Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
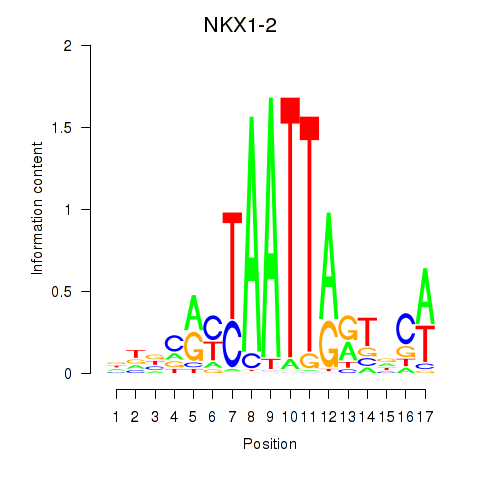
Results for NKX1-2_RAX
Z-value: 0.53
Transcription factors associated with NKX1-2_RAX
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NKX1-2
|
ENSG00000229544.6 | NKX1-2 |
|
RAX
|
ENSG00000134438.9 | RAX |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| RAX | hg19_v2_chr18_-_56940611_56940660 | 0.43 | 9.5e-02 | Click! |
Activity profile of NKX1-2_RAX motif
Sorted Z-values of NKX1-2_RAX motif
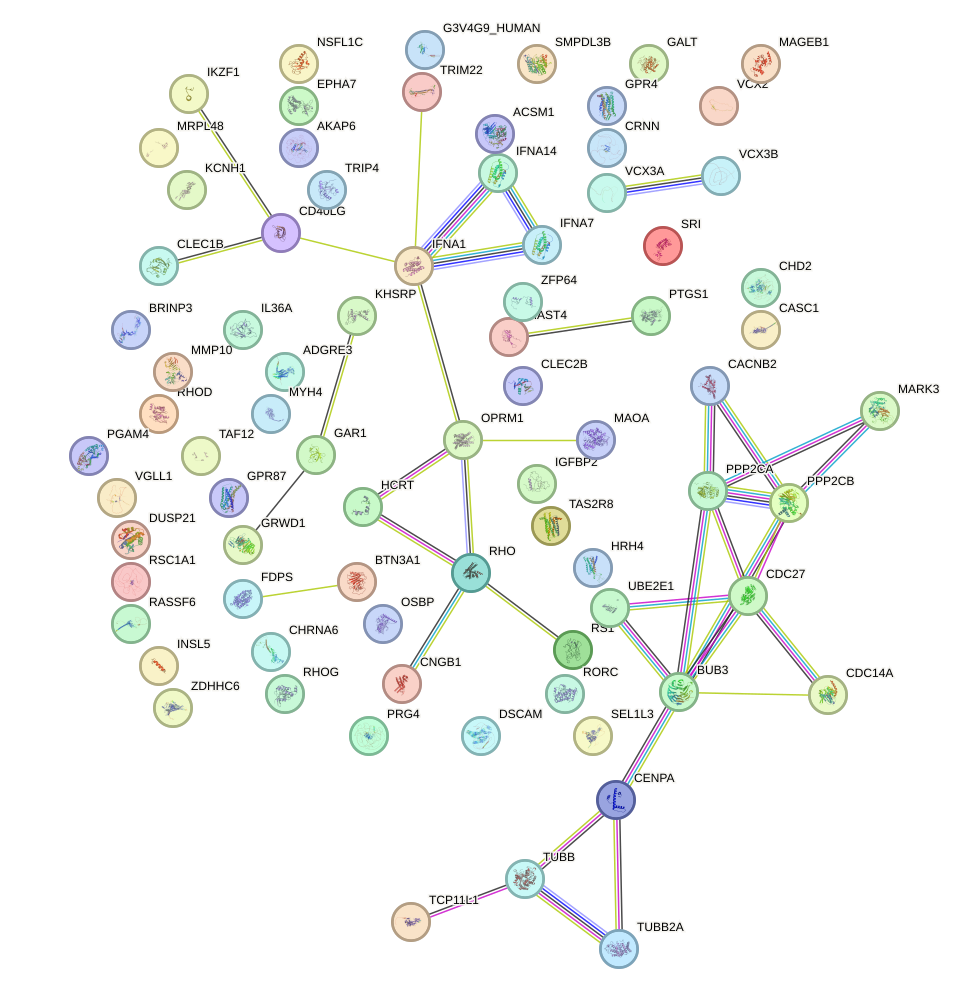
Network of associatons between targets according to the STRING database.
First level regulatory network of NKX1-2_RAX
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_62521614 | 1.87 |
ENST00000527994.1 ENST00000394807.3 |
ZBTB3 |
zinc finger and BTB domain containing 3 |
| chr7_-_87856280 | 1.56 |
ENST00000490437.1 ENST00000431660.1 |
SRI |
sorcin |
| chr7_-_87856303 | 1.55 |
ENST00000394641.3 |
SRI |
sorcin |
| chr4_-_25865159 | 1.44 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr3_-_151034734 | 1.41 |
ENST00000260843.4 |
GPR87 |
G protein-coupled receptor 87 |
| chr5_+_66300446 | 0.75 |
ENST00000261569.7 |
MAST4 |
microtubule associated serine/threonine kinase family member 4 |
| chr6_+_34204642 | 0.58 |
ENST00000347617.6 ENST00000401473.3 ENST00000311487.5 ENST00000447654.1 ENST00000395004.3 |
HMGA1 |
high mobility group AT-hook 1 |
| chr12_-_10022735 | 0.53 |
ENST00000228438.2 |
CLEC2B |
C-type lectin domain family 2, member B |
| chr10_-_105845674 | 0.45 |
ENST00000353479.5 ENST00000369733.3 |
COL17A1 |
collagen, type XVII, alpha 1 |
| chr12_+_32832203 | 0.40 |
ENST00000553257.1 ENST00000549701.1 ENST00000358214.5 ENST00000266481.6 ENST00000551476.1 ENST00000550154.1 ENST00000547312.1 ENST00000414834.2 ENST00000381000.4 ENST00000548750.1 |
DNM1L |
dynamin 1-like |
| chr11_+_5710919 | 0.38 |
ENST00000379965.3 ENST00000425490.1 |
TRIM22 |
tripartite motif containing 22 |
| chr11_-_102651343 | 0.37 |
ENST00000279441.4 ENST00000539681.1 |
MMP10 |
matrix metallopeptidase 10 (stromelysin 2) |
| chr2_+_90077680 | 0.34 |
ENST00000390270.2 |
IGKV3D-20 |
immunoglobulin kappa variable 3D-20 |
| chr17_-_41466555 | 0.29 |
ENST00000586231.1 |
LINC00910 |
long intergenic non-protein coding RNA 910 |
| chr1_+_155278539 | 0.28 |
ENST00000447866.1 |
FDPS |
farnesyl diphosphate synthase |
| chr3_+_183853052 | 0.26 |
ENST00000273783.3 ENST00000432569.1 ENST00000444495.1 |
EIF2B5 |
eukaryotic translation initiation factor 2B, subunit 5 epsilon, 82kDa |
| chr6_-_111927062 | 0.26 |
ENST00000359831.4 |
TRAF3IP2 |
TRAF3 interacting protein 2 |
| chr1_+_155278625 | 0.25 |
ENST00000368356.4 ENST00000356657.6 |
FDPS |
farnesyl diphosphate synthase |
| chr12_-_112279694 | 0.24 |
ENST00000443596.1 ENST00000442119.1 |
MAPKAPK5-AS1 |
MAPKAPK5 antisense RNA 1 |
| chrX_+_43515467 | 0.24 |
ENST00000338702.3 ENST00000542639.1 |
MAOA |
monoamine oxidase A |
| chr10_+_124914285 | 0.23 |
ENST00000407911.2 |
BUB3 |
BUB3 mitotic checkpoint protein |
| chr20_-_50722183 | 0.23 |
ENST00000371523.4 |
ZFP64 |
ZFP64 zinc finger protein |
| chr2_+_217524323 | 0.22 |
ENST00000456764.1 |
IGFBP2 |
insulin-like growth factor binding protein 2, 36kDa |
| chrX_-_77225135 | 0.21 |
ENST00000458128.1 |
PGAM4 |
phosphoglycerate mutase family member 4 |
| chr11_-_59383617 | 0.20 |
ENST00000263847.1 |
OSBP |
oxysterol binding protein |
| chr15_+_64680003 | 0.20 |
ENST00000261884.3 |
TRIP4 |
thyroid hormone receptor interactor 4 |
| chr6_+_26402517 | 0.19 |
ENST00000414912.2 |
BTN3A1 |
butyrophilin, subfamily 3, member A1 |
| chr1_-_190446759 | 0.16 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr8_-_102803163 | 0.16 |
ENST00000523645.1 ENST00000520346.1 ENST00000220931.6 ENST00000522448.1 ENST00000522951.1 ENST00000522252.1 ENST00000519098.1 |
NCALD |
neurocalcin delta |
| chr19_+_48949030 | 0.16 |
ENST00000253237.5 |
GRWD1 |
glutamate-rich WD repeat containing 1 |
| chr12_-_12837423 | 0.16 |
ENST00000540510.1 |
GPR19 |
G protein-coupled receptor 19 |
| chr1_-_67266939 | 0.15 |
ENST00000304526.2 |
INSL5 |
insulin-like 5 |
| chr17_-_45266542 | 0.15 |
ENST00000531206.1 ENST00000527547.1 ENST00000446365.2 ENST00000575483.1 ENST00000066544.3 |
CDC27 |
cell division cycle 27 |
| chr3_+_23851928 | 0.15 |
ENST00000467766.1 ENST00000424381.1 |
UBE2E1 |
ubiquitin-conjugating enzyme E2E 1 |
| chrX_+_30261847 | 0.15 |
ENST00000378981.3 ENST00000397550.1 |
MAGEB1 |
melanoma antigen family B, 1 |
| chr14_+_103851712 | 0.15 |
ENST00000440884.3 ENST00000416682.2 ENST00000429436.2 ENST00000303622.9 |
MARK3 |
MAP/microtubule affinity-regulating kinase 3 |
| chrX_+_44703249 | 0.14 |
ENST00000339042.4 |
DUSP21 |
dual specificity phosphatase 21 |
| chrX_+_77154935 | 0.13 |
ENST00000481445.1 |
COX7B |
cytochrome c oxidase subunit VIIb |
| chr6_-_94129244 | 0.13 |
ENST00000369303.4 ENST00000369297.1 |
EPHA7 |
EPH receptor A7 |
| chr11_+_73498898 | 0.13 |
ENST00000535529.1 ENST00000497094.2 ENST00000411840.2 ENST00000535277.1 ENST00000398483.3 ENST00000542303.1 |
MRPL48 |
mitochondrial ribosomal protein L48 |
| chr11_-_3859089 | 0.13 |
ENST00000396979.1 |
RHOG |
ras homolog family member G |
| chr19_-_6424783 | 0.13 |
ENST00000398148.3 |
KHSRP |
KH-type splicing regulatory protein |
| chr8_-_42623747 | 0.12 |
ENST00000534622.1 |
CHRNA6 |
cholinergic receptor, nicotinic, alpha 6 (neuronal) |
| chr1_+_153747746 | 0.11 |
ENST00000368661.3 |
SLC27A3 |
solute carrier family 27 (fatty acid transporter), member 3 |
| chrX_+_135730373 | 0.11 |
ENST00000370628.2 |
CD40LG |
CD40 ligand |
| chr1_+_186265399 | 0.11 |
ENST00000367486.3 ENST00000367484.3 ENST00000533951.1 ENST00000367482.4 ENST00000367483.4 ENST00000367485.4 ENST00000445192.2 |
PRG4 |
proteoglycan 4 |
| chr6_-_111927449 | 0.11 |
ENST00000368761.5 ENST00000392556.4 ENST00000340026.6 |
TRAF3IP2 |
TRAF3 interacting protein 2 |
| chr1_+_62439037 | 0.10 |
ENST00000545929.1 |
INADL |
InaD-like (Drosophila) |
| chr10_+_124913930 | 0.08 |
ENST00000368858.5 |
BUB3 |
BUB3 mitotic checkpoint protein |
| chrX_+_135618258 | 0.08 |
ENST00000440515.1 ENST00000456412.1 |
VGLL1 |
vestigial like 1 (Drosophila) |
| chr5_-_20575959 | 0.08 |
ENST00000507958.1 |
CDH18 |
cadherin 18, type 2 |
| chr8_+_105235572 | 0.08 |
ENST00000523362.1 |
RIMS2 |
regulating synaptic membrane exocytosis 2 |
| chr1_+_100818156 | 0.08 |
ENST00000336454.3 |
CDC14A |
cell division cycle 14A |
| chr9_+_125133315 | 0.07 |
ENST00000223423.4 ENST00000362012.2 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr1_+_153746683 | 0.07 |
ENST00000271857.2 |
SLC27A3 |
solute carrier family 27 (fatty acid transporter), member 3 |
| chrX_-_6453159 | 0.07 |
ENST00000381089.3 ENST00000398729.1 |
VCX3A |
variable charge, X-linked 3A |
| chr17_-_40337470 | 0.07 |
ENST00000293330.1 |
HCRT |
hypocretin (orexin) neuropeptide precursor |
| chr19_-_14785622 | 0.07 |
ENST00000443157.2 |
EMR3 |
egf-like module containing, mucin-like, hormone receptor-like 3 |
| chr9_-_21202204 | 0.07 |
ENST00000239347.3 |
IFNA7 |
interferon, alpha 7 |
| chr7_+_50348268 | 0.07 |
ENST00000438033.1 ENST00000439701.1 |
IKZF1 |
IKAROS family zinc finger 1 (Ikaros) |
| chr19_-_14785698 | 0.07 |
ENST00000344373.4 ENST00000595472.1 |
EMR3 |
egf-like module containing, mucin-like, hormone receptor-like 3 |
| chr1_-_152386732 | 0.06 |
ENST00000271835.3 |
CRNN |
cornulin |
| chr8_-_30670384 | 0.06 |
ENST00000221138.4 ENST00000518243.1 |
PPP2CB |
protein phosphatase 2, catalytic subunit, beta isozyme |
| chr2_+_27008865 | 0.06 |
ENST00000335756.4 ENST00000233505.8 |
CENPA |
centromere protein A |
| chr1_-_211307404 | 0.06 |
ENST00000367007.4 |
KCNH1 |
potassium voltage-gated channel, subfamily H (eag-related), member 1 |
| chr19_+_11071546 | 0.06 |
ENST00000358026.2 |
SMARCA4 |
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 |
| chr2_-_169887827 | 0.06 |
ENST00000263817.6 |
ABCB11 |
ATP-binding cassette, sub-family B (MDR/TAP), member 11 |
| chr2_+_166095898 | 0.06 |
ENST00000424833.1 ENST00000375437.2 ENST00000357398.3 |
SCN2A |
sodium channel, voltage-gated, type II, alpha subunit |
| chr22_+_22730353 | 0.06 |
ENST00000390296.2 |
IGLV5-45 |
immunoglobulin lambda variable 5-45 |
| chr19_-_14785674 | 0.06 |
ENST00000253673.5 |
EMR3 |
egf-like module containing, mucin-like, hormone receptor-like 3 |
| chr1_-_28969517 | 0.05 |
ENST00000263974.4 ENST00000373824.4 |
TAF12 |
TAF12 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 20kDa |
| chr9_-_21239978 | 0.05 |
ENST00000380222.2 |
IFNA14 |
interferon, alpha 14 |
| chr4_-_66536196 | 0.05 |
ENST00000511294.1 |
EPHA5 |
EPH receptor A5 |
| chr3_+_129247479 | 0.05 |
ENST00000296271.3 |
RHO |
rhodopsin |
| chr3_+_186560462 | 0.05 |
ENST00000412955.2 |
ADIPOQ |
adiponectin, C1Q and collagen domain containing |
| chr3_+_186560476 | 0.05 |
ENST00000320741.2 ENST00000444204.2 |
ADIPOQ |
adiponectin, C1Q and collagen domain containing |
| chr2_-_89597542 | 0.05 |
ENST00000465170.1 |
IGKV1-37 |
immunoglobulin kappa variable 1-37 (non-functional) |
| chr9_-_21368075 | 0.04 |
ENST00000449498.1 |
IFNA13 |
interferon, alpha 13 |
| chr6_+_30687978 | 0.04 |
ENST00000327892.8 ENST00000435534.1 |
TUBB |
tubulin, beta class I |
| chrX_-_18690210 | 0.04 |
ENST00000379984.3 |
RS1 |
retinoschisin 1 |
| chr4_+_80584903 | 0.04 |
ENST00000506460.1 |
RP11-452C8.1 |
RP11-452C8.1 |
| chr17_-_10372875 | 0.04 |
ENST00000255381.2 |
MYH4 |
myosin, heavy chain 4, skeletal muscle |
| chr10_-_114206649 | 0.04 |
ENST00000369404.3 ENST00000369405.3 |
ZDHHC6 |
zinc finger, DHHC-type containing 6 |
| chr14_+_32798547 | 0.04 |
ENST00000557354.1 ENST00000557102.1 ENST00000557272.1 |
AKAP6 |
A kinase (PRKA) anchor protein 6 |
| chr1_+_28261492 | 0.03 |
ENST00000373894.3 |
SMPDL3B |
sphingomyelin phosphodiesterase, acid-like 3B |
| chr4_-_74486347 | 0.03 |
ENST00000342081.3 |
RASSF6 |
Ras association (RalGDS/AF-6) domain family member 6 |
| chr6_+_154360357 | 0.03 |
ENST00000330432.7 ENST00000360422.4 |
OPRM1 |
opioid receptor, mu 1 |
| chr1_+_100818009 | 0.03 |
ENST00000370125.2 ENST00000361544.6 ENST00000370124.3 |
CDC14A |
cell division cycle 14A |
| chr4_+_110736659 | 0.03 |
ENST00000394631.3 ENST00000226796.6 |
GAR1 |
GAR1 ribonucleoprotein |
| chr12_-_25348007 | 0.03 |
ENST00000354189.5 ENST00000545133.1 ENST00000554347.1 ENST00000395987.3 ENST00000320267.9 ENST00000395990.2 ENST00000537577.1 |
CASC1 |
cancer susceptibility candidate 1 |
| chr15_+_93443419 | 0.03 |
ENST00000557381.1 ENST00000420239.2 |
CHD2 |
chromodomain helicase DNA binding protein 2 |
| chr4_-_66536057 | 0.02 |
ENST00000273854.3 |
EPHA5 |
EPH receptor A5 |
| chr16_-_20702578 | 0.02 |
ENST00000307493.4 ENST00000219151.4 |
ACSM1 |
acyl-CoA synthetase medium-chain family member 1 |
| chr10_+_124913793 | 0.02 |
ENST00000368865.4 ENST00000538238.1 ENST00000368859.2 |
BUB3 |
BUB3 mitotic checkpoint protein |
| chr11_+_33061543 | 0.02 |
ENST00000432887.1 ENST00000528898.1 ENST00000531632.2 |
TCP11L1 |
t-complex 11, testis-specific-like 1 |
| chr12_-_10959892 | 0.02 |
ENST00000240615.2 |
TAS2R8 |
taste receptor, type 2, member 8 |
| chr10_+_18549645 | 0.02 |
ENST00000396576.2 |
CACNB2 |
calcium channel, voltage-dependent, beta 2 subunit |
| chr2_+_113763031 | 0.02 |
ENST00000259211.6 |
IL36A |
interleukin 36, alpha |
| chr9_+_34646651 | 0.02 |
ENST00000378842.3 |
GALT |
galactose-1-phosphate uridylyltransferase |
| chrX_-_8139308 | 0.02 |
ENST00000317103.4 |
VCX2 |
variable charge, X-linked 2 |
| chr7_-_86849883 | 0.01 |
ENST00000433078.1 |
TMEM243 |
transmembrane protein 243, mitochondrial |
| chr1_-_151804314 | 0.01 |
ENST00000318247.6 |
RORC |
RAR-related orphan receptor C |
| chr18_+_22040620 | 0.01 |
ENST00000426880.2 |
HRH4 |
histamine receptor H4 |
| chr9_+_34646624 | 0.01 |
ENST00000450095.2 ENST00000556278.1 |
GALT GALT |
galactose-1-phosphate uridylyltransferase Uncharacterized protein |
| chr20_-_1447467 | 0.00 |
ENST00000353088.2 ENST00000350991.4 |
NSFL1C |
NSFL1 (p97) cofactor (p47) |
| chrX_-_13835147 | 0.00 |
ENST00000493677.1 ENST00000355135.2 |
GPM6B |
glycoprotein M6B |
| chr1_+_192605252 | 0.00 |
ENST00000391995.2 ENST00000543215.1 |
RGS13 |
regulator of G-protein signaling 13 |
| chr18_+_22040593 | 0.00 |
ENST00000256906.4 |
HRH4 |
histamine receptor H4 |
| chr2_+_74682150 | 0.00 |
ENST00000233331.7 ENST00000431187.1 ENST00000409917.1 ENST00000409493.2 |
INO80B |
INO80 complex subunit B |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 3.1 | GO:0006880 | intracellular sequestering of iron ion(GO:0006880) sequestering of iron ion(GO:0097577) |
| 0.1 | 0.6 | GO:0090402 | oncogene-induced cell senescence(GO:0090402) |
| 0.1 | 0.4 | GO:0036466 | synaptic vesicle recycling via endosome(GO:0036466) mitochondrial membrane fission(GO:0090149) |
| 0.1 | 0.5 | GO:0033384 | geranyl diphosphate metabolic process(GO:0033383) geranyl diphosphate biosynthetic process(GO:0033384) farnesyl diphosphate biosynthetic process(GO:0045337) |
| 0.1 | 1.4 | GO:0035589 | G-protein coupled purinergic nucleotide receptor signaling pathway(GO:0035589) |
| 0.1 | 0.2 | GO:0043456 | regulation of pentose-phosphate shunt(GO:0043456) |
| 0.0 | 0.3 | GO:0032057 | negative regulation of translational initiation in response to stress(GO:0032057) |
| 0.0 | 0.1 | GO:0097018 | renal albumin absorption(GO:0097018) regulation of renal albumin absorption(GO:2000532) regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000583) negative regulation of platelet-derived growth factor receptor-alpha signaling pathway(GO:2000584) |
| 0.0 | 0.2 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.0 | 0.1 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.0 | 0.1 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.0 | 0.4 | GO:0002230 | positive regulation of defense response to virus by host(GO:0002230) |
| 0.0 | 0.4 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.2 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.0 | 0.1 | GO:2000628 | regulation of miRNA metabolic process(GO:2000628) |
| 0.0 | 0.1 | GO:0007068 | negative regulation of transcription during mitosis(GO:0007068) negative regulation of transcription from RNA polymerase II promoter during mitosis(GO:0007070) |
| 0.0 | 0.1 | GO:0015722 | canalicular bile acid transport(GO:0015722) |
| 0.0 | 0.1 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.0 | 0.1 | GO:0051256 | mitotic spindle midzone assembly(GO:0051256) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.0 | 0.1 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 0.6 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.0 | 0.5 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.2 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.4 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0004337 | dimethylallyltranstransferase activity(GO:0004161) geranyltranstransferase activity(GO:0004337) |
| 0.1 | 3.1 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 1.4 | GO:0045028 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.1 | 0.2 | GO:0004082 | bisphosphoglycerate mutase activity(GO:0004082) phosphoglycerate mutase activity(GO:0004619) 2,3-bisphosphoglycerate-dependent phosphoglycerate mutase activity(GO:0046538) |
| 0.0 | 0.6 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.0 | 0.1 | GO:0005174 | CD40 receptor binding(GO:0005174) |
| 0.0 | 0.2 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.0 | 0.2 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.0 | 0.1 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 0.0 | 0.1 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.0 | 0.2 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.4 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 0.2 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.0 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.0 | 0.1 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 3.1 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.1 | 0.3 | GO:0033597 | mitotic checkpoint complex(GO:0033597) bub1-bub3 complex(GO:1990298) |
| 0.1 | 0.6 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.0 | 0.3 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.0 | 0.1 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.0 | 0.1 | GO:0000939 | condensed chromosome inner kinetochore(GO:0000939) |
| 0.0 | 0.4 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.1 | GO:0060091 | kinocilium(GO:0060091) |
| 0.0 | 0.2 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |