Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
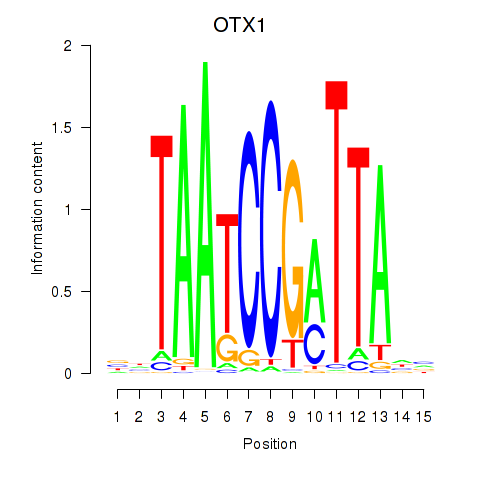
Results for OTX1
Z-value: 0.49
Transcription factors associated with OTX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
OTX1
|
ENSG00000115507.5 | OTX1 |
Activity profile of OTX1 motif
Sorted Z-values of OTX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of OTX1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_92351769 | 1.36 |
ENST00000212355.4 |
TGFBR3 |
transforming growth factor, beta receptor III |
| chr12_-_117537240 | 0.60 |
ENST00000392545.4 ENST00000541210.1 ENST00000335209.7 |
TESC |
tescalcin |
| chr15_-_61521495 | 0.56 |
ENST00000335670.6 |
RORA |
RAR-related orphan receptor A |
| chr2_-_160654745 | 0.53 |
ENST00000259053.4 ENST00000429078.2 |
CD302 |
CD302 molecule |
| chr10_+_114133773 | 0.48 |
ENST00000354655.4 |
ACSL5 |
acyl-CoA synthetase long-chain family member 5 |
| chr1_-_247335269 | 0.46 |
ENST00000543802.2 ENST00000491356.1 ENST00000472531.1 ENST00000340684.6 |
ZNF124 |
zinc finger protein 124 |
| chr16_+_33605231 | 0.43 |
ENST00000570121.2 |
IGHV3OR16-12 |
immunoglobulin heavy variable 3/OR16-12 (non-functional) |
| chr12_-_54071181 | 0.42 |
ENST00000338662.5 |
ATP5G2 |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C2 (subunit 9) |
| chr6_+_26251835 | 0.42 |
ENST00000356350.2 |
HIST1H2BH |
histone cluster 1, H2bh |
| chr20_+_48884002 | 0.40 |
ENST00000425497.1 ENST00000445003.1 |
RP11-290F20.3 |
RP11-290F20.3 |
| chr11_-_78052923 | 0.37 |
ENST00000340149.2 |
GAB2 |
GRB2-associated binding protein 2 |
| chr1_+_28764653 | 0.35 |
ENST00000373836.3 |
PHACTR4 |
phosphatase and actin regulator 4 |
| chr1_-_146696901 | 0.35 |
ENST00000369272.3 ENST00000441068.2 |
FMO5 |
flavin containing monooxygenase 5 |
| chr21_-_43816052 | 0.34 |
ENST00000398405.1 |
TMPRSS3 |
transmembrane protease, serine 3 |
| chr7_-_100239132 | 0.34 |
ENST00000223051.3 ENST00000431692.1 |
TFR2 |
transferrin receptor 2 |
| chr7_+_120702819 | 0.34 |
ENST00000423795.1 |
CPED1 |
cadherin-like and PC-esterase domain containing 1 |
| chr3_-_141747950 | 0.33 |
ENST00000497579.1 |
TFDP2 |
transcription factor Dp-2 (E2F dimerization partner 2) |
| chr12_+_6603253 | 0.33 |
ENST00000382457.4 ENST00000545962.1 |
NCAPD2 |
non-SMC condensin I complex, subunit D2 |
| chr15_+_58702742 | 0.30 |
ENST00000356113.6 ENST00000414170.3 |
LIPC |
lipase, hepatic |
| chr11_+_128563652 | 0.30 |
ENST00000527786.2 |
FLI1 |
Fli-1 proto-oncogene, ETS transcription factor |
| chr6_+_26158343 | 0.29 |
ENST00000377777.4 ENST00000289316.2 |
HIST1H2BD |
histone cluster 1, H2bd |
| chr19_+_17448348 | 0.29 |
ENST00000324894.8 ENST00000358792.7 ENST00000600625.1 |
GTPBP3 |
GTP binding protein 3 (mitochondrial) |
| chr6_+_106546808 | 0.29 |
ENST00000369089.3 |
PRDM1 |
PR domain containing 1, with ZNF domain |
| chr12_+_124997766 | 0.26 |
ENST00000543970.1 |
RP11-83B20.1 |
RP11-83B20.1 |
| chr19_+_10381769 | 0.26 |
ENST00000423829.2 ENST00000588645.1 |
ICAM1 |
intercellular adhesion molecule 1 |
| chr12_+_49740700 | 0.25 |
ENST00000549441.2 ENST00000395069.3 |
DNAJC22 |
DnaJ (Hsp40) homolog, subfamily C, member 22 |
| chr6_+_96025341 | 0.25 |
ENST00000369293.1 ENST00000358812.4 |
MANEA |
mannosidase, endo-alpha |
| chr4_-_122744998 | 0.24 |
ENST00000274026.5 |
CCNA2 |
cyclin A2 |
| chr9_-_35665165 | 0.24 |
ENST00000343259.3 ENST00000378387.3 |
ARHGEF39 |
Rho guanine nucleotide exchange factor (GEF) 39 |
| chr5_-_111091948 | 0.22 |
ENST00000447165.2 |
NREP |
neuronal regeneration related protein |
| chr17_-_34329084 | 0.21 |
ENST00000354059.4 ENST00000536149.1 |
CCL15 CCL14 |
chemokine (C-C motif) ligand 15 chemokine (C-C motif) ligand 14 |
| chr22_-_36903101 | 0.21 |
ENST00000397224.4 |
FOXRED2 |
FAD-dependent oxidoreductase domain containing 2 |
| chr15_+_42787452 | 0.20 |
ENST00000249647.3 |
SNAP23 |
synaptosomal-associated protein, 23kDa |
| chr19_-_41903161 | 0.20 |
ENST00000602129.1 ENST00000593771.1 ENST00000596905.1 ENST00000221233.4 |
EXOSC5 |
exosome component 5 |
| chr2_-_74619152 | 0.20 |
ENST00000440727.1 ENST00000409240.1 |
DCTN1 |
dynactin 1 |
| chr2_-_112642267 | 0.20 |
ENST00000341068.3 |
ANAPC1 |
anaphase promoting complex subunit 1 |
| chr8_-_79717750 | 0.19 |
ENST00000263851.4 ENST00000379113.2 |
IL7 |
interleukin 7 |
| chr1_-_216596738 | 0.19 |
ENST00000307340.3 ENST00000366943.2 ENST00000366942.3 |
USH2A |
Usher syndrome 2A (autosomal recessive, mild) |
| chr19_-_47616992 | 0.18 |
ENST00000253048.5 |
ZC3H4 |
zinc finger CCCH-type containing 4 |
| chr1_+_74701062 | 0.18 |
ENST00000326637.3 |
TNNI3K |
TNNI3 interacting kinase |
| chr17_+_41158742 | 0.17 |
ENST00000415816.2 ENST00000438323.2 |
IFI35 |
interferon-induced protein 35 |
| chrX_+_149861836 | 0.17 |
ENST00000542156.1 ENST00000370390.3 ENST00000490316.2 ENST00000445323.2 ENST00000544228.1 ENST00000451863.2 |
MTMR1 |
myotubularin related protein 1 |
| chr12_-_71533055 | 0.16 |
ENST00000552128.1 |
TSPAN8 |
tetraspanin 8 |
| chr3_+_58223228 | 0.16 |
ENST00000478253.1 ENST00000295962.4 |
ABHD6 |
abhydrolase domain containing 6 |
| chr2_-_74618964 | 0.16 |
ENST00000417090.1 ENST00000409868.1 |
DCTN1 |
dynactin 1 |
| chr1_+_174844645 | 0.16 |
ENST00000486220.1 |
RABGAP1L |
RAB GTPase activating protein 1-like |
| chr2_+_122513109 | 0.15 |
ENST00000389682.3 ENST00000536142.1 |
TSN |
translin |
| chr20_-_3140490 | 0.15 |
ENST00000449731.1 ENST00000380266.3 |
UBOX5 FASTKD5 |
U-box domain containing 5 FAST kinase domains 5 |
| chr1_+_229440129 | 0.15 |
ENST00000366688.3 |
SPHAR |
S-phase response (cyclin related) |
| chr14_-_106471723 | 0.15 |
ENST00000390595.2 |
IGHV1-3 |
immunoglobulin heavy variable 1-3 |
| chr4_-_70626314 | 0.15 |
ENST00000510821.1 |
SULT1B1 |
sulfotransferase family, cytosolic, 1B, member 1 |
| chr12_+_113354341 | 0.15 |
ENST00000553152.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr14_-_24911448 | 0.14 |
ENST00000555355.1 ENST00000553343.1 ENST00000556523.1 ENST00000556249.1 ENST00000538105.2 ENST00000555225.1 |
SDR39U1 |
short chain dehydrogenase/reductase family 39U, member 1 |
| chr14_-_106539557 | 0.14 |
ENST00000390599.2 |
IGHV1-8 |
immunoglobulin heavy variable 1-8 |
| chr9_+_125132803 | 0.13 |
ENST00000540753.1 |
PTGS1 |
prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) |
| chr16_-_21868978 | 0.13 |
ENST00000357370.5 ENST00000451409.1 ENST00000341400.7 ENST00000518761.4 |
NPIPB4 |
nuclear pore complex interacting protein family, member B4 |
| chr1_-_204116078 | 0.13 |
ENST00000367198.2 ENST00000452983.1 |
ETNK2 |
ethanolamine kinase 2 |
| chr11_+_113258495 | 0.13 |
ENST00000303941.3 |
ANKK1 |
ankyrin repeat and kinase domain containing 1 |
| chr6_+_26204825 | 0.13 |
ENST00000360441.4 |
HIST1H4E |
histone cluster 1, H4e |
| chr20_+_3052264 | 0.13 |
ENST00000217386.2 |
OXT |
oxytocin/neurophysin I prepropeptide |
| chr15_+_43425672 | 0.12 |
ENST00000260403.2 |
TMEM62 |
transmembrane protein 62 |
| chr19_+_41903709 | 0.12 |
ENST00000542943.1 ENST00000457836.2 |
BCKDHA |
branched chain keto acid dehydrogenase E1, alpha polypeptide |
| chr6_-_32731243 | 0.12 |
ENST00000427449.1 ENST00000411527.1 |
HLA-DQB2 |
major histocompatibility complex, class II, DQ beta 2 |
| chr12_-_14956396 | 0.12 |
ENST00000535328.1 ENST00000261167.2 |
WBP11 |
WW domain binding protein 11 |
| chr6_-_41039567 | 0.12 |
ENST00000468811.1 |
OARD1 |
O-acyl-ADP-ribose deacylase 1 |
| chr2_-_201936302 | 0.11 |
ENST00000453765.1 ENST00000452799.1 ENST00000446678.1 ENST00000418596.3 |
FAM126B |
family with sequence similarity 126, member B |
| chr14_+_21458127 | 0.11 |
ENST00000382985.4 ENST00000556670.2 ENST00000553564.1 ENST00000554751.1 ENST00000554283.1 ENST00000555670.1 |
METTL17 |
methyltransferase like 17 |
| chrX_-_100307043 | 0.11 |
ENST00000372939.1 ENST00000372935.1 ENST00000372936.3 |
TRMT2B |
tRNA methyltransferase 2 homolog B (S. cerevisiae) |
| chr5_+_59783941 | 0.11 |
ENST00000506884.1 ENST00000504876.2 |
PART1 |
prostate androgen-regulated transcript 1 (non-protein coding) |
| chr8_-_125577940 | 0.11 |
ENST00000519168.1 ENST00000395508.2 |
MTSS1 |
metastasis suppressor 1 |
| chr8_-_33370607 | 0.11 |
ENST00000360742.5 ENST00000523305.1 |
TTI2 |
TELO2 interacting protein 2 |
| chr16_-_8962200 | 0.11 |
ENST00000562843.1 ENST00000561530.1 ENST00000396593.2 |
CARHSP1 |
calcium regulated heat stable protein 1, 24kDa |
| chr16_+_69373323 | 0.11 |
ENST00000254940.5 |
NIP7 |
NIP7, nucleolar pre-rRNA processing protein |
| chr7_+_55980331 | 0.11 |
ENST00000429591.2 |
ZNF713 |
zinc finger protein 713 |
| chr1_-_26372602 | 0.10 |
ENST00000374278.3 ENST00000374276.3 |
SLC30A2 |
solute carrier family 30 (zinc transporter), member 2 |
| chr16_+_33204156 | 0.10 |
ENST00000398667.4 |
TP53TG3C |
TP53 target 3C |
| chr14_-_107131560 | 0.10 |
ENST00000390632.2 |
IGHV3-66 |
immunoglobulin heavy variable 3-66 |
| chr5_-_59783882 | 0.10 |
ENST00000505507.2 ENST00000502484.2 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr2_+_102721023 | 0.09 |
ENST00000409589.1 ENST00000409329.1 |
IL1R1 |
interleukin 1 receptor, type I |
| chrM_+_12331 | 0.09 |
ENST00000361567.2 |
MT-ND5 |
mitochondrially encoded NADH dehydrogenase 5 |
| chr16_-_29874211 | 0.09 |
ENST00000563415.1 |
CDIPT |
CDP-diacylglycerol--inositol 3-phosphatidyltransferase |
| chr1_+_150293973 | 0.09 |
ENST00000414970.2 ENST00000543398.1 |
PRPF3 |
pre-mRNA processing factor 3 |
| chr16_-_29415350 | 0.09 |
ENST00000524087.1 |
NPIPB11 |
nuclear pore complex interacting protein family, member B11 |
| chr1_-_36930012 | 0.08 |
ENST00000373116.5 |
MRPS15 |
mitochondrial ribosomal protein S15 |
| chr2_+_201936458 | 0.08 |
ENST00000237889.4 |
NDUFB3 |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 3, 12kDa |
| chr1_+_150293921 | 0.07 |
ENST00000324862.6 |
PRPF3 |
pre-mRNA processing factor 3 |
| chr16_-_21436459 | 0.07 |
ENST00000448012.2 ENST00000504841.2 ENST00000419180.2 |
NPIPB3 |
nuclear pore complex interacting protein family, member B3 |
| chr8_-_110988070 | 0.07 |
ENST00000524391.1 |
KCNV1 |
potassium channel, subfamily V, member 1 |
| chr14_-_102552659 | 0.07 |
ENST00000441629.2 |
HSP90AA1 |
heat shock protein 90kDa alpha (cytosolic), class A member 1 |
| chr16_-_29875057 | 0.07 |
ENST00000219789.6 |
CDIPT |
CDP-diacylglycerol--inositol 3-phosphatidyltransferase |
| chr1_+_45805342 | 0.07 |
ENST00000372090.5 |
TOE1 |
target of EGR1, member 1 (nuclear) |
| chr16_-_15950868 | 0.07 |
ENST00000396324.3 ENST00000452625.2 ENST00000576790.2 ENST00000300036.5 |
MYH11 |
myosin, heavy chain 11, smooth muscle |
| chrX_-_15333736 | 0.07 |
ENST00000380470.3 |
ASB11 |
ankyrin repeat and SOCS box containing 11 |
| chr11_-_124767693 | 0.06 |
ENST00000533054.1 |
ROBO4 |
roundabout, axon guidance receptor, homolog 4 (Drosophila) |
| chr3_+_195447738 | 0.06 |
ENST00000447234.2 ENST00000320736.6 ENST00000436408.1 |
MUC20 |
mucin 20, cell surface associated |
| chrX_+_100645812 | 0.06 |
ENST00000427805.2 ENST00000553110.3 ENST00000392994.3 ENST00000409338.1 ENST00000409170.3 |
RPL36A RPL36A-HNRNPH2 |
ribosomal protein L36a RPL36A-HNRNPH2 readthrough |
| chr1_-_151300160 | 0.06 |
ENST00000368874.4 |
PI4KB |
phosphatidylinositol 4-kinase, catalytic, beta |
| chr9_-_100935043 | 0.06 |
ENST00000343933.5 |
CORO2A |
coronin, actin binding protein, 2A |
| chr1_-_160832642 | 0.06 |
ENST00000368034.4 |
CD244 |
CD244 molecule, natural killer cell receptor 2B4 |
| chr1_+_17531614 | 0.06 |
ENST00000375471.4 |
PADI1 |
peptidyl arginine deiminase, type I |
| chr5_-_154230130 | 0.06 |
ENST00000519501.1 ENST00000518651.1 ENST00000517938.1 ENST00000520461.1 |
FAXDC2 |
fatty acid hydroxylase domain containing 2 |
| chr1_-_153029980 | 0.06 |
ENST00000392653.2 |
SPRR2A |
small proline-rich protein 2A |
| chr6_-_144416737 | 0.06 |
ENST00000367569.2 |
SF3B5 |
splicing factor 3b, subunit 5, 10kDa |
| chr11_-_67275542 | 0.05 |
ENST00000531506.1 |
CDK2AP2 |
cyclin-dependent kinase 2 associated protein 2 |
| chr14_-_27066636 | 0.05 |
ENST00000267422.7 ENST00000344429.5 ENST00000574031.1 ENST00000465357.2 ENST00000547619.1 |
NOVA1 |
neuro-oncological ventral antigen 1 |
| chr6_-_137539651 | 0.05 |
ENST00000543628.1 |
IFNGR1 |
interferon gamma receptor 1 |
| chr16_-_29874462 | 0.05 |
ENST00000566113.1 ENST00000569956.1 ENST00000570016.1 |
CDIPT |
CDP-diacylglycerol--inositol 3-phosphatidyltransferase |
| chr2_+_16080659 | 0.05 |
ENST00000281043.3 |
MYCN |
v-myc avian myelocytomatosis viral oncogene neuroblastoma derived homolog |
| chrX_+_76709648 | 0.05 |
ENST00000439435.1 |
FGF16 |
fibroblast growth factor 16 |
| chr5_+_59783540 | 0.05 |
ENST00000515734.2 |
PART1 |
prostate androgen-regulated transcript 1 (non-protein coding) |
| chr19_+_21265028 | 0.05 |
ENST00000291770.7 |
ZNF714 |
zinc finger protein 714 |
| chr14_-_106967788 | 0.05 |
ENST00000390622.2 |
IGHV1-46 |
immunoglobulin heavy variable 1-46 |
| chr3_+_11314099 | 0.04 |
ENST00000446450.2 ENST00000354956.5 ENST00000354449.3 ENST00000419112.1 |
ATG7 |
autophagy related 7 |
| chr8_-_56986768 | 0.04 |
ENST00000523936.1 |
RPS20 |
ribosomal protein S20 |
| chr15_-_34629922 | 0.04 |
ENST00000559484.1 ENST00000354181.3 ENST00000558589.1 ENST00000458406.2 |
SLC12A6 |
solute carrier family 12 (potassium/chloride transporter), member 6 |
| chr19_-_3761673 | 0.04 |
ENST00000316757.3 |
APBA3 |
amyloid beta (A4) precursor protein-binding, family A, member 3 |
| chr5_+_147443534 | 0.04 |
ENST00000398454.1 ENST00000359874.3 ENST00000508733.1 ENST00000256084.7 |
SPINK5 |
serine peptidase inhibitor, Kazal type 5 |
| chr14_-_107170409 | 0.04 |
ENST00000390633.2 |
IGHV1-69 |
immunoglobulin heavy variable 1-69 |
| chr15_-_43876702 | 0.04 |
ENST00000348806.6 |
PPIP5K1 |
diphosphoinositol pentakisphosphate kinase 1 |
| chr11_-_62420757 | 0.04 |
ENST00000330574.2 |
INTS5 |
integrator complex subunit 5 |
| chr19_-_16008880 | 0.04 |
ENST00000011989.7 ENST00000221700.6 |
CYP4F2 |
cytochrome P450, family 4, subfamily F, polypeptide 2 |
| chr16_-_66583701 | 0.04 |
ENST00000527800.1 ENST00000525974.1 ENST00000563369.2 |
TK2 |
thymidine kinase 2, mitochondrial |
| chr11_-_65626797 | 0.04 |
ENST00000525451.2 |
CFL1 |
cofilin 1 (non-muscle) |
| chr5_+_177631497 | 0.04 |
ENST00000358344.3 |
HNRNPAB |
heterogeneous nuclear ribonucleoprotein A/B |
| chr10_+_60028818 | 0.04 |
ENST00000333926.5 |
CISD1 |
CDGSH iron sulfur domain 1 |
| chr17_+_43971643 | 0.04 |
ENST00000344290.5 ENST00000262410.5 ENST00000351559.5 ENST00000340799.5 ENST00000535772.1 ENST00000347967.5 |
MAPT |
microtubule-associated protein tau |
| chr6_-_34393825 | 0.04 |
ENST00000605528.1 ENST00000326199.8 |
RPS10-NUDT3 RPS10 |
RPS10-NUDT3 readthrough ribosomal protein S10 |
| chr17_+_56769924 | 0.03 |
ENST00000461271.1 ENST00000583539.1 ENST00000337432.4 ENST00000421782.2 |
RAD51C |
RAD51 paralog C |
| chr5_-_77072085 | 0.03 |
ENST00000518338.2 ENST00000520039.1 ENST00000306388.6 ENST00000520361.1 |
TBCA |
tubulin folding cofactor A |
| chr19_+_35940486 | 0.03 |
ENST00000246549.2 |
FFAR2 |
free fatty acid receptor 2 |
| chr15_-_89755034 | 0.03 |
ENST00000563254.1 |
RLBP1 |
retinaldehyde binding protein 1 |
| chr4_+_100737954 | 0.03 |
ENST00000296414.7 ENST00000512369.1 |
DAPP1 |
dual adaptor of phosphotyrosine and 3-phosphoinositides |
| chr15_-_43877062 | 0.03 |
ENST00000381885.1 ENST00000396923.3 |
PPIP5K1 |
diphosphoinositol pentakisphosphate kinase 1 |
| chr22_+_30163340 | 0.03 |
ENST00000330029.6 ENST00000401406.3 |
UQCR10 |
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr3_-_58572760 | 0.03 |
ENST00000447756.2 |
FAM107A |
family with sequence similarity 107, member A |
| chrX_-_49089771 | 0.03 |
ENST00000376251.1 ENST00000323022.5 ENST00000376265.2 |
CACNA1F |
calcium channel, voltage-dependent, L type, alpha 1F subunit |
| chr8_+_22132847 | 0.03 |
ENST00000521356.1 |
PIWIL2 |
piwi-like RNA-mediated gene silencing 2 |
| chr19_-_42916499 | 0.03 |
ENST00000601189.1 ENST00000599211.1 |
LIPE |
lipase, hormone-sensitive |
| chr3_+_142315225 | 0.03 |
ENST00000457734.2 ENST00000483373.1 ENST00000475296.1 ENST00000495744.1 ENST00000476044.1 ENST00000461644.1 |
PLS1 |
plastin 1 |
| chr10_-_115423792 | 0.03 |
ENST00000369360.3 ENST00000360478.3 ENST00000359988.3 ENST00000369358.4 |
NRAP |
nebulin-related anchoring protein |
| chr3_+_42544084 | 0.02 |
ENST00000543411.1 ENST00000438259.2 ENST00000439731.1 ENST00000325123.4 |
VIPR1 |
vasoactive intestinal peptide receptor 1 |
| chr2_+_234580499 | 0.02 |
ENST00000354728.4 |
UGT1A9 |
UDP glucuronosyltransferase 1 family, polypeptide A9 |
| chr2_+_234580525 | 0.02 |
ENST00000609637.1 |
UGT1A1 |
UDP glucuronosyltransferase 1 family, polypeptide A8 |
| chr11_-_67210930 | 0.02 |
ENST00000453768.2 ENST00000545016.1 ENST00000341356.5 |
CORO1B |
coronin, actin binding protein, 1B |
| chr7_-_120498357 | 0.02 |
ENST00000415871.1 ENST00000222747.3 ENST00000430985.1 |
TSPAN12 |
tetraspanin 12 |
| chr2_+_232135245 | 0.02 |
ENST00000446447.1 |
ARMC9 |
armadillo repeat containing 9 |
| chr6_+_35420091 | 0.02 |
ENST00000229769.2 |
FANCE |
Fanconi anemia, complementation group E |
| chr14_-_89878369 | 0.01 |
ENST00000553840.1 ENST00000556916.1 |
FOXN3 |
forkhead box N3 |
| chr3_+_121311966 | 0.01 |
ENST00000338040.4 |
FBXO40 |
F-box protein 40 |
| chr8_+_101162812 | 0.01 |
ENST00000353107.3 ENST00000522439.1 |
POLR2K |
polymerase (RNA) II (DNA directed) polypeptide K, 7.0kDa |
| chr5_-_9712312 | 0.01 |
ENST00000506620.1 ENST00000514078.1 ENST00000606744.1 |
TAS2R1 CTD-2143L24.1 |
taste receptor, type 2, member 1 CTD-2143L24.1 |
| chr14_-_27066960 | 0.01 |
ENST00000539517.2 |
NOVA1 |
neuro-oncological ventral antigen 1 |
| chr9_-_100684845 | 0.01 |
ENST00000375119.3 |
C9orf156 |
chromosome 9 open reading frame 156 |
| chr1_+_241695670 | 0.01 |
ENST00000366557.4 |
KMO |
kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) |
| chr13_+_23755099 | 0.01 |
ENST00000537476.1 |
SGCG |
sarcoglycan, gamma (35kDa dystrophin-associated glycoprotein) |
| chr1_+_117910047 | 0.01 |
ENST00000356554.3 |
MAN1A2 |
mannosidase, alpha, class 1A, member 2 |
| chr21_+_44073916 | 0.00 |
ENST00000349112.3 ENST00000398224.3 |
PDE9A |
phosphodiesterase 9A |
| chr1_-_78444738 | 0.00 |
ENST00000436586.2 ENST00000370768.2 |
FUBP1 |
far upstream element (FUSE) binding protein 1 |
| chr12_+_100594557 | 0.00 |
ENST00000546902.1 ENST00000552376.1 ENST00000551617.1 |
ACTR6 |
ARP6 actin-related protein 6 homolog (yeast) |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | GO:0005114 | type II transforming growth factor beta receptor binding(GO:0005114) |
| 0.1 | 0.3 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 0.1 | 0.6 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.1 | 0.2 | GO:0003881 | CDP-diacylglycerol-inositol 3-phosphatidyltransferase activity(GO:0003881) |
| 0.1 | 0.4 | GO:0072542 | protein phosphatase activator activity(GO:0072542) |
| 0.0 | 0.1 | GO:0003826 | alpha-ketoacid dehydrogenase activity(GO:0003826) 3-methyl-2-oxobutanoate dehydrogenase (2-methylpropanoyl-transferring) activity(GO:0003863) |
| 0.0 | 0.3 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.0 | 0.1 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.0 | 0.5 | GO:0102391 | decanoate--CoA ligase activity(GO:0102391) |
| 0.0 | 0.1 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 0.0 | 0.1 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.0 | 0.4 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.0 | 0.3 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.0 | 0.2 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.0 | 0.4 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 0.3 | GO:0004559 | alpha-mannosidase activity(GO:0004559) |
| 0.0 | 0.1 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.0 | 0.1 | GO:0004305 | ethanolamine kinase activity(GO:0004305) |
| 0.0 | 0.1 | GO:0033857 | diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.0 | 0.2 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.0 | 0.0 | GO:0018685 | alkane 1-monooxygenase activity(GO:0018685) tocopherol omega-hydroxylase activity(GO:0052870) alpha-tocopherol omega-hydroxylase activity(GO:0052871) 20-hydroxy-leukotriene B4 omega oxidase activity(GO:0097258) 20-aldehyde-leukotriene B4 20-monooxygenase activity(GO:0097259) |
| 0.0 | 0.0 | GO:0004137 | deoxycytidine kinase activity(GO:0004137) thymidine kinase activity(GO:0004797) |
| 0.0 | 0.2 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.0 | 0.1 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.0 | 0.6 | GO:0019212 | phosphatase inhibitor activity(GO:0019212) |
| 0.0 | 0.1 | GO:0004906 | interferon-gamma receptor activity(GO:0004906) |
| 0.0 | 0.1 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.0 | 0.4 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.0 | 0.2 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | GO:0034673 | inhibin-betaglycan-ActRII complex(GO:0034673) |
| 0.1 | 0.3 | GO:0000799 | nuclear condensin complex(GO:0000799) |
| 0.1 | 0.2 | GO:0097124 | cyclin A2-CDK2 complex(GO:0097124) |
| 0.1 | 0.2 | GO:0002139 | stereocilia coupling link(GO:0002139) |
| 0.0 | 0.3 | GO:1990712 | HFE-transferrin receptor complex(GO:1990712) |
| 0.0 | 0.1 | GO:0005947 | mitochondrial alpha-ketoglutarate dehydrogenase complex(GO:0005947) |
| 0.0 | 0.4 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 0.4 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.0 | 0.2 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 0.7 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.1 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.1 | 0.4 | GO:0061386 | closure of optic fissure(GO:0061386) negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.1 | 0.2 | GO:0071314 | cellular response to cocaine(GO:0071314) |
| 0.1 | 0.6 | GO:0032417 | positive regulation of sodium:proton antiporter activity(GO:0032417) |
| 0.1 | 0.2 | GO:0071051 | polyadenylation-dependent snoRNA 3'-end processing(GO:0071051) |
| 0.1 | 0.3 | GO:0097368 | establishment of Sertoli cell barrier(GO:0097368) |
| 0.1 | 0.3 | GO:0033078 | extrathymic T cell differentiation(GO:0033078) |
| 0.1 | 0.3 | GO:0097460 | ferrous iron import into cell(GO:0097460) ferrous iron import across plasma membrane(GO:0098707) |
| 0.0 | 0.2 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.0 | 0.4 | GO:0090063 | positive regulation of microtubule nucleation(GO:0090063) |
| 0.0 | 0.1 | GO:0002125 | maternal aggressive behavior(GO:0002125) positive regulation of female receptivity(GO:0045925) |
| 0.0 | 0.3 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.0 | 0.6 | GO:0021702 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.0 | 0.1 | GO:2000661 | positive regulation of interleukin-1-mediated signaling pathway(GO:2000661) |
| 0.0 | 0.2 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 0.0 | 0.2 | GO:0002349 | histamine production involved in inflammatory response(GO:0002349) histamine secretion involved in inflammatory response(GO:0002441) histamine secretion by mast cell(GO:0002553) |
| 0.0 | 0.1 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) |
| 0.0 | 0.3 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) phosphatidylcholine catabolic process(GO:0034638) |
| 0.0 | 0.5 | GO:0035338 | long-chain fatty-acyl-CoA biosynthetic process(GO:0035338) |
| 0.0 | 0.1 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.1 | GO:2000566 | positive regulation of CD8-positive, alpha-beta T cell proliferation(GO:2000566) |
| 0.0 | 0.4 | GO:0033008 | positive regulation of mast cell activation involved in immune response(GO:0033008) positive regulation of mast cell degranulation(GO:0043306) |
| 0.0 | 0.0 | GO:0032304 | negative regulation of icosanoid secretion(GO:0032304) |
| 0.0 | 0.1 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.0 | 0.1 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.0 | 0.3 | GO:0006883 | cellular sodium ion homeostasis(GO:0006883) |
| 0.0 | 0.0 | GO:1901859 | late nucleophagy(GO:0044805) negative regulation of mitochondrial DNA replication(GO:0090298) negative regulation of mitochondrial DNA metabolic process(GO:1901859) single-organism membrane invagination(GO:1902534) |
| 0.0 | 0.3 | GO:0035855 | megakaryocyte development(GO:0035855) |
| 0.0 | 0.4 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.0 | 0.1 | GO:0036414 | protein citrullination(GO:0018101) histone citrullination(GO:0036414) |
| 0.0 | 0.2 | GO:0046341 | CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.0 | 0.2 | GO:0030422 | production of siRNA involved in RNA interference(GO:0030422) |
| 0.0 | 0.0 | GO:0002752 | leukocyte chemotaxis involved in inflammatory response(GO:0002232) cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 0.0 | 0.1 | GO:0086024 | adrenergic receptor signaling pathway involved in positive regulation of heart rate(GO:0086024) |
| 0.0 | 0.0 | GO:1900425 | negative regulation of defense response to bacterium(GO:1900425) |
| 0.0 | 0.1 | GO:0070341 | fat cell proliferation(GO:0070341) regulation of fat cell proliferation(GO:0070344) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.2 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.0 | 0.6 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 0.8 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.0 | 0.3 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.2 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |