Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
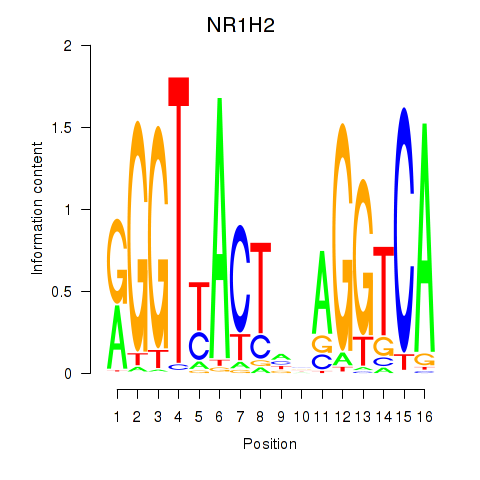
Results for NR1H2
Z-value: 0.75
Transcription factors associated with NR1H2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR1H2
|
ENSG00000131408.9 | NR1H2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR1H2 | hg19_v2_chr19_+_50879705_50879775 | -0.28 | 2.9e-01 | Click! |
Activity profile of NR1H2 motif
Sorted Z-values of NR1H2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR1H2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_+_10326612 | 1.72 |
ENST00000534464.1 ENST00000530439.1 ENST00000524948.1 ENST00000528655.1 ENST00000526492.1 ENST00000525063.1 |
ADM |
adrenomedullin |
| chrX_+_102631248 | 1.14 |
ENST00000361298.4 ENST00000372645.3 ENST00000372635.1 |
NGFRAP1 |
nerve growth factor receptor (TNFRSF16) associated protein 1 |
| chr12_-_91573249 | 1.11 |
ENST00000550099.1 ENST00000546391.1 ENST00000551354.1 |
DCN |
decorin |
| chr12_-_91573132 | 1.09 |
ENST00000550563.1 ENST00000546370.1 |
DCN |
decorin |
| chr1_+_169075554 | 1.07 |
ENST00000367815.4 |
ATP1B1 |
ATPase, Na+/K+ transporting, beta 1 polypeptide |
| chr9_-_107690420 | 0.97 |
ENST00000423487.2 ENST00000374733.1 ENST00000374736.3 |
ABCA1 |
ATP-binding cassette, sub-family A (ABC1), member 1 |
| chr8_-_48651648 | 0.97 |
ENST00000408965.3 |
CEBPD |
CCAAT/enhancer binding protein (C/EBP), delta |
| chr17_-_76975925 | 0.91 |
ENST00000591274.1 ENST00000589906.1 ENST00000591778.1 ENST00000589775.2 ENST00000585407.1 ENST00000262776.3 |
LGALS3BP |
lectin, galactoside-binding, soluble, 3 binding protein |
| chr5_+_35856951 | 0.89 |
ENST00000303115.3 ENST00000343305.4 ENST00000506850.1 ENST00000511982.1 |
IL7R |
interleukin 7 receptor |
| chr7_-_76247617 | 0.80 |
ENST00000441393.1 |
POMZP3 |
POM121 and ZP3 fusion |
| chr1_-_237167718 | 0.77 |
ENST00000464121.2 |
MT1HL1 |
metallothionein 1H-like 1 |
| chr20_+_43343517 | 0.73 |
ENST00000372865.4 |
WISP2 |
WNT1 inducible signaling pathway protein 2 |
| chr20_+_34680620 | 0.72 |
ENST00000430276.1 ENST00000373950.2 ENST00000452261.1 |
EPB41L1 |
erythrocyte membrane protein band 4.1-like 1 |
| chr15_+_96876340 | 0.71 |
ENST00000453270.2 |
NR2F2 |
nuclear receptor subfamily 2, group F, member 2 |
| chr20_+_43343886 | 0.71 |
ENST00000190983.4 |
WISP2 |
WNT1 inducible signaling pathway protein 2 |
| chr8_-_93115445 | 0.70 |
ENST00000523629.1 |
RUNX1T1 |
runt-related transcription factor 1; translocated to, 1 (cyclin D-related) |
| chr11_+_844406 | 0.70 |
ENST00000397404.1 |
TSPAN4 |
tetraspanin 4 |
| chr17_-_26694979 | 0.66 |
ENST00000438614.1 |
VTN |
vitronectin |
| chr17_-_26695013 | 0.64 |
ENST00000555059.2 |
CTB-96E2.2 |
Homeobox protein SEBOX |
| chr8_+_74903580 | 0.61 |
ENST00000284818.2 ENST00000518893.1 |
LY96 |
lymphocyte antigen 96 |
| chr15_+_96875657 | 0.59 |
ENST00000559679.1 ENST00000394171.2 |
NR2F2 |
nuclear receptor subfamily 2, group F, member 2 |
| chr17_+_4675175 | 0.57 |
ENST00000270560.3 |
TM4SF5 |
transmembrane 4 L six family member 5 |
| chr10_-_70287172 | 0.55 |
ENST00000539557.1 |
SLC25A16 |
solute carrier family 25 (mitochondrial carrier; Graves disease autoantigen), member 16 |
| chr11_+_66276550 | 0.55 |
ENST00000419755.3 |
CTD-3074O7.11 |
Bardet-Biedl syndrome 1 protein |
| chr11_-_82611448 | 0.54 |
ENST00000393399.2 ENST00000313010.3 |
PRCP |
prolylcarboxypeptidase (angiotensinase C) |
| chr2_+_223725652 | 0.53 |
ENST00000357430.3 ENST00000392066.3 |
ACSL3 |
acyl-CoA synthetase long-chain family member 3 |
| chrX_+_23801280 | 0.51 |
ENST00000379251.3 ENST00000379253.3 ENST00000379254.1 ENST00000379270.4 |
SAT1 |
spermidine/spermine N1-acetyltransferase 1 |
| chr11_-_35441524 | 0.51 |
ENST00000395750.1 ENST00000449068.1 |
SLC1A2 |
solute carrier family 1 (glial high affinity glutamate transporter), member 2 |
| chr1_+_87595433 | 0.49 |
ENST00000469312.2 ENST00000490006.2 |
RP5-1052I5.1 |
long intergenic non-protein coding RNA 1140 |
| chr7_-_22259845 | 0.47 |
ENST00000420196.1 |
RAPGEF5 |
Rap guanine nucleotide exchange factor (GEF) 5 |
| chr1_+_87458692 | 0.46 |
ENST00000370548.2 ENST00000356813.4 |
RP5-1052I5.2 HS2ST1 |
Heparan sulfate 2-O-sulfotransferase 1 heparan sulfate 2-O-sulfotransferase 1 |
| chr9_-_130829588 | 0.42 |
ENST00000373078.4 |
NAIF1 |
nuclear apoptosis inducing factor 1 |
| chr4_-_7941596 | 0.42 |
ENST00000420658.1 ENST00000358461.2 |
AFAP1 |
actin filament associated protein 1 |
| chr10_+_81838411 | 0.41 |
ENST00000372281.3 ENST00000372277.3 ENST00000372275.1 ENST00000372274.1 |
TMEM254 |
transmembrane protein 254 |
| chr16_-_18468926 | 0.40 |
ENST00000545114.1 |
RP11-1212A22.4 |
LOC339047 protein; Nuclear pore complex-interacting protein family member A3; Nuclear pore complex-interacting protein family member A5; Protein PKD1P1 |
| chr8_+_132916318 | 0.37 |
ENST00000254624.5 ENST00000522709.1 |
EFR3A |
EFR3 homolog A (S. cerevisiae) |
| chr11_+_119039414 | 0.36 |
ENST00000409991.1 ENST00000292199.2 ENST00000409265.4 ENST00000409109.1 |
NLRX1 |
NLR family member X1 |
| chr2_-_165697920 | 0.36 |
ENST00000342193.4 ENST00000375458.2 |
COBLL1 |
cordon-bleu WH2 repeat protein-like 1 |
| chr10_-_70287231 | 0.36 |
ENST00000609923.1 |
SLC25A16 |
solute carrier family 25 (mitochondrial carrier; Graves disease autoantigen), member 16 |
| chr11_-_35440796 | 0.36 |
ENST00000278379.3 |
SLC1A2 |
solute carrier family 1 (glial high affinity glutamate transporter), member 2 |
| chr15_-_63674218 | 0.35 |
ENST00000178638.3 |
CA12 |
carbonic anhydrase XII |
| chr4_-_39640700 | 0.35 |
ENST00000295958.5 |
SMIM14 |
small integral membrane protein 14 |
| chr19_-_7968427 | 0.34 |
ENST00000539278.1 |
AC010336.1 |
Uncharacterized protein |
| chr11_-_35441597 | 0.32 |
ENST00000395753.1 |
SLC1A2 |
solute carrier family 1 (glial high affinity glutamate transporter), member 2 |
| chr2_+_27255806 | 0.31 |
ENST00000238788.9 ENST00000404032.3 |
TMEM214 |
transmembrane protein 214 |
| chr15_-_90892669 | 0.31 |
ENST00000412799.2 |
GABARAPL3 |
GABA(A) receptors associated protein like 3, pseudogene |
| chr6_+_36097992 | 0.30 |
ENST00000211287.4 |
MAPK13 |
mitogen-activated protein kinase 13 |
| chr11_+_86748863 | 0.30 |
ENST00000340353.7 |
TMEM135 |
transmembrane protein 135 |
| chr19_+_2977444 | 0.29 |
ENST00000246112.4 ENST00000453329.1 ENST00000482627.1 ENST00000452088.1 |
TLE6 |
transducin-like enhancer of split 6 (E(sp1) homolog, Drosophila) |
| chr12_-_125399573 | 0.27 |
ENST00000339647.5 |
UBC |
ubiquitin C |
| chr3_-_51994694 | 0.27 |
ENST00000395014.2 |
PCBP4 |
poly(rC) binding protein 4 |
| chr16_-_1661988 | 0.26 |
ENST00000426508.2 |
IFT140 |
intraflagellar transport 140 homolog (Chlamydomonas) |
| chr7_+_100136811 | 0.25 |
ENST00000300176.4 ENST00000262935.4 |
AGFG2 |
ArfGAP with FG repeats 2 |
| chr1_+_156338993 | 0.24 |
ENST00000368249.1 ENST00000368246.2 ENST00000537040.1 ENST00000400992.2 ENST00000255013.3 ENST00000451864.2 |
RHBG |
Rh family, B glycoprotein (gene/pseudogene) |
| chr14_-_73925225 | 0.24 |
ENST00000356296.4 ENST00000355058.3 ENST00000359560.3 ENST00000557597.1 ENST00000554394.1 ENST00000555238.1 ENST00000535282.1 ENST00000555987.1 ENST00000555394.1 ENST00000554546.1 |
NUMB |
numb homolog (Drosophila) |
| chr11_+_64692143 | 0.23 |
ENST00000164133.2 ENST00000532850.1 |
PPP2R5B |
protein phosphatase 2, regulatory subunit B', beta |
| chr2_-_216946500 | 0.23 |
ENST00000265322.7 |
PECR |
peroxisomal trans-2-enoyl-CoA reductase |
| chr13_+_49684445 | 0.22 |
ENST00000398316.3 |
FNDC3A |
fibronectin type III domain containing 3A |
| chr19_+_15751689 | 0.22 |
ENST00000586182.2 ENST00000591058.1 ENST00000221307.8 |
CYP4F3 |
cytochrome P450, family 4, subfamily F, polypeptide 3 |
| chr1_-_67142710 | 0.22 |
ENST00000502413.2 |
AL139147.1 |
Uncharacterized protein |
| chr19_+_1026298 | 0.21 |
ENST00000263097.4 |
CNN2 |
calponin 2 |
| chrX_+_49020121 | 0.21 |
ENST00000415364.1 ENST00000376338.3 ENST00000425285.1 |
MAGIX |
MAGI family member, X-linked |
| chr17_-_42580738 | 0.21 |
ENST00000585614.1 ENST00000591680.1 ENST00000434000.1 ENST00000588554.1 ENST00000592154.1 |
GPATCH8 |
G patch domain containing 8 |
| chr1_+_150337100 | 0.20 |
ENST00000401000.4 |
RPRD2 |
regulation of nuclear pre-mRNA domain containing 2 |
| chr12_+_57914742 | 0.20 |
ENST00000551351.1 |
MBD6 |
methyl-CpG binding domain protein 6 |
| chr16_+_21312170 | 0.19 |
ENST00000338573.5 ENST00000561968.1 |
CRYM-AS1 |
CRYM antisense RNA 1 |
| chr16_+_4674787 | 0.18 |
ENST00000262370.7 |
MGRN1 |
mahogunin ring finger 1, E3 ubiquitin protein ligase |
| chr19_+_39881951 | 0.18 |
ENST00000315588.5 ENST00000594368.1 ENST00000599213.2 ENST00000596297.1 |
MED29 |
mediator complex subunit 29 |
| chr19_+_15752088 | 0.18 |
ENST00000585846.1 |
CYP4F3 |
cytochrome P450, family 4, subfamily F, polypeptide 3 |
| chr19_+_17622415 | 0.18 |
ENST00000252603.2 ENST00000600923.1 |
PGLS |
6-phosphogluconolactonase |
| chr16_-_68034470 | 0.18 |
ENST00000412757.2 |
DPEP2 |
dipeptidase 2 |
| chr16_+_4674814 | 0.18 |
ENST00000415496.1 ENST00000587747.1 ENST00000399577.5 ENST00000588994.1 ENST00000586183.1 |
MGRN1 |
mahogunin ring finger 1, E3 ubiquitin protein ligase |
| chr2_-_32390801 | 0.17 |
ENST00000608489.1 |
RP11-563N4.1 |
RP11-563N4.1 |
| chr22_-_24096562 | 0.17 |
ENST00000398465.3 |
VPREB3 |
pre-B lymphocyte 3 |
| chr19_-_45873642 | 0.17 |
ENST00000485403.2 ENST00000586856.1 ENST00000586131.1 ENST00000391940.4 ENST00000221481.6 ENST00000391944.3 ENST00000391945.4 |
ERCC2 |
excision repair cross-complementing rodent repair deficiency, complementation group 2 |
| chr11_-_68518910 | 0.17 |
ENST00000544963.1 ENST00000443940.2 |
MTL5 |
metallothionein-like 5, testis-specific (tesmin) |
| chr20_-_39928705 | 0.17 |
ENST00000436099.2 ENST00000309060.3 ENST00000373261.1 ENST00000436440.2 ENST00000540170.1 ENST00000557816.1 ENST00000560361.1 |
ZHX3 |
zinc fingers and homeoboxes 3 |
| chr3_+_9404526 | 0.17 |
ENST00000452837.2 ENST00000417036.1 ENST00000419437.1 ENST00000345094.3 ENST00000515662.2 |
THUMPD3 |
THUMP domain containing 3 |
| chr20_-_44600810 | 0.16 |
ENST00000322927.2 ENST00000426788.1 |
ZNF335 |
zinc finger protein 335 |
| chr2_+_71357744 | 0.15 |
ENST00000498451.2 |
MPHOSPH10 |
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr1_+_26503894 | 0.15 |
ENST00000361530.6 ENST00000374253.5 |
CNKSR1 |
connector enhancer of kinase suppressor of Ras 1 |
| chr9_+_34179003 | 0.14 |
ENST00000545103.1 ENST00000543944.1 ENST00000536252.1 ENST00000540348.1 ENST00000297661.4 ENST00000379186.4 |
UBAP1 |
ubiquitin associated protein 1 |
| chr22_-_24096630 | 0.14 |
ENST00000248948.3 |
VPREB3 |
pre-B lymphocyte 3 |
| chr3_-_49466686 | 0.13 |
ENST00000273598.3 ENST00000436744.2 |
NICN1 |
nicolin 1 |
| chr19_+_7968728 | 0.13 |
ENST00000397981.3 ENST00000545011.1 ENST00000397983.3 ENST00000397979.3 |
MAP2K7 |
mitogen-activated protein kinase kinase 7 |
| chr4_-_140216948 | 0.13 |
ENST00000265500.4 |
NDUFC1 |
NADH dehydrogenase (ubiquinone) 1, subcomplex unknown, 1, 6kDa |
| chr12_-_125398850 | 0.12 |
ENST00000535859.1 ENST00000546271.1 ENST00000540700.1 ENST00000546120.1 |
UBC |
ubiquitin C |
| chr19_-_38806560 | 0.12 |
ENST00000591755.1 ENST00000337679.8 ENST00000339413.6 |
YIF1B |
Yip1 interacting factor homolog B (S. cerevisiae) |
| chr17_-_17480779 | 0.12 |
ENST00000395782.1 |
PEMT |
phosphatidylethanolamine N-methyltransferase |
| chr17_-_35716019 | 0.12 |
ENST00000591148.1 ENST00000394406.2 ENST00000394403.1 |
ACACA |
acetyl-CoA carboxylase alpha |
| chr12_-_113841678 | 0.12 |
ENST00000552280.1 ENST00000257549.4 |
SDS |
serine dehydratase |
| chr8_-_27941380 | 0.12 |
ENST00000413272.2 ENST00000341513.6 |
NUGGC |
nuclear GTPase, germinal center associated |
| chr16_+_29467780 | 0.11 |
ENST00000395400.3 |
SULT1A4 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 4 |
| chr14_-_70826438 | 0.11 |
ENST00000389912.6 |
COX16 |
COX16 cytochrome c oxidase assembly homolog (S. cerevisiae) |
| chr21_-_37432764 | 0.11 |
ENST00000399212.1 ENST00000446166.1 |
SETD4 |
SET domain containing 4 |
| chr19_-_46296011 | 0.11 |
ENST00000377735.3 ENST00000270223.6 |
DMWD |
dystrophia myotonica, WD repeat containing |
| chr2_-_107084826 | 0.11 |
ENST00000304514.7 ENST00000409886.3 |
RGPD3 |
RANBP2-like and GRIP domain containing 3 |
| chr18_+_21693306 | 0.10 |
ENST00000540918.2 |
TTC39C |
tetratricopeptide repeat domain 39C |
| chr17_+_32683456 | 0.10 |
ENST00000225844.2 |
CCL13 |
chemokine (C-C motif) ligand 13 |
| chr21_-_37432832 | 0.10 |
ENST00000332131.4 |
SETD4 |
SET domain containing 4 |
| chr20_-_39928756 | 0.09 |
ENST00000432768.2 |
ZHX3 |
zinc fingers and homeoboxes 3 |
| chr10_+_126490354 | 0.09 |
ENST00000298492.5 |
FAM175B |
family with sequence similarity 175, member B |
| chr12_+_120972606 | 0.09 |
ENST00000413266.2 |
RNF10 |
ring finger protein 10 |
| chr11_-_506316 | 0.08 |
ENST00000532055.1 ENST00000531540.1 |
RNH1 |
ribonuclease/angiogenin inhibitor 1 |
| chr13_-_33112823 | 0.08 |
ENST00000504114.1 |
N4BP2L2 |
NEDD4 binding protein 2-like 2 |
| chr16_+_30710462 | 0.08 |
ENST00000262518.4 ENST00000395059.2 ENST00000344771.4 |
SRCAP |
Snf2-related CREBBP activator protein |
| chr9_-_86955657 | 0.08 |
ENST00000537648.1 |
SLC28A3 |
solute carrier family 28 (concentrative nucleoside transporter), member 3 |
| chr4_-_83483395 | 0.08 |
ENST00000515780.2 |
TMEM150C |
transmembrane protein 150C |
| chr12_+_22778009 | 0.08 |
ENST00000266517.4 ENST00000335148.3 |
ETNK1 |
ethanolamine kinase 1 |
| chr9_-_36400213 | 0.08 |
ENST00000259605.6 ENST00000353739.4 |
RNF38 |
ring finger protein 38 |
| chr7_+_123485102 | 0.08 |
ENST00000488323.1 ENST00000223026.4 |
HYAL4 |
hyaluronoglucosaminidase 4 |
| chr16_+_3550924 | 0.08 |
ENST00000576634.1 ENST00000574369.1 ENST00000341633.5 ENST00000417763.2 ENST00000571025.1 |
CLUAP1 |
clusterin associated protein 1 |
| chr14_+_23790655 | 0.07 |
ENST00000397276.2 |
PABPN1 |
poly(A) binding protein, nuclear 1 |
| chr19_-_58427944 | 0.07 |
ENST00000312026.5 ENST00000536263.1 ENST00000598629.1 ENST00000595559.1 ENST00000597515.1 ENST00000599251.1 ENST00000598526.1 |
ZNF417 |
zinc finger protein 417 |
| chr19_-_2042065 | 0.07 |
ENST00000591588.1 ENST00000591142.1 |
MKNK2 |
MAP kinase interacting serine/threonine kinase 2 |
| chr19_+_57631411 | 0.07 |
ENST00000254181.4 ENST00000600940.1 |
USP29 |
ubiquitin specific peptidase 29 |
| chr17_+_66243715 | 0.07 |
ENST00000359904.3 |
AMZ2 |
archaelysin family metallopeptidase 2 |
| chr6_+_31674639 | 0.07 |
ENST00000556581.1 ENST00000375832.4 ENST00000503322.1 |
LY6G6F MEGT1 |
lymphocyte antigen 6 complex, locus G6F HCG43720, isoform CRA_a; Lymphocyte antigen 6 complex locus protein G6f; Megakaryocyte-enhanced gene transcript 1 protein; Uncharacterized protein |
| chr19_+_41768401 | 0.07 |
ENST00000352456.3 ENST00000595018.1 ENST00000597725.1 |
HNRNPUL1 |
heterogeneous nuclear ribonucleoprotein U-like 1 |
| chr8_-_53626974 | 0.07 |
ENST00000435644.2 ENST00000518710.1 ENST00000025008.5 ENST00000517963.1 |
RB1CC1 |
RB1-inducible coiled-coil 1 |
| chr17_-_76183111 | 0.07 |
ENST00000405273.1 ENST00000590862.1 ENST00000590430.1 ENST00000586613.1 |
TK1 |
thymidine kinase 1, soluble |
| chr11_+_75110530 | 0.07 |
ENST00000531188.1 ENST00000530164.1 ENST00000422465.2 ENST00000278572.6 ENST00000534440.1 ENST00000527446.1 ENST00000526608.1 ENST00000527273.1 ENST00000524851.1 |
RPS3 |
ribosomal protein S3 |
| chr19_+_58361185 | 0.06 |
ENST00000339656.5 ENST00000423137.1 |
ZNF587 |
zinc finger protein 587 |
| chr2_+_113735575 | 0.06 |
ENST00000376489.2 ENST00000259205.4 |
IL36G |
interleukin 36, gamma |
| chr16_+_56995854 | 0.06 |
ENST00000566128.1 |
CETP |
cholesteryl ester transfer protein, plasma |
| chr12_-_118797475 | 0.06 |
ENST00000541786.1 ENST00000419821.2 ENST00000541878.1 |
TAOK3 |
TAO kinase 3 |
| chr6_+_33378738 | 0.06 |
ENST00000374512.3 ENST00000374516.3 |
PHF1 |
PHD finger protein 1 |
| chr13_-_33112956 | 0.06 |
ENST00000505213.1 |
N4BP2L2 |
NEDD4 binding protein 2-like 2 |
| chr19_+_44220247 | 0.05 |
ENST00000596627.1 |
IRGC |
immunity-related GTPase family, cinema |
| chr22_-_38577782 | 0.05 |
ENST00000430886.1 ENST00000332509.3 ENST00000447598.2 ENST00000435484.1 ENST00000402064.1 ENST00000436218.1 |
PLA2G6 |
phospholipase A2, group VI (cytosolic, calcium-independent) |
| chr5_-_180671172 | 0.05 |
ENST00000512805.1 |
GNB2L1 |
guanine nucleotide binding protein (G protein), beta polypeptide 2-like 1 |
| chr9_-_86955598 | 0.05 |
ENST00000376238.4 |
SLC28A3 |
solute carrier family 28 (concentrative nucleoside transporter), member 3 |
| chr17_-_79604075 | 0.05 |
ENST00000374747.5 ENST00000539314.1 ENST00000331134.6 |
NPLOC4 |
nuclear protein localization 4 homolog (S. cerevisiae) |
| chr19_+_16254488 | 0.05 |
ENST00000588246.1 ENST00000593031.1 |
HSH2D |
hematopoietic SH2 domain containing |
| chr7_+_153584166 | 0.05 |
ENST00000404039.1 |
DPP6 |
dipeptidyl-peptidase 6 |
| chr22_-_38577731 | 0.05 |
ENST00000335539.3 |
PLA2G6 |
phospholipase A2, group VI (cytosolic, calcium-independent) |
| chr13_-_33112899 | 0.04 |
ENST00000267068.3 ENST00000357505.6 ENST00000399396.3 |
N4BP2L2 |
NEDD4 binding protein 2-like 2 |
| chr5_+_172410757 | 0.04 |
ENST00000519374.1 ENST00000519911.1 ENST00000265093.4 ENST00000517669.1 |
ATP6V0E1 |
ATPase, H+ transporting, lysosomal 9kDa, V0 subunit e1 |
| chr9_+_135937365 | 0.04 |
ENST00000372080.4 ENST00000351304.7 |
CEL |
carboxyl ester lipase |
| chr2_+_108443388 | 0.04 |
ENST00000354986.4 ENST00000408999.3 |
RGPD4 |
RANBP2-like and GRIP domain containing 4 |
| chr3_+_46283863 | 0.04 |
ENST00000545097.1 ENST00000541018.1 |
CCR3 |
chemokine (C-C motif) receptor 3 |
| chrX_+_30248553 | 0.03 |
ENST00000361644.2 |
MAGEB3 |
melanoma antigen family B, 3 |
| chr22_-_36031181 | 0.03 |
ENST00000594060.1 |
AL049747.1 |
AL049747.1 |
| chrX_+_21874105 | 0.03 |
ENST00000429584.2 |
YY2 |
YY2 transcription factor |
| chr19_+_15904761 | 0.03 |
ENST00000308940.8 |
OR10H5 |
olfactory receptor, family 10, subfamily H, member 5 |
| chr12_-_56122761 | 0.03 |
ENST00000552164.1 ENST00000420846.3 ENST00000257857.4 |
CD63 |
CD63 molecule |
| chr19_+_41768561 | 0.03 |
ENST00000599719.1 ENST00000601309.1 |
HNRNPUL1 |
heterogeneous nuclear ribonucleoprotein U-like 1 |
| chrX_-_9001351 | 0.03 |
ENST00000362066.3 |
FAM9B |
family with sequence similarity 9, member B |
| chr1_+_24117627 | 0.03 |
ENST00000400061.1 |
LYPLA2 |
lysophospholipase II |
| chr12_-_56123444 | 0.02 |
ENST00000546457.1 ENST00000549117.1 |
CD63 |
CD63 molecule |
| chr12_+_122064398 | 0.02 |
ENST00000330079.7 |
ORAI1 |
ORAI calcium release-activated calcium modulator 1 |
| chr22_+_36113919 | 0.02 |
ENST00000249044.2 |
APOL5 |
apolipoprotein L, 5 |
| chr19_-_54850417 | 0.02 |
ENST00000291759.4 |
LILRA4 |
leukocyte immunoglobulin-like receptor, subfamily A (with TM domain), member 4 |
| chr14_-_77787198 | 0.02 |
ENST00000261534.4 |
POMT2 |
protein-O-mannosyltransferase 2 |
| chr4_-_39367949 | 0.02 |
ENST00000503784.1 ENST00000349703.2 ENST00000381897.1 |
RFC1 |
replication factor C (activator 1) 1, 145kDa |
| chr1_-_13115578 | 0.02 |
ENST00000414205.2 |
PRAMEF6 |
PRAME family member 6 |
| chr3_+_39371255 | 0.02 |
ENST00000414803.1 ENST00000545843.1 |
CCR8 |
chemokine (C-C motif) receptor 8 |
| chr8_-_145652336 | 0.01 |
ENST00000529182.1 ENST00000526054.1 |
VPS28 |
vacuolar protein sorting 28 homolog (S. cerevisiae) |
| chr1_+_13516066 | 0.01 |
ENST00000332192.6 |
PRAMEF21 |
PRAME family member 21 |
| chr15_+_64386261 | 0.01 |
ENST00000560829.1 |
SNX1 |
sorting nexin 1 |
| chr3_+_9691117 | 0.01 |
ENST00000353332.5 ENST00000420925.1 ENST00000296003.4 ENST00000351233.5 |
MTMR14 |
myotubularin related protein 14 |
| chr3_-_127309879 | 0.00 |
ENST00000489960.1 ENST00000490290.1 |
TPRA1 |
transmembrane protein, adipocyte asscociated 1 |
| chr11_-_68519026 | 0.00 |
ENST00000255087.5 |
MTL5 |
metallothionein-like 5, testis-specific (tesmin) |
| chr1_+_169337172 | 0.00 |
ENST00000367807.3 ENST00000367808.3 ENST00000329281.2 ENST00000420531.1 |
BLZF1 |
basic leucine zipper nuclear factor 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.7 | GO:0045906 | negative regulation of vasoconstriction(GO:0045906) |
| 0.3 | 1.3 | GO:0009956 | radial pattern formation(GO:0009956) |
| 0.3 | 1.1 | GO:0044861 | protein transport into plasma membrane raft(GO:0044861) positive regulation of potassium ion import(GO:1903288) |
| 0.2 | 1.0 | GO:0090107 | aminophospholipid transport(GO:0015917) regulation of high-density lipoprotein particle assembly(GO:0090107) |
| 0.2 | 2.2 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.2 | 0.5 | GO:2001245 | regulation of phosphatidylcholine biosynthetic process(GO:2001245) |
| 0.2 | 0.5 | GO:0009447 | putrescine catabolic process(GO:0009447) |
| 0.1 | 1.2 | GO:0070777 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.1 | 0.5 | GO:0002254 | regulation of thyroid hormone mediated signaling pathway(GO:0002155) kinin cascade(GO:0002254) plasma kallikrein-kinin cascade(GO:0002353) |
| 0.1 | 0.8 | GO:0035803 | egg coat formation(GO:0035803) |
| 0.1 | 0.9 | GO:0033089 | positive regulation of T cell differentiation in thymus(GO:0033089) positive regulation of thymocyte aggregation(GO:2000400) |
| 0.1 | 0.6 | GO:0032497 | detection of lipopolysaccharide(GO:0032497) |
| 0.1 | 0.2 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.1 | 0.4 | GO:0036101 | leukotriene catabolic process(GO:0036100) leukotriene B4 catabolic process(GO:0036101) leukotriene B4 metabolic process(GO:0036102) icosanoid catabolic process(GO:1901523) fatty acid derivative catabolic process(GO:1901569) |
| 0.1 | 0.2 | GO:0030860 | regulation of polarized epithelial cell differentiation(GO:0030860) |
| 0.1 | 0.6 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 | 0.4 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.1 | 0.2 | GO:0070317 | negative regulation of G0 to G1 transition(GO:0070317) |
| 0.1 | 0.2 | GO:0032289 | central nervous system myelin formation(GO:0032289) |
| 0.0 | 0.3 | GO:0060136 | embryonic process involved in female pregnancy(GO:0060136) |
| 0.0 | 0.4 | GO:0039536 | negative regulation of RIG-I signaling pathway(GO:0039536) |
| 0.0 | 0.4 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.0 | 0.1 | GO:2001295 | malonyl-CoA biosynthetic process(GO:2001295) |
| 0.0 | 0.3 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.0 | 0.2 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 0.0 | 0.2 | GO:1902035 | positive regulation of hematopoietic stem cell proliferation(GO:1902035) |
| 0.0 | 0.1 | GO:0015860 | pyrimidine nucleobase transport(GO:0015855) purine nucleoside transmembrane transport(GO:0015860) purine nucleobase transmembrane transport(GO:1904823) |
| 0.0 | 0.1 | GO:0032049 | cardiolipin biosynthetic process(GO:0032049) |
| 0.0 | 0.1 | GO:1904247 | positive regulation of polynucleotide adenylyltransferase activity(GO:1904247) |
| 0.0 | 0.1 | GO:0090264 | immune complex clearance(GO:0002434) immune complex clearance by monocytes and macrophages(GO:0002436) T cell extravasation(GO:0072683) regulation of immune complex clearance by monocytes and macrophages(GO:0090264) |
| 0.0 | 0.4 | GO:0051639 | actin filament network formation(GO:0051639) |
| 0.0 | 0.4 | GO:0010992 | ubiquitin homeostasis(GO:0010992) |
| 0.0 | 0.1 | GO:0042866 | pyruvate biosynthetic process(GO:0042866) |
| 0.0 | 0.3 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 0.1 | GO:0030242 | pexophagy(GO:0030242) |
| 0.0 | 0.1 | GO:0035897 | proteolysis in other organism(GO:0035897) |
| 0.0 | 0.1 | GO:0046104 | thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.0 | 0.1 | GO:1902544 | regulation of DNA N-glycosylase activity(GO:1902544) |
| 0.0 | 0.1 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.0 | 0.7 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 0.7 | GO:0045599 | negative regulation of fat cell differentiation(GO:0045599) |
| 0.0 | 1.2 | GO:0045669 | positive regulation of osteoblast differentiation(GO:0045669) |
| 0.0 | 0.1 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.0 | 0.2 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.0 | 0.1 | GO:0010624 | regulation of Schwann cell proliferation(GO:0010624) negative regulation of Schwann cell proliferation(GO:0010626) |
| 0.0 | 0.2 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.0 | 0.3 | GO:0090140 | regulation of mitochondrial fission(GO:0090140) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0090556 | apolipoprotein A-I receptor activity(GO:0034188) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.3 | 0.9 | GO:0004917 | interleukin-7 receptor activity(GO:0004917) |
| 0.2 | 1.2 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.1 | 0.4 | GO:0052871 | tocopherol omega-hydroxylase activity(GO:0052870) alpha-tocopherol omega-hydroxylase activity(GO:0052871) 20-hydroxy-leukotriene B4 omega oxidase activity(GO:0097258) 20-aldehyde-leukotriene B4 20-monooxygenase activity(GO:0097259) |
| 0.1 | 0.8 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.1 | 0.2 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 0.1 | 0.6 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 0.1 | 1.1 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.1 | 0.2 | GO:0017057 | 6-phosphogluconolactonase activity(GO:0017057) |
| 0.0 | 0.1 | GO:0015389 | pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |
| 0.0 | 0.7 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 0.1 | GO:0004608 | phosphatidyl-N-methylethanolamine N-methyltransferase activity(GO:0000773) phosphatidylethanolamine N-methyltransferase activity(GO:0004608) phosphatidyl-N-dimethylethanolamine N-methyltransferase activity(GO:0080101) |
| 0.0 | 1.4 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 0.5 | GO:0017034 | Rap guanyl-nucleotide exchange factor activity(GO:0017034) |
| 0.0 | 1.3 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 0.8 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 2.2 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.1 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.0 | 0.5 | GO:0034483 | heparan sulfate sulfotransferase activity(GO:0034483) |
| 0.0 | 0.1 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 0.0 | 0.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.0 | 0.4 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.5 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 0.1 | GO:0004797 | thymidine kinase activity(GO:0004797) |
| 0.0 | 0.2 | GO:0043139 | 5'-3' DNA helicase activity(GO:0043139) |
| 0.0 | 0.9 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.2 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.0 | 0.3 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.2 | GO:0016423 | tRNA (guanine) methyltransferase activity(GO:0016423) |
| 0.0 | 0.0 | GO:0017129 | triglyceride binding(GO:0017129) |
| 0.0 | 1.6 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 0.1 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.0 | 0.2 | GO:0030506 | ankyrin binding(GO:0030506) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0071062 | rough endoplasmic reticulum lumen(GO:0048237) alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 0.2 | 2.2 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.1 | 0.6 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.1 | 1.1 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.1 | 0.9 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.0 | 0.3 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.0 | 1.2 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 0.1 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.0 | 0.2 | GO:0071817 | MMXD complex(GO:0071817) |
| 0.0 | 1.0 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.0 | 0.2 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 0.1 | GO:0071159 | NF-kappaB complex(GO:0071159) |
| 0.0 | 0.2 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.0 | 0.1 | GO:0032311 | angiogenin-PRI complex(GO:0032311) |
| 0.0 | 1.0 | GO:0031903 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 0.4 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.0 | 0.1 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.2 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 0.9 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 0.6 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.0 | 0.9 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.9 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.5 | REACTOME INTRINSIC PATHWAY | Genes involved in Intrinsic Pathway |
| 0.0 | 1.1 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 2.4 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.4 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.1 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.0 | 0.3 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.0 | 0.8 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 1.1 | REACTOME CELL DEATH SIGNALLING VIA NRAGE NRIF AND NADE | Genes involved in Cell death signalling via NRAGE, NRIF and NADE |
| 0.0 | 1.6 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.7 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.0 | 0.5 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.3 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.0 | 0.2 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.2 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 2.2 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 0.7 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 1.0 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.6 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.3 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 1.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 1.7 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 0.5 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.0 | 0.7 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.7 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 1.1 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 2.2 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |