Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
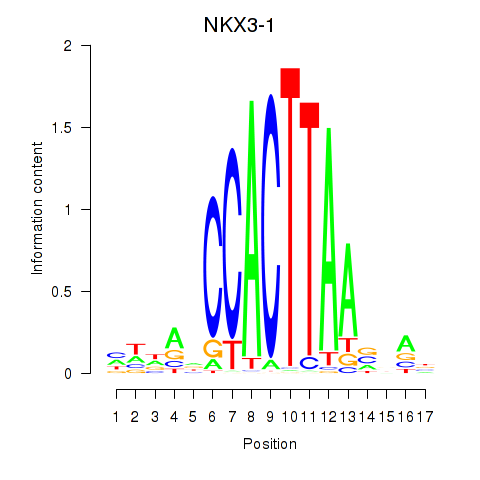
Results for NKX3-1
Z-value: 1.29
Transcription factors associated with NKX3-1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NKX3-1
|
ENSG00000167034.9 | NKX3-1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NKX3-1 | hg19_v2_chr8_-_23540402_23540466 | 0.28 | 2.9e-01 | Click! |
Activity profile of NKX3-1 motif
Sorted Z-values of NKX3-1 motif
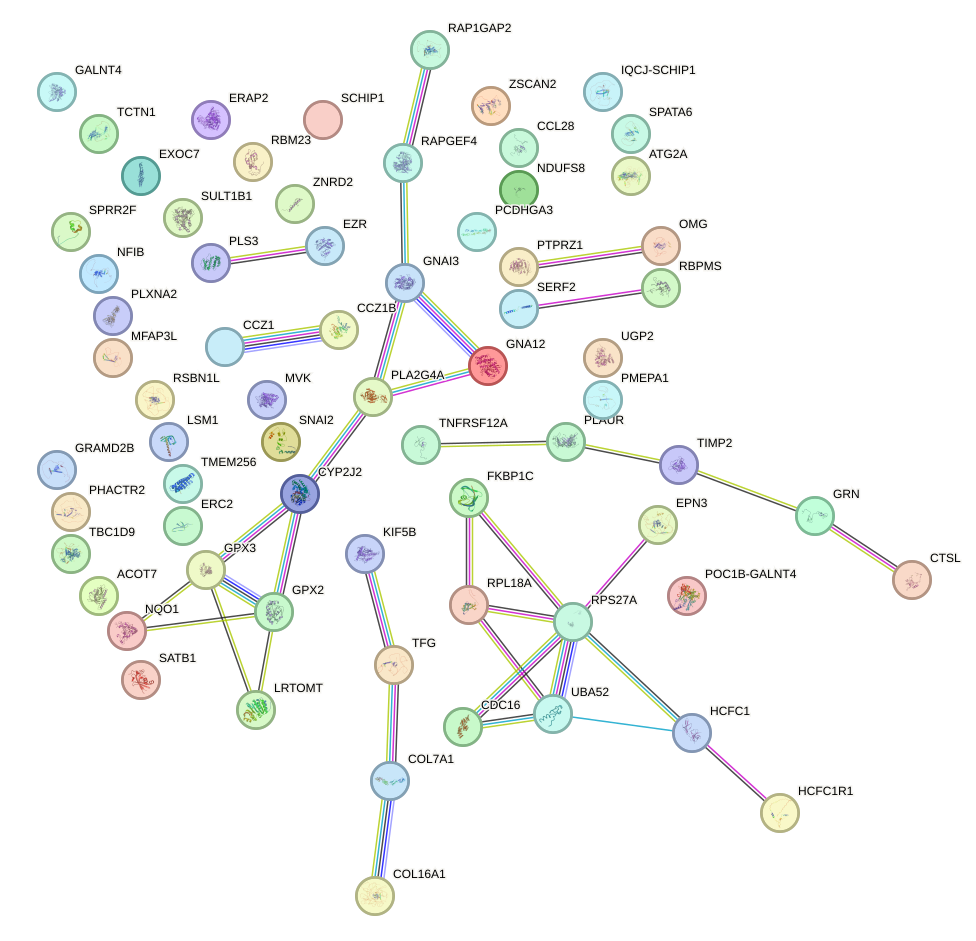
Network of associatons between targets according to the STRING database.
First level regulatory network of NKX3-1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_14180778 | 3.80 |
ENST00000380924.1 ENST00000543693.1 |
NFIB |
nuclear factor I/B |
| chr18_+_61445007 | 3.73 |
ENST00000447428.1 ENST00000546027.1 |
SERPINB7 |
serpin peptidase inhibitor, clade B (ovalbumin), member 7 |
| chr20_-_56285595 | 3.60 |
ENST00000395816.3 ENST00000347215.4 |
PMEPA1 |
prostate transmembrane protein, androgen induced 1 |
| chr9_+_706842 | 3.38 |
ENST00000382293.3 |
KANK1 |
KN motif and ankyrin repeat domains 1 |
| chr20_-_56286479 | 3.26 |
ENST00000265626.4 |
PMEPA1 |
prostate transmembrane protein, androgen induced 1 |
| chr8_-_49834299 | 3.26 |
ENST00000396822.1 |
SNAI2 |
snail family zinc finger 2 |
| chr16_+_3068393 | 3.07 |
ENST00000573001.1 |
TNFRSF12A |
tumor necrosis factor receptor superfamily, member 12A |
| chr8_-_49833978 | 3.07 |
ENST00000020945.1 |
SNAI2 |
snail family zinc finger 2 |
| chr8_+_30300119 | 2.69 |
ENST00000520191.1 |
RBPMS |
RNA binding protein with multiple splicing |
| chr17_+_48609903 | 2.62 |
ENST00000268933.3 |
EPN3 |
epsin 3 |
| chr3_+_158991025 | 2.42 |
ENST00000337808.6 |
IQCJ-SCHIP1 |
IQCJ-SCHIP1 readthrough |
| chr7_+_79765071 | 2.39 |
ENST00000457358.2 |
GNAI1 |
guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 |
| chr7_+_121513143 | 2.11 |
ENST00000393386.2 |
PTPRZ1 |
protein tyrosine phosphatase, receptor-type, Z polypeptide 1 |
| chr1_+_186798073 | 2.06 |
ENST00000367466.3 ENST00000442353.2 |
PLA2G4A |
phospholipase A2, group IVA (cytosolic, calcium-dependent) |
| chrX_-_107681633 | 1.89 |
ENST00000394872.2 ENST00000334504.7 |
COL4A6 |
collagen, type IV, alpha 6 |
| chr11_+_101983176 | 1.88 |
ENST00000524575.1 |
YAP1 |
Yes-associated protein 1 |
| chr11_+_114168085 | 1.83 |
ENST00000541754.1 |
NNMT |
nicotinamide N-methyltransferase |
| chrX_+_114795489 | 1.74 |
ENST00000355899.3 ENST00000537301.1 ENST00000289290.3 |
PLS3 |
plastin 3 |
| chr9_+_90341024 | 1.68 |
ENST00000340342.6 ENST00000342020.5 |
CTSL |
cathepsin L |
| chr19_-_44174330 | 1.64 |
ENST00000340093.3 |
PLAUR |
plasminogen activator, urokinase receptor |
| chr4_-_90757364 | 1.63 |
ENST00000508895.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr2_+_102456277 | 1.57 |
ENST00000421882.1 |
MAP4K4 |
mitogen-activated protein kinase kinase kinase kinase 4 |
| chr1_-_32169920 | 1.56 |
ENST00000373672.3 ENST00000373668.3 |
COL16A1 |
collagen, type XVI, alpha 1 |
| chrX_-_153637612 | 1.56 |
ENST00000369807.1 ENST00000369808.3 |
DNASE1L1 |
deoxyribonuclease I-like 1 |
| chr1_-_32169761 | 1.54 |
ENST00000271069.6 |
COL16A1 |
collagen, type XVI, alpha 1 |
| chr5_+_125758865 | 1.24 |
ENST00000542322.1 ENST00000544396.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr5_+_125758813 | 1.21 |
ENST00000285689.3 ENST00000515200.1 |
GRAMD3 |
GRAM domain containing 3 |
| chr1_-_208417620 | 1.17 |
ENST00000367033.3 |
PLXNA2 |
plexin A2 |
| chr4_-_90756769 | 1.15 |
ENST00000345009.4 ENST00000505199.1 ENST00000502987.1 |
SNCA |
synuclein, alpha (non A4 component of amyloid precursor) |
| chr17_-_7307358 | 1.09 |
ENST00000576017.1 ENST00000302422.3 ENST00000535512.1 |
TMEM256 TMEM256-PLSCR3 |
transmembrane protein 256 TMEM256-PLSCR3 readthrough (NMD candidate) |
| chr5_+_96211643 | 1.03 |
ENST00000437043.3 ENST00000510373.1 |
ERAP2 |
endoplasmic reticulum aminopeptidase 2 |
| chr17_+_48610074 | 1.03 |
ENST00000503690.1 ENST00000514874.1 ENST00000537145.1 ENST00000541226.1 |
EPN3 |
epsin 3 |
| chr3_-_48632593 | 0.97 |
ENST00000454817.1 ENST00000328333.8 |
COL7A1 |
collagen, type VII, alpha 1 |
| chr4_-_170924888 | 0.96 |
ENST00000502832.1 ENST00000393704.3 |
MFAP3L |
microfibrillar-associated protein 3-like |
| chr17_+_2699697 | 0.95 |
ENST00000254695.8 ENST00000366401.4 ENST00000542807.1 |
RAP1GAP2 |
RAP1 GTPase activating protein 2 |
| chr19_-_44174305 | 0.94 |
ENST00000601723.1 ENST00000339082.3 |
PLAUR |
plasminogen activator, urokinase receptor |
| chr5_+_96038476 | 0.87 |
ENST00000511049.1 ENST00000309190.5 ENST00000510156.1 ENST00000509903.1 ENST00000511782.1 ENST00000504465.1 |
CAST |
calpastatin |
| chr3_-_18480260 | 0.85 |
ENST00000454909.2 |
SATB1 |
SATB homeobox 1 |
| chrX_-_114253536 | 0.84 |
ENST00000371936.1 |
IL13RA2 |
interleukin 13 receptor, alpha 2 |
| chr4_-_141677267 | 0.82 |
ENST00000442267.2 |
TBC1D9 |
TBC1 domain family, member 9 (with GRAM domain) |
| chr9_+_116263639 | 0.79 |
ENST00000343817.5 |
RGS3 |
regulator of G-protein signaling 3 |
| chr1_-_6445809 | 0.70 |
ENST00000377855.2 |
ACOT7 |
acyl-CoA thioesterase 7 |
| chr3_+_100428188 | 0.70 |
ENST00000418917.2 ENST00000490574.1 |
TFG |
TRK-fused gene |
| chr4_+_56719782 | 0.69 |
ENST00000381295.2 ENST00000346134.7 ENST00000349598.6 |
EXOC1 |
exocyst complex component 1 |
| chr1_-_153066998 | 0.68 |
ENST00000368750.3 |
SPRR2E |
small proline-rich protein 2E |
| chr1_-_153113927 | 0.66 |
ENST00000368752.4 |
SPRR2B |
small proline-rich protein 2B |
| chr16_-_69760409 | 0.63 |
ENST00000561500.1 ENST00000439109.2 ENST00000564043.1 ENST00000379046.2 ENST00000379047.3 |
NQO1 |
NAD(P)H dehydrogenase, quinone 1 |
| chr6_+_143999072 | 0.60 |
ENST00000440869.2 ENST00000367582.3 ENST00000451827.2 |
PHACTR2 |
phosphatase and actin regulator 2 |
| chr3_+_100428316 | 0.58 |
ENST00000479672.1 ENST00000476228.1 ENST00000463568.1 |
TFG |
TRK-fused gene |
| chr12_+_110011571 | 0.57 |
ENST00000539696.1 ENST00000228510.3 ENST00000392727.3 |
MVK |
mevalonate kinase |
| chr3_+_100428268 | 0.57 |
ENST00000240851.4 |
TFG |
TRK-fused gene |
| chr17_-_76870126 | 0.56 |
ENST00000586057.1 |
TIMP2 |
TIMP metallopeptidase inhibitor 2 |
| chr6_-_159240415 | 0.56 |
ENST00000367075.3 |
EZR |
ezrin |
| chr5_-_176889381 | 0.54 |
ENST00000393563.4 ENST00000512501.1 |
DBN1 |
drebrin 1 |
| chr17_-_76870222 | 0.53 |
ENST00000585421.1 |
TIMP2 |
TIMP metallopeptidase inhibitor 2 |
| chr13_+_115000556 | 0.53 |
ENST00000252458.6 |
CDC16 |
cell division cycle 16 |
| chr3_+_52489503 | 0.53 |
ENST00000345716.4 |
NISCH |
nischarin |
| chr8_-_38034192 | 0.52 |
ENST00000520755.1 |
LSM1 |
LSM1 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr6_+_63921399 | 0.50 |
ENST00000356170.3 |
FKBP1C |
FK506 binding protein 1C |
| chr1_-_21377447 | 0.48 |
ENST00000374937.3 ENST00000264211.8 |
EIF4G3 |
eukaryotic translation initiation factor 4 gamma, 3 |
| chr1_-_60392452 | 0.48 |
ENST00000371204.3 |
CYP2J2 |
cytochrome P450, family 2, subfamily J, polypeptide 2 |
| chr12_-_53298841 | 0.47 |
ENST00000293308.6 |
KRT8 |
keratin 8 |
| chr7_-_2883928 | 0.46 |
ENST00000275364.3 |
GNA12 |
guanine nucleotide binding protein (G protein) alpha 12 |
| chr10_-_32345305 | 0.44 |
ENST00000302418.4 |
KIF5B |
kinesin family member 5B |
| chr4_-_70626314 | 0.43 |
ENST00000510821.1 |
SULT1B1 |
sulfotransferase family, cytosolic, 1B, member 1 |
| chr7_+_5938351 | 0.43 |
ENST00000325974.6 |
CCZ1 |
CCZ1 vacuolar protein trafficking and biogenesis associated homolog (S. cerevisiae) |
| chr1_-_153085984 | 0.42 |
ENST00000468739.1 |
SPRR2F |
small proline-rich protein 2F |
| chr14_-_23388338 | 0.42 |
ENST00000555209.1 ENST00000554256.1 ENST00000557403.1 ENST00000557549.1 ENST00000555676.1 ENST00000557571.1 ENST00000557464.1 ENST00000554618.1 ENST00000556862.1 ENST00000555722.1 ENST00000346528.5 ENST00000542016.2 ENST00000399922.2 ENST00000557227.1 ENST00000359890.3 |
RBM23 |
RNA binding motif protein 23 |
| chr8_-_38034234 | 0.42 |
ENST00000311351.4 |
LSM1 |
LSM1 homolog, U6 small nuclear RNA associated (S. cerevisiae) |
| chr13_+_50570019 | 0.41 |
ENST00000442421.1 |
TRIM13 |
tripartite motif containing 13 |
| chr11_-_64684672 | 0.40 |
ENST00000377264.3 ENST00000421419.2 |
ATG2A |
autophagy related 2A |
| chr19_+_17970677 | 0.40 |
ENST00000222247.5 |
RPL18A |
ribosomal protein L18a |
| chr19_+_17970693 | 0.39 |
ENST00000600147.1 ENST00000599898.1 |
RPL18A |
ribosomal protein L18a |
| chr17_+_42422662 | 0.38 |
ENST00000593167.1 ENST00000585512.1 ENST00000591740.1 ENST00000592783.1 ENST00000587387.1 ENST00000588237.1 ENST00000589265.1 |
GRN |
granulin |
| chrX_-_153236819 | 0.37 |
ENST00000354233.3 |
HCFC1 |
host cell factor C1 (VP16-accessory protein) |
| chr17_-_37009882 | 0.36 |
ENST00000378096.3 ENST00000394332.1 ENST00000394333.1 ENST00000577407.1 ENST00000479035.2 |
RPL23 |
ribosomal protein L23 |
| chr5_+_150404904 | 0.36 |
ENST00000521632.1 |
GPX3 |
glutathione peroxidase 3 (plasma) |
| chr12_-_89920030 | 0.35 |
ENST00000413530.1 ENST00000547474.1 |
GALNT4 POC1B-GALNT4 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 4 (GalNAc-T4) POC1B-GALNT4 readthrough |
| chr15_+_44084503 | 0.35 |
ENST00000409960.2 ENST00000409646.1 ENST00000594896.1 ENST00000339624.5 ENST00000409291.1 ENST00000402131.1 ENST00000403425.1 ENST00000430901.1 |
SERF2 |
small EDRK-rich factor 2 |
| chr6_-_144385698 | 0.35 |
ENST00000444202.1 ENST00000437412.1 |
PLAGL1 |
pleiomorphic adenoma gene-like 1 |
| chr6_-_33285505 | 0.34 |
ENST00000431845.2 |
ZBTB22 |
zinc finger and BTB domain containing 22 |
| chr5_-_43412418 | 0.34 |
ENST00000537013.1 ENST00000361115.4 |
CCL28 |
chemokine (C-C motif) ligand 28 |
| chr14_-_65409438 | 0.34 |
ENST00000557049.1 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr5_+_140723601 | 0.34 |
ENST00000253812.6 |
PCDHGA3 |
protocadherin gamma subfamily A, 3 |
| chr11_+_71791693 | 0.33 |
ENST00000289488.2 ENST00000447974.1 |
LRTOMT |
leucine rich transmembrane and O-methyltransferase domain containing |
| chr7_+_5938386 | 0.33 |
ENST00000537980.1 |
CCZ1 |
CCZ1 vacuolar protein trafficking and biogenesis associated homolog (S. cerevisiae) |
| chr12_-_89919965 | 0.33 |
ENST00000548729.1 |
POC1B-GALNT4 |
POC1B-GALNT4 readthrough |
| chr17_+_66521936 | 0.33 |
ENST00000592800.1 |
PRKAR1A |
protein kinase, cAMP-dependent, regulatory, type I, alpha |
| chr1_-_48866517 | 0.33 |
ENST00000371841.1 |
SPATA6 |
spermatogenesis associated 6 |
| chr2_+_64069459 | 0.33 |
ENST00000445915.2 ENST00000475462.1 |
UGP2 |
UDP-glucose pyrophosphorylase 2 |
| chr2_+_64068844 | 0.33 |
ENST00000337130.5 |
UGP2 |
UDP-glucose pyrophosphorylase 2 |
| chr2_+_173686303 | 0.32 |
ENST00000397087.3 |
RAPGEF4 |
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr3_+_98482175 | 0.32 |
ENST00000485391.1 ENST00000492254.1 |
ST3GAL6 |
ST3 beta-galactoside alpha-2,3-sialyltransferase 6 |
| chr16_-_3074231 | 0.31 |
ENST00000572355.1 ENST00000248089.3 ENST00000574980.1 ENST00000354679.3 ENST00000396916.1 ENST00000573842.1 |
HCFC1R1 |
host cell factor C1 regulator 1 (XPO1 dependent) |
| chr6_-_57087042 | 0.31 |
ENST00000317483.3 |
RAB23 |
RAB23, member RAS oncogene family |
| chr7_+_77325738 | 0.30 |
ENST00000334955.8 |
RSBN1L |
round spermatid basic protein 1-like |
| chr12_-_125398654 | 0.30 |
ENST00000541645.1 ENST00000540351.1 |
UBC |
ubiquitin C |
| chr11_+_65337901 | 0.30 |
ENST00000309328.3 ENST00000531405.1 ENST00000527920.1 ENST00000526877.1 ENST00000533115.1 ENST00000526433.1 |
SSSCA1 |
Sjogren syndrome/scleroderma autoantigen 1 |
| chr12_+_111051902 | 0.30 |
ENST00000397655.3 ENST00000471804.2 ENST00000377654.3 ENST00000397659.4 |
TCTN1 |
tectonic family member 1 |
| chr17_-_29624343 | 0.30 |
ENST00000247271.4 |
OMG |
oligodendrocyte myelin glycoprotein |
| chr11_+_67798090 | 0.29 |
ENST00000313468.5 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chrX_-_153237258 | 0.28 |
ENST00000310441.7 |
HCFC1 |
host cell factor C1 (VP16-accessory protein) |
| chr7_-_6865826 | 0.28 |
ENST00000538180.1 |
CCZ1B |
CCZ1 vacuolar protein trafficking and biogenesis associated homolog B (S. cerevisiae) |
| chr17_+_42422637 | 0.28 |
ENST00000053867.3 ENST00000588143.1 |
GRN |
granulin |
| chr15_+_85147127 | 0.27 |
ENST00000541040.1 ENST00000538076.1 ENST00000485222.2 |
ZSCAN2 |
zinc finger and SCAN domain containing 2 |
| chr17_-_64187973 | 0.27 |
ENST00000583358.1 ENST00000392769.2 |
CEP112 |
centrosomal protein 112kDa |
| chr19_-_15544099 | 0.27 |
ENST00000599910.2 |
WIZ |
widely interspaced zinc finger motifs |
| chr4_-_122791583 | 0.27 |
ENST00000506636.1 ENST00000264499.4 |
BBS7 |
Bardet-Biedl syndrome 7 |
| chr12_-_10562356 | 0.26 |
ENST00000309384.1 |
KLRC4 |
killer cell lectin-like receptor subfamily C, member 4 |
| chr1_+_158223923 | 0.26 |
ENST00000289429.5 |
CD1A |
CD1a molecule |
| chrX_+_153237740 | 0.26 |
ENST00000369982.4 |
TMEM187 |
transmembrane protein 187 |
| chr16_-_66864806 | 0.26 |
ENST00000566336.1 ENST00000394074.2 ENST00000563185.2 ENST00000359087.4 ENST00000379463.2 ENST00000565535.1 ENST00000290810.3 |
NAE1 |
NEDD8 activating enzyme E1 subunit 1 |
| chr11_+_8704748 | 0.24 |
ENST00000526562.1 ENST00000525981.1 |
RPL27A |
ribosomal protein L27a |
| chr16_+_1832902 | 0.24 |
ENST00000262302.9 ENST00000563136.1 ENST00000565987.1 ENST00000543305.1 ENST00000568287.1 ENST00000565134.1 |
NUBP2 |
nucleotide binding protein 2 |
| chr1_+_22962948 | 0.24 |
ENST00000374642.3 |
C1QA |
complement component 1, q subcomponent, A chain |
| chr9_+_36572851 | 0.23 |
ENST00000298048.2 ENST00000538311.1 ENST00000536987.1 ENST00000545008.1 ENST00000536860.1 ENST00000536329.1 ENST00000541717.1 ENST00000543751.1 |
MELK |
maternal embryonic leucine zipper kinase |
| chr4_-_10118469 | 0.23 |
ENST00000499869.2 |
WDR1 |
WD repeat domain 1 |
| chr16_+_22501658 | 0.23 |
ENST00000415833.2 |
NPIPB5 |
nuclear pore complex interacting protein family, member B5 |
| chr11_+_67798114 | 0.23 |
ENST00000453471.2 ENST00000528492.1 ENST00000526339.1 ENST00000525419.1 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr20_+_123010 | 0.22 |
ENST00000382398.3 |
DEFB126 |
defensin, beta 126 |
| chr20_+_2821340 | 0.22 |
ENST00000380445.3 ENST00000380469.3 |
VPS16 |
vacuolar protein sorting 16 homolog (S. cerevisiae) |
| chr15_+_44084040 | 0.21 |
ENST00000249786.4 |
SERF2 |
small EDRK-rich factor 2 |
| chr10_-_104953009 | 0.20 |
ENST00000470299.1 ENST00000343289.5 |
NT5C2 |
5'-nucleotidase, cytosolic II |
| chr12_+_111051832 | 0.20 |
ENST00000550703.2 ENST00000551590.1 |
TCTN1 |
tectonic family member 1 |
| chr17_-_7145475 | 0.19 |
ENST00000571129.1 ENST00000571253.1 ENST00000573928.1 |
GABARAP |
GABA(A) receptor-associated protein |
| chr12_-_125398602 | 0.19 |
ENST00000541272.1 ENST00000535131.1 |
UBC |
ubiquitin C |
| chr2_+_113672770 | 0.19 |
ENST00000311328.2 |
IL37 |
interleukin 37 |
| chr1_+_158323244 | 0.19 |
ENST00000434258.1 |
CD1E |
CD1e molecule |
| chr7_+_7222233 | 0.19 |
ENST00000436587.2 |
C1GALT1 |
core 1 synthase, glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase, 1 |
| chr6_+_43266063 | 0.18 |
ENST00000372574.3 |
SLC22A7 |
solute carrier family 22 (organic anion transporter), member 7 |
| chr8_-_93029865 | 0.18 |
ENST00000422361.2 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr17_-_2966901 | 0.18 |
ENST00000575751.1 |
OR1D5 |
olfactory receptor, family 1, subfamily D, member 5 |
| chr13_+_115000521 | 0.18 |
ENST00000252457.5 ENST00000375308.1 |
CDC16 |
cell division cycle 16 |
| chr1_-_77685084 | 0.18 |
ENST00000370812.3 ENST00000359130.1 ENST00000445065.1 ENST00000370813.5 |
PIGK |
phosphatidylinositol glycan anchor biosynthesis, class K |
| chr19_-_1592652 | 0.17 |
ENST00000156825.1 ENST00000434436.3 |
MBD3 |
methyl-CpG binding domain protein 3 |
| chr1_-_216978709 | 0.17 |
ENST00000360012.3 |
ESRRG |
estrogen-related receptor gamma |
| chr1_+_173793777 | 0.17 |
ENST00000239457.5 |
DARS2 |
aspartyl-tRNA synthetase 2, mitochondrial |
| chr2_-_79313973 | 0.17 |
ENST00000454188.1 |
REG1B |
regenerating islet-derived 1 beta |
| chrX_-_24665353 | 0.16 |
ENST00000379144.2 |
PCYT1B |
phosphate cytidylyltransferase 1, choline, beta |
| chr17_+_80674559 | 0.15 |
ENST00000269373.6 ENST00000535965.1 ENST00000577128.1 ENST00000573158.1 |
FN3KRP |
fructosamine 3 kinase related protein |
| chr14_-_74551172 | 0.15 |
ENST00000553458.1 |
ALDH6A1 |
aldehyde dehydrogenase 6 family, member A1 |
| chr10_+_135204338 | 0.14 |
ENST00000468317.2 |
RP11-108K14.8 |
Mitochondrial GTPase 1 |
| chr20_-_34330129 | 0.14 |
ENST00000397370.3 ENST00000528062.3 ENST00000407261.4 ENST00000374038.3 ENST00000361162.6 |
RBM39 |
RNA binding motif protein 39 |
| chr7_+_110731062 | 0.14 |
ENST00000308478.5 ENST00000451085.1 ENST00000422987.3 ENST00000421101.1 |
LRRN3 |
leucine rich repeat neuronal 3 |
| chr21_-_46964306 | 0.14 |
ENST00000443742.1 ENST00000528477.1 ENST00000567670.1 |
SLC19A1 |
solute carrier family 19 (folate transporter), member 1 |
| chr16_+_56969284 | 0.14 |
ENST00000568358.1 |
HERPUD1 |
homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 |
| chr4_+_88529681 | 0.14 |
ENST00000399271.1 |
DSPP |
dentin sialophosphoprotein |
| chr3_+_132316081 | 0.14 |
ENST00000249887.2 |
ACKR4 |
atypical chemokine receptor 4 |
| chr1_+_171217622 | 0.14 |
ENST00000433267.1 ENST00000367750.3 |
FMO1 |
flavin containing monooxygenase 1 |
| chrX_+_13671225 | 0.13 |
ENST00000545566.1 ENST00000544987.1 ENST00000314720.4 |
TCEANC |
transcription elongation factor A (SII) N-terminal and central domain containing |
| chr7_-_138794394 | 0.13 |
ENST00000242351.5 ENST00000471652.1 |
ZC3HAV1 |
zinc finger CCCH-type, antiviral 1 |
| chr11_+_67798363 | 0.13 |
ENST00000525628.1 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr9_-_104145795 | 0.12 |
ENST00000259407.2 |
BAAT |
bile acid CoA: amino acid N-acyltransferase (glycine N-choloyltransferase) |
| chr2_-_187367356 | 0.12 |
ENST00000595956.1 |
AC018867.2 |
AC018867.2 |
| chr1_-_160232197 | 0.11 |
ENST00000419626.1 ENST00000610139.1 ENST00000475733.1 ENST00000407642.2 ENST00000368073.3 ENST00000326837.2 |
DCAF8 |
DDB1 and CUL4 associated factor 8 |
| chr11_+_7506713 | 0.11 |
ENST00000329293.3 ENST00000534244.1 |
OLFML1 |
olfactomedin-like 1 |
| chr11_+_9482551 | 0.10 |
ENST00000438144.2 ENST00000526657.1 ENST00000299606.2 ENST00000534265.1 ENST00000412390.2 |
ZNF143 |
zinc finger protein 143 |
| chr7_-_112430647 | 0.10 |
ENST00000312814.6 |
TMEM168 |
transmembrane protein 168 |
| chr10_-_28571015 | 0.09 |
ENST00000375719.3 ENST00000375732.1 |
MPP7 |
membrane protein, palmitoylated 7 (MAGUK p55 subfamily member 7) |
| chr1_-_160232291 | 0.09 |
ENST00000368074.1 ENST00000447377.1 |
DCAF8 |
DDB1 and CUL4 associated factor 8 |
| chr19_-_14889349 | 0.09 |
ENST00000315576.3 ENST00000392967.2 ENST00000346057.1 ENST00000353876.1 ENST00000353005.1 |
EMR2 |
egf-like module containing, mucin-like, hormone receptor-like 2 |
| chr1_+_151372010 | 0.09 |
ENST00000290541.6 |
PSMB4 |
proteasome (prosome, macropain) subunit, beta type, 4 |
| chr7_+_141463897 | 0.09 |
ENST00000247879.2 |
TAS2R3 |
taste receptor, type 2, member 3 |
| chr1_+_158323486 | 0.09 |
ENST00000444681.2 ENST00000368167.3 |
CD1E |
CD1e molecule |
| chr12_+_4829507 | 0.09 |
ENST00000252318.2 |
GALNT8 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 8 (GalNAc-T8) |
| chr12_+_81110684 | 0.08 |
ENST00000228644.3 |
MYF5 |
myogenic factor 5 |
| chr14_-_65409502 | 0.08 |
ENST00000389614.5 |
GPX2 |
glutathione peroxidase 2 (gastrointestinal) |
| chr2_+_216974020 | 0.08 |
ENST00000392132.2 ENST00000417391.1 |
XRCC5 |
X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining) |
| chr16_-_21436459 | 0.07 |
ENST00000448012.2 ENST00000504841.2 ENST00000419180.2 |
NPIPB3 |
nuclear pore complex interacting protein family, member B3 |
| chr19_-_55791563 | 0.07 |
ENST00000588971.1 ENST00000255631.5 ENST00000587551.1 |
HSPBP1 |
HSPA (heat shock 70kDa) binding protein, cytoplasmic cochaperone 1 |
| chr16_-_69373396 | 0.06 |
ENST00000562595.1 ENST00000562081.1 ENST00000306875.4 |
COG8 |
component of oligomeric golgi complex 8 |
| chr13_+_27998681 | 0.06 |
ENST00000381140.4 |
GTF3A |
general transcription factor IIIA |
| chrX_-_38186775 | 0.06 |
ENST00000339363.3 ENST00000309513.3 ENST00000338898.3 ENST00000342811.3 ENST00000378505.2 |
RPGR |
retinitis pigmentosa GTPase regulator |
| chr6_+_52930237 | 0.06 |
ENST00000323557.7 |
FBXO9 |
F-box protein 9 |
| chr15_-_31393910 | 0.06 |
ENST00000397795.2 ENST00000256552.6 ENST00000559179.1 |
TRPM1 |
transient receptor potential cation channel, subfamily M, member 1 |
| chr12_-_4553385 | 0.05 |
ENST00000543077.1 |
FGF6 |
fibroblast growth factor 6 |
| chr8_-_97247759 | 0.05 |
ENST00000518406.1 ENST00000523920.1 ENST00000287022.5 |
UQCRB |
ubiquinol-cytochrome c reductase binding protein |
| chr16_+_69373323 | 0.05 |
ENST00000254940.5 |
NIP7 |
NIP7, nucleolar pre-rRNA processing protein |
| chr5_+_64920826 | 0.05 |
ENST00000438419.2 ENST00000231526.4 ENST00000505553.1 ENST00000545191.1 |
TRAPPC13 |
trafficking protein particle complex 13 |
| chr22_-_50523760 | 0.04 |
ENST00000395876.2 |
MLC1 |
megalencephalic leukoencephalopathy with subcortical cysts 1 |
| chr12_-_57039739 | 0.04 |
ENST00000552959.1 ENST00000551020.1 ENST00000553007.2 ENST00000552919.1 ENST00000552104.1 ENST00000262030.3 |
ATP5B |
ATP synthase, H+ transporting, mitochondrial F1 complex, beta polypeptide |
| chr17_-_34257731 | 0.04 |
ENST00000431884.2 ENST00000425909.3 ENST00000394528.3 ENST00000430160.2 |
RDM1 |
RAD52 motif 1 |
| chr12_-_54978086 | 0.04 |
ENST00000553113.1 |
PPP1R1A |
protein phosphatase 1, regulatory (inhibitor) subunit 1A |
| chr7_+_100303676 | 0.03 |
ENST00000303151.4 |
POP7 |
processing of precursor 7, ribonuclease P/MRP subunit (S. cerevisiae) |
| chr12_+_12966250 | 0.03 |
ENST00000352940.4 ENST00000358007.3 ENST00000544400.1 |
DDX47 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 |
| chr12_-_125398850 | 0.03 |
ENST00000535859.1 ENST00000546271.1 ENST00000540700.1 ENST00000546120.1 |
UBC |
ubiquitin C |
| chr12_+_113229452 | 0.03 |
ENST00000389385.4 |
RPH3A |
rabphilin 3A homolog (mouse) |
| chr11_-_71823715 | 0.03 |
ENST00000545944.1 ENST00000502597.2 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr11_-_101454234 | 0.03 |
ENST00000348423.4 ENST00000360497.4 ENST00000532133.1 |
TRPC6 |
transient receptor potential cation channel, subfamily C, member 6 |
| chr9_+_117350009 | 0.02 |
ENST00000374050.3 |
ATP6V1G1 |
ATPase, H+ transporting, lysosomal 13kDa, V1 subunit G1 |
| chr11_-_71823796 | 0.02 |
ENST00000545680.1 ENST00000543587.1 ENST00000538393.1 ENST00000535234.1 ENST00000227618.4 ENST00000535503.1 |
ANAPC15 |
anaphase promoting complex subunit 15 |
| chr1_+_171217677 | 0.02 |
ENST00000402921.2 |
FMO1 |
flavin containing monooxygenase 1 |
| chr1_-_35497506 | 0.02 |
ENST00000317538.5 ENST00000373340.2 ENST00000357182.4 |
ZMYM6 |
zinc finger, MYM-type 6 |
| chr4_+_159131630 | 0.02 |
ENST00000504569.1 ENST00000509278.1 ENST00000514558.1 ENST00000503200.1 |
TMEM144 |
transmembrane protein 144 |
| chr11_+_8040739 | 0.01 |
ENST00000534099.1 |
TUB |
tubby bipartite transcription factor |
| chr6_+_42928485 | 0.01 |
ENST00000372808.3 |
GNMT |
glycine N-methyltransferase |
| chr1_+_173793641 | 0.01 |
ENST00000361951.4 |
DARS2 |
aspartyl-tRNA synthetase 2, mitochondrial |
| chr17_+_7358889 | 0.01 |
ENST00000575379.1 |
CHRNB1 |
cholinergic receptor, nicotinic, beta 1 (muscle) |
| chr6_-_31648150 | 0.01 |
ENST00000375858.3 ENST00000383237.4 |
LY6G5C |
lymphocyte antigen 6 complex, locus G5C |
| chrX_-_102757802 | 0.01 |
ENST00000372633.1 |
RAB40A |
RAB40A, member RAS oncogene family |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 2.6 | GO:0030377 | urokinase plasminogen activator receptor activity(GO:0030377) |
| 0.6 | 2.8 | GO:0060961 | phospholipase D inhibitor activity(GO:0060961) |
| 0.5 | 1.8 | GO:0030760 | nicotinamide N-methyltransferase activity(GO:0008112) pyridine N-methyltransferase activity(GO:0030760) |
| 0.4 | 2.7 | GO:0035614 | snRNA stem-loop binding(GO:0035614) |
| 0.2 | 0.7 | GO:0003983 | UTP:glucose-1-phosphate uridylyltransferase activity(GO:0003983) UTP-monosaccharide-1-phosphate uridylyltransferase activity(GO:0051748) |
| 0.2 | 2.1 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.2 | 2.4 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.2 | 1.6 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.2 | 6.8 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.1 | 0.9 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.1 | 0.4 | GO:0099609 | microtubule lateral binding(GO:0099609) |
| 0.1 | 0.4 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.3 | GO:0070984 | SET domain binding(GO:0070984) |
| 0.1 | 0.3 | GO:0052798 | beta-galactoside alpha-2,3-sialyltransferase activity(GO:0052798) |
| 0.1 | 2.1 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 0.8 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.1 | 0.6 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.1 | 1.2 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.1 | 1.7 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.1 | 1.4 | GO:0000339 | RNA cap binding(GO:0000339) |
| 0.1 | 0.2 | GO:0003923 | GPI-anchor transamidase activity(GO:0003923) |
| 0.1 | 1.9 | GO:0070064 | proline-rich region binding(GO:0070064) |
| 0.1 | 0.7 | GO:0043996 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
| 0.1 | 0.5 | GO:0030884 | lipid antigen binding(GO:0030882) endogenous lipid antigen binding(GO:0030883) exogenous lipid antigen binding(GO:0030884) |
| 0.0 | 0.3 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.0 | 1.0 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 6.4 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 1.1 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.6 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.0 | 0.9 | GO:0034236 | protein kinase A catalytic subunit binding(GO:0034236) |
| 0.0 | 0.7 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.2 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.0 | 0.1 | GO:0015350 | reduced folate carrier activity(GO:0008518) methotrexate transporter activity(GO:0015350) |
| 0.0 | 0.4 | GO:1904264 | ubiquitin protein ligase activity involved in ERAD pathway(GO:1904264) |
| 0.0 | 3.6 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.4 | GO:0008430 | selenium binding(GO:0008430) |
| 0.0 | 0.2 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.0 | 3.4 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.5 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.3 | GO:0032027 | myosin light chain binding(GO:0032027) |
| 0.0 | 0.4 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.5 | GO:0070330 | arachidonic acid monooxygenase activity(GO:0008391) arachidonic acid epoxygenase activity(GO:0008392) aromatase activity(GO:0070330) |
| 0.0 | 0.2 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.0 | 0.5 | GO:0050780 | dopamine receptor binding(GO:0050780) |
| 0.0 | 1.1 | GO:0005198 | structural molecule activity(GO:0005198) |
| 0.0 | 0.3 | GO:0008641 | small protein activating enzyme activity(GO:0008641) |
| 0.0 | 1.9 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.5 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.4 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.0 | 0.2 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 0.2 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.0 | 0.9 | GO:0051539 | 4 iron, 4 sulfur cluster binding(GO:0051539) |
| 0.0 | 1.3 | GO:0004536 | deoxyribonuclease activity(GO:0004536) |
| 0.0 | 0.4 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.0 | 0.6 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 2.2 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.2 | GO:0048531 | beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.0 | 0.3 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 0.2 | GO:0005149 | interleukin-1 receptor binding(GO:0005149) |
| 0.0 | 3.8 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.8 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.1 | 6.0 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 2.6 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.1 | 0.8 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.1 | 6.3 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 2.4 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 3.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 2.5 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 0.8 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 1.6 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.0 | 1.9 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 5.4 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.7 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.0 | 1.1 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 0.6 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 0.4 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 6.3 | GO:0070563 | negative regulation of vitamin D receptor signaling pathway(GO:0070563) |
| 1.1 | 3.4 | GO:2000393 | negative regulation of lamellipodium morphogenesis(GO:2000393) |
| 0.9 | 3.7 | GO:0090362 | positive regulation of platelet-derived growth factor production(GO:0090362) |
| 0.6 | 2.6 | GO:0038195 | urokinase plasminogen activator signaling pathway(GO:0038195) |
| 0.6 | 6.9 | GO:0010991 | negative regulation of SMAD protein complex assembly(GO:0010991) |
| 0.6 | 2.8 | GO:0032227 | negative regulation of synaptic transmission, dopaminergic(GO:0032227) negative regulation of dopamine uptake involved in synaptic transmission(GO:0051585) norepinephrine uptake(GO:0051620) regulation of norepinephrine uptake(GO:0051621) negative regulation of norepinephrine uptake(GO:0051622) negative regulation of catecholamine uptake involved in synaptic transmission(GO:0051945) regulation of glutathione peroxidase activity(GO:1903282) positive regulation of glutathione peroxidase activity(GO:1903284) positive regulation of hydrogen peroxide catabolic process(GO:1903285) regulation of peroxidase activity(GO:2000468) positive regulation of peroxidase activity(GO:2000470) |
| 0.5 | 3.8 | GO:0021740 | trigeminal sensory nucleus development(GO:0021730) principal sensory nucleus of trigeminal nerve development(GO:0021740) negative regulation of epithelial cell proliferation involved in lung morphogenesis(GO:2000795) |
| 0.5 | 2.1 | GO:0006663 | platelet activating factor biosynthetic process(GO:0006663) |
| 0.4 | 3.1 | GO:0006931 | substrate-dependent cell migration, cell attachment to substrate(GO:0006931) |
| 0.3 | 2.1 | GO:0070444 | oligodendrocyte progenitor proliferation(GO:0070444) regulation of oligodendrocyte progenitor proliferation(GO:0070445) |
| 0.2 | 1.1 | GO:0032487 | regulation of Rap protein signal transduction(GO:0032487) |
| 0.2 | 1.9 | GO:1902459 | positive regulation of stem cell population maintenance(GO:1902459) |
| 0.2 | 2.4 | GO:1904321 | response to forskolin(GO:1904321) cellular response to forskolin(GO:1904322) |
| 0.2 | 0.6 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.2 | 0.7 | GO:0036114 | medium-chain fatty-acyl-CoA catabolic process(GO:0036114) long-chain fatty-acyl-CoA catabolic process(GO:0036116) palmitic acid metabolic process(GO:1900533) palmitic acid biosynthetic process(GO:1900535) |
| 0.2 | 1.8 | GO:0034356 | NAD biosynthesis via nicotinamide riboside salvage pathway(GO:0034356) |
| 0.2 | 0.7 | GO:0019046 | release from viral latency(GO:0019046) |
| 0.1 | 0.7 | GO:0019255 | glucose 1-phosphate metabolic process(GO:0019255) |
| 0.1 | 1.7 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.1 | 0.5 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 0.1 | 0.4 | GO:0006982 | response to lipid hydroperoxide(GO:0006982) |
| 0.1 | 0.6 | GO:1900041 | negative regulation of interleukin-2 secretion(GO:1900041) terminal web assembly(GO:1902896) |
| 0.1 | 0.5 | GO:0010643 | cell communication by chemical coupling(GO:0010643) |
| 0.1 | 1.2 | GO:0021785 | branchiomotor neuron axon guidance(GO:0021785) |
| 0.1 | 2.1 | GO:0060391 | positive regulation of SMAD protein import into nucleus(GO:0060391) |
| 0.1 | 2.4 | GO:0033622 | integrin activation(GO:0033622) |
| 0.1 | 0.8 | GO:0043305 | negative regulation of mast cell degranulation(GO:0043305) |
| 0.1 | 0.4 | GO:0035617 | stress granule disassembly(GO:0035617) |
| 0.1 | 1.0 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.1 | 0.4 | GO:0072717 | cellular response to actinomycin D(GO:0072717) |
| 0.1 | 1.6 | GO:0071888 | macrophage apoptotic process(GO:0071888) |
| 0.1 | 1.6 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.1 | 0.3 | GO:1903237 | negative regulation of leukocyte tethering or rolling(GO:1903237) |
| 0.1 | 0.7 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.1 | 0.9 | GO:2000675 | negative regulation of type B pancreatic cell apoptotic process(GO:2000675) |
| 0.1 | 0.2 | GO:0006421 | asparaginyl-tRNA aminoacylation(GO:0006421) |
| 0.1 | 0.2 | GO:0016333 | morphogenesis of follicular epithelium(GO:0016333) establishment or maintenance of polarity of follicular epithelium(GO:0016334) establishment of planar polarity of follicular epithelium(GO:0042247) |
| 0.1 | 0.8 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.1 | 0.3 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.0 | 0.1 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.5 | GO:0048007 | antigen processing and presentation of lipid antigen via MHC class Ib(GO:0048003) antigen processing and presentation, exogenous lipid antigen via MHC class Ib(GO:0048007) |
| 0.0 | 0.2 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 0.5 | GO:0051599 | response to hydrostatic pressure(GO:0051599) |
| 0.0 | 0.2 | GO:0035542 | regulation of SNARE complex assembly(GO:0035542) |
| 0.0 | 2.9 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.0 | 0.1 | GO:0098838 | methotrexate transport(GO:0051958) reduced folate transmembrane transport(GO:0098838) |
| 0.0 | 0.9 | GO:1904380 | endoplasmic reticulum mannose trimming(GO:1904380) |
| 0.0 | 0.1 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.0 | 0.3 | GO:0010571 | positive regulation of nuclear cell cycle DNA replication(GO:0010571) |
| 0.0 | 0.5 | GO:0043651 | linoleic acid metabolic process(GO:0043651) |
| 0.0 | 0.1 | GO:1904430 | negative regulation of t-circle formation(GO:1904430) |
| 0.0 | 0.3 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 0.3 | GO:2000480 | negative regulation of activated T cell proliferation(GO:0046007) negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.0 | 0.2 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 0.0 | 1.3 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.0 | 1.1 | GO:0006308 | DNA catabolic process(GO:0006308) |
| 0.0 | 0.1 | GO:1903071 | positive regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903071) |
| 0.0 | 0.3 | GO:0060117 | auditory receptor cell development(GO:0060117) |
| 0.0 | 0.2 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.0 | 0.8 | GO:0000188 | inactivation of MAPK activity(GO:0000188) |
| 0.0 | 0.4 | GO:0044804 | nucleophagy(GO:0044804) |
| 0.0 | 1.6 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.0 | 0.4 | GO:0006068 | ethanol catabolic process(GO:0006068) |
| 0.0 | 0.6 | GO:0019430 | removal of superoxide radicals(GO:0019430) |
| 0.0 | 0.2 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 0.7 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 0.5 | GO:0032228 | regulation of synaptic transmission, GABAergic(GO:0032228) |
| 0.0 | 0.2 | GO:0006657 | CDP-choline pathway(GO:0006657) |
| 0.0 | 0.0 | GO:0006933 | negative regulation of cell adhesion involved in substrate-bound cell migration(GO:0006933) |
| 0.0 | 0.2 | GO:0060576 | intestinal epithelial cell development(GO:0060576) |
| 0.0 | 0.1 | GO:0019530 | taurine metabolic process(GO:0019530) |
| 0.0 | 0.3 | GO:0097503 | sialylation(GO:0097503) |
| 0.0 | 0.3 | GO:0044458 | motile cilium assembly(GO:0044458) |
| 0.0 | 0.1 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.0 | 0.5 | GO:0010762 | regulation of fibroblast migration(GO:0010762) |
| 0.0 | 0.2 | GO:0006346 | methylation-dependent chromatin silencing(GO:0006346) |
| 0.0 | 0.2 | GO:0016226 | iron-sulfur cluster assembly(GO:0016226) metallo-sulfur cluster assembly(GO:0031163) |
| 0.0 | 0.8 | GO:0031338 | regulation of vesicle fusion(GO:0031338) |
| 0.0 | 0.5 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.4 | GO:0016266 | O-glycan processing(GO:0016266) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.1 | GO:0072534 | perineuronal net(GO:0072534) |
| 0.4 | 3.1 | GO:0005593 | FACIT collagen trimer(GO:0005593) |
| 0.4 | 1.9 | GO:0071148 | TEAD-1-YAP complex(GO:0071148) |
| 0.3 | 3.8 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.3 | 1.2 | GO:0098592 | cytoplasmic side of apical plasma membrane(GO:0098592) |
| 0.2 | 1.7 | GO:0036021 | endolysosome lumen(GO:0036021) |
| 0.2 | 1.9 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.2 | 2.0 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.1 | 0.9 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.1 | 3.4 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.1 | 1.2 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.1 | 0.3 | GO:0097224 | sperm connecting piece(GO:0097224) |
| 0.1 | 2.3 | GO:0005685 | U1 snRNP(GO:0005685) |
| 0.1 | 1.8 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.1 | 0.2 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.1 | 2.7 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 0.7 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.2 | GO:0005602 | complement component C1 complex(GO:0005602) |
| 0.0 | 0.2 | GO:0042643 | actomyosin, actin portion(GO:0042643) |
| 0.0 | 6.6 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.5 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 1.0 | GO:0098644 | complex of collagen trimers(GO:0098644) |
| 0.0 | 3.3 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.4 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.0 | 1.7 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.0 | 0.3 | GO:0034464 | BBSome(GO:0034464) |
| 0.0 | 0.1 | GO:0035838 | growing cell tip(GO:0035838) |
| 0.0 | 2.3 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 0.9 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.0 | 6.5 | GO:0001726 | ruffle(GO:0001726) |
| 0.0 | 0.8 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 0.1 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.0 | 0.3 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.0 | 0.5 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 1.0 | GO:0016235 | aggresome(GO:0016235) |
| 0.0 | 0.5 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 0.2 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.0 | 1.5 | GO:0035580 | specific granule lumen(GO:0035580) |
| 0.0 | 0.3 | GO:0031588 | cAMP-dependent protein kinase complex(GO:0005952) nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 1.7 | GO:0032432 | actin filament bundle(GO:0032432) |
| 0.0 | 5.3 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.0 | 1.2 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.2 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 1.2 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.4 | GO:0045095 | keratin filament(GO:0045095) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 6.3 | REACTOME DOWNREGULATION OF TGF BETA RECEPTOR SIGNALING | Genes involved in Downregulation of TGF-beta receptor signaling |
| 0.1 | 2.4 | REACTOME ADENYLATE CYCLASE INHIBITORY PATHWAY | Genes involved in Adenylate cyclase inhibitory pathway |
| 0.1 | 2.1 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.1 | 3.8 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.1 | 6.0 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 2.4 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
| 0.1 | 1.9 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 1.2 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 1.1 | REACTOME EXTRACELLULAR MATRIX ORGANIZATION | Genes involved in Extracellular matrix organization |
| 0.0 | 1.3 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.0 | 0.7 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 0.7 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 0.2 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.0 | 0.2 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.0 | 0.3 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.0 | 1.6 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.0 | 0.6 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.2 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.0 | 0.4 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 0.3 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.5 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.8 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 0.1 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 0.5 | REACTOME THROMBIN SIGNALLING THROUGH PROTEINASE ACTIVATED RECEPTORS PARS | Genes involved in Thrombin signalling through proteinase activated receptors (PARs) |