043486) |
| 0.1 | 0.2 | GO:2000854 | positive regulation of corticosterone secretion(GO:2000854) |
| 0.1 | 0.2 | GO:1901842 | negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.1 | 1.8 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.1 | 0.8 | GO:0043569 | negative regulation of insulin-like growth factor receptor signaling pathway(GO:0043569) |
| 0.1 | 1.2 | GO:0009756 | carbohydrate mediated signaling(GO:0009756) |
| 0.1 | 0.2 | GO:1990637 | response to prolactin(GO:1990637) |
| 0.1 | 0.3 | GO:0021759 | globus pallidus development(GO:0021759) |
| 0.1 | 0.2 | GO:0071934 | thiamine transport(GO:0015888) thiamine transmembrane transport(GO:0071934) |
| 0.0 | 0.2 | GO:0032661 | regulation of interleukin-18 production(GO:0032661) |
| 0.0 | 0.8 | GO:0030277 | maintenance of gastrointestinal epithelium(GO:0030277) |
| 0.0 | 0.1 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.0 | 1.4 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.0 | 1.4 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.0 | 0.2 | GO:0030950 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) |
| 0.0 | 0.2 | GO:0032277 | negative regulation of gonadotropin secretion(GO:0032277) |
| 0.0 | 0.5 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.0 | 0.1 | GO:0003172 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.0 | 0.4 | GO:0060252 | positive regulation of glial cell proliferation(GO:0060252) |
| 0.0 | 2.8 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 0.6 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.0 | 0.9 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.7 | GO:2000272 | acylglycerol homeostasis(GO:0055090) triglyceride homeostasis(GO:0070328) negative regulation of receptor activity(GO:2000272) |
| 0.0 | 0.3 | GO:1900103 | positive regulation of endoplasmic reticulum unfolded protein response(GO:1900103) |
| 0.0 | 0.4 | GO:0071481 | cellular response to X-ray(GO:0071481) |
| 0.0 | 0.1 | GO:0042354 | fucose catabolic process(GO:0019317) L-fucose metabolic process(GO:0042354) L-fucose catabolic process(GO:0042355) |
| 0.0 | 0.7 | GO:0071431 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.0 | 1.3 | GO:0009798 | axis specification(GO:0009798) |
| 0.0 | 0.7 | GO:0046677 | response to antibiotic(GO:0046677) |
| 0.0 | 0.3 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.5 | GO:2000144 | positive regulation of DNA-templated transcription, initiation(GO:2000144) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) |
| 0.2 | 3.0 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.2 | 4.3 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.1 | 0.9 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.1 | 1.4 | GO:0043203 | axon hillock(GO:0043203) |
| 0.1 | 2.0 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.1 | 1.1 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.1 | 0.3 | GO:1990578 | perinuclear endoplasmic reticulum membrane(GO:1990578) |
| 0.1 | 0.4 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.1 | 1.1 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.0 | 1.7 | GO:0043034 | costamere(GO:0043034) |
| 0.0 | 0.2 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.0 | 0.2 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.0 | 2.8 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 2.8 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.0 | 0.7 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.5 | GO:0032982 | myosin filament(GO:0032982) |
| 0.0 | 2.0 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 0.4 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 0.5 | GO:0034399 | nuclear periphery(GO:0034399) |
| 0.0 | 0.7 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.0 | 0.2 | GO:0031932 | TORC2 complex(GO:0031932) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0003870 | 5-aminolevulinate synthase activity(GO:0003870) N-succinyltransferase activity(GO:0016749) |
| 0.4 | 2.0 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.4 | 1.2 | GO:0003978 | UDP-N-acetylglucosamine 4-epimerase activity(GO:0003974) UDP-glucose 4-epimerase activity(GO:0003978) |
| 0.3 | 1.7 | GO:0090554 | phosphatidylcholine-translocating ATPase activity(GO:0090554) |
| 0.3 | 1.8 | GO:0047894 | flavonol 3-sulfotransferase activity(GO:0047894) steroid sulfotransferase activity(GO:0050294) |
| 0.2 | 1.8 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.2 | 1.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.1 | 1.7 | GO:0035256 | G-protein coupled glutamate receptor binding(GO:0035256) |
| 0.1 | 1.4 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.1 | 4.3 | GO:0019212 | phosphatase inhibitor activity(GO:0019212) |
| 0.1 | 1.1 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 0.2 | GO:0015234 | thiamine transmembrane transporter activity(GO:0015234) thiamine uptake transmembrane transporter activity(GO:0015403) |
| 0.0 | 0.7 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.0 | 1.1 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.0 | 1.2 | GO:0008329 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.0 | 0.3 | GO:0043559 | insulin binding(GO:0043559) |
| 0.0 | 0.1 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.0 | 0.5 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.0 | 3.0 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.0 | 0.5 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.0 | 0.6 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.6 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.0 | 0.3 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.1 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 0.3 | GO:0001161 | intronic transcription regulatory region sequence-specific DNA binding(GO:0001161) intronic transcription regulatory region DNA binding(GO:0044213) |
| 0.0 | 0.7 | GO:0008536 | Ran GTPase binding(GO:0008536) |
| 0.0 | 1.5 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.7 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.4 | GO:0043236 | laminin binding(GO:0043236) |
| 0.0 | 0.5 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.3 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 2.7 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.1 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 0.6 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 1.4 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.7 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.8 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 1.7 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 1.4 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 1.5 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.0 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.1 | 1.7 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.1 | 1.8 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.1 | 1.8 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.1 | 1.4 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.1 | 1.5 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 1.1 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.6 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.0 | 1.2 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.3 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.0 | 0.9 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 0.3 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |