Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
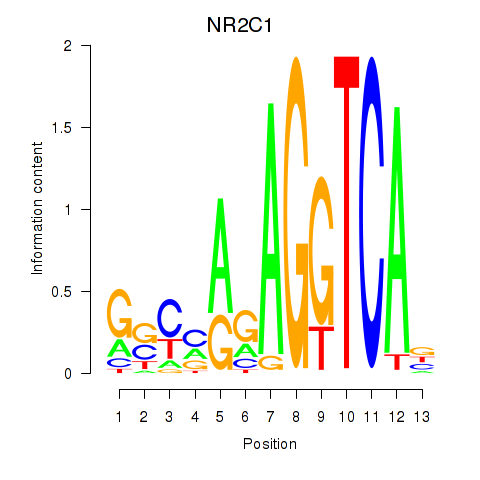
Results for NR2C1
Z-value: 0.81
Transcription factors associated with NR2C1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR2C1
|
ENSG00000120798.12 | NR2C1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR2C1 | hg19_v2_chr12_-_95467267_95467350 | -0.02 | 9.3e-01 | Click! |
Activity profile of NR2C1 motif
Sorted Z-values of NR2C1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR2C1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_35685452 | 2.17 |
ENST00000607559.1 |
TPM2 |
tropomyosin 2 (beta) |
| chr8_+_70404996 | 1.46 |
ENST00000402687.4 ENST00000419716.3 |
SULF1 |
sulfatase 1 |
| chr19_+_48828582 | 1.43 |
ENST00000270221.6 ENST00000596315.1 |
EMP3 |
epithelial membrane protein 3 |
| chr5_+_1801503 | 1.37 |
ENST00000274137.5 ENST00000469176.1 |
NDUFS6 |
NADH dehydrogenase (ubiquinone) Fe-S protein 6, 13kDa (NADH-coenzyme Q reductase) |
| chr19_+_48828788 | 1.36 |
ENST00000594198.1 ENST00000597279.1 ENST00000593437.1 |
EMP3 |
epithelial membrane protein 3 |
| chr2_-_163099885 | 1.32 |
ENST00000443424.1 |
FAP |
fibroblast activation protein, alpha |
| chr4_-_111544254 | 1.27 |
ENST00000306732.3 |
PITX2 |
paired-like homeodomain 2 |
| chr2_-_163100045 | 1.27 |
ENST00000188790.4 |
FAP |
fibroblast activation protein, alpha |
| chr11_+_131781290 | 1.23 |
ENST00000425719.2 ENST00000374784.1 |
NTM |
neurotrimin |
| chr17_-_39743139 | 1.22 |
ENST00000167586.6 |
KRT14 |
keratin 14 |
| chr1_-_229569834 | 1.14 |
ENST00000366684.3 ENST00000366683.2 |
ACTA1 |
actin, alpha 1, skeletal muscle |
| chr3_+_159557637 | 1.05 |
ENST00000445224.2 |
SCHIP1 |
schwannomin interacting protein 1 |
| chr12_-_90049828 | 0.93 |
ENST00000261173.2 ENST00000348959.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr12_-_90049878 | 0.88 |
ENST00000359142.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr8_-_145028013 | 0.87 |
ENST00000354958.2 |
PLEC |
plectin |
| chr11_-_64013663 | 0.82 |
ENST00000392210.2 |
PPP1R14B |
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr6_+_160148593 | 0.77 |
ENST00000337387.4 |
WTAP |
Wilms tumor 1 associated protein |
| chr13_-_33760216 | 0.72 |
ENST00000255486.4 |
STARD13 |
StAR-related lipid transfer (START) domain containing 13 |
| chr18_+_54318616 | 0.72 |
ENST00000254442.3 |
WDR7 |
WD repeat domain 7 |
| chr11_+_10471836 | 0.71 |
ENST00000444303.2 |
AMPD3 |
adenosine monophosphate deaminase 3 |
| chr18_+_54318566 | 0.67 |
ENST00000589935.1 ENST00000357574.3 |
WDR7 |
WD repeat domain 7 |
| chr11_-_64013288 | 0.61 |
ENST00000542235.1 |
PPP1R14B |
protein phosphatase 1, regulatory (inhibitor) subunit 14B |
| chr15_-_82555000 | 0.60 |
ENST00000557844.1 ENST00000359445.3 ENST00000268206.7 |
EFTUD1 |
elongation factor Tu GTP binding domain containing 1 |
| chr19_+_676385 | 0.55 |
ENST00000166139.4 |
FSTL3 |
follistatin-like 3 (secreted glycoprotein) |
| chr1_+_25071848 | 0.55 |
ENST00000374379.4 |
CLIC4 |
chloride intracellular channel 4 |
| chr7_-_38948774 | 0.55 |
ENST00000395969.2 ENST00000414632.1 ENST00000310301.4 |
VPS41 |
vacuolar protein sorting 41 homolog (S. cerevisiae) |
| chr1_-_155881156 | 0.53 |
ENST00000539040.1 ENST00000368323.3 |
RIT1 |
Ras-like without CAAX 1 |
| chr4_+_26321284 | 0.50 |
ENST00000506956.1 ENST00000512671.1 ENST00000345843.3 ENST00000342295.1 |
RBPJ |
recombination signal binding protein for immunoglobulin kappa J region |
| chr11_-_73687997 | 0.49 |
ENST00000545212.1 |
UCP2 |
uncoupling protein 2 (mitochondrial, proton carrier) |
| chr4_+_159593418 | 0.47 |
ENST00000507475.1 ENST00000307738.5 |
ETFDH |
electron-transferring-flavoprotein dehydrogenase |
| chr19_+_39616410 | 0.47 |
ENST00000602004.1 ENST00000599470.1 ENST00000321944.4 ENST00000593480.1 ENST00000358301.3 ENST00000593690.1 ENST00000599386.1 |
PAK4 |
p21 protein (Cdc42/Rac)-activated kinase 4 |
| chr2_-_38303218 | 0.47 |
ENST00000407341.1 ENST00000260630.3 |
CYP1B1 |
cytochrome P450, family 1, subfamily B, polypeptide 1 |
| chr15_-_43802769 | 0.46 |
ENST00000263801.3 |
TP53BP1 |
tumor protein p53 binding protein 1 |
| chr7_+_121513143 | 0.45 |
ENST00000393386.2 |
PTPRZ1 |
protein tyrosine phosphatase, receptor-type, Z polypeptide 1 |
| chr11_+_67798363 | 0.41 |
ENST00000525628.1 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr15_+_45422131 | 0.40 |
ENST00000321429.4 |
DUOX1 |
dual oxidase 1 |
| chr21_-_32931290 | 0.39 |
ENST00000286827.3 |
TIAM1 |
T-cell lymphoma invasion and metastasis 1 |
| chr12_-_118810688 | 0.38 |
ENST00000542532.1 ENST00000392533.3 |
TAOK3 |
TAO kinase 3 |
| chr16_-_4292071 | 0.38 |
ENST00000399609.3 |
SRL |
sarcalumenin |
| chr3_+_179065474 | 0.38 |
ENST00000471841.1 ENST00000280653.7 |
MFN1 |
mitofusin 1 |
| chr9_+_33795533 | 0.37 |
ENST00000379405.3 |
PRSS3 |
protease, serine, 3 |
| chr1_+_160097462 | 0.37 |
ENST00000447527.1 |
ATP1A2 |
ATPase, Na+/K+ transporting, alpha 2 polypeptide |
| chr15_+_45422178 | 0.36 |
ENST00000389037.3 ENST00000558322.1 |
DUOX1 |
dual oxidase 1 |
| chr13_+_21714711 | 0.35 |
ENST00000607003.1 ENST00000492245.1 |
SAP18 |
Sin3A-associated protein, 18kDa |
| chr11_-_45939565 | 0.35 |
ENST00000525192.1 ENST00000378750.5 |
PEX16 |
peroxisomal biogenesis factor 16 |
| chr16_+_2255841 | 0.34 |
ENST00000301725.7 |
MLST8 |
MTOR associated protein, LST8 homolog (S. cerevisiae) |
| chr20_-_44485835 | 0.34 |
ENST00000457981.1 ENST00000426915.1 ENST00000217455.4 |
ACOT8 |
acyl-CoA thioesterase 8 |
| chr2_+_201936707 | 0.34 |
ENST00000433898.1 ENST00000454214.1 |
NDUFB3 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 3, 12kDa |
| chr12_-_42632016 | 0.34 |
ENST00000442791.3 ENST00000327791.4 ENST00000534854.2 ENST00000380788.3 ENST00000380790.4 |
YAF2 |
YY1 associated factor 2 |
| chr19_+_44100632 | 0.33 |
ENST00000533118.1 |
ZNF576 |
zinc finger protein 576 |
| chr6_+_57037089 | 0.33 |
ENST00000370693.5 |
BAG2 |
BCL2-associated athanogene 2 |
| chr11_-_26743546 | 0.33 |
ENST00000280467.6 ENST00000396005.3 |
SLC5A12 |
solute carrier family 5 (sodium/monocarboxylate cotransporter), member 12 |
| chr16_+_2255710 | 0.33 |
ENST00000397124.1 ENST00000565250.1 |
MLST8 |
MTOR associated protein, LST8 homolog (S. cerevisiae) |
| chr3_-_50605077 | 0.32 |
ENST00000426034.1 ENST00000441239.1 |
C3orf18 |
chromosome 3 open reading frame 18 |
| chr1_-_155880672 | 0.30 |
ENST00000609492.1 ENST00000368322.3 |
RIT1 |
Ras-like without CAAX 1 |
| chr11_+_64073699 | 0.30 |
ENST00000405666.1 ENST00000468670.1 |
ESRRA |
estrogen-related receptor alpha |
| chr10_-_103815874 | 0.29 |
ENST00000370033.4 ENST00000311122.5 |
C10orf76 |
chromosome 10 open reading frame 76 |
| chr3_-_113465065 | 0.28 |
ENST00000497255.1 ENST00000478020.1 ENST00000240922.3 ENST00000493900.1 |
NAA50 |
N(alpha)-acetyltransferase 50, NatE catalytic subunit |
| chr13_+_21714653 | 0.28 |
ENST00000382533.4 |
SAP18 |
Sin3A-associated protein, 18kDa |
| chr8_-_80942467 | 0.28 |
ENST00000518271.1 ENST00000276585.4 ENST00000521605.1 |
MRPS28 |
mitochondrial ribosomal protein S28 |
| chr6_+_149068464 | 0.27 |
ENST00000367463.4 |
UST |
uronyl-2-sulfotransferase |
| chr18_-_54318353 | 0.27 |
ENST00000590954.1 ENST00000540155.1 |
TXNL1 |
thioredoxin-like 1 |
| chr19_+_44100727 | 0.27 |
ENST00000528387.1 ENST00000529930.1 ENST00000336564.4 ENST00000607544.1 ENST00000526798.1 |
ZNF576 SRRM5 |
zinc finger protein 576 serine/arginine repetitive matrix 5 |
| chr2_+_109237717 | 0.26 |
ENST00000409441.1 |
LIMS1 |
LIM and senescent cell antigen-like domains 1 |
| chr12_-_6665200 | 0.25 |
ENST00000336604.4 ENST00000396840.2 ENST00000356896.4 |
IFFO1 |
intermediate filament family orphan 1 |
| chr3_+_37284824 | 0.25 |
ENST00000431105.1 |
GOLGA4 |
golgin A4 |
| chr2_+_201936458 | 0.25 |
ENST00000237889.4 |
NDUFB3 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 3, 12kDa |
| chr2_-_202316260 | 0.24 |
ENST00000332624.3 |
TRAK2 |
trafficking protein, kinesin binding 2 |
| chr10_+_26986582 | 0.24 |
ENST00000376215.5 ENST00000376203.5 |
PDSS1 |
prenyl (decaprenyl) diphosphate synthase, subunit 1 |
| chr12_+_67663056 | 0.24 |
ENST00000545606.1 |
CAND1 |
cullin-associated and neddylation-dissociated 1 |
| chr6_+_30594619 | 0.23 |
ENST00000318999.7 ENST00000376485.4 ENST00000376478.2 ENST00000319027.5 ENST00000376483.4 ENST00000329992.8 ENST00000330083.5 |
ATAT1 |
alpha tubulin acetyltransferase 1 |
| chr11_+_67798090 | 0.22 |
ENST00000313468.5 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr4_-_76598296 | 0.22 |
ENST00000395719.3 |
G3BP2 |
GTPase activating protein (SH3 domain) binding protein 2 |
| chr22_+_39745930 | 0.21 |
ENST00000318801.4 ENST00000216155.7 ENST00000406293.3 ENST00000328933.5 |
SYNGR1 |
synaptogyrin 1 |
| chr3_-_50605150 | 0.20 |
ENST00000357203.3 |
C3orf18 |
chromosome 3 open reading frame 18 |
| chr11_-_45939374 | 0.20 |
ENST00000533151.1 ENST00000241041.3 |
PEX16 |
peroxisomal biogenesis factor 16 |
| chr11_+_67798114 | 0.19 |
ENST00000453471.2 ENST00000528492.1 ENST00000526339.1 ENST00000525419.1 |
NDUFS8 |
NADH dehydrogenase (ubiquinone) Fe-S protein 8, 23kDa (NADH-coenzyme Q reductase) |
| chr3_+_37284668 | 0.19 |
ENST00000361924.2 ENST00000444882.1 ENST00000356847.4 ENST00000450863.2 ENST00000429018.1 |
GOLGA4 |
golgin A4 |
| chr3_-_116163830 | 0.19 |
ENST00000333617.4 |
LSAMP |
limbic system-associated membrane protein |
| chr14_-_65289812 | 0.18 |
ENST00000389720.3 ENST00000389721.5 ENST00000389722.3 |
SPTB |
spectrin, beta, erythrocytic |
| chr22_-_29784519 | 0.18 |
ENST00000357586.2 ENST00000356015.2 ENST00000432560.2 ENST00000317368.7 |
AP1B1 |
adaptor-related protein complex 1, beta 1 subunit |
| chr19_+_10197463 | 0.18 |
ENST00000590378.1 ENST00000397881.3 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr19_-_10679697 | 0.17 |
ENST00000335766.2 |
CDKN2D |
cyclin-dependent kinase inhibitor 2D (p19, inhibits CDK4) |
| chr10_+_64564469 | 0.16 |
ENST00000373783.1 |
ADO |
2-aminoethanethiol (cysteamine) dioxygenase |
| chr3_+_101546827 | 0.16 |
ENST00000461724.1 ENST00000483180.1 ENST00000394054.2 |
NFKBIZ |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta |
| chr2_+_46926326 | 0.15 |
ENST00000394861.2 |
SOCS5 |
suppressor of cytokine signaling 5 |
| chr17_-_15903002 | 0.15 |
ENST00000399277.1 |
ZSWIM7 |
zinc finger, SWIM-type containing 7 |
| chr19_+_44100544 | 0.15 |
ENST00000391965.2 ENST00000525771.1 |
ZNF576 |
zinc finger protein 576 |
| chr19_+_10196981 | 0.14 |
ENST00000591813.1 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr1_-_226374373 | 0.14 |
ENST00000366812.5 |
ACBD3 |
acyl-CoA binding domain containing 3 |
| chr2_+_219433281 | 0.13 |
ENST00000273064.6 ENST00000509807.2 ENST00000542068.1 |
RQCD1 |
RCD1 required for cell differentiation1 homolog (S. pombe) |
| chr16_-_70729496 | 0.13 |
ENST00000567648.1 |
VAC14 |
Vac14 homolog (S. cerevisiae) |
| chr3_-_116164306 | 0.12 |
ENST00000490035.2 |
LSAMP |
limbic system-associated membrane protein |
| chr1_+_165796753 | 0.12 |
ENST00000367879.4 |
UCK2 |
uridine-cytidine kinase 2 |
| chr10_-_118502070 | 0.12 |
ENST00000369209.3 |
HSPA12A |
heat shock 70kDa protein 12A |
| chr17_+_37821593 | 0.11 |
ENST00000578283.1 |
TCAP |
titin-cap |
| chr22_+_30163340 | 0.10 |
ENST00000330029.6 ENST00000401406.3 |
UQCR10 |
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr22_+_31518938 | 0.10 |
ENST00000412985.1 ENST00000331075.5 ENST00000412277.2 ENST00000420017.1 ENST00000400294.2 ENST00000405300.1 ENST00000404390.3 |
INPP5J |
inositol polyphosphate-5-phosphatase J |
| chr6_-_31138439 | 0.09 |
ENST00000259915.8 |
POU5F1 |
POU class 5 homeobox 1 |
| chr3_-_33700589 | 0.09 |
ENST00000461133.3 ENST00000496954.2 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr3_+_183903811 | 0.09 |
ENST00000429586.2 ENST00000292808.5 |
ABCF3 |
ATP-binding cassette, sub-family F (GCN20), member 3 |
| chr11_-_67275542 | 0.08 |
ENST00000531506.1 |
CDK2AP2 |
cyclin-dependent kinase 2 associated protein 2 |
| chr19_-_49371711 | 0.08 |
ENST00000355496.5 ENST00000263265.6 |
PLEKHA4 |
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 4 |
| chr2_+_46926048 | 0.07 |
ENST00000306503.5 |
SOCS5 |
suppressor of cytokine signaling 5 |
| chr7_-_72936608 | 0.07 |
ENST00000404251.1 |
BAZ1B |
bromodomain adjacent to zinc finger domain, 1B |
| chr22_-_31536480 | 0.07 |
ENST00000215885.3 |
PLA2G3 |
phospholipase A2, group III |
| chr22_+_41777927 | 0.07 |
ENST00000266304.4 |
TEF |
thyrotrophic embryonic factor |
| chr11_+_111896320 | 0.06 |
ENST00000531306.1 ENST00000537636.1 |
DLAT |
dihydrolipoamide S-acetyltransferase |
| chr12_+_132413739 | 0.04 |
ENST00000443358.2 |
PUS1 |
pseudouridylate synthase 1 |
| chr19_-_51472031 | 0.03 |
ENST00000391808.1 |
KLK6 |
kallikrein-related peptidase 6 |
| chr10_+_104005272 | 0.03 |
ENST00000369983.3 |
GBF1 |
golgi brefeldin A resistant guanine nucleotide exchange factor 1 |
| chr12_-_95467267 | 0.03 |
ENST00000330677.7 |
NR2C1 |
nuclear receptor subfamily 2, group C, member 1 |
| chr1_-_101491319 | 0.03 |
ENST00000342173.7 ENST00000488176.1 ENST00000370109.3 |
DPH5 |
diphthamide biosynthesis 5 |
| chr1_-_120439112 | 0.02 |
ENST00000369400.1 |
ADAM30 |
ADAM metallopeptidase domain 30 |
| chr3_+_113465866 | 0.02 |
ENST00000273398.3 ENST00000538620.1 ENST00000496747.1 ENST00000475322.1 |
ATP6V1A |
ATPase, H+ transporting, lysosomal 70kDa, V1 subunit A |
| chr6_-_109703663 | 0.01 |
ENST00000368961.5 |
CD164 |
CD164 molecule, sialomucin |
| chr6_+_29364416 | 0.01 |
ENST00000383555.2 |
OR12D2 |
olfactory receptor, family 12, subfamily D, member 2 (gene/pseudogene) |
| chr9_+_140513438 | 0.01 |
ENST00000462484.1 ENST00000334856.6 ENST00000460843.1 |
EHMT1 |
euchromatic histone-lysine N-methyltransferase 1 |
| chr8_-_30706608 | 0.00 |
ENST00000256246.2 |
TEX15 |
testis expressed 15 |
| chr22_-_38851205 | 0.00 |
ENST00000303592.3 |
KCNJ4 |
potassium inwardly-rectifying channel, subfamily J, member 4 |
| chr17_+_73997419 | 0.00 |
ENST00000425876.2 |
CDK3 |
cyclin-dependent kinase 3 |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.5 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.2 | 1.2 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.1 | 2.6 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 0.5 | GO:0048039 | ubiquinone binding(GO:0048039) |
| 0.1 | 0.7 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.1 | 0.5 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.1 | 0.5 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 0.1 | 1.8 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.1 | 2.8 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.1 | 0.8 | GO:0016174 | NAD(P)H oxidase activity(GO:0016174) |
| 0.0 | 0.2 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.0 | 0.5 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.0 | 0.3 | GO:0047134 | protein-disulfide reductase activity(GO:0047134) |
| 0.0 | 1.1 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.3 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.6 | GO:0048185 | activin binding(GO:0048185) |
| 0.0 | 1.4 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 0.4 | GO:1990239 | steroid hormone binding(GO:1990239) |
| 0.0 | 0.3 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.0 | 0.5 | GO:0070330 | aromatase activity(GO:0070330) |
| 0.0 | 2.3 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.3 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.0 | 1.3 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.0 | 1.1 | GO:0043531 | ADP binding(GO:0043531) |
| 0.0 | 0.5 | GO:0098641 | cadherin binding involved in cell-cell adhesion(GO:0098641) |
| 0.0 | 0.1 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.0 | 0.7 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.4 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.0 | 0.4 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.1 | GO:0016679 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors(GO:0016679) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.0 | 0.0 | GO:0004730 | pseudouridylate synthase activity(GO:0004730) |
| 0.0 | 0.2 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.4 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 0.6 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 0.6 | GO:0004407 | histone deacetylase activity(GO:0004407) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.3 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.0 | 1.7 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.9 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 0.4 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.0 | 1.2 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 0.4 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.8 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.0 | 1.8 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.7 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.0 | 0.6 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.5 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.5 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.5 | PID ATM PATHWAY | ATM pathway |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0036396 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 0.2 | 2.6 | GO:0071438 | invadopodium membrane(GO:0071438) |
| 0.1 | 0.4 | GO:0072534 | perineuronal net(GO:0072534) |
| 0.1 | 2.2 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.1 | 0.5 | GO:0033263 | CORVET complex(GO:0033263) |
| 0.1 | 0.5 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.1 | 0.7 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.1 | 1.8 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.1 | 0.6 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.0 | 2.8 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.3 | GO:0031415 | NatA complex(GO:0031415) |
| 0.0 | 0.9 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.0 | 0.4 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.0 | 1.1 | GO:0005865 | striated muscle thin filament(GO:0005865) |
| 0.0 | 0.4 | GO:0098799 | outer mitochondrial membrane protein complex(GO:0098799) |
| 0.0 | 1.2 | GO:0045095 | keratin filament(GO:0045095) |
| 0.0 | 0.2 | GO:0097129 | cyclin D2-CDK4 complex(GO:0097129) |
| 0.0 | 0.2 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.0 | 0.6 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 0.1 | GO:0035032 | phosphatidylinositol 3-kinase complex, class III(GO:0035032) |
| 0.0 | 0.4 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.0 | 0.4 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.0 | 0.5 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.5 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 0.2 | GO:0008091 | spectrin(GO:0008091) |
| 0.0 | 0.3 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.6 | GO:0097325 | melanocyte proliferation(GO:0097325) |
| 0.4 | 1.3 | GO:0060578 | subthalamic nucleus development(GO:0021763) prolactin secreting cell differentiation(GO:0060127) left lung morphogenesis(GO:0060460) pulmonary vein morphogenesis(GO:0060577) superior vena cava morphogenesis(GO:0060578) |
| 0.4 | 1.8 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.2 | 1.5 | GO:0060686 | negative regulation of prostatic bud formation(GO:0060686) |
| 0.2 | 1.2 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 0.2 | 1.1 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 0.2 | 0.8 | GO:0042335 | cuticle development(GO:0042335) |
| 0.2 | 0.6 | GO:0032581 | ER-dependent peroxisome organization(GO:0032581) |
| 0.2 | 0.5 | GO:1901297 | arterial endothelial cell fate commitment(GO:0060844) blood vessel lumenization(GO:0072554) positive regulation of ephrin receptor signaling pathway(GO:1901189) positive regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901297) positive regulation of canonical Wnt signaling pathway involved in heart development(GO:1905068) |
| 0.2 | 0.8 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.1 | 0.7 | GO:0097498 | endothelial tube lumen extension(GO:0097498) |
| 0.1 | 2.8 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.1 | 0.7 | GO:0006196 | AMP catabolic process(GO:0006196) |
| 0.1 | 0.5 | GO:0061304 | retinal blood vessel morphogenesis(GO:0061304) |
| 0.1 | 0.5 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.1 | 0.2 | GO:0098935 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) |
| 0.1 | 0.4 | GO:0051941 | regulation of amino acid uptake involved in synaptic transmission(GO:0051941) regulation of glutamate uptake involved in transmission of nerve impulse(GO:0051946) regulation of L-glutamate import(GO:1900920) |
| 0.1 | 0.6 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.1 | 0.6 | GO:0061299 | retina vasculature morphogenesis in camera-type eye(GO:0061299) |
| 0.1 | 0.4 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.1 | 0.5 | GO:2000042 | negative regulation of double-strand break repair via homologous recombination(GO:2000042) |
| 0.1 | 0.3 | GO:0016559 | peroxisome fission(GO:0016559) |
| 0.1 | 0.4 | GO:0070444 | oligodendrocyte progenitor proliferation(GO:0070444) regulation of oligodendrocyte progenitor proliferation(GO:0070445) |
| 0.0 | 0.5 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.0 | 1.4 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.0 | 0.1 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) |
| 0.0 | 0.2 | GO:0045629 | negative regulation of T-helper 2 cell differentiation(GO:0045629) |
| 0.0 | 0.3 | GO:0015727 | lactate transport(GO:0015727) lactate transmembrane transport(GO:0035873) |
| 0.0 | 0.3 | GO:0070601 | centromeric sister chromatid cohesion(GO:0070601) |
| 0.0 | 0.5 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.0 | 0.9 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.3 | GO:0006477 | protein sulfation(GO:0006477) |
| 0.0 | 0.1 | GO:0006238 | CMP salvage(GO:0006238) CMP biosynthetic process(GO:0009224) CMP metabolic process(GO:0046035) |
| 0.0 | 2.2 | GO:0033275 | muscle filament sliding(GO:0030049) actin-myosin filament sliding(GO:0033275) |
| 0.0 | 1.2 | GO:0008038 | neuron recognition(GO:0008038) |
| 0.0 | 0.2 | GO:0010265 | SCF complex assembly(GO:0010265) |
| 0.0 | 0.4 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.0 | 0.2 | GO:1900227 | positive regulation of NLRP3 inflammasome complex assembly(GO:1900227) |
| 0.0 | 0.1 | GO:2000035 | regulation of stem cell division(GO:2000035) |
| 0.0 | 1.4 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.7 | GO:0038202 | TORC1 signaling(GO:0038202) |
| 0.0 | 0.1 | GO:0001714 | endodermal cell fate specification(GO:0001714) |
| 0.0 | 0.2 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 0.3 | GO:0035641 | locomotory exploration behavior(GO:0035641) |
| 0.0 | 0.2 | GO:0048102 | autophagic cell death(GO:0048102) |
| 0.0 | 0.4 | GO:0007095 | mitotic G2 DNA damage checkpoint(GO:0007095) |
| 0.0 | 0.5 | GO:0048025 | negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.0 | 0.4 | GO:0000042 | protein targeting to Golgi(GO:0000042) |
| 0.0 | 0.2 | GO:0006744 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.0 | 0.0 | GO:0070902 | mitochondrial tRNA pseudouridine synthesis(GO:0070902) |
| 0.0 | 0.4 | GO:0009235 | cobalamin metabolic process(GO:0009235) |
| 0.0 | 0.1 | GO:0017062 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.0 | 0.2 | GO:0000098 | sulfur amino acid catabolic process(GO:0000098) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 3.3 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 0.8 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.0 | 2.2 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.7 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.7 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 0.9 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.5 | REACTOME ENDOGENOUS STEROLS | Genes involved in Endogenous sterols |
| 0.0 | 0.5 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 0.5 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.0 | 0.5 | REACTOME RESPIRATORY ELECTRON TRANSPORT ATP SYNTHESIS BY CHEMIOSMOTIC COUPLING AND HEAT PRODUCTION BY UNCOUPLING PROTEINS | Genes involved in Respiratory electron transport, ATP synthesis by chemiosmotic coupling, and heat production by uncoupling proteins. |
| 0.0 | 0.2 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.0 | 0.1 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |