Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
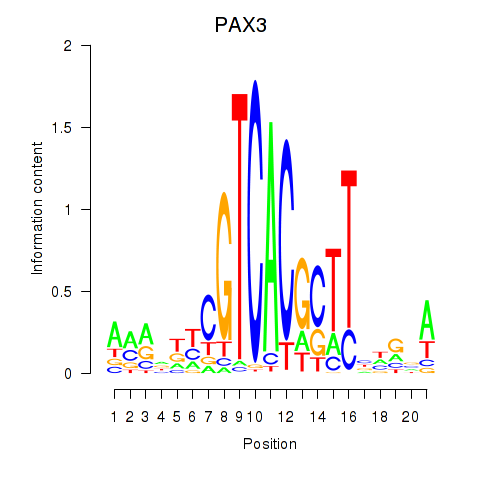
Results for PAX3
Z-value: 0.78
Transcription factors associated with PAX3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PAX3
|
ENSG00000135903.14 | PAX3 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PAX3 | hg19_v2_chr2_-_223163465_223163730 | -0.01 | 9.6e-01 | Click! |
Activity profile of PAX3 motif
Sorted Z-values of PAX3 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PAX3
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_+_7060432 | 2.43 |
ENST00000318974.9 ENST00000456013.1 |
PTPN6 |
protein tyrosine phosphatase, non-receptor type 6 |
| chr11_-_73694346 | 2.32 |
ENST00000310473.3 |
UCP2 |
uncoupling protein 2 (mitochondrial, proton carrier) |
| chr11_+_313503 | 2.14 |
ENST00000528780.1 ENST00000328221.5 |
IFITM1 |
interferon induced transmembrane protein 1 |
| chr16_+_23847339 | 1.32 |
ENST00000303531.7 |
PRKCB |
protein kinase C, beta |
| chr6_-_32908792 | 1.31 |
ENST00000418107.2 |
HLA-DMB |
major histocompatibility complex, class II, DM beta |
| chr6_+_14117872 | 1.28 |
ENST00000379153.3 |
CD83 |
CD83 molecule |
| chr12_-_53594227 | 1.23 |
ENST00000550743.2 |
ITGB7 |
integrin, beta 7 |
| chr7_+_99816859 | 1.16 |
ENST00000317271.2 |
PVRIG |
poliovirus receptor related immunoglobulin domain containing |
| chr15_-_80263506 | 1.14 |
ENST00000335661.6 |
BCL2A1 |
BCL2-related protein A1 |
| chr17_-_34417479 | 1.02 |
ENST00000225245.5 |
CCL3 |
chemokine (C-C motif) ligand 3 |
| chr1_+_101185290 | 0.97 |
ENST00000370119.4 ENST00000347652.2 ENST00000294728.2 ENST00000370115.1 |
VCAM1 |
vascular cell adhesion molecule 1 |
| chr11_-_615570 | 0.93 |
ENST00000525445.1 ENST00000348655.6 ENST00000397566.1 |
IRF7 |
interferon regulatory factor 7 |
| chrX_-_53461288 | 0.87 |
ENST00000375298.4 ENST00000375304.5 |
HSD17B10 |
hydroxysteroid (17-beta) dehydrogenase 10 |
| chrX_-_53461305 | 0.86 |
ENST00000168216.6 |
HSD17B10 |
hydroxysteroid (17-beta) dehydrogenase 10 |
| chr6_-_109804412 | 0.79 |
ENST00000230122.3 |
ZBTB24 |
zinc finger and BTB domain containing 24 |
| chr15_-_43622736 | 0.76 |
ENST00000544735.1 ENST00000567039.1 ENST00000305641.5 |
LCMT2 |
leucine carboxyl methyltransferase 2 |
| chrX_-_15872914 | 0.75 |
ENST00000380291.1 ENST00000545766.1 ENST00000421527.2 ENST00000329235.2 |
AP1S2 |
adaptor-related protein complex 1, sigma 2 subunit |
| chr11_-_615942 | 0.74 |
ENST00000397562.3 ENST00000330243.5 ENST00000397570.1 ENST00000397574.2 |
IRF7 |
interferon regulatory factor 7 |
| chr15_+_81591757 | 0.71 |
ENST00000558332.1 |
IL16 |
interleukin 16 |
| chr5_+_49961727 | 0.70 |
ENST00000505697.2 ENST00000503750.2 ENST00000514342.2 |
PARP8 |
poly (ADP-ribose) polymerase family, member 8 |
| chr9_-_35650900 | 0.69 |
ENST00000259608.3 |
SIT1 |
signaling threshold regulating transmembrane adaptor 1 |
| chr6_-_32908765 | 0.51 |
ENST00000416244.2 |
HLA-DMB |
major histocompatibility complex, class II, DM beta |
| chr2_+_152214098 | 0.50 |
ENST00000243347.3 |
TNFAIP6 |
tumor necrosis factor, alpha-induced protein 6 |
| chr7_-_148580563 | 0.46 |
ENST00000476773.1 |
EZH2 |
enhancer of zeste homolog 2 (Drosophila) |
| chr4_+_156680153 | 0.44 |
ENST00000502959.1 ENST00000505764.1 ENST00000507146.1 ENST00000264424.8 ENST00000503520.1 |
GUCY1B3 |
guanylate cyclase 1, soluble, beta 3 |
| chr10_-_75226166 | 0.41 |
ENST00000544628.1 |
PPP3CB |
protein phosphatase 3, catalytic subunit, beta isozyme |
| chr19_+_35168567 | 0.40 |
ENST00000457781.2 ENST00000505163.1 ENST00000505242.1 ENST00000423823.2 ENST00000507959.1 ENST00000446502.2 |
ZNF302 |
zinc finger protein 302 |
| chr1_+_99729813 | 0.40 |
ENST00000457765.1 |
LPPR4 |
Lipid phosphate phosphatase-related protein type 4 |
| chr7_+_116502605 | 0.39 |
ENST00000458284.2 ENST00000490693.1 |
CAPZA2 |
capping protein (actin filament) muscle Z-line, alpha 2 |
| chr6_-_169654139 | 0.39 |
ENST00000366787.3 |
THBS2 |
thrombospondin 2 |
| chr5_-_180688105 | 0.38 |
ENST00000327767.4 |
TRIM52 |
tripartite motif containing 52 |
| chr6_+_127588020 | 0.35 |
ENST00000309649.3 ENST00000610162.1 ENST00000610153.1 ENST00000608991.1 ENST00000480444.1 |
RNF146 |
ring finger protein 146 |
| chr4_+_156680143 | 0.34 |
ENST00000505154.1 |
GUCY1B3 |
guanylate cyclase 1, soluble, beta 3 |
| chr19_+_35168633 | 0.34 |
ENST00000505365.2 |
ZNF302 |
zinc finger protein 302 |
| chr8_+_107670064 | 0.32 |
ENST00000312046.6 |
OXR1 |
oxidation resistance 1 |
| chr12_+_112563335 | 0.31 |
ENST00000549358.1 ENST00000257604.5 ENST00000548092.1 ENST00000552896.1 |
TRAFD1 |
TRAF-type zinc finger domain containing 1 |
| chr3_-_119379427 | 0.31 |
ENST00000264231.3 ENST00000468801.1 ENST00000538678.1 |
POPDC2 |
popeye domain containing 2 |
| chr1_-_205180664 | 0.30 |
ENST00000367161.3 ENST00000367162.3 ENST00000367160.4 |
DSTYK |
dual serine/threonine and tyrosine protein kinase |
| chr12_+_112563303 | 0.30 |
ENST00000412615.2 |
TRAFD1 |
TRAF-type zinc finger domain containing 1 |
| chr3_-_119379719 | 0.30 |
ENST00000493094.1 |
POPDC2 |
popeye domain containing 2 |
| chr6_+_127587755 | 0.29 |
ENST00000368314.1 ENST00000476956.1 ENST00000609447.1 ENST00000356799.2 ENST00000477776.1 ENST00000609944.1 |
RNF146 |
ring finger protein 146 |
| chr10_+_91087651 | 0.28 |
ENST00000371818.4 |
IFIT3 |
interferon-induced protein with tetratricopeptide repeats 3 |
| chrX_+_30265256 | 0.27 |
ENST00000397548.2 |
MAGEB1 |
melanoma antigen family B, 1 |
| chr11_-_104972158 | 0.26 |
ENST00000598974.1 ENST00000593315.1 ENST00000594519.1 ENST00000415981.2 ENST00000525374.1 ENST00000375707.1 |
CASP1 CARD16 CARD17 |
caspase 1, apoptosis-related cysteine peptidase caspase recruitment domain family, member 16 caspase recruitment domain family, member 17 |
| chr3_+_183892635 | 0.26 |
ENST00000427072.1 ENST00000411763.2 ENST00000292807.5 ENST00000448139.1 ENST00000455925.1 |
AP2M1 |
adaptor-related protein complex 2, mu 1 subunit |
| chr5_+_178977546 | 0.25 |
ENST00000319449.4 ENST00000377001.2 |
RUFY1 |
RUN and FYVE domain containing 1 |
| chr14_+_19553365 | 0.25 |
ENST00000409832.3 |
POTEG |
POTE ankyrin domain family, member G |
| chr2_-_68479614 | 0.25 |
ENST00000234310.3 |
PPP3R1 |
protein phosphatase 3, regulatory subunit B, alpha |
| chr15_-_52861157 | 0.25 |
ENST00000564163.1 |
ARPP19 |
cAMP-regulated phosphoprotein, 19kDa |
| chr1_+_203444887 | 0.24 |
ENST00000343110.2 |
PRELP |
proline/arginine-rich end leucine-rich repeat protein |
| chrX_+_102883620 | 0.22 |
ENST00000372626.3 |
TCEAL1 |
transcription elongation factor A (SII)-like 1 |
| chr9_+_100395891 | 0.22 |
ENST00000375147.3 |
NCBP1 |
nuclear cap binding protein subunit 1, 80kDa |
| chr19_-_41903161 | 0.21 |
ENST00000602129.1 ENST00000593771.1 ENST00000596905.1 ENST00000221233.4 |
EXOSC5 |
exosome component 5 |
| chr2_-_44588893 | 0.20 |
ENST00000409272.1 ENST00000410081.1 ENST00000541738.1 |
PREPL |
prolyl endopeptidase-like |
| chr5_+_70220768 | 0.20 |
ENST00000351205.4 ENST00000503079.2 ENST00000380707.4 ENST00000514951.1 ENST00000506163.1 |
SMN1 |
survival of motor neuron 1, telomeric |
| chr3_+_160117418 | 0.19 |
ENST00000465903.1 ENST00000485645.1 ENST00000360111.2 ENST00000472991.1 ENST00000467468.1 ENST00000469762.1 ENST00000489573.1 ENST00000462787.1 ENST00000490207.1 ENST00000485867.1 |
SMC4 |
structural maintenance of chromosomes 4 |
| chr1_-_24151903 | 0.18 |
ENST00000436439.2 ENST00000374490.3 |
HMGCL |
3-hydroxymethyl-3-methylglutaryl-CoA lyase |
| chr2_-_11810284 | 0.18 |
ENST00000306928.5 |
NTSR2 |
neurotensin receptor 2 |
| chr10_-_104178857 | 0.18 |
ENST00000020673.5 |
PSD |
pleckstrin and Sec7 domain containing |
| chr17_+_4855053 | 0.18 |
ENST00000518175.1 |
ENO3 |
enolase 3 (beta, muscle) |
| chr18_+_23806437 | 0.18 |
ENST00000578121.1 |
TAF4B |
TAF4b RNA polymerase II, TATA box binding protein (TBP)-associated factor, 105kDa |
| chr10_+_118305435 | 0.18 |
ENST00000369221.2 |
PNLIP |
pancreatic lipase |
| chr10_+_104178946 | 0.17 |
ENST00000432590.1 |
FBXL15 |
F-box and leucine-rich repeat protein 15 |
| chr12_-_57400227 | 0.17 |
ENST00000300101.2 |
ZBTB39 |
zinc finger and BTB domain containing 39 |
| chr2_-_44588694 | 0.17 |
ENST00000409957.1 |
PREPL |
prolyl endopeptidase-like |
| chr1_-_155214475 | 0.17 |
ENST00000428024.3 |
GBA |
glucosidase, beta, acid |
| chr2_-_44588679 | 0.16 |
ENST00000409411.1 |
PREPL |
prolyl endopeptidase-like |
| chr8_+_125463048 | 0.15 |
ENST00000328599.3 |
TRMT12 |
tRNA methyltransferase 12 homolog (S. cerevisiae) |
| chr5_+_69345350 | 0.15 |
ENST00000380741.4 ENST00000380743.4 ENST00000511812.1 ENST00000380742.4 |
SMN2 |
survival of motor neuron 2, centromeric |
| chr6_-_24721054 | 0.15 |
ENST00000378119.4 |
C6orf62 |
chromosome 6 open reading frame 62 |
| chr16_+_27214802 | 0.14 |
ENST00000380948.2 ENST00000286096.4 |
KDM8 |
lysine (K)-specific demethylase 8 |
| chr19_+_2841433 | 0.14 |
ENST00000334241.4 ENST00000585966.1 ENST00000591539.1 |
ZNF555 |
zinc finger protein 555 |
| chr2_-_44588624 | 0.14 |
ENST00000438314.1 ENST00000409936.1 |
PREPL |
prolyl endopeptidase-like |
| chr9_+_104161123 | 0.14 |
ENST00000374861.3 ENST00000339664.2 ENST00000259395.4 |
ZNF189 |
zinc finger protein 189 |
| chr2_-_183106641 | 0.13 |
ENST00000346717.4 |
PDE1A |
phosphodiesterase 1A, calmodulin-dependent |
| chr7_+_116502527 | 0.13 |
ENST00000361183.3 |
CAPZA2 |
capping protein (actin filament) muscle Z-line, alpha 2 |
| chr1_-_24151892 | 0.13 |
ENST00000235958.4 |
HMGCL |
3-hydroxymethyl-3-methylglutaryl-CoA lyase |
| chr15_+_51973680 | 0.13 |
ENST00000542355.2 |
SCG3 |
secretogranin III |
| chr9_-_88897426 | 0.12 |
ENST00000375991.4 ENST00000326094.4 |
ISCA1 |
iron-sulfur cluster assembly 1 |
| chr22_+_18593507 | 0.11 |
ENST00000330423.3 |
TUBA8 |
tubulin, alpha 8 |
| chr3_+_160117087 | 0.11 |
ENST00000357388.3 |
SMC4 |
structural maintenance of chromosomes 4 |
| chr1_-_205091115 | 0.11 |
ENST00000264515.6 ENST00000367164.1 |
RBBP5 |
retinoblastoma binding protein 5 |
| chr22_-_44258360 | 0.11 |
ENST00000330884.4 ENST00000249130.5 |
SULT4A1 |
sulfotransferase family 4A, member 1 |
| chr8_-_81083731 | 0.10 |
ENST00000379096.5 |
TPD52 |
tumor protein D52 |
| chr15_+_51973550 | 0.10 |
ENST00000220478.3 |
SCG3 |
secretogranin III |
| chr21_-_37451680 | 0.10 |
ENST00000399201.1 |
SETD4 |
SET domain containing 4 |
| chr8_-_101718991 | 0.09 |
ENST00000517990.1 |
PABPC1 |
poly(A) binding protein, cytoplasmic 1 |
| chr11_+_66624527 | 0.09 |
ENST00000393952.3 |
LRFN4 |
leucine rich repeat and fibronectin type III domain containing 4 |
| chr1_+_12524965 | 0.09 |
ENST00000471923.1 |
VPS13D |
vacuolar protein sorting 13 homolog D (S. cerevisiae) |
| chr19_-_41388657 | 0.09 |
ENST00000301146.4 ENST00000291764.3 |
CYP2A7 |
cytochrome P450, family 2, subfamily A, polypeptide 7 |
| chr17_-_39538550 | 0.08 |
ENST00000394001.1 |
KRT34 |
keratin 34 |
| chr6_+_89674246 | 0.08 |
ENST00000369474.1 |
AL079342.1 |
Uncharacterized protein; cDNA FLJ27030 fis, clone SLV07741 |
| chr14_-_24711806 | 0.08 |
ENST00000540705.1 ENST00000538777.1 ENST00000558566.1 ENST00000559019.1 |
TINF2 |
TERF1 (TRF1)-interacting nuclear factor 2 |
| chr1_+_26496362 | 0.07 |
ENST00000374266.5 ENST00000270812.5 |
ZNF593 |
zinc finger protein 593 |
| chr11_+_33563821 | 0.07 |
ENST00000321505.4 ENST00000265654.5 ENST00000389726.3 |
KIAA1549L |
KIAA1549-like |
| chr6_+_35227449 | 0.07 |
ENST00000373953.3 ENST00000440666.2 ENST00000339411.5 |
ZNF76 |
zinc finger protein 76 |
| chr9_-_91793675 | 0.07 |
ENST00000375835.4 ENST00000375830.1 |
SHC3 |
SHC (Src homology 2 domain containing) transforming protein 3 |
| chr1_+_154244987 | 0.06 |
ENST00000328703.7 ENST00000457918.2 ENST00000483970.2 ENST00000435087.1 ENST00000532105.1 |
HAX1 |
HCLS1 associated protein X-1 |
| chr12_+_10460549 | 0.05 |
ENST00000543420.1 ENST00000543777.1 |
KLRD1 |
killer cell lectin-like receptor subfamily D, member 1 |
| chr22_+_20105259 | 0.05 |
ENST00000416427.1 ENST00000421656.1 ENST00000423859.1 ENST00000418705.2 |
RANBP1 |
RAN binding protein 1 |
| chr1_-_155214436 | 0.04 |
ENST00000327247.5 |
GBA |
glucosidase, beta, acid |
| chr1_-_52344471 | 0.04 |
ENST00000352171.7 ENST00000354831.7 |
NRD1 |
nardilysin (N-arginine dibasic convertase) |
| chr12_+_69080734 | 0.04 |
ENST00000378905.2 |
NUP107 |
nucleoporin 107kDa |
| chr13_+_96085847 | 0.03 |
ENST00000376873.3 |
CLDN10 |
claudin 10 |
| chr19_-_54784353 | 0.03 |
ENST00000391746.1 |
LILRB2 |
leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 2 |
| chr20_-_17511962 | 0.03 |
ENST00000377873.3 |
BFSP1 |
beaded filament structural protein 1, filensin |
| chr2_-_219524193 | 0.03 |
ENST00000450560.1 ENST00000449707.1 ENST00000432460.1 ENST00000411696.2 |
ZNF142 |
zinc finger protein 142 |
| chr22_+_39101728 | 0.03 |
ENST00000216044.5 ENST00000484657.1 |
GTPBP1 |
GTP binding protein 1 |
| chr3_+_112709804 | 0.02 |
ENST00000383677.3 |
GTPBP8 |
GTP-binding protein 8 (putative) |
| chr3_+_112709755 | 0.02 |
ENST00000383678.2 |
GTPBP8 |
GTP-binding protein 8 (putative) |
| chr5_+_159343688 | 0.02 |
ENST00000306675.3 |
ADRA1B |
adrenoceptor alpha 1B |
| chr15_+_99791567 | 0.02 |
ENST00000558879.1 ENST00000301981.3 ENST00000422500.2 ENST00000447360.2 ENST00000442993.2 |
LRRC28 |
leucine rich repeat containing 28 |
| chr5_-_157079428 | 0.02 |
ENST00000265007.6 |
SOX30 |
SRY (sex determining region Y)-box 30 |
| chr4_-_168155417 | 0.02 |
ENST00000511269.1 ENST00000506697.1 ENST00000512042.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chr2_-_217724767 | 0.02 |
ENST00000236979.2 |
TNP1 |
transition protein 1 (during histone to protamine replacement) |
| chrX_+_70364667 | 0.02 |
ENST00000536169.1 ENST00000395855.2 ENST00000374051.3 ENST00000358741.3 |
NLGN3 |
neuroligin 3 |
| chr20_-_23807358 | 0.01 |
ENST00000304725.2 |
CST2 |
cystatin SA |
| chr17_+_3118915 | 0.01 |
ENST00000304094.1 |
OR1A1 |
olfactory receptor, family 1, subfamily A, member 1 |
| chrX_+_1734051 | 0.00 |
ENST00000381229.4 ENST00000381233.3 |
ASMT |
acetylserotonin O-methyltransferase |
| chr10_-_102289611 | 0.00 |
ENST00000299166.4 ENST00000370320.4 ENST00000531258.1 ENST00000370322.1 ENST00000535773.1 |
NDUFB8 SEC31B |
NADH dehydrogenase (ubiquinone) 1 beta subcomplex, 8, 19kDa SEC31 homolog B (S. cerevisiae) |
| chr19_+_55105085 | 0.00 |
ENST00000251372.3 ENST00000453777.1 |
LILRA1 |
leukocyte immunoglobulin-like receptor, subfamily A (with TM domain), member 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.8 | GO:2001190 | positive regulation of T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:2001190) |
| 0.6 | 2.4 | GO:0033277 | abortive mitotic cell cycle(GO:0033277) |
| 0.6 | 1.7 | GO:0070901 | mitochondrial tRNA methylation(GO:0070901) |
| 0.4 | 1.3 | GO:0035408 | histone H3-T6 phosphorylation(GO:0035408) |
| 0.3 | 1.7 | GO:0034124 | regulation of MyD88-dependent toll-like receptor signaling pathway(GO:0034124) negative regulation of macrophage apoptotic process(GO:2000110) |
| 0.3 | 1.2 | GO:0003366 | cell-matrix adhesion involved in ameboidal cell migration(GO:0003366) |
| 0.3 | 0.8 | GO:0038060 | nitric oxide-cGMP-mediated signaling pathway(GO:0038060) |
| 0.2 | 2.3 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.2 | 1.0 | GO:0022614 | membrane to membrane docking(GO:0022614) |
| 0.2 | 1.3 | GO:0032713 | negative regulation of interleukin-4 production(GO:0032713) |
| 0.2 | 0.5 | GO:1904772 | hepatocyte homeostasis(GO:0036333) response to tetrachloromethane(GO:1904772) |
| 0.1 | 0.8 | GO:2000503 | positive regulation of natural killer cell chemotaxis(GO:2000503) |
| 0.1 | 2.4 | GO:0035455 | response to interferon-alpha(GO:0035455) |
| 0.1 | 0.2 | GO:0071051 | polyadenylation-dependent snoRNA 3'-end processing(GO:0071051) |
| 0.1 | 0.3 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 0.1 | 0.2 | GO:1901804 | beta-glucoside metabolic process(GO:1901804) beta-glucoside catabolic process(GO:1901805) positive regulation of neuronal action potential(GO:1904457) |
| 0.1 | 1.2 | GO:0050860 | negative regulation of T cell receptor signaling pathway(GO:0050860) |
| 0.1 | 0.3 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.0 | 1.1 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.0 | 0.3 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.0 | 0.4 | GO:0001915 | negative regulation of T cell mediated cytotoxicity(GO:0001915) |
| 0.0 | 0.3 | GO:0006552 | leucine catabolic process(GO:0006552) |
| 0.0 | 0.7 | GO:0050930 | induction of positive chemotaxis(GO:0050930) |
| 0.0 | 0.8 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 0.2 | GO:1905216 | positive regulation of RNA binding(GO:1905216) |
| 0.0 | 1.0 | GO:0050690 | regulation of defense response to virus by virus(GO:0050690) |
| 0.0 | 0.3 | GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation(GO:1902083) |
| 0.0 | 0.1 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.0 | 0.1 | GO:0070544 | histone H3-K36 demethylation(GO:0070544) |
| 0.0 | 0.7 | GO:0043029 | T cell homeostasis(GO:0043029) |
| 0.0 | 0.5 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 0.7 | GO:2000300 | regulation of synaptic vesicle exocytosis(GO:2000300) |
| 0.0 | 0.2 | GO:0061365 | positive regulation of triglyceride lipase activity(GO:0061365) |
| 0.0 | 0.5 | GO:0051016 | barbed-end actin filament capping(GO:0051016) |
| 0.0 | 0.1 | GO:0045722 | positive regulation of gluconeogenesis(GO:0045722) |
| 0.0 | 0.1 | GO:0010836 | negative regulation of protein ADP-ribosylation(GO:0010836) |
| 0.0 | 0.2 | GO:0042340 | keratan sulfate catabolic process(GO:0042340) |
| 0.0 | 0.6 | GO:0045824 | negative regulation of innate immune response(GO:0045824) |
| 0.0 | 0.1 | GO:2000622 | regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.7 | GO:0030678 | mitochondrial ribonuclease P complex(GO:0030678) |
| 0.3 | 1.0 | GO:0071065 | alpha9-beta1 integrin-vascular cell adhesion molecule-1 complex(GO:0071065) |
| 0.3 | 1.2 | GO:0034669 | integrin alpha4-beta7 complex(GO:0034669) |
| 0.2 | 2.4 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.1 | 1.8 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.1 | 0.7 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.1 | 0.2 | GO:0005846 | nuclear cap binding complex(GO:0005846) |
| 0.1 | 0.3 | GO:0097179 | protease inhibitor complex(GO:0097179) |
| 0.0 | 0.5 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.0 | 0.5 | GO:0045120 | pronucleus(GO:0045120) |
| 0.0 | 0.8 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.0 | 0.3 | GO:0000796 | condensin complex(GO:0000796) |
| 0.0 | 0.2 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 0.2 | GO:0097504 | Gemini of coiled bodies(GO:0097504) |
| 0.0 | 0.2 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.0 | 1.0 | GO:0030669 | clathrin-coated endocytic vesicle membrane(GO:0030669) |
| 0.0 | 0.1 | GO:0010370 | perinucleolar chromocenter(GO:0010370) |
| 0.0 | 0.1 | GO:0044666 | MLL3/4 complex(GO:0044666) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.2 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 1.3 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.0 | 3.9 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 2.3 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.3 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.0 | 1.7 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 0.3 | PID ARF 3PATHWAY | Arf1 pathway |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.7 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION IN TLR7 8 OR 9 SIGNALING | Genes involved in TRAF6 mediated IRF7 activation in TLR7/8 or 9 signaling |
| 0.1 | 2.4 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.1 | 1.7 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 1.6 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.1 | 0.7 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.0 | 4.4 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.2 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.0 | 2.3 | REACTOME RESPIRATORY ELECTRON TRANSPORT ATP SYNTHESIS BY CHEMIOSMOTIC COUPLING AND HEAT PRODUCTION BY UNCOUPLING PROTEINS | Genes involved in Respiratory electron transport, ATP synthesis by chemiosmotic coupling, and heat production by uncoupling proteins. |
| 0.0 | 0.3 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.0 | 0.5 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.0 | 0.9 | REACTOME NITRIC OXIDE STIMULATES GUANYLATE CYCLASE | Genes involved in Nitric oxide stimulates guanylate cyclase |
| 0.0 | 1.8 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.2 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.0 | 0.2 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.0 | 0.8 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.2 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.3 | GO:0004698 | calcium-dependent protein kinase C activity(GO:0004698) |
| 0.3 | 2.3 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 0.1 | 0.3 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.1 | 1.7 | GO:0003857 | 3-hydroxyacyl-CoA dehydrogenase activity(GO:0003857) |
| 0.1 | 2.4 | GO:0019198 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.1 | 1.8 | GO:0023026 | MHC class II protein complex binding(GO:0023026) |
| 0.1 | 1.1 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.1 | 0.2 | GO:0004348 | glucosylceramidase activity(GO:0004348) |
| 0.1 | 0.5 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.0 | 0.7 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) |
| 0.0 | 0.7 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.0 | 0.8 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.0 | 0.2 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.8 | GO:0008175 | tRNA methyltransferase activity(GO:0008175) |
| 0.0 | 0.7 | GO:0070008 | serine-type exopeptidase activity(GO:0070008) |
| 0.0 | 0.2 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.0 | 0.1 | GO:0000035 | acyl binding(GO:0000035) phosphopantetheine binding(GO:0031177) |
| 0.0 | 0.7 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.3 | GO:0089720 | caspase binding(GO:0089720) |
| 0.0 | 0.1 | GO:0048101 | calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.0 | 0.1 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.0 | 0.5 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 1.7 | GO:0000979 | RNA polymerase II core promoter sequence-specific DNA binding(GO:0000979) |
| 0.0 | 0.2 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.0 | 0.7 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 1.2 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.0 | 0.3 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.0 | 0.1 | GO:0023024 | MHC class I protein complex binding(GO:0023024) |