Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
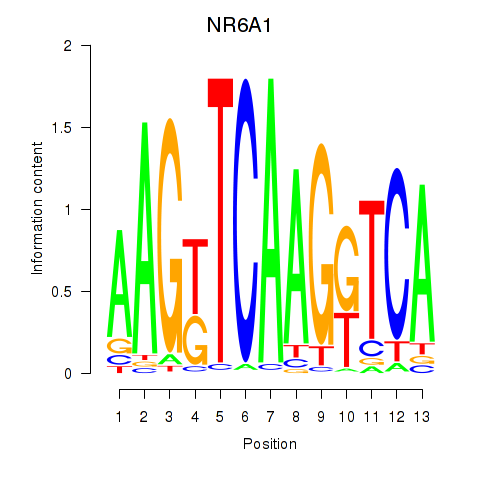
Results for NR6A1
Z-value: 1.07
Transcription factors associated with NR6A1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
NR6A1
|
ENSG00000148200.12 | NR6A1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| NR6A1 | hg19_v2_chr9_-_127533582_127533615 | 0.41 | 1.1e-01 | Click! |
Activity profile of NR6A1 motif
Sorted Z-values of NR6A1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of NR6A1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_5271122 | 1.67 |
ENST00000330597.3 |
HBG1 |
hemoglobin, gamma A |
| chr19_-_10446449 | 1.49 |
ENST00000592439.1 |
ICAM3 |
intercellular adhesion molecule 3 |
| chr2_+_219283815 | 1.33 |
ENST00000248444.5 ENST00000454069.1 ENST00000392114.2 |
VIL1 |
villin 1 |
| chr8_-_27472198 | 1.22 |
ENST00000519472.1 ENST00000523589.1 ENST00000522413.1 ENST00000523396.1 ENST00000560366.1 |
CLU |
clusterin |
| chr17_+_47448102 | 1.21 |
ENST00000576461.1 |
RP11-81K2.1 |
Uncharacterized protein |
| chr19_-_43383789 | 1.20 |
ENST00000595356.1 |
PSG1 |
pregnancy specific beta-1-glycoprotein 1 |
| chr3_-_148804275 | 1.15 |
ENST00000392912.2 ENST00000465259.1 ENST00000310053.5 ENST00000494055.1 |
HLTF |
helicase-like transcription factor |
| chr19_-_43383819 | 1.11 |
ENST00000312439.6 ENST00000403380.3 |
PSG1 |
pregnancy specific beta-1-glycoprotein 1 |
| chr1_+_111415757 | 1.08 |
ENST00000429072.2 ENST00000271324.5 |
CD53 |
CD53 molecule |
| chrX_-_118827333 | 1.02 |
ENST00000360156.7 ENST00000354228.4 ENST00000489216.1 ENST00000354416.3 ENST00000394610.1 ENST00000343984.5 |
SEPT6 |
septin 6 |
| chr14_-_106539557 | 1.00 |
ENST00000390599.2 |
IGHV1-8 |
immunoglobulin heavy variable 1-8 |
| chr19_+_42381337 | 1.00 |
ENST00000597454.1 ENST00000444740.2 |
CD79A |
CD79a molecule, immunoglobulin-associated alpha |
| chr14_-_96180435 | 0.99 |
ENST00000556450.1 ENST00000555202.1 ENST00000554012.1 ENST00000402399.1 |
TCL1A |
T-cell leukemia/lymphoma 1A |
| chr12_-_91574142 | 0.96 |
ENST00000547937.1 |
DCN |
decorin |
| chr19_-_43690674 | 0.93 |
ENST00000342951.6 ENST00000366175.3 |
PSG5 |
pregnancy specific beta-1-glycoprotein 5 |
| chrX_-_43741594 | 0.93 |
ENST00000536181.1 ENST00000378069.4 |
MAOB |
monoamine oxidase B |
| chrX_-_70329118 | 0.91 |
ENST00000374188.3 |
IL2RG |
interleukin 2 receptor, gamma |
| chr6_+_31620191 | 0.88 |
ENST00000375918.2 ENST00000375920.4 |
APOM |
apolipoprotein M |
| chr14_-_106471723 | 0.87 |
ENST00000390595.2 |
IGHV1-3 |
immunoglobulin heavy variable 1-3 |
| chr1_+_212738676 | 0.86 |
ENST00000366981.4 ENST00000366987.2 |
ATF3 |
activating transcription factor 3 |
| chr19_+_42381173 | 0.82 |
ENST00000221972.3 |
CD79A |
CD79a molecule, immunoglobulin-associated alpha |
| chr19_-_43383850 | 0.82 |
ENST00000436291.2 ENST00000595124.1 ENST00000244296.2 |
PSG1 |
pregnancy specific beta-1-glycoprotein 1 |
| chr1_+_225997791 | 0.81 |
ENST00000445856.1 ENST00000272167.5 |
EPHX1 |
epoxide hydrolase 1, microsomal (xenobiotic) |
| chr18_-_10701979 | 0.81 |
ENST00000538948.1 ENST00000285141.4 |
PIEZO2 |
piezo-type mechanosensitive ion channel component 2 |
| chr19_-_43709703 | 0.80 |
ENST00000599391.1 ENST00000244295.9 |
PSG4 |
pregnancy specific beta-1-glycoprotein 4 |
| chr19_-_43708378 | 0.79 |
ENST00000599746.1 |
PSG4 |
pregnancy specific beta-1-glycoprotein 4 |
| chr19_-_43709817 | 0.79 |
ENST00000433626.2 ENST00000405312.3 |
PSG4 |
pregnancy specific beta-1-glycoprotein 4 |
| chr14_-_106963409 | 0.78 |
ENST00000390621.2 |
IGHV1-45 |
immunoglobulin heavy variable 1-45 |
| chr15_+_75182346 | 0.78 |
ENST00000569931.1 ENST00000352410.4 ENST00000566377.1 ENST00000569233.1 ENST00000567132.1 ENST00000564633.1 ENST00000568907.1 ENST00000563422.1 ENST00000564003.1 ENST00000562800.1 ENST00000563786.1 ENST00000535694.1 ENST00000323744.6 ENST00000568828.1 ENST00000562606.1 ENST00000565576.1 ENST00000567570.1 |
MPI |
mannose phosphate isomerase |
| chr15_+_41057818 | 0.77 |
ENST00000558467.1 |
GCHFR |
GTP cyclohydrolase I feedback regulator |
| chr19_-_43690642 | 0.77 |
ENST00000407356.1 ENST00000407568.1 ENST00000404580.1 ENST00000599812.1 |
PSG5 |
pregnancy specific beta-1-glycoprotein 5 |
| chr5_-_138725594 | 0.75 |
ENST00000302125.8 |
MZB1 |
marginal zone B and B1 cell-specific protein |
| chr5_-_138725560 | 0.75 |
ENST00000412103.2 ENST00000457570.2 |
MZB1 |
marginal zone B and B1 cell-specific protein |
| chr17_-_26903900 | 0.75 |
ENST00000395319.3 ENST00000581807.1 ENST00000584086.1 ENST00000395321.2 |
ALDOC |
aldolase C, fructose-bisphosphate |
| chr22_+_23229960 | 0.74 |
ENST00000526893.1 ENST00000532223.2 ENST00000531372.1 |
IGLL5 |
immunoglobulin lambda-like polypeptide 5 |
| chr7_-_150038704 | 0.74 |
ENST00000466675.1 ENST00000482669.1 ENST00000467793.1 ENST00000223271.3 |
RARRES2 |
retinoic acid receptor responder (tazarotene induced) 2 |
| chr10_+_114135004 | 0.72 |
ENST00000393081.1 |
ACSL5 |
acyl-CoA synthetase long-chain family member 5 |
| chr11_-_78052923 | 0.72 |
ENST00000340149.2 |
GAB2 |
GRB2-associated binding protein 2 |
| chr19_-_43269809 | 0.72 |
ENST00000406636.3 ENST00000404209.4 ENST00000306511.4 |
PSG8 |
pregnancy specific beta-1-glycoprotein 8 |
| chr10_+_70480963 | 0.71 |
ENST00000265872.6 ENST00000535016.1 ENST00000538031.1 ENST00000543719.1 ENST00000539539.1 ENST00000543225.1 ENST00000536012.1 ENST00000494903.2 |
CCAR1 |
cell division cycle and apoptosis regulator 1 |
| chr19_-_43382142 | 0.71 |
ENST00000597058.1 |
PSG1 |
pregnancy specific beta-1-glycoprotein 1 |
| chr14_-_106733624 | 0.69 |
ENST00000390610.2 |
IGHV1-24 |
immunoglobulin heavy variable 1-24 |
| chr15_-_22448819 | 0.66 |
ENST00000604066.1 |
IGHV1OR15-1 |
immunoglobulin heavy variable 1/OR15-1 (non-functional) |
| chr7_+_43803790 | 0.65 |
ENST00000424330.1 |
BLVRA |
biliverdin reductase A |
| chr19_-_43709772 | 0.63 |
ENST00000596907.1 ENST00000451895.1 |
PSG4 |
pregnancy specific beta-1-glycoprotein 4 |
| chr2_-_175547571 | 0.62 |
ENST00000409415.3 ENST00000359761.3 ENST00000272746.5 |
WIPF1 |
WAS/WASL interacting protein family, member 1 |
| chr18_+_29171689 | 0.62 |
ENST00000237014.3 |
TTR |
transthyretin |
| chr8_-_27468842 | 0.62 |
ENST00000523500.1 |
CLU |
clusterin |
| chr7_+_94536898 | 0.61 |
ENST00000433360.1 ENST00000340694.4 ENST00000424654.1 |
PPP1R9A |
protein phosphatase 1, regulatory subunit 9A |
| chr2_+_201994208 | 0.60 |
ENST00000440180.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr1_-_184943610 | 0.60 |
ENST00000367511.3 |
FAM129A |
family with sequence similarity 129, member A |
| chr1_+_45212051 | 0.59 |
ENST00000372222.3 |
KIF2C |
kinesin family member 2C |
| chr1_+_45212074 | 0.59 |
ENST00000372217.1 |
KIF2C |
kinesin family member 2C |
| chr19_-_7293942 | 0.59 |
ENST00000341500.5 ENST00000302850.5 |
INSR |
insulin receptor |
| chr12_+_102091400 | 0.57 |
ENST00000229266.3 ENST00000549872.1 |
CHPT1 |
choline phosphotransferase 1 |
| chr2_+_68592305 | 0.57 |
ENST00000234313.7 |
PLEK |
pleckstrin |
| chr11_-_35287243 | 0.55 |
ENST00000464522.2 |
SLC1A2 |
solute carrier family 1 (glial high affinity glutamate transporter), member 2 |
| chr7_-_97501733 | 0.55 |
ENST00000444334.1 ENST00000422745.1 ENST00000394308.3 ENST00000451771.1 ENST00000175506.4 |
ASNS |
asparagine synthetase (glutamine-hydrolyzing) |
| chr6_-_160166218 | 0.55 |
ENST00000537657.1 |
SOD2 |
superoxide dismutase 2, mitochondrial |
| chr11_+_125496619 | 0.54 |
ENST00000532669.1 ENST00000278916.3 |
CHEK1 |
checkpoint kinase 1 |
| chr10_+_81107271 | 0.53 |
ENST00000448165.1 |
PPIF |
peptidylprolyl isomerase F |
| chr2_+_219575653 | 0.50 |
ENST00000442769.1 ENST00000424644.1 |
TTLL4 |
tubulin tyrosine ligase-like family, member 4 |
| chr6_-_43027105 | 0.50 |
ENST00000230413.5 ENST00000487429.1 ENST00000489623.1 ENST00000468957.1 |
MRPL2 |
mitochondrial ribosomal protein L2 |
| chr21_-_46330545 | 0.50 |
ENST00000320216.6 ENST00000397852.1 |
ITGB2 |
integrin, beta 2 (complement component 3 receptor 3 and 4 subunit) |
| chr16_+_30212378 | 0.49 |
ENST00000569485.1 |
SULT1A3 |
sulfotransferase family, cytosolic, 1A, phenol-preferring, member 3 |
| chr2_-_26467465 | 0.48 |
ENST00000457468.2 |
HADHA |
hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase/enoyl-CoA hydratase (trifunctional protein), alpha subunit |
| chr2_+_134877740 | 0.47 |
ENST00000409645.1 |
MGAT5 |
mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase |
| chr11_+_111957497 | 0.47 |
ENST00000375549.3 ENST00000528182.1 ENST00000528048.1 ENST00000528021.1 ENST00000526592.1 ENST00000525291.1 |
SDHD |
succinate dehydrogenase complex, subunit D, integral membrane protein |
| chr13_-_24463530 | 0.47 |
ENST00000382172.3 |
MIPEP |
mitochondrial intermediate peptidase |
| chrX_+_30233668 | 0.47 |
ENST00000378988.4 |
MAGEB2 |
melanoma antigen family B, 2 |
| chr3_+_73045936 | 0.47 |
ENST00000356692.5 ENST00000488810.1 ENST00000394284.3 ENST00000295862.9 ENST00000495566.1 |
PPP4R2 |
protein phosphatase 4, regulatory subunit 2 |
| chr19_-_4540486 | 0.46 |
ENST00000306390.6 |
LRG1 |
leucine-rich alpha-2-glycoprotein 1 |
| chr12_-_53594227 | 0.45 |
ENST00000550743.2 |
ITGB7 |
integrin, beta 7 |
| chr16_-_30122717 | 0.45 |
ENST00000566613.1 |
GDPD3 |
glycerophosphodiester phosphodiesterase domain containing 3 |
| chr19_-_6720686 | 0.45 |
ENST00000245907.6 |
C3 |
complement component 3 |
| chrX_+_30261847 | 0.44 |
ENST00000378981.3 ENST00000397550.1 |
MAGEB1 |
melanoma antigen family B, 1 |
| chr6_+_106534192 | 0.44 |
ENST00000369091.2 ENST00000369096.4 |
PRDM1 |
PR domain containing 1, with ZNF domain |
| chr12_-_7261772 | 0.43 |
ENST00000545280.1 ENST00000543933.1 ENST00000545337.1 ENST00000544702.1 ENST00000266542.4 |
C1RL |
complement component 1, r subcomponent-like |
| chr22_+_18593446 | 0.43 |
ENST00000316027.6 |
TUBA8 |
tubulin, alpha 8 |
| chr12_-_54691668 | 0.43 |
ENST00000553198.1 |
NFE2 |
nuclear factor, erythroid 2 |
| chr19_-_43586820 | 0.42 |
ENST00000406487.1 |
PSG2 |
pregnancy specific beta-1-glycoprotein 2 |
| chr3_+_151986709 | 0.42 |
ENST00000495875.2 ENST00000493459.1 ENST00000324210.5 ENST00000459747.1 |
MBNL1 |
muscleblind-like splicing regulator 1 |
| chr14_-_106642049 | 0.41 |
ENST00000390605.2 |
IGHV1-18 |
immunoglobulin heavy variable 1-18 |
| chrX_+_48644962 | 0.41 |
ENST00000376670.3 ENST00000376665.3 |
GATA1 |
GATA binding protein 1 (globin transcription factor 1) |
| chr5_-_130970723 | 0.41 |
ENST00000308008.6 ENST00000296859.6 ENST00000507093.1 ENST00000510071.1 ENST00000509018.1 ENST00000307984.5 |
RAPGEF6 |
Rap guanine nucleotide exchange factor (GEF) 6 |
| chr3_-_72496035 | 0.40 |
ENST00000477973.2 |
RYBP |
RING1 and YY1 binding protein |
| chrX_-_6453159 | 0.40 |
ENST00000381089.3 ENST00000398729.1 |
VCX3A |
variable charge, X-linked 3A |
| chr14_-_90085458 | 0.40 |
ENST00000345097.4 ENST00000555855.1 ENST00000555353.1 |
FOXN3 |
forkhead box N3 |
| chr10_+_16478942 | 0.39 |
ENST00000535784.2 ENST00000423462.2 ENST00000378000.1 |
PTER |
phosphotriesterase related |
| chr3_-_46249878 | 0.39 |
ENST00000296140.3 |
CCR1 |
chemokine (C-C motif) receptor 1 |
| chr3_+_46618727 | 0.39 |
ENST00000296145.5 |
TDGF1 |
teratocarcinoma-derived growth factor 1 |
| chr2_-_60780536 | 0.38 |
ENST00000538214.1 |
BCL11A |
B-cell CLL/lymphoma 11A (zinc finger protein) |
| chr1_+_156338993 | 0.38 |
ENST00000368249.1 ENST00000368246.2 ENST00000537040.1 ENST00000400992.2 ENST00000255013.3 ENST00000451864.2 |
RHBG |
Rh family, B glycoprotein (gene/pseudogene) |
| chr21_+_10862622 | 0.37 |
ENST00000302092.5 ENST00000559480.1 |
IGHV1OR21-1 |
immunoglobulin heavy variable 1/OR21-1 (non-functional) |
| chr10_+_11207438 | 0.36 |
ENST00000609692.1 ENST00000354897.3 |
CELF2 |
CUGBP, Elav-like family member 2 |
| chr3_-_194072019 | 0.36 |
ENST00000429275.1 ENST00000323830.3 |
CPN2 |
carboxypeptidase N, polypeptide 2 |
| chr19_+_15160130 | 0.36 |
ENST00000427043.3 |
CASP14 |
caspase 14, apoptosis-related cysteine peptidase |
| chr7_-_87104963 | 0.36 |
ENST00000359206.3 ENST00000358400.3 ENST00000265723.4 |
ABCB4 |
ATP-binding cassette, sub-family B (MDR/TAP), member 4 |
| chr11_+_125496400 | 0.35 |
ENST00000524737.1 |
CHEK1 |
checkpoint kinase 1 |
| chr12_+_53399942 | 0.35 |
ENST00000262056.9 |
EIF4B |
eukaryotic translation initiation factor 4B |
| chr17_-_36413133 | 0.35 |
ENST00000523089.1 ENST00000312412.4 ENST00000520237.1 |
RP11-1407O15.2 |
TBC1 domain family member 3 |
| chr6_+_25652432 | 0.34 |
ENST00000377961.2 |
SCGN |
secretagogin, EF-hand calcium binding protein |
| chr11_+_125496124 | 0.34 |
ENST00000533778.2 ENST00000534070.1 |
CHEK1 |
checkpoint kinase 1 |
| chr9_-_130679257 | 0.33 |
ENST00000361444.3 ENST00000335791.5 ENST00000343609.2 |
ST6GALNAC4 |
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 4 |
| chr2_-_26467557 | 0.33 |
ENST00000380649.3 |
HADHA |
hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase/enoyl-CoA hydratase (trifunctional protein), alpha subunit |
| chr16_+_20817761 | 0.33 |
ENST00000568046.1 ENST00000261377.6 |
AC004381.6 |
Putative RNA exonuclease NEF-sp |
| chr7_-_100844193 | 0.32 |
ENST00000440203.2 ENST00000379423.3 ENST00000223114.4 |
MOGAT3 |
monoacylglycerol O-acyltransferase 3 |
| chrX_+_8432871 | 0.32 |
ENST00000381032.1 ENST00000453306.1 ENST00000444481.1 |
VCX3B |
variable charge, X-linked 3B |
| chr2_-_60780607 | 0.31 |
ENST00000537768.1 ENST00000335712.6 ENST00000356842.4 |
BCL11A |
B-cell CLL/lymphoma 11A (zinc finger protein) |
| chr7_-_43769066 | 0.31 |
ENST00000223336.6 ENST00000310564.6 ENST00000431651.1 ENST00000415798.1 |
COA1 |
cytochrome c oxidase assembly factor 1 homolog (S. cerevisiae) |
| chr7_-_75115548 | 0.31 |
ENST00000453279.2 |
POM121C |
POM121 transmembrane nucleoporin C |
| chr19_-_43244610 | 0.31 |
ENST00000595140.1 ENST00000327495.5 |
PSG3 |
pregnancy specific beta-1-glycoprotein 3 |
| chr7_-_43769051 | 0.31 |
ENST00000395880.3 |
COA1 |
cytochrome c oxidase assembly factor 1 homolog (S. cerevisiae) |
| chr15_-_65477637 | 0.30 |
ENST00000300107.3 |
CLPX |
caseinolytic mitochondrial matrix peptidase chaperone subunit |
| chr13_+_33160553 | 0.30 |
ENST00000315596.10 |
PDS5B |
PDS5, regulator of cohesion maintenance, homolog B (S. cerevisiae) |
| chr11_-_45940343 | 0.30 |
ENST00000532681.1 |
PEX16 |
peroxisomal biogenesis factor 16 |
| chr1_+_214776516 | 0.30 |
ENST00000366955.3 |
CENPF |
centromere protein F, 350/400kDa |
| chr11_-_64511575 | 0.29 |
ENST00000431822.1 ENST00000377486.3 ENST00000394432.3 |
RASGRP2 |
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr5_+_157158205 | 0.29 |
ENST00000231198.7 |
THG1L |
tRNA-histidine guanylyltransferase 1-like (S. cerevisiae) |
| chr2_+_219575543 | 0.29 |
ENST00000457313.1 ENST00000415717.1 ENST00000392102.1 |
TTLL4 |
tubulin tyrosine ligase-like family, member 4 |
| chr14_-_106453155 | 0.29 |
ENST00000390594.2 |
IGHV1-2 |
immunoglobulin heavy variable 1-2 |
| chr11_-_104480019 | 0.28 |
ENST00000536529.1 ENST00000545630.1 ENST00000538641.1 |
RP11-886D15.1 |
RP11-886D15.1 |
| chr19_-_8373173 | 0.28 |
ENST00000537716.2 ENST00000301458.5 |
CD320 |
CD320 molecule |
| chr12_+_57623477 | 0.27 |
ENST00000557487.1 ENST00000555634.1 ENST00000556689.1 |
SHMT2 |
serine hydroxymethyltransferase 2 (mitochondrial) |
| chr19_-_43773589 | 0.27 |
ENST00000291752.5 ENST00000244293.7 ENST00000596730.1 ENST00000593948.1 ENST00000270077.3 ENST00000443718.3 ENST00000418820.2 |
PSG9 |
pregnancy specific beta-1-glycoprotein 9 |
| chr15_-_20170354 | 0.26 |
ENST00000338912.5 |
IGHV1OR15-9 |
immunoglobulin heavy variable 1/OR15-9 (non-functional) |
| chr19_-_59023348 | 0.25 |
ENST00000601355.1 ENST00000263093.2 |
SLC27A5 |
solute carrier family 27 (fatty acid transporter), member 5 |
| chrX_-_69479654 | 0.25 |
ENST00000374519.2 |
P2RY4 |
pyrimidinergic receptor P2Y, G-protein coupled, 4 |
| chr7_+_142985308 | 0.25 |
ENST00000310447.5 |
CASP2 |
caspase 2, apoptosis-related cysteine peptidase |
| chr16_+_30759700 | 0.25 |
ENST00000328273.7 |
PHKG2 |
phosphorylase kinase, gamma 2 (testis) |
| chrY_+_16636354 | 0.25 |
ENST00000339174.5 |
NLGN4Y |
neuroligin 4, Y-linked |
| chr2_+_26467762 | 0.24 |
ENST00000317799.5 ENST00000405867.3 ENST00000537713.1 |
HADHB |
hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase/enoyl-CoA hydratase (trifunctional protein), beta subunit |
| chr3_-_114790179 | 0.24 |
ENST00000462705.1 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chrX_-_8139308 | 0.23 |
ENST00000317103.4 |
VCX2 |
variable charge, X-linked 2 |
| chr1_-_153517473 | 0.23 |
ENST00000368715.1 |
S100A4 |
S100 calcium binding protein A4 |
| chr14_-_47120956 | 0.23 |
ENST00000298283.3 |
RPL10L |
ribosomal protein L10-like |
| chr13_+_100741269 | 0.23 |
ENST00000376286.4 ENST00000376279.3 ENST00000376285.1 |
PCCA |
propionyl CoA carboxylase, alpha polypeptide |
| chr2_-_25564750 | 0.22 |
ENST00000321117.5 |
DNMT3A |
DNA (cytosine-5-)-methyltransferase 3 alpha |
| chr5_-_86708833 | 0.22 |
ENST00000256897.4 |
CCNH |
cyclin H |
| chr6_-_25874440 | 0.22 |
ENST00000361703.6 ENST00000397060.4 |
SLC17A3 |
solute carrier family 17 (organic anion transporter), member 3 |
| chr19_-_43422019 | 0.21 |
ENST00000402603.4 ENST00000594375.1 |
PSG6 |
pregnancy specific beta-1-glycoprotein 6 |
| chr2_-_60780702 | 0.21 |
ENST00000359629.5 |
BCL11A |
B-cell CLL/lymphoma 11A (zinc finger protein) |
| chrX_-_138724677 | 0.21 |
ENST00000370573.4 ENST00000338585.6 ENST00000370576.4 |
MCF2 |
MCF.2 cell line derived transforming sequence |
| chr7_+_72349920 | 0.21 |
ENST00000395270.1 ENST00000446813.1 ENST00000257622.4 |
POM121 |
POM121 transmembrane nucleoporin |
| chr17_-_39093672 | 0.21 |
ENST00000209718.3 ENST00000436344.3 ENST00000485751.1 |
KRT23 |
keratin 23 (histone deacetylase inducible) |
| chr1_-_207224307 | 0.21 |
ENST00000315927.4 |
YOD1 |
YOD1 deubiquitinase |
| chr16_+_30759563 | 0.21 |
ENST00000563588.1 ENST00000565924.1 ENST00000424889.3 |
PHKG2 |
phosphorylase kinase, gamma 2 (testis) |
| chr1_+_29213584 | 0.21 |
ENST00000343067.4 ENST00000356093.2 ENST00000398863.2 ENST00000373800.3 ENST00000349460.4 |
EPB41 |
erythrocyte membrane protein band 4.1 (elliptocytosis 1, RH-linked) |
| chr7_+_102004322 | 0.21 |
ENST00000496391.1 |
PRKRIP1 |
PRKR interacting protein 1 (IL11 inducible) |
| chr8_-_72268968 | 0.20 |
ENST00000388740.3 |
EYA1 |
eyes absent homolog 1 (Drosophila) |
| chr20_-_35402123 | 0.20 |
ENST00000373740.3 ENST00000426836.1 ENST00000373745.3 ENST00000448110.2 ENST00000438549.1 ENST00000447406.1 ENST00000373750.4 ENST00000373734.4 |
DSN1 |
DSN1, MIS12 kinetochore complex component |
| chr5_-_140027175 | 0.19 |
ENST00000512088.1 |
NDUFA2 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 2, 8kDa |
| chr10_+_114710516 | 0.19 |
ENST00000542695.1 ENST00000346198.4 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr19_+_45542773 | 0.19 |
ENST00000544944.2 |
CLASRP |
CLK4-associating serine/arginine rich protein |
| chr5_-_86708670 | 0.19 |
ENST00000504878.1 |
CCNH |
cyclin H |
| chr6_-_31620149 | 0.19 |
ENST00000435080.1 ENST00000375976.4 ENST00000441054.1 |
BAG6 |
BCL2-associated athanogene 6 |
| chr9_+_131447342 | 0.19 |
ENST00000409104.3 |
SET |
SET nuclear oncogene |
| chr11_-_64512273 | 0.18 |
ENST00000377497.3 ENST00000377487.1 ENST00000430645.1 |
RASGRP2 |
RAS guanyl releasing protein 2 (calcium and DAG-regulated) |
| chr2_+_26467825 | 0.18 |
ENST00000545822.1 ENST00000425035.1 |
HADHB |
hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase/enoyl-CoA hydratase (trifunctional protein), beta subunit |
| chr17_+_55173933 | 0.18 |
ENST00000539273.1 |
AKAP1 |
A kinase (PRKA) anchor protein 1 |
| chr19_+_41117770 | 0.18 |
ENST00000601032.1 |
LTBP4 |
latent transforming growth factor beta binding protein 4 |
| chr18_-_74728998 | 0.18 |
ENST00000359645.3 ENST00000397875.3 ENST00000397869.3 ENST00000578193.1 ENST00000578873.1 ENST00000397866.4 ENST00000528160.1 ENST00000527041.1 ENST00000526111.1 ENST00000397865.5 ENST00000382582.3 |
MBP |
myelin basic protein |
| chr16_-_18801582 | 0.18 |
ENST00000565420.1 |
RPS15A |
ribosomal protein S15a |
| chr16_-_420514 | 0.17 |
ENST00000199706.8 |
MRPL28 |
mitochondrial ribosomal protein L28 |
| chr6_+_25652501 | 0.17 |
ENST00000334979.6 |
SCGN |
secretagogin, EF-hand calcium binding protein |
| chr5_-_140027357 | 0.17 |
ENST00000252102.4 |
NDUFA2 |
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 2, 8kDa |
| chr16_+_31366536 | 0.17 |
ENST00000562522.1 |
ITGAX |
integrin, alpha X (complement component 3 receptor 4 subunit) |
| chr22_+_40390930 | 0.17 |
ENST00000333407.6 |
FAM83F |
family with sequence similarity 83, member F |
| chr20_+_3052264 | 0.16 |
ENST00000217386.2 |
OXT |
oxytocin/neurophysin I prepropeptide |
| chr11_+_65407331 | 0.16 |
ENST00000527525.1 |
SIPA1 |
signal-induced proliferation-associated 1 |
| chr12_+_122242597 | 0.16 |
ENST00000267197.5 |
SETD1B |
SET domain containing 1B |
| chr1_+_6684918 | 0.16 |
ENST00000054650.4 |
THAP3 |
THAP domain containing, apoptosis associated protein 3 |
| chr2_+_234296792 | 0.16 |
ENST00000409813.3 |
DGKD |
diacylglycerol kinase, delta 130kDa |
| chr10_+_47894023 | 0.16 |
ENST00000358474.5 |
FAM21B |
family with sequence similarity 21, member B |
| chr19_+_10197463 | 0.16 |
ENST00000590378.1 ENST00000397881.3 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr17_+_76210267 | 0.15 |
ENST00000301633.4 ENST00000350051.3 ENST00000374948.2 ENST00000590449.1 |
BIRC5 |
baculoviral IAP repeat containing 5 |
| chr6_-_42690312 | 0.15 |
ENST00000230381.5 |
PRPH2 |
peripherin 2 (retinal degeneration, slow) |
| chr2_-_175711133 | 0.15 |
ENST00000409597.1 ENST00000413882.1 |
CHN1 |
chimerin 1 |
| chr22_+_45680822 | 0.15 |
ENST00000216211.4 ENST00000396082.2 |
UPK3A |
uroplakin 3A |
| chr4_+_159593271 | 0.15 |
ENST00000512251.1 ENST00000511912.1 |
ETFDH |
electron-transferring-flavoprotein dehydrogenase |
| chr3_+_51976338 | 0.14 |
ENST00000417220.2 ENST00000431474.1 ENST00000398755.3 |
PARP3 |
poly (ADP-ribose) polymerase family, member 3 |
| chr15_+_64752927 | 0.14 |
ENST00000416172.1 |
ZNF609 |
zinc finger protein 609 |
| chr16_+_31128978 | 0.14 |
ENST00000448516.2 ENST00000219797.4 |
KAT8 |
K(lysine) acetyltransferase 8 |
| chr12_+_69186125 | 0.14 |
ENST00000399333.3 |
AC124890.1 |
HCG1774533, isoform CRA_a; PRO2268; Uncharacterized protein |
| chr2_+_24163281 | 0.13 |
ENST00000309033.4 |
UBXN2A |
UBX domain protein 2A |
| chr3_+_180630090 | 0.13 |
ENST00000357559.4 ENST00000305586.7 |
FXR1 |
fragile X mental retardation, autosomal homolog 1 |
| chr17_-_40729681 | 0.13 |
ENST00000590760.1 ENST00000587209.1 ENST00000393795.3 ENST00000253789.5 |
PSMC3IP |
PSMC3 interacting protein |
| chrX_+_22050546 | 0.13 |
ENST00000379374.4 |
PHEX |
phosphate regulating endopeptidase homolog, X-linked |
| chr13_+_98086445 | 0.13 |
ENST00000245304.4 |
RAP2A |
RAP2A, member of RAS oncogene family |
| chr11_-_68518910 | 0.13 |
ENST00000544963.1 ENST00000443940.2 |
MTL5 |
metallothionein-like 5, testis-specific (tesmin) |
| chr2_-_176046391 | 0.13 |
ENST00000392541.3 ENST00000409194.1 |
ATP5G3 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C3 (subunit 9) |
| chr19_-_49956728 | 0.13 |
ENST00000601825.1 ENST00000596049.1 ENST00000599366.1 ENST00000597415.1 |
PIH1D1 |
PIH1 domain containing 1 |
| chr16_-_5147743 | 0.12 |
ENST00000587133.1 ENST00000458008.4 ENST00000427587.4 |
FAM86A |
family with sequence similarity 86, member A |
| chr3_+_113465866 | 0.12 |
ENST00000273398.3 ENST00000538620.1 ENST00000496747.1 ENST00000475322.1 |
ATP6V1A |
ATPase, H+ transporting, lysosomal 70kDa, V1 subunit A |
| chr7_-_55640176 | 0.12 |
ENST00000285279.5 |
VOPP1 |
vesicular, overexpressed in cancer, prosurvival protein 1 |
| chr16_-_12897642 | 0.12 |
ENST00000433677.2 ENST00000261660.4 ENST00000381774.4 |
CPPED1 |
calcineurin-like phosphoesterase domain containing 1 |
| chr17_-_37308824 | 0.12 |
ENST00000415163.1 ENST00000441877.1 ENST00000444911.2 |
PLXDC1 |
plexin domain containing 1 |
| chr17_+_7452189 | 0.12 |
ENST00000293825.6 |
TNFSF12 |
tumor necrosis factor (ligand) superfamily, member 12 |
| chr22_+_30163340 | 0.12 |
ENST00000330029.6 ENST00000401406.3 |
UQCR10 |
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr6_-_31763721 | 0.12 |
ENST00000375663.3 |
VARS |
valyl-tRNA synthetase |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.8 | GO:1902847 | macrophage proliferation(GO:0061517) microglial cell proliferation(GO:0061518) regulation of neuronal signal transduction(GO:1902847) positive regulation of neurofibrillary tangle assembly(GO:1902998) |
| 0.3 | 0.9 | GO:1903860 | negative regulation of dendrite extension(GO:1903860) regulation of neuron remodeling(GO:1904799) negative regulation of neuron remodeling(GO:1904800) negative regulation of branching morphogenesis of a nerve(GO:2000173) |
| 0.3 | 1.5 | GO:0002378 | immunoglobulin biosynthetic process(GO:0002378) negative regulation of glucose import in response to insulin stimulus(GO:2001274) |
| 0.3 | 0.9 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.3 | 1.3 | GO:1902896 | terminal web assembly(GO:1902896) |
| 0.3 | 0.8 | GO:0043105 | regulation of GTP cyclohydrolase I activity(GO:0043095) negative regulation of GTP cyclohydrolase I activity(GO:0043105) |
| 0.2 | 0.7 | GO:0061760 | antifungal innate immune response(GO:0061760) |
| 0.2 | 0.9 | GO:0034444 | regulation of plasma lipoprotein particle oxidation(GO:0034444) negative regulation of plasma lipoprotein particle oxidation(GO:0034445) |
| 0.2 | 1.2 | GO:0035965 | cardiolipin acyl-chain remodeling(GO:0035965) |
| 0.2 | 1.2 | GO:0035407 | histone H3-T11 phosphorylation(GO:0035407) |
| 0.2 | 0.8 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 0.2 | 0.6 | GO:0006663 | platelet activating factor biosynthetic process(GO:0006663) |
| 0.2 | 0.4 | GO:1905232 | cellular response to L-glutamate(GO:1905232) |
| 0.2 | 0.5 | GO:1902445 | regulation of mitochondrial membrane permeability involved in programmed necrotic cell death(GO:1902445) negative regulation of oxidative phosphorylation uncoupler activity(GO:2000276) |
| 0.2 | 0.5 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.2 | 0.6 | GO:1904048 | regulation of spontaneous neurotransmitter secretion(GO:1904048) |
| 0.1 | 0.4 | GO:0001798 | positive regulation of type IIa hypersensitivity(GO:0001798) positive regulation of type II hypersensitivity(GO:0002894) |
| 0.1 | 0.6 | GO:1990535 | transformation of host cell by virus(GO:0019087) neuron projection maintenance(GO:1990535) |
| 0.1 | 0.5 | GO:0070981 | L-asparagine biosynthetic process(GO:0070981) L-asparagine metabolic process(GO:0070982) |
| 0.1 | 0.4 | GO:0010725 | regulation of primitive erythrocyte differentiation(GO:0010725) eosinophil fate commitment(GO:0035854) |
| 0.1 | 0.9 | GO:0010269 | response to selenium ion(GO:0010269) |
| 0.1 | 0.8 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.1 | 0.9 | GO:0038110 | interleukin-2-mediated signaling pathway(GO:0038110) |
| 0.1 | 0.4 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.1 | 1.2 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.1 | 0.5 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.1 | 0.5 | GO:0003366 | cell-matrix adhesion involved in ameboidal cell migration(GO:0003366) |
| 0.1 | 1.7 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.1 | 0.5 | GO:0007571 | age-dependent response to oxidative stress(GO:0001306) age-dependent response to reactive oxygen species(GO:0001315) regulation of systemic arterial blood pressure by acetylcholine(GO:0003068) vasodilation by acetylcholine involved in regulation of systemic arterial blood pressure(GO:0003069) regulation of systemic arterial blood pressure by neurotransmitter(GO:0003070) age-dependent general metabolic decline(GO:0007571) |
| 0.1 | 0.3 | GO:0032581 | ER-dependent peroxisome organization(GO:0032581) |
| 0.1 | 1.0 | GO:0010918 | positive regulation of mitochondrial membrane potential(GO:0010918) |
| 0.1 | 0.4 | GO:1904379 | maintenance of unfolded protein(GO:0036506) protein localization to cytosolic proteasome complex(GO:1904327) maintenance of unfolded protein involved in ERAD pathway(GO:1904378) protein localization to cytosolic proteasome complex involved in ERAD pathway(GO:1904379) |
| 0.1 | 0.6 | GO:0070560 | protein secretion by platelet(GO:0070560) |
| 0.1 | 0.4 | GO:0097010 | eukaryotic translation initiation factor 4F complex assembly(GO:0097010) |
| 0.1 | 0.4 | GO:0033078 | extrathymic T cell differentiation(GO:0033078) |
| 0.1 | 1.0 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.1 | 0.1 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.1 | 0.5 | GO:0045819 | positive regulation of glycogen catabolic process(GO:0045819) |
| 0.1 | 0.2 | GO:0019626 | short-chain fatty acid catabolic process(GO:0019626) |
| 0.1 | 0.2 | GO:0035523 | protein K29-linked deubiquitination(GO:0035523) protein K33-linked deubiquitination(GO:1990168) |
| 0.1 | 0.5 | GO:0098989 | NMDA selective glutamate receptor signaling pathway(GO:0098989) |
| 0.1 | 0.6 | GO:0070779 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.1 | 4.5 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.1 | 0.2 | GO:0048203 | vesicle targeting, trans-Golgi to endosome(GO:0048203) |
| 0.1 | 0.2 | GO:0015911 | plasma membrane long-chain fatty acid transport(GO:0015911) |
| 0.1 | 0.6 | GO:1903944 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 0.2 | GO:0002125 | maternal aggressive behavior(GO:0002125) positive regulation of female receptivity(GO:0045925) |
| 0.1 | 0.7 | GO:0033008 | positive regulation of mast cell activation involved in immune response(GO:0033008) positive regulation of mast cell degranulation(GO:0043306) |
| 0.1 | 0.2 | GO:0060041 | retina development in camera-type eye(GO:0060041) |
| 0.0 | 0.6 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.0 | 0.1 | GO:0051106 | positive regulation of DNA ligation(GO:0051106) |
| 0.0 | 0.6 | GO:0097034 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.0 | 1.1 | GO:1901741 | positive regulation of myoblast fusion(GO:1901741) |
| 0.0 | 0.1 | GO:1903939 | negative regulation of histone H3-K9 dimethylation(GO:1900110) regulation of TORC2 signaling(GO:1903939) |
| 0.0 | 0.7 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.0 | 0.4 | GO:1901409 | positive regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901409) |
| 0.0 | 0.2 | GO:0042631 | cellular response to water deprivation(GO:0042631) |
| 0.0 | 0.3 | GO:0019264 | glycine biosynthetic process from serine(GO:0019264) |
| 0.0 | 0.1 | GO:0006438 | valyl-tRNA aminoacylation(GO:0006438) |
| 0.0 | 0.2 | GO:0090116 | C-5 methylation of cytosine(GO:0090116) |
| 0.0 | 0.1 | GO:0032237 | activation of store-operated calcium channel activity(GO:0032237) positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.0 | 0.2 | GO:1904207 | regulation of chemokine (C-C motif) ligand 2 secretion(GO:1904207) positive regulation of chemokine (C-C motif) ligand 2 secretion(GO:1904209) |
| 0.0 | 0.2 | GO:0035234 | luteolysis(GO:0001554) ectopic germ cell programmed cell death(GO:0035234) |
| 0.0 | 0.8 | GO:0050974 | detection of mechanical stimulus involved in sensory perception(GO:0050974) |
| 0.0 | 0.3 | GO:0072513 | positive regulation of secondary heart field cardioblast proliferation(GO:0072513) |
| 0.0 | 0.2 | GO:0045144 | meiotic sister chromatid segregation(GO:0045144) |
| 0.0 | 0.1 | GO:1990418 | response to insulin-like growth factor stimulus(GO:1990418) |
| 0.0 | 0.2 | GO:0015712 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.0 | 0.7 | GO:0035338 | long-chain fatty-acyl-CoA biosynthetic process(GO:0035338) |
| 0.0 | 7.9 | GO:0007565 | female pregnancy(GO:0007565) |
| 0.0 | 0.3 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.0 | 0.2 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 0.0 | 0.1 | GO:0042377 | menaquinone catabolic process(GO:0042361) vitamin K catabolic process(GO:0042377) |
| 0.0 | 0.6 | GO:0006337 | nucleosome disassembly(GO:0006337) |
| 0.0 | 1.8 | GO:0050853 | B cell receptor signaling pathway(GO:0050853) |
| 0.0 | 0.6 | GO:0010569 | regulation of double-strand break repair via homologous recombination(GO:0010569) |
| 0.0 | 0.1 | GO:0002457 | T cell antigen processing and presentation(GO:0002457) |
| 0.0 | 0.4 | GO:0007351 | tripartite regional subdivision(GO:0007351) anterior/posterior axis specification, embryo(GO:0008595) |
| 0.0 | 0.2 | GO:0060157 | urinary bladder development(GO:0060157) |
| 0.0 | 0.5 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.0 | 0.4 | GO:0097094 | craniofacial suture morphogenesis(GO:0097094) |
| 0.0 | 0.4 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 0.2 | GO:0048625 | myoblast fate commitment(GO:0048625) |
| 0.0 | 0.4 | GO:0090026 | positive regulation of monocyte chemotaxis(GO:0090026) |
| 0.0 | 0.1 | GO:1900194 | negative regulation of oocyte maturation(GO:1900194) |
| 0.0 | 0.3 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.0 | 0.4 | GO:0035518 | histone H2A monoubiquitination(GO:0035518) |
| 0.0 | 0.1 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.0 | 0.2 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.0 | 0.3 | GO:0006071 | glycerol metabolic process(GO:0006071) |
| 0.0 | 1.2 | GO:0006301 | postreplication repair(GO:0006301) |
| 0.0 | 0.5 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway(GO:0030511) positive regulation of cellular response to transforming growth factor beta stimulus(GO:1903846) |
| 0.0 | 0.2 | GO:1904776 | regulation of protein localization to cell cortex(GO:1904776) positive regulation of protein localization to cell cortex(GO:1904778) |
| 0.0 | 0.0 | GO:0002541 | activation of plasma proteins involved in acute inflammatory response(GO:0002541) |
| 0.0 | 0.1 | GO:0052199 | negative regulation of catalytic activity in other organism involved in symbiotic interaction(GO:0052199) |
| 0.0 | 0.3 | GO:0097503 | sialylation(GO:0097503) |
| 0.0 | 0.1 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.0 | 0.5 | GO:0006409 | tRNA export from nucleus(GO:0006409) tRNA-containing ribonucleoprotein complex export from nucleus(GO:0071431) |
| 0.0 | 0.8 | GO:0070126 | mitochondrial translational elongation(GO:0070125) mitochondrial translational termination(GO:0070126) |
| 0.0 | 0.2 | GO:1990001 | inhibition of cysteine-type endopeptidase activity involved in apoptotic process(GO:1990001) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.0 | 2.5 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 1.6 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.0 | 0.6 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 1.2 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 0.5 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 0.9 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.0 | 0.2 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 1.0 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.7 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 2.0 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.5 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.4 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 0.4 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 0.4 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 0.6 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0004913 | interleukin-4 receptor activity(GO:0004913) interleukin-7 receptor activity(GO:0004917) |
| 0.3 | 0.8 | GO:0044549 | GTP cyclohydrolase binding(GO:0044549) |
| 0.2 | 1.2 | GO:0016509 | long-chain-3-hydroxyacyl-CoA dehydrogenase activity(GO:0016509) |
| 0.2 | 1.2 | GO:0035402 | histone kinase activity (H3-T11 specific)(GO:0035402) |
| 0.2 | 0.7 | GO:0004074 | biliverdin reductase activity(GO:0004074) |
| 0.2 | 0.8 | GO:0070740 | tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.2 | 0.5 | GO:0008177 | succinate dehydrogenase (ubiquinone) activity(GO:0008177) |
| 0.1 | 0.6 | GO:0004142 | diacylglycerol cholinephosphotransferase activity(GO:0004142) |
| 0.1 | 0.4 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.1 | 0.8 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.1 | 1.3 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.1 | 1.7 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.1 | 1.2 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.1 | 0.8 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.1 | 2.3 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.1 | 0.3 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.1 | 0.5 | GO:0004066 | asparagine synthase (glutamine-hydrolyzing) activity(GO:0004066) |
| 0.1 | 0.3 | GO:0003846 | 2-acylglycerol O-acyltransferase activity(GO:0003846) |
| 0.1 | 0.8 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.1 | 0.3 | GO:0008192 | RNA guanylyltransferase activity(GO:0008192) |
| 0.1 | 0.3 | GO:0015235 | cobalamin transporter activity(GO:0015235) |
| 0.1 | 0.4 | GO:0034057 | RNA strand-exchange activity(GO:0034057) |
| 0.1 | 4.5 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 0.1 | 0.5 | GO:0047685 | amine sulfotransferase activity(GO:0047685) |
| 0.1 | 0.6 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.1 | 0.6 | GO:0043559 | insulin binding(GO:0043559) |
| 0.1 | 0.4 | GO:0090554 | phosphatidylcholine-translocating ATPase activity(GO:0090554) |
| 0.1 | 0.8 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.1 | 0.5 | GO:0030369 | ICAM-3 receptor activity(GO:0030369) |
| 0.1 | 0.2 | GO:1904455 | ubiquitin-specific protease activity involved in negative regulation of ERAD pathway(GO:1904455) |
| 0.1 | 0.7 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.1 | 0.5 | GO:0016721 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.1 | 1.3 | GO:0043325 | phosphatidylinositol-3,4-bisphosphate binding(GO:0043325) |
| 0.0 | 0.7 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 1.2 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.3 | GO:0008732 | glycine hydroxymethyltransferase activity(GO:0004372) threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.0 | 0.1 | GO:0004832 | valine-tRNA ligase activity(GO:0004832) |
| 0.0 | 0.5 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.3 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.0 | 0.2 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.0 | 0.6 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 0.1 | GO:0048039 | ubiquinone binding(GO:0048039) |
| 0.0 | 0.2 | GO:0004467 | long-chain fatty acid-CoA ligase activity(GO:0004467) very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.2 | GO:0071553 | uridine nucleotide receptor activity(GO:0015065) G-protein coupled pyrimidinergic nucleotide receptor activity(GO:0071553) |
| 0.0 | 0.6 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.4 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.0 | 0.2 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.0 | 0.2 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.0 | 1.0 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.1 | GO:0019961 | interferon binding(GO:0019961) |
| 0.0 | 0.4 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.0 | 0.2 | GO:0009374 | biotin carboxylase activity(GO:0004075) biotin binding(GO:0009374) |
| 0.0 | 0.1 | GO:0032138 | single base insertion or deletion binding(GO:0032138) |
| 0.0 | 0.6 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 0.1 | GO:0033592 | RNA strand annealing activity(GO:0033592) |
| 0.0 | 0.1 | GO:0001165 | RNA polymerase I upstream control element sequence-specific DNA binding(GO:0001165) |
| 0.0 | 0.5 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.4 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.0 | 0.2 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 0.1 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.0 | 0.1 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.1 | GO:0017060 | 3-galactosyl-N-acetylglucosaminide 4-alpha-L-fucosyltransferase activity(GO:0017060) |
| 0.0 | 2.7 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.1 | GO:0032453 | histone demethylase activity (H3-K4 specific)(GO:0032453) |
| 0.0 | 0.1 | GO:0003910 | DNA ligase (ATP) activity(GO:0003910) |
| 0.0 | 0.2 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.4 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.0 | 0.1 | GO:0005497 | androgen binding(GO:0005497) |
| 0.0 | 0.8 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.4 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.0 | 0.3 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.0 | 0.2 | GO:0050786 | RAGE receptor binding(GO:0050786) |
| 0.0 | 0.1 | GO:0043995 | histone acetyltransferase activity (H4-K5 specific)(GO:0043995) histone acetyltransferase activity (H4-K8 specific)(GO:0043996) histone acetyltransferase activity (H4-K16 specific)(GO:0046972) |
| 0.0 | 0.2 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 1.6 | GO:0047485 | protein N-terminus binding(GO:0047485) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0034365 | discoidal high-density lipoprotein particle(GO:0034365) |
| 0.3 | 0.9 | GO:1990622 | CHOP-ATF3 complex(GO:1990622) |
| 0.3 | 1.8 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.2 | 0.5 | GO:0030289 | protein phosphatase 4 complex(GO:0030289) |
| 0.1 | 1.8 | GO:0034366 | spherical high-density lipoprotein particle(GO:0034366) |
| 0.1 | 1.7 | GO:0005833 | hemoglobin complex(GO:0005833) |
| 0.1 | 0.6 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.1 | 0.5 | GO:0045283 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.1 | 0.5 | GO:0034669 | integrin alpha4-beta7 complex(GO:0034669) |
| 0.1 | 0.4 | GO:0070985 | TFIIK complex(GO:0070985) |
| 0.1 | 0.5 | GO:0034688 | integrin alphaM-beta2 complex(GO:0034688) |
| 0.1 | 0.4 | GO:0071818 | BAT3 complex(GO:0071818) ER membrane insertion complex(GO:0072379) |
| 0.1 | 4.5 | GO:0042571 | immunoglobulin complex(GO:0019814) immunoglobulin complex, circulating(GO:0042571) |
| 0.1 | 0.6 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.1 | 1.0 | GO:0097227 | sperm annulus(GO:0097227) |
| 0.1 | 1.0 | GO:0098647 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.1 | 1.3 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.1 | 0.2 | GO:0000818 | nuclear MIS12/MIND complex(GO:0000818) condensed chromosome inner kinetochore(GO:0000939) |
| 0.1 | 0.3 | GO:0009368 | endopeptidase Clp complex(GO:0009368) |
| 0.1 | 0.4 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.5 | GO:0044615 | nuclear pore nuclear basket(GO:0044615) |
| 0.0 | 0.3 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.0 | 0.7 | GO:0031089 | platelet dense granule lumen(GO:0031089) |
| 0.0 | 1.2 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.0 | 0.3 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 0.6 | GO:0097342 | ripoptosome(GO:0097342) |
| 0.0 | 0.6 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.0 | 0.2 | GO:0033269 | internode region of axon(GO:0033269) |
| 0.0 | 0.2 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.0 | 1.2 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.1 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.1 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.0 | 0.6 | GO:0030673 | axolemma(GO:0030673) |
| 0.0 | 1.2 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 0.1 | GO:0072487 | MSL complex(GO:0072487) |
| 0.0 | 0.8 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.2 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 0.0 | 0.1 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.0 | 0.5 | GO:0031305 | intrinsic component of mitochondrial inner membrane(GO:0031304) integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.4 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.0 | 0.1 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 0.0 | 0.2 | GO:0001741 | XY body(GO:0001741) |
| 0.0 | 0.1 | GO:0000836 | Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.0 | 1.2 | GO:0000794 | condensed nuclear chromosome(GO:0000794) |
| 0.0 | 0.2 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.3 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 1.2 | GO:0035578 | azurophil granule lumen(GO:0035578) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 1.2 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.0 | 0.6 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.0 | 0.8 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 0.8 | REACTOME TETRAHYDROBIOPTERIN BH4 SYNTHESIS RECYCLING SALVAGE AND REGULATION | Genes involved in Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation |
| 0.0 | 1.2 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 1.4 | REACTOME ACTIVATION OF GENES BY ATF4 | Genes involved in Activation of Genes by ATF4 |
| 0.0 | 0.9 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 1.9 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 0.7 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 1.0 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.8 | REACTOME IL RECEPTOR SHC SIGNALING | Genes involved in Interleukin receptor SHC signaling |
| 0.0 | 2.7 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.2 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.0 | 0.4 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.0 | 0.6 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.5 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.4 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.7 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 0.4 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.6 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 2.2 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 0.2 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.4 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.2 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 0.3 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |