Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
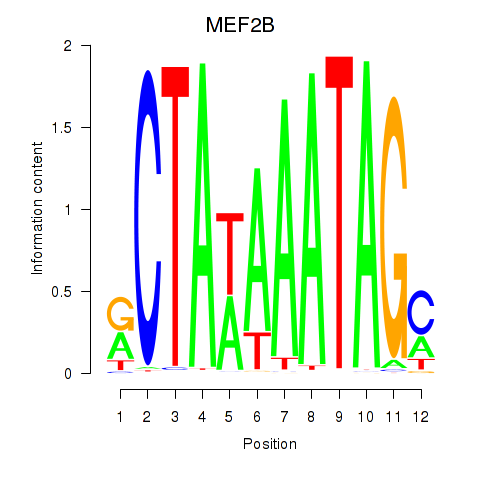
Results for MEF2B
Z-value: 1.11
Transcription factors associated with MEF2B
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
MEF2B
|
ENSG00000213999.11 | MEF2B |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| MEF2B | hg19_v2_chr19_-_19281054_19281098 | 0.32 | 2.3e-01 | Click! |
Activity profile of MEF2B motif
Sorted Z-values of MEF2B motif
Network of associatons between targets according to the STRING database.
First level regulatory network of MEF2B
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_-_111091948 | 3.96 |
ENST00000447165.2 |
NREP |
neuronal regeneration related protein |
| chr2_+_170366203 | 3.02 |
ENST00000284669.1 |
KLHL41 |
kelch-like family member 41 |
| chr5_+_53751445 | 2.73 |
ENST00000302005.1 |
HSPB3 |
heat shock 27kDa protein 3 |
| chr5_-_150521192 | 2.72 |
ENST00000523714.1 ENST00000521749.1 |
ANXA6 |
annexin A6 |
| chr12_+_119616447 | 2.40 |
ENST00000281938.2 |
HSPB8 |
heat shock 22kDa protein 8 |
| chr1_-_116311402 | 2.37 |
ENST00000261448.5 |
CASQ2 |
calsequestrin 2 (cardiac muscle) |
| chr2_-_211168332 | 2.26 |
ENST00000341685.4 |
MYL1 |
myosin, light chain 1, alkali; skeletal, fast |
| chr3_-_52486841 | 2.09 |
ENST00000496590.1 |
TNNC1 |
troponin C type 1 (slow) |
| chr1_-_201391149 | 2.04 |
ENST00000555948.1 ENST00000556362.1 |
TNNI1 |
troponin I type 1 (skeletal, slow) |
| chr9_+_100263912 | 1.92 |
ENST00000259365.4 |
TMOD1 |
tropomodulin 1 |
| chr3_+_138067521 | 1.87 |
ENST00000494949.1 |
MRAS |
muscle RAS oncogene homolog |
| chr10_-_75415825 | 1.55 |
ENST00000394810.2 |
SYNPO2L |
synaptopodin 2-like |
| chr3_+_138067666 | 1.53 |
ENST00000475711.1 ENST00000464896.1 |
MRAS |
muscle RAS oncogene homolog |
| chr3_+_138067314 | 1.52 |
ENST00000423968.2 |
MRAS |
muscle RAS oncogene homolog |
| chr2_-_86564776 | 1.49 |
ENST00000165698.5 ENST00000541910.1 ENST00000535845.1 |
REEP1 |
receptor accessory protein 1 |
| chr16_-_88717423 | 1.45 |
ENST00000568278.1 ENST00000569359.1 ENST00000567174.1 |
CYBA |
cytochrome b-245, alpha polypeptide |
| chr3_+_8543393 | 1.41 |
ENST00000157600.3 ENST00000415597.1 ENST00000535732.1 |
LMCD1 |
LIM and cysteine-rich domains 1 |
| chr3_+_8543561 | 1.37 |
ENST00000397386.3 |
LMCD1 |
LIM and cysteine-rich domains 1 |
| chr16_-_88717482 | 1.31 |
ENST00000261623.3 |
CYBA |
cytochrome b-245, alpha polypeptide |
| chrX_-_11284095 | 1.17 |
ENST00000303025.6 ENST00000534860.1 |
ARHGAP6 |
Rho GTPase activating protein 6 |
| chr2_+_33661382 | 1.16 |
ENST00000402538.3 |
RASGRP3 |
RAS guanyl releasing protein 3 (calcium and DAG-regulated) |
| chr3_+_8543533 | 1.15 |
ENST00000454244.1 |
LMCD1 |
LIM and cysteine-rich domains 1 |
| chr4_+_41362796 | 1.14 |
ENST00000508501.1 ENST00000512946.1 ENST00000313860.7 ENST00000512632.1 ENST00000512820.1 |
LIMCH1 |
LIM and calponin homology domains 1 |
| chr1_+_212782012 | 1.13 |
ENST00000341491.4 ENST00000366985.1 |
ATF3 |
activating transcription factor 3 |
| chr16_+_12058961 | 1.11 |
ENST00000053243.1 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr12_+_25205568 | 1.10 |
ENST00000548766.1 ENST00000556887.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr16_+_12059050 | 1.09 |
ENST00000396495.3 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr17_-_34417479 | 1.09 |
ENST00000225245.5 |
CCL3 |
chemokine (C-C motif) ligand 3 |
| chr12_-_111358372 | 1.08 |
ENST00000548438.1 ENST00000228841.8 |
MYL2 |
myosin, light chain 2, regulatory, cardiac, slow |
| chr2_-_190044480 | 1.05 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr8_+_11351876 | 1.02 |
ENST00000529894.1 |
BLK |
B lymphoid tyrosine kinase |
| chr8_+_11351494 | 1.02 |
ENST00000259089.4 |
BLK |
B lymphoid tyrosine kinase |
| chr12_+_25205666 | 1.01 |
ENST00000547044.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr11_+_7597639 | 0.99 |
ENST00000533792.1 |
PPFIBP2 |
PTPRF interacting protein, binding protein 2 (liprin beta 2) |
| chr4_+_120056939 | 0.97 |
ENST00000307128.5 |
MYOZ2 |
myozenin 2 |
| chr3_+_179370517 | 0.94 |
ENST00000263966.3 |
USP13 |
ubiquitin specific peptidase 13 (isopeptidase T-3) |
| chr2_+_233497931 | 0.93 |
ENST00000264059.3 |
EFHD1 |
EF-hand domain family, member D1 |
| chr1_+_145524891 | 0.83 |
ENST00000369304.3 |
ITGA10 |
integrin, alpha 10 |
| chr12_+_25205446 | 0.79 |
ENST00000557489.1 ENST00000354454.3 ENST00000536173.1 |
LRMP |
lymphoid-restricted membrane protein |
| chr16_+_84328429 | 0.76 |
ENST00000568638.1 |
WFDC1 |
WAP four-disulfide core domain 1 |
| chr16_+_84328252 | 0.74 |
ENST00000219454.5 |
WFDC1 |
WAP four-disulfide core domain 1 |
| chr16_+_12059091 | 0.73 |
ENST00000562385.1 |
TNFRSF17 |
tumor necrosis factor receptor superfamily, member 17 |
| chr5_+_155753745 | 0.67 |
ENST00000435422.3 ENST00000337851.4 ENST00000447401.1 |
SGCD |
sarcoglycan, delta (35kDa dystrophin-associated glycoprotein) |
| chr14_-_22005343 | 0.61 |
ENST00000327430.3 |
SALL2 |
spalt-like transcription factor 2 |
| chr15_+_100106155 | 0.57 |
ENST00000557785.1 ENST00000558049.1 ENST00000449277.2 |
MEF2A |
myocyte enhancer factor 2A |
| chr5_-_42811986 | 0.56 |
ENST00000511224.1 ENST00000507920.1 ENST00000510965.1 |
SEPP1 |
selenoprotein P, plasma, 1 |
| chrX_+_70521584 | 0.56 |
ENST00000373829.3 ENST00000538820.1 |
ITGB1BP2 |
integrin beta 1 binding protein (melusin) 2 |
| chr15_+_100106126 | 0.53 |
ENST00000558812.1 ENST00000338042.6 |
MEF2A |
myocyte enhancer factor 2A |
| chr8_+_1993152 | 0.53 |
ENST00000262113.4 |
MYOM2 |
myomesin 2 |
| chr15_+_100106244 | 0.53 |
ENST00000557942.1 |
MEF2A |
myocyte enhancer factor 2A |
| chr7_-_37488834 | 0.53 |
ENST00000310758.4 |
ELMO1 |
engulfment and cell motility 1 |
| chr5_-_42812143 | 0.49 |
ENST00000514985.1 |
SEPP1 |
selenoprotein P, plasma, 1 |
| chr8_+_107282389 | 0.49 |
ENST00000577661.1 ENST00000445937.1 |
RP11-395G23.3 OXR1 |
RP11-395G23.3 oxidation resistance 1 |
| chr8_+_1993173 | 0.47 |
ENST00000523438.1 |
MYOM2 |
myomesin 2 |
| chr7_+_120629653 | 0.46 |
ENST00000450913.2 ENST00000340646.5 |
CPED1 |
cadherin-like and PC-esterase domain containing 1 |
| chr17_-_5322786 | 0.41 |
ENST00000225696.4 |
NUP88 |
nucleoporin 88kDa |
| chr22_-_30642782 | 0.40 |
ENST00000249075.3 |
LIF |
leukemia inhibitory factor |
| chr17_+_80186908 | 0.37 |
ENST00000582743.1 ENST00000578684.1 ENST00000577650.1 ENST00000582715.1 |
SLC16A3 |
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr14_-_102704783 | 0.36 |
ENST00000522534.1 |
MOK |
MOK protein kinase |
| chr5_+_138609782 | 0.34 |
ENST00000361059.2 ENST00000514694.1 ENST00000504203.1 ENST00000502929.1 ENST00000394800.2 ENST00000509644.1 ENST00000505016.1 |
MATR3 |
matrin 3 |
| chr4_-_25865159 | 0.34 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr6_+_139094657 | 0.34 |
ENST00000332797.6 |
CCDC28A |
coiled-coil domain containing 28A |
| chr8_-_70745575 | 0.33 |
ENST00000524945.1 |
SLCO5A1 |
solute carrier organic anion transporter family, member 5A1 |
| chr15_+_68346501 | 0.33 |
ENST00000249636.6 |
PIAS1 |
protein inhibitor of activated STAT, 1 |
| chr1_-_204329013 | 0.32 |
ENST00000272203.3 ENST00000414478.1 |
PLEKHA6 |
pleckstrin homology domain containing, family A member 6 |
| chr6_+_42984723 | 0.27 |
ENST00000332245.8 |
KLHDC3 |
kelch domain containing 3 |
| chr3_+_186739636 | 0.25 |
ENST00000440338.1 ENST00000448044.1 |
ST6GAL1 |
ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 |
| chr10_+_118350522 | 0.24 |
ENST00000530319.1 ENST00000527980.1 ENST00000471549.1 ENST00000534537.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr3_-_176914238 | 0.23 |
ENST00000430069.1 ENST00000428970.1 |
TBL1XR1 |
transducin (beta)-like 1 X-linked receptor 1 |
| chr3_+_127634312 | 0.22 |
ENST00000407609.3 |
KBTBD12 |
kelch repeat and BTB (POZ) domain containing 12 |
| chr6_+_44194762 | 0.22 |
ENST00000371708.1 |
SLC29A1 |
solute carrier family 29 (equilibrative nucleoside transporter), member 1 |
| chr16_-_31439735 | 0.21 |
ENST00000287490.4 |
COX6A2 |
cytochrome c oxidase subunit VIa polypeptide 2 |
| chr19_-_40324767 | 0.20 |
ENST00000601972.1 ENST00000430012.2 ENST00000323039.5 ENST00000348817.3 |
DYRK1B |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1B |
| chr10_+_118350468 | 0.20 |
ENST00000358834.4 ENST00000528052.1 ENST00000442761.1 |
PNLIPRP1 |
pancreatic lipase-related protein 1 |
| chr9_-_74980113 | 0.18 |
ENST00000376962.5 ENST00000376960.4 ENST00000237937.3 |
ZFAND5 |
zinc finger, AN1-type domain 5 |
| chr7_+_143013198 | 0.18 |
ENST00000343257.2 |
CLCN1 |
chloride channel, voltage-sensitive 1 |
| chr3_+_54157480 | 0.18 |
ENST00000490478.1 |
CACNA2D3 |
calcium channel, voltage-dependent, alpha 2/delta subunit 3 |
| chr19_+_56713670 | 0.17 |
ENST00000534327.1 |
ZSCAN5C |
zinc finger and SCAN domain containing 5C |
| chr2_-_157198860 | 0.14 |
ENST00000409572.1 |
NR4A2 |
nuclear receptor subfamily 4, group A, member 2 |
| chr1_+_170633047 | 0.14 |
ENST00000239461.6 ENST00000497230.2 |
PRRX1 |
paired related homeobox 1 |
| chr17_-_42200996 | 0.13 |
ENST00000587135.1 ENST00000225983.6 ENST00000393622.2 ENST00000588703.1 |
HDAC5 |
histone deacetylase 5 |
| chr8_-_27115903 | 0.10 |
ENST00000350889.4 ENST00000519997.1 ENST00000519614.1 ENST00000522908.1 ENST00000265770.7 |
STMN4 |
stathmin-like 4 |
| chr3_+_127634069 | 0.09 |
ENST00000405109.1 |
KBTBD12 |
kelch repeat and BTB (POZ) domain containing 12 |
| chr5_-_66492562 | 0.09 |
ENST00000256447.4 |
CD180 |
CD180 molecule |
| chr9_+_116327326 | 0.07 |
ENST00000342620.5 |
RGS3 |
regulator of G-protein signaling 3 |
| chr7_+_30951461 | 0.07 |
ENST00000311813.4 |
AQP1 |
aquaporin 1 (Colton blood group) |
| chr4_+_44680429 | 0.06 |
ENST00000281543.5 |
GUF1 |
GUF1 GTPase homolog (S. cerevisiae) |
| chr3_+_46924829 | 0.04 |
ENST00000313049.5 |
PTH1R |
parathyroid hormone 1 receptor |
| chr19_-_40324255 | 0.04 |
ENST00000593685.1 ENST00000600611.1 |
DYRK1B |
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1B |
| chr17_-_42200958 | 0.04 |
ENST00000336057.5 |
HDAC5 |
histone deacetylase 5 |
| chr7_+_86781847 | 0.03 |
ENST00000432366.2 ENST00000423590.2 ENST00000394703.5 |
DMTF1 |
cyclin D binding myb-like transcription factor 1 |
| chr8_-_27115931 | 0.02 |
ENST00000523048.1 |
STMN4 |
stathmin-like 4 |
| chr8_+_95565947 | 0.01 |
ENST00000523011.1 |
RP11-267M23.4 |
RP11-267M23.4 |
| chr11_-_47400078 | 0.01 |
ENST00000378538.3 |
SPI1 |
spleen focus forming virus (SFFV) proviral integration oncogene |
| chr12_+_15699286 | 0.00 |
ENST00000442921.2 ENST00000542557.1 ENST00000445537.2 ENST00000544244.1 |
PTPRO |
protein tyrosine phosphatase, receptor type, O |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:1904288 | BAT3 complex binding(GO:1904288) |
| 0.2 | 1.1 | GO:0004698 | calcium-dependent protein kinase C activity(GO:0004698) |
| 0.2 | 1.0 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.2 | 0.8 | GO:0098639 | collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.2 | 1.8 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.1 | 4.9 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.1 | 2.8 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.1 | 1.9 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 0.3 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.1 | 1.2 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.1 | 0.3 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.1 | 5.0 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.9 | GO:0097493 | muscle alpha-actinin binding(GO:0051371) structural molecule activity conferring elasticity(GO:0097493) |
| 0.0 | 0.2 | GO:0001025 | RNA polymerase III transcription factor binding(GO:0001025) |
| 0.0 | 0.4 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.6 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.0 | 0.4 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.0 | 1.6 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 0.1 | GO:0035379 | carbon dioxide transmembrane transporter activity(GO:0035379) |
| 0.0 | 2.0 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.4 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 4.8 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 1.5 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 2.0 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 0.2 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.0 | 2.0 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 4.7 | ST INTEGRIN SIGNALING PATHWAY | Integrin Signaling Pathway |
| 0.0 | 0.3 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.0 | 2.0 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 1.2 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 1.2 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 2.7 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.5 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 1.1 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.0 | 0.9 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.8 | GO:1904845 | response to L-glutamine(GO:1904844) cellular response to L-glutamine(GO:1904845) |
| 0.5 | 2.1 | GO:0032972 | diaphragm contraction(GO:0002086) regulation of muscle filament sliding speed(GO:0032972) |
| 0.4 | 1.1 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.3 | 1.9 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.3 | 1.6 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 0.3 | 1.1 | GO:0098735 | positive regulation of the force of heart contraction(GO:0098735) |
| 0.3 | 1.1 | GO:1903224 | regulation of endodermal cell differentiation(GO:1903224) |
| 0.2 | 2.4 | GO:0071313 | cellular response to caffeine(GO:0071313) |
| 0.2 | 0.9 | GO:0035523 | protein K29-linked deubiquitination(GO:0035523) maintenance of unfolded protein(GO:0036506) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) |
| 0.2 | 3.0 | GO:0031272 | regulation of pseudopodium assembly(GO:0031272) |
| 0.2 | 3.9 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade(GO:0070886) |
| 0.2 | 1.1 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.1 | 2.0 | GO:0014883 | transition between fast and slow fiber(GO:0014883) |
| 0.1 | 0.4 | GO:0043974 | histone H3-K27 acetylation(GO:0043974) regulation of histone H3-K27 acetylation(GO:1901674) |
| 0.1 | 0.9 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.1 | 2.7 | GO:0051560 | mitochondrial calcium ion homeostasis(GO:0051560) |
| 0.1 | 1.5 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.1 | 0.4 | GO:0035879 | plasma membrane lactate transport(GO:0035879) |
| 0.1 | 0.4 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.1 | 2.0 | GO:0050855 | regulation of B cell receptor signaling pathway(GO:0050855) |
| 0.0 | 0.1 | GO:0021986 | epithalamus development(GO:0021538) habenula development(GO:0021986) |
| 0.0 | 0.2 | GO:0015862 | uridine transport(GO:0015862) |
| 0.0 | 1.2 | GO:0051895 | negative regulation of focal adhesion assembly(GO:0051895) |
| 0.0 | 0.2 | GO:0060613 | fat pad development(GO:0060613) |
| 0.0 | 2.9 | GO:0002260 | lymphocyte homeostasis(GO:0002260) |
| 0.0 | 0.3 | GO:0051152 | positive regulation of smooth muscle cell differentiation(GO:0051152) |
| 0.0 | 4.9 | GO:1990823 | response to leukemia inhibitory factor(GO:1990823) cellular response to leukemia inhibitory factor(GO:1990830) |
| 0.0 | 0.5 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.0 | 0.1 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 0.0 | 2.9 | GO:0006903 | vesicle targeting(GO:0006903) |
| 0.0 | 0.1 | GO:0072229 | carbon dioxide transmembrane transport(GO:0035378) proximal convoluted tubule development(GO:0072019) metanephric proximal convoluted tubule development(GO:0072229) |
| 0.0 | 0.3 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.0 | 2.4 | GO:1900034 | regulation of cellular response to heat(GO:1900034) |
| 0.0 | 1.6 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.0 | 0.1 | GO:0048664 | neuron fate determination(GO:0048664) |
| 0.0 | 1.5 | GO:0061045 | negative regulation of wound healing(GO:0061045) |
| 0.0 | 0.9 | GO:0033275 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.0 | 0.2 | GO:0019227 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.0 | 0.5 | GO:0016601 | Rac protein signal transduction(GO:0016601) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 8.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 2.8 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.0 | 1.6 | REACTOME ERK MAPK TARGETS | Genes involved in ERK/MAPK targets |
| 0.0 | 2.0 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 1.1 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 0.4 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 1.1 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 1.1 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.3 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.0 | 0.3 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 0.5 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.0 | 0.4 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.4 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 0.4 | 1.1 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.4 | 1.1 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.3 | 2.1 | GO:1990584 | cardiac Troponin complex(GO:1990584) |
| 0.3 | 0.8 | GO:0034680 | integrin alpha10-beta1 complex(GO:0034680) |
| 0.2 | 2.8 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.1 | 1.7 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.1 | 3.0 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.1 | 1.1 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.1 | 0.7 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.1 | 2.7 | GO:0005859 | muscle myosin complex(GO:0005859) |
| 0.1 | 3.1 | GO:0036379 | myofilament(GO:0036379) |
| 0.0 | 2.9 | GO:0035577 | azurophil granule membrane(GO:0035577) |
| 0.0 | 0.8 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) |
| 0.0 | 0.1 | GO:0020005 | symbiont-containing vacuole(GO:0020003) symbiont-containing vacuole membrane(GO:0020005) |
| 0.0 | 2.7 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.0 | 0.6 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 0.2 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.0 | 2.0 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 2.0 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |