Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
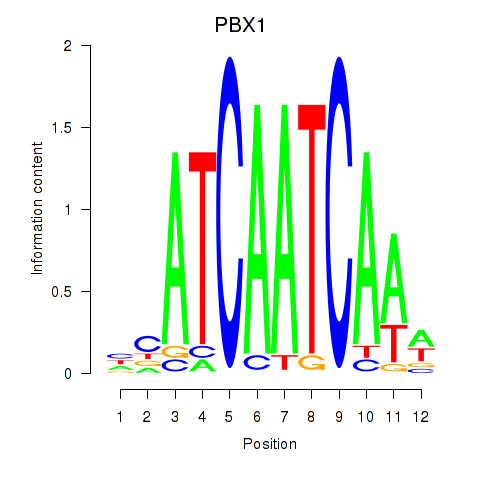
Results for PBX1
Z-value: 1.21
Transcription factors associated with PBX1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PBX1
|
ENSG00000185630.14 | PBX1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PBX1 | hg19_v2_chr1_+_164529004_164529060 | -0.30 | 2.6e-01 | Click! |
Activity profile of PBX1 motif
Sorted Z-values of PBX1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PBX1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_2162162 | 2.50 |
ENST00000381389.1 |
IGF2 |
insulin-like growth factor 2 (somatomedin A) |
| chr13_-_46679185 | 2.11 |
ENST00000439329.3 |
CPB2 |
carboxypeptidase B2 (plasma) |
| chr13_-_46679144 | 2.08 |
ENST00000181383.4 |
CPB2 |
carboxypeptidase B2 (plasma) |
| chr7_+_120629653 | 1.98 |
ENST00000450913.2 ENST00000340646.5 |
CPED1 |
cadherin-like and PC-esterase domain containing 1 |
| chr17_+_1665345 | 1.87 |
ENST00000576406.1 ENST00000571149.1 |
SERPINF1 |
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 |
| chr4_+_69962185 | 1.85 |
ENST00000305231.7 |
UGT2B7 |
UDP glucuronosyltransferase 2 family, polypeptide B7 |
| chr18_+_29171689 | 1.83 |
ENST00000237014.3 |
TTR |
transthyretin |
| chr12_+_81471816 | 1.78 |
ENST00000261206.3 |
ACSS3 |
acyl-CoA synthetase short-chain family member 3 |
| chr4_+_69962212 | 1.76 |
ENST00000508661.1 |
UGT2B7 |
UDP glucuronosyltransferase 2 family, polypeptide B7 |
| chr5_-_42825983 | 1.76 |
ENST00000506577.1 |
SEPP1 |
selenoprotein P, plasma, 1 |
| chr17_-_26694979 | 1.71 |
ENST00000438614.1 |
VTN |
vitronectin |
| chr2_-_200323414 | 1.69 |
ENST00000443023.1 |
SATB2 |
SATB homeobox 2 |
| chr17_-_26695013 | 1.67 |
ENST00000555059.2 |
CTB-96E2.2 |
Homeobox protein SEBOX |
| chr2_-_200322723 | 1.52 |
ENST00000417098.1 |
SATB2 |
SATB homeobox 2 |
| chr2_-_88427568 | 1.51 |
ENST00000393750.3 ENST00000295834.3 |
FABP1 |
fatty acid binding protein 1, liver |
| chr8_-_108510224 | 1.49 |
ENST00000517746.1 ENST00000297450.3 |
ANGPT1 |
angiopoietin 1 |
| chr8_-_91095099 | 1.35 |
ENST00000265431.3 |
CALB1 |
calbindin 1, 28kDa |
| chr1_-_173176452 | 1.33 |
ENST00000281834.3 |
TNFSF4 |
tumor necrosis factor (ligand) superfamily, member 4 |
| chr22_-_29137771 | 1.28 |
ENST00000439200.1 ENST00000405598.1 ENST00000398017.2 ENST00000425190.2 ENST00000348295.3 ENST00000382578.1 ENST00000382565.1 ENST00000382566.1 ENST00000382580.2 ENST00000328354.6 |
CHEK2 |
checkpoint kinase 2 |
| chr6_+_26199737 | 1.25 |
ENST00000359985.1 |
HIST1H2BF |
histone cluster 1, H2bf |
| chr4_+_69681710 | 1.22 |
ENST00000265403.7 ENST00000458688.2 |
UGT2B10 |
UDP glucuronosyltransferase 2 family, polypeptide B10 |
| chr4_-_70080449 | 1.22 |
ENST00000446444.1 |
UGT2B11 |
UDP glucuronosyltransferase 2 family, polypeptide B11 |
| chr6_-_26199499 | 1.06 |
ENST00000377831.5 |
HIST1H3D |
histone cluster 1, H3d |
| chr11_+_22689648 | 1.01 |
ENST00000278187.3 |
GAS2 |
growth arrest-specific 2 |
| chr12_+_12870055 | 0.99 |
ENST00000228872.4 |
CDKN1B |
cyclin-dependent kinase inhibitor 1B (p27, Kip1) |
| chr11_-_34937858 | 0.95 |
ENST00000278359.5 |
APIP |
APAF1 interacting protein |
| chrX_-_70473957 | 0.95 |
ENST00000373984.3 ENST00000314425.5 ENST00000373982.1 |
ZMYM3 |
zinc finger, MYM-type 3 |
| chr6_-_26124138 | 0.95 |
ENST00000314332.5 ENST00000396984.1 |
HIST1H2BC |
histone cluster 1, H2bc |
| chrX_-_6453159 | 0.94 |
ENST00000381089.3 ENST00000398729.1 |
VCX3A |
variable charge, X-linked 3A |
| chr10_+_124320156 | 0.93 |
ENST00000338354.3 ENST00000344338.3 ENST00000330163.4 ENST00000368909.3 ENST00000368955.3 ENST00000368956.2 |
DMBT1 |
deleted in malignant brain tumors 1 |
| chr15_-_37393406 | 0.87 |
ENST00000338564.5 ENST00000558313.1 ENST00000340545.5 |
MEIS2 |
Meis homeobox 2 |
| chr10_+_124320195 | 0.85 |
ENST00000359586.6 |
DMBT1 |
deleted in malignant brain tumors 1 |
| chr21_+_44313375 | 0.84 |
ENST00000354250.2 ENST00000340344.4 |
NDUFV3 |
NADH dehydrogenase (ubiquinone) flavoprotein 3, 10kDa |
| chrX_-_70474499 | 0.84 |
ENST00000353904.2 |
ZMYM3 |
zinc finger, MYM-type 3 |
| chr4_-_73434498 | 0.83 |
ENST00000286657.4 |
ADAMTS3 |
ADAM metallopeptidase with thrombospondin type 1 motif, 3 |
| chr6_+_26124373 | 0.83 |
ENST00000377791.2 ENST00000602637.1 |
HIST1H2AC |
histone cluster 1, H2ac |
| chr5_+_162864575 | 0.82 |
ENST00000512163.1 ENST00000393929.1 ENST00000340828.2 ENST00000511683.2 ENST00000510097.1 ENST00000511490.2 ENST00000510664.1 |
CCNG1 |
cyclin G1 |
| chr10_+_5488564 | 0.80 |
ENST00000449083.1 ENST00000380359.3 |
NET1 |
neuroepithelial cell transforming 1 |
| chr3_+_139063372 | 0.78 |
ENST00000478464.1 |
MRPS22 |
mitochondrial ribosomal protein S22 |
| chr17_+_58018269 | 0.77 |
ENST00000591035.1 |
RP11-178C3.1 |
Uncharacterized protein |
| chr6_-_26199471 | 0.73 |
ENST00000341023.1 |
HIST1H2AD |
histone cluster 1, H2ad |
| chr2_+_44396000 | 0.72 |
ENST00000409895.4 ENST00000409432.3 ENST00000282412.4 ENST00000378551.2 ENST00000345249.4 |
PPM1B |
protein phosphatase, Mg2+/Mn2+ dependent, 1B |
| chr4_-_140005443 | 0.71 |
ENST00000510408.1 ENST00000420916.2 ENST00000358635.3 |
ELF2 |
E74-like factor 2 (ets domain transcription factor) |
| chr2_+_201997492 | 0.68 |
ENST00000494258.1 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr2_-_96971232 | 0.67 |
ENST00000323853.5 |
SNRNP200 |
small nuclear ribonucleoprotein 200kDa (U5) |
| chr4_-_140005341 | 0.65 |
ENST00000379549.2 ENST00000512627.1 |
ELF2 |
E74-like factor 2 (ets domain transcription factor) |
| chr6_+_27775899 | 0.65 |
ENST00000358739.3 |
HIST1H2AI |
histone cluster 1, H2ai |
| chr14_+_23938891 | 0.64 |
ENST00000408901.3 ENST00000397154.3 ENST00000555128.1 |
NGDN |
neuroguidin, EIF4E binding protein |
| chr5_+_43603229 | 0.64 |
ENST00000344920.4 ENST00000512996.2 |
NNT |
nicotinamide nucleotide transhydrogenase |
| chr10_+_48189612 | 0.63 |
ENST00000453919.1 |
AGAP9 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 9 |
| chr11_-_34938039 | 0.62 |
ENST00000395787.3 |
APIP |
APAF1 interacting protein |
| chr16_+_19079215 | 0.61 |
ENST00000544894.2 ENST00000561858.1 |
COQ7 |
coenzyme Q7 homolog, ubiquinone (yeast) |
| chr11_+_95523621 | 0.59 |
ENST00000325542.5 ENST00000325486.5 ENST00000544522.1 ENST00000541365.1 |
CEP57 |
centrosomal protein 57kDa |
| chr22_-_26961328 | 0.59 |
ENST00000398110.2 |
TPST2 |
tyrosylprotein sulfotransferase 2 |
| chr10_-_101190202 | 0.59 |
ENST00000543866.1 ENST00000370508.5 |
GOT1 |
glutamic-oxaloacetic transaminase 1, soluble |
| chr19_-_49955050 | 0.59 |
ENST00000262265.5 |
PIH1D1 |
PIH1 domain containing 1 |
| chr2_-_27886460 | 0.58 |
ENST00000404798.2 ENST00000405491.1 ENST00000464789.2 ENST00000406540.1 |
SUPT7L |
suppressor of Ty 7 (S. cerevisiae)-like |
| chr11_-_57158109 | 0.58 |
ENST00000525955.1 ENST00000533605.1 ENST00000311862.5 |
PRG2 |
proteoglycan 2, bone marrow (natural killer cell activator, eosinophil granule major basic protein) |
| chr4_+_120056939 | 0.57 |
ENST00000307128.5 |
MYOZ2 |
myozenin 2 |
| chr5_+_65440032 | 0.57 |
ENST00000334121.6 |
SREK1 |
splicing regulatory glutamine/lysine-rich protein 1 |
| chr5_+_43602750 | 0.57 |
ENST00000505678.2 ENST00000512422.1 ENST00000264663.5 |
NNT |
nicotinamide nucleotide transhydrogenase |
| chr2_+_201997595 | 0.53 |
ENST00000470178.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr2_+_27886330 | 0.52 |
ENST00000326019.6 |
SLC4A1AP |
solute carrier family 4 (anion exchanger), member 1, adaptor protein |
| chr16_+_19078960 | 0.52 |
ENST00000568985.1 ENST00000566110.1 |
COQ7 |
coenzyme Q7 homolog, ubiquinone (yeast) |
| chr2_+_20646824 | 0.51 |
ENST00000272233.4 |
RHOB |
ras homolog family member B |
| chr14_+_24583836 | 0.48 |
ENST00000559115.1 ENST00000558215.1 ENST00000557810.1 ENST00000561375.1 ENST00000446197.3 ENST00000559796.1 ENST00000560713.1 ENST00000560901.1 ENST00000559382.1 |
DCAF11 |
DDB1 and CUL4 associated factor 11 |
| chr12_-_121342170 | 0.47 |
ENST00000353487.2 |
SPPL3 |
signal peptide peptidase like 3 |
| chr11_+_34938119 | 0.47 |
ENST00000227868.4 ENST00000430469.2 ENST00000533262.1 |
PDHX |
pyruvate dehydrogenase complex, component X |
| chr2_+_264913 | 0.47 |
ENST00000439645.2 ENST00000405233.1 |
ACP1 |
acid phosphatase 1, soluble |
| chr1_+_226250379 | 0.47 |
ENST00000366815.3 ENST00000366814.3 |
H3F3A |
H3 histone, family 3A |
| chr11_+_95523823 | 0.45 |
ENST00000538658.1 |
CEP57 |
centrosomal protein 57kDa |
| chr5_-_133702761 | 0.44 |
ENST00000521118.1 ENST00000265334.4 ENST00000435211.1 |
CDKL3 |
cyclin-dependent kinase-like 3 |
| chr12_-_76478686 | 0.44 |
ENST00000261182.8 |
NAP1L1 |
nucleosome assembly protein 1-like 1 |
| chr2_+_264869 | 0.42 |
ENST00000272067.6 ENST00000272065.5 ENST00000407983.3 |
ACP1 |
acid phosphatase 1, soluble |
| chrX_+_7810303 | 0.41 |
ENST00000381059.3 ENST00000341408.4 |
VCX |
variable charge, X-linked |
| chr6_+_167704838 | 0.40 |
ENST00000366829.2 |
UNC93A |
unc-93 homolog A (C. elegans) |
| chr9_-_101471479 | 0.37 |
ENST00000259455.2 |
GABBR2 |
gamma-aminobutyric acid (GABA) B receptor, 2 |
| chr11_+_71903169 | 0.37 |
ENST00000393676.3 |
FOLR1 |
folate receptor 1 (adult) |
| chr6_-_26032288 | 0.37 |
ENST00000244661.2 |
HIST1H3B |
histone cluster 1, H3b |
| chr6_-_31782813 | 0.37 |
ENST00000375654.4 |
HSPA1L |
heat shock 70kDa protein 1-like |
| chr22_-_22337146 | 0.36 |
ENST00000398793.2 |
TOP3B |
topoisomerase (DNA) III beta |
| chr15_-_64673630 | 0.35 |
ENST00000558008.1 ENST00000559519.1 ENST00000380258.2 |
KIAA0101 |
KIAA0101 |
| chr16_-_70323422 | 0.34 |
ENST00000261772.8 |
AARS |
alanyl-tRNA synthetase |
| chr3_+_157827841 | 0.34 |
ENST00000295930.3 ENST00000471994.1 ENST00000464171.1 ENST00000312179.6 ENST00000475278.2 |
RSRC1 |
arginine/serine-rich coiled-coil 1 |
| chr2_-_27886676 | 0.34 |
ENST00000337768.5 |
SUPT7L |
suppressor of Ty 7 (S. cerevisiae)-like |
| chr15_-_64673665 | 0.34 |
ENST00000300035.4 |
KIAA0101 |
KIAA0101 |
| chr19_-_3801789 | 0.33 |
ENST00000590849.1 ENST00000395045.2 |
MATK |
megakaryocyte-associated tyrosine kinase |
| chr5_-_179045199 | 0.32 |
ENST00000523921.1 |
HNRNPH1 |
heterogeneous nuclear ribonucleoprotein H1 (H) |
| chr6_-_110736742 | 0.32 |
ENST00000368924.3 ENST00000368923.3 |
DDO |
D-aspartate oxidase |
| chr14_-_36789783 | 0.32 |
ENST00000605579.1 ENST00000604336.1 ENST00000359527.7 ENST00000603139.1 ENST00000318473.7 |
MBIP |
MAP3K12 binding inhibitory protein 1 |
| chr5_-_10761206 | 0.32 |
ENST00000432074.2 ENST00000230895.6 |
DAP |
death-associated protein |
| chr19_+_19030497 | 0.31 |
ENST00000438170.2 |
DDX49 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 49 |
| chr3_+_120461484 | 0.31 |
ENST00000484715.1 ENST00000469772.1 ENST00000283875.5 ENST00000492959.1 |
GTF2E1 |
general transcription factor IIE, polypeptide 1, alpha 56kDa |
| chrX_+_120181457 | 0.31 |
ENST00000328078.1 |
GLUD2 |
glutamate dehydrogenase 2 |
| chr10_+_5238793 | 0.29 |
ENST00000263126.1 |
AKR1C4 |
aldo-keto reductase family 1, member C4 |
| chr13_+_97928395 | 0.29 |
ENST00000445661.2 |
MBNL2 |
muscleblind-like splicing regulator 2 |
| chr1_-_28241024 | 0.28 |
ENST00000313433.7 ENST00000444045.1 |
RPA2 |
replication protein A2, 32kDa |
| chr19_+_782755 | 0.28 |
ENST00000606242.1 ENST00000586061.1 |
AC006273.5 |
AC006273.5 |
| chr19_+_19030478 | 0.27 |
ENST00000247003.4 |
DDX49 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 49 |
| chr12_-_122985067 | 0.27 |
ENST00000540586.1 ENST00000543897.1 |
ZCCHC8 |
zinc finger, CCHC domain containing 8 |
| chr22_-_22337204 | 0.25 |
ENST00000430142.1 ENST00000357179.5 |
TOP3B |
topoisomerase (DNA) III beta |
| chr14_+_50234309 | 0.25 |
ENST00000298307.5 |
KLHDC2 |
kelch domain containing 2 |
| chr5_-_86708833 | 0.23 |
ENST00000256897.4 |
CCNH |
cyclin H |
| chr16_+_19078911 | 0.23 |
ENST00000321998.5 |
COQ7 |
coenzyme Q7 homolog, ubiquinone (yeast) |
| chr19_-_33360647 | 0.22 |
ENST00000590341.1 ENST00000587772.1 ENST00000023064.4 |
SLC7A9 |
solute carrier family 7 (amino acid transporter light chain, bo,+ system), member 9 |
| chr12_+_56401268 | 0.22 |
ENST00000262032.5 |
IKZF4 |
IKAROS family zinc finger 4 (Eos) |
| chr8_-_95487331 | 0.22 |
ENST00000336148.5 |
RAD54B |
RAD54 homolog B (S. cerevisiae) |
| chr2_-_109605663 | 0.22 |
ENST00000409271.1 ENST00000258443.2 ENST00000376651.1 |
EDAR |
ectodysplasin A receptor |
| chr4_-_168155169 | 0.22 |
ENST00000534949.1 ENST00000535728.1 |
SPOCK3 |
sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 3 |
| chrX_+_140677562 | 0.22 |
ENST00000370518.3 |
SPANXA2 |
SPANX family, member A2 |
| chr8_+_98788003 | 0.21 |
ENST00000521545.2 |
LAPTM4B |
lysosomal protein transmembrane 4 beta |
| chr1_-_1690014 | 0.20 |
ENST00000400922.2 ENST00000342348.5 |
NADK |
NAD kinase |
| chr15_+_44069276 | 0.20 |
ENST00000381359.1 |
SERF2 |
small EDRK-rich factor 2 |
| chr2_+_201997676 | 0.20 |
ENST00000462763.1 ENST00000479953.2 |
CFLAR |
CASP8 and FADD-like apoptosis regulator |
| chr19_-_49121054 | 0.19 |
ENST00000546623.1 ENST00000084795.5 |
RPL18 |
ribosomal protein L18 |
| chr11_+_31531291 | 0.19 |
ENST00000350638.5 ENST00000379163.5 ENST00000395934.2 |
ELP4 |
elongator acetyltransferase complex subunit 4 |
| chr19_-_16008880 | 0.19 |
ENST00000011989.7 ENST00000221700.6 |
CYP4F2 |
cytochrome P450, family 4, subfamily F, polypeptide 2 |
| chr4_-_70626430 | 0.19 |
ENST00000310613.3 |
SULT1B1 |
sulfotransferase family, cytosolic, 1B, member 1 |
| chr5_-_86708670 | 0.19 |
ENST00000504878.1 |
CCNH |
cyclin H |
| chrM_+_4431 | 0.19 |
ENST00000361453.3 |
MT-ND2 |
mitochondrially encoded NADH dehydrogenase 2 |
| chr14_+_22985251 | 0.18 |
ENST00000390510.1 |
TRAJ27 |
T cell receptor alpha joining 27 |
| chrX_-_92928557 | 0.18 |
ENST00000373079.3 ENST00000475430.2 |
NAP1L3 |
nucleosome assembly protein 1-like 3 |
| chr3_+_148508845 | 0.18 |
ENST00000491148.1 |
CPB1 |
carboxypeptidase B1 (tissue) |
| chr9_-_73736511 | 0.18 |
ENST00000377110.3 ENST00000377111.2 |
TRPM3 |
transient receptor potential cation channel, subfamily M, member 3 |
| chr3_+_155860751 | 0.17 |
ENST00000471742.1 |
KCNAB1 |
potassium voltage-gated channel, shaker-related subfamily, beta member 1 |
| chr11_-_104035088 | 0.17 |
ENST00000302251.5 |
PDGFD |
platelet derived growth factor D |
| chrX_-_102757802 | 0.17 |
ENST00000372633.1 |
RAB40A |
RAB40A, member RAS oncogene family |
| chr3_+_129159039 | 0.17 |
ENST00000507564.1 ENST00000431818.2 ENST00000504021.1 ENST00000349441.2 ENST00000348417.2 ENST00000440957.2 |
IFT122 |
intraflagellar transport 122 homolog (Chlamydomonas) |
| chr2_+_169926047 | 0.17 |
ENST00000428522.1 ENST00000450153.1 ENST00000421653.1 |
DHRS9 |
dehydrogenase/reductase (SDR family) member 9 |
| chr3_+_120626919 | 0.17 |
ENST00000273666.6 ENST00000471454.1 ENST00000472879.1 ENST00000497029.1 ENST00000492541.1 |
STXBP5L |
syntaxin binding protein 5-like |
| chr12_-_56615693 | 0.16 |
ENST00000394013.2 ENST00000345093.4 ENST00000551711.1 ENST00000552656.1 |
RNF41 |
ring finger protein 41 |
| chr14_+_29236269 | 0.16 |
ENST00000313071.4 |
FOXG1 |
forkhead box G1 |
| chr5_+_54398463 | 0.16 |
ENST00000274306.6 |
GZMA |
granzyme A (granzyme 1, cytotoxic T-lymphocyte-associated serine esterase 3) |
| chr11_+_66384053 | 0.16 |
ENST00000310137.4 ENST00000393979.3 ENST00000409372.1 ENST00000443702.1 ENST00000409738.4 ENST00000514361.3 ENST00000503028.2 ENST00000412278.2 ENST00000500635.2 |
RBM14 RBM4 RBM14-RBM4 |
RNA binding motif protein 14 RNA binding motif protein 4 RBM14-RBM4 readthrough |
| chr10_+_114709999 | 0.15 |
ENST00000355995.4 ENST00000545257.1 ENST00000543371.1 ENST00000536810.1 ENST00000355717.4 ENST00000538897.1 ENST00000534894.1 |
TCF7L2 |
transcription factor 7-like 2 (T-cell specific, HMG-box) |
| chr17_+_46125707 | 0.15 |
ENST00000584137.1 ENST00000362042.3 ENST00000585291.1 ENST00000357480.5 |
NFE2L1 |
nuclear factor, erythroid 2-like 1 |
| chr5_+_140207536 | 0.15 |
ENST00000529310.1 ENST00000527624.1 |
PCDHA6 |
protocadherin alpha 6 |
| chr3_+_129158926 | 0.15 |
ENST00000347300.2 ENST00000296266.3 |
IFT122 |
intraflagellar transport 122 homolog (Chlamydomonas) |
| chr1_+_205473720 | 0.15 |
ENST00000429964.2 ENST00000506784.1 ENST00000360066.2 |
CDK18 |
cyclin-dependent kinase 18 |
| chr22_+_35695793 | 0.14 |
ENST00000456128.1 ENST00000449058.2 ENST00000411850.1 ENST00000425375.1 ENST00000436462.2 ENST00000382034.5 |
TOM1 |
target of myb1 (chicken) |
| chr9_-_6015607 | 0.14 |
ENST00000259569.5 |
RANBP6 |
RAN binding protein 6 |
| chr6_-_7313381 | 0.14 |
ENST00000489567.1 ENST00000479365.1 ENST00000462112.1 ENST00000397511.2 ENST00000534851.1 ENST00000474597.1 ENST00000244763.4 |
SSR1 |
signal sequence receptor, alpha |
| chr17_+_45973516 | 0.14 |
ENST00000376741.4 |
SP2 |
Sp2 transcription factor |
| chr1_-_115323245 | 0.14 |
ENST00000060969.5 ENST00000369528.5 |
SIKE1 |
suppressor of IKBKE 1 |
| chr16_-_20364030 | 0.13 |
ENST00000396134.2 ENST00000573567.1 ENST00000570757.1 ENST00000424589.1 ENST00000302509.4 ENST00000571174.1 ENST00000576688.1 |
UMOD |
uromodulin |
| chrX_-_140673133 | 0.13 |
ENST00000370519.3 |
SPANXA1 |
sperm protein associated with the nucleus, X-linked, family member A1 |
| chr14_-_60097297 | 0.13 |
ENST00000395090.1 |
RTN1 |
reticulon 1 |
| chr2_+_12857015 | 0.13 |
ENST00000155926.4 |
TRIB2 |
tribbles pseudokinase 2 |
| chr3_+_193853927 | 0.12 |
ENST00000232424.3 |
HES1 |
hes family bHLH transcription factor 1 |
| chr6_-_27775694 | 0.12 |
ENST00000377401.2 |
HIST1H2BL |
histone cluster 1, H2bl |
| chr3_-_33481835 | 0.12 |
ENST00000283629.3 |
UBP1 |
upstream binding protein 1 (LBP-1a) |
| chr19_-_3978083 | 0.12 |
ENST00000600794.1 |
EEF2 |
eukaryotic translation elongation factor 2 |
| chr11_+_68228186 | 0.12 |
ENST00000393799.2 ENST00000393800.2 ENST00000528635.1 ENST00000533127.1 ENST00000529907.1 ENST00000529344.1 ENST00000534534.1 ENST00000524845.1 ENST00000265637.4 ENST00000524904.1 ENST00000393801.3 ENST00000265636.5 ENST00000529710.1 |
PPP6R3 |
protein phosphatase 6, regulatory subunit 3 |
| chr1_+_156589051 | 0.11 |
ENST00000255039.1 |
HAPLN2 |
hyaluronan and proteoglycan link protein 2 |
| chr6_-_26250835 | 0.11 |
ENST00000446824.2 |
HIST1H3F |
histone cluster 1, H3f |
| chr12_-_54653313 | 0.11 |
ENST00000550411.1 ENST00000439541.2 |
CBX5 |
chromobox homolog 5 |
| chr2_+_27255806 | 0.11 |
ENST00000238788.9 ENST00000404032.3 |
TMEM214 |
transmembrane protein 214 |
| chr11_-_114466477 | 0.11 |
ENST00000375478.3 |
NXPE4 |
neurexophilin and PC-esterase domain family, member 4 |
| chr1_-_3713042 | 0.11 |
ENST00000378251.1 |
LRRC47 |
leucine rich repeat containing 47 |
| chr12_-_56848426 | 0.11 |
ENST00000257979.4 |
MIP |
major intrinsic protein of lens fiber |
| chr11_+_61717336 | 0.11 |
ENST00000378042.3 |
BEST1 |
bestrophin 1 |
| chr12_+_101988627 | 0.11 |
ENST00000547405.1 ENST00000452455.2 ENST00000441232.1 ENST00000360610.2 ENST00000392934.3 ENST00000547509.1 ENST00000361685.2 ENST00000549145.1 ENST00000553190.1 |
MYBPC1 |
myosin binding protein C, slow type |
| chr15_+_69222909 | 0.10 |
ENST00000455873.3 |
NOX5 |
NADPH oxidase, EF-hand calcium binding domain 5 |
| chrX_+_140084756 | 0.10 |
ENST00000449283.1 |
SPANXB2 |
SPANX family, member B2 |
| chr6_+_151561506 | 0.10 |
ENST00000253332.1 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr11_+_61717279 | 0.09 |
ENST00000378043.4 |
BEST1 |
bestrophin 1 |
| chr6_+_27114861 | 0.09 |
ENST00000377459.1 |
HIST1H2AH |
histone cluster 1, H2ah |
| chr12_-_57472522 | 0.09 |
ENST00000379391.3 ENST00000300128.4 |
TMEM194A |
transmembrane protein 194A |
| chr6_-_27114577 | 0.08 |
ENST00000356950.1 ENST00000396891.4 |
HIST1H2BK |
histone cluster 1, H2bk |
| chr5_-_59481406 | 0.08 |
ENST00000546160.1 |
PDE4D |
phosphodiesterase 4D, cAMP-specific |
| chr13_-_33760216 | 0.08 |
ENST00000255486.4 |
STARD13 |
StAR-related lipid transfer (START) domain containing 13 |
| chr16_+_16484691 | 0.07 |
ENST00000344087.4 |
NPIPA7 |
nuclear pore complex interacting protein family, member A7 |
| chr12_-_91573249 | 0.07 |
ENST00000550099.1 ENST00000546391.1 ENST00000551354.1 |
DCN |
decorin |
| chr1_-_28241226 | 0.07 |
ENST00000373912.3 ENST00000373909.3 |
RPA2 |
replication protein A2, 32kDa |
| chr16_+_50099852 | 0.07 |
ENST00000299192.7 ENST00000285767.4 |
HEATR3 |
HEAT repeat containing 3 |
| chr19_-_54804173 | 0.07 |
ENST00000391744.3 ENST00000251390.3 |
LILRA3 |
leukocyte immunoglobulin-like receptor, subfamily A (without TM domain), member 3 |
| chr16_-_20364122 | 0.07 |
ENST00000396138.4 ENST00000577168.1 |
UMOD |
uromodulin |
| chr12_+_101988774 | 0.07 |
ENST00000545503.2 ENST00000536007.1 ENST00000541119.1 ENST00000361466.2 ENST00000551300.1 ENST00000550270.1 |
MYBPC1 |
myosin binding protein C, slow type |
| chr9_-_118417 | 0.07 |
ENST00000382500.2 |
FOXD4 |
forkhead box D4 |
| chr19_+_11485333 | 0.07 |
ENST00000312423.2 |
SWSAP1 |
SWIM-type zinc finger 7 associated protein 1 |
| chr12_+_21168630 | 0.07 |
ENST00000421593.2 |
SLCO1B7 |
solute carrier organic anion transporter family, member 1B7 (non-functional) |
| chr7_+_107110488 | 0.06 |
ENST00000304402.4 |
GPR22 |
G protein-coupled receptor 22 |
| chr3_-_187388173 | 0.06 |
ENST00000287641.3 |
SST |
somatostatin |
| chr2_+_211342432 | 0.06 |
ENST00000430249.2 |
CPS1 |
carbamoyl-phosphate synthase 1, mitochondrial |
| chr4_-_185776789 | 0.06 |
ENST00000510284.1 |
RP11-701P16.5 |
RP11-701P16.5 |
| chr4_+_37962018 | 0.06 |
ENST00000504686.1 |
PTTG2 |
pituitary tumor-transforming 2 |
| chr9_+_106856831 | 0.05 |
ENST00000303219.8 ENST00000374787.3 |
SMC2 |
structural maintenance of chromosomes 2 |
| chr4_+_175839551 | 0.05 |
ENST00000404450.4 ENST00000514159.1 |
ADAM29 |
ADAM metallopeptidase domain 29 |
| chr2_+_12857043 | 0.04 |
ENST00000381465.2 |
TRIB2 |
tribbles pseudokinase 2 |
| chr18_+_22040593 | 0.04 |
ENST00000256906.4 |
HRH4 |
histamine receptor H4 |
| chr5_-_177210399 | 0.04 |
ENST00000510276.1 |
FAM153A |
family with sequence similarity 153, member A |
| chr4_-_113437328 | 0.04 |
ENST00000313341.3 |
NEUROG2 |
neurogenin 2 |
| chr7_-_7782204 | 0.03 |
ENST00000418534.2 |
AC007161.5 |
AC007161.5 |
| chr21_-_35014027 | 0.03 |
ENST00000399442.1 ENST00000413017.2 ENST00000445393.1 ENST00000417979.1 ENST00000426935.1 ENST00000381540.3 ENST00000361534.2 ENST00000381554.3 |
CRYZL1 |
crystallin, zeta (quinone reductase)-like 1 |
| chr10_+_60028818 | 0.03 |
ENST00000333926.5 |
CISD1 |
CDGSH iron sulfur domain 1 |
| chr19_+_10828724 | 0.03 |
ENST00000585892.1 ENST00000314646.5 ENST00000359692.6 |
DNM2 |
dynamin 2 |
| chr4_-_105416039 | 0.02 |
ENST00000394767.2 |
CXXC4 |
CXXC finger protein 4 |
| chr2_+_233897382 | 0.02 |
ENST00000233840.3 |
NEU2 |
sialidase 2 (cytosolic sialidase) |
| chr1_-_13673511 | 0.02 |
ENST00000344998.3 ENST00000334600.6 |
PRAMEF14 |
PRAME family member 14 |
| chr17_-_26941179 | 0.01 |
ENST00000301037.5 ENST00000530121.1 ENST00000525510.1 ENST00000577790.1 ENST00000531839.1 ENST00000534850.1 |
SGK494 RP11-192H23.4 |
uncharacterized serine/threonine-protein kinase SgK494 Uncharacterized protein |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.5 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.2 | 0.6 | GO:0008476 | protein-tyrosine sulfotransferase activity(GO:0008476) |
| 0.2 | 0.6 | GO:0070546 | phosphatidylserine decarboxylase activity(GO:0004609) L-phenylalanine aminotransferase activity(GO:0070546) L-phenylalanine:2-oxoglutarate aminotransferase activity(GO:0080130) |
| 0.2 | 1.8 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.2 | 0.3 | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor(GO:0016639) |
| 0.1 | 4.2 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.1 | 0.4 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.1 | 0.7 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.1 | 0.5 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.1 | 0.6 | GO:0051373 | FATZ binding(GO:0051373) |
| 0.1 | 0.3 | GO:0004813 | alanine-tRNA ligase activity(GO:0004813) |
| 0.1 | 0.3 | GO:0030366 | molybdopterin synthase activity(GO:0030366) |
| 0.1 | 1.7 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.1 | 1.7 | GO:0016405 | CoA-ligase activity(GO:0016405) |
| 0.1 | 0.6 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.1 | 0.4 | GO:0061714 | folic acid receptor activity(GO:0061714) |
| 0.1 | 0.6 | GO:0001165 | RNA polymerase I upstream control element sequence-specific DNA binding(GO:0001165) |
| 0.1 | 0.3 | GO:0047023 | androsterone dehydrogenase activity(GO:0047023) |
| 0.1 | 0.2 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.1 | 0.2 | GO:0018685 | alkane 1-monooxygenase activity(GO:0018685) tocopherol omega-hydroxylase activity(GO:0052870) alpha-tocopherol omega-hydroxylase activity(GO:0052871) 20-hydroxy-leukotriene B4 omega oxidase activity(GO:0097258) 20-aldehyde-leukotriene B4 20-monooxygenase activity(GO:0097259) |
| 0.1 | 0.9 | GO:0003993 | acid phosphatase activity(GO:0003993) non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.1 | 2.5 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.1 | 0.5 | GO:0004738 | pyruvate dehydrogenase activity(GO:0004738) pyruvate dehydrogenase [NAD(P)+] activity(GO:0034603) pyruvate dehydrogenase (NAD+) activity(GO:0034604) |
| 0.1 | 2.2 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 1.5 | GO:0008329 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.1 | 1.0 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.1 | 0.2 | GO:0097158 | pre-mRNA intronic pyrimidine-rich binding(GO:0097158) |
| 0.0 | 1.4 | GO:0097200 | cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:0097200) |
| 0.0 | 0.8 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.2 | GO:0015184 | L-cystine transmembrane transporter activity(GO:0015184) |
| 0.0 | 1.0 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.4 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.0 | 0.1 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.0 | 0.4 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 1.6 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 0.3 | GO:0070513 | death domain binding(GO:0070513) |
| 0.0 | 1.7 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.9 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.1 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.0 | 0.3 | GO:0016641 | oxidoreductase activity, acting on the CH-NH2 group of donors, oxygen as acceptor(GO:0016641) |
| 0.0 | 0.2 | GO:0008607 | phosphorylase kinase regulator activity(GO:0008607) |
| 0.0 | 1.0 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 1.9 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.2 | GO:0047035 | alcohol dehydrogenase (NAD) activity(GO:0004022) testosterone dehydrogenase (NAD+) activity(GO:0047035) |
| 0.0 | 1.2 | GO:0050661 | NADP binding(GO:0050661) |
| 0.0 | 0.2 | GO:0031432 | titin binding(GO:0031432) |
| 0.0 | 1.3 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.0 | 0.9 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.0 | 0.2 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.0 | 0.3 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.0 | 0.0 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.7 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 0.6 | GO:0004693 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) |
| 0.0 | 0.2 | GO:0019864 | IgG binding(GO:0019864) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.6 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.1 | 5.8 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.1 | 2.0 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.1 | 1.9 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 1.3 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.1 | 1.0 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.1 | 1.2 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 1.4 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 1.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 1.8 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 1.0 | REACTOME CASPASE MEDIATED CLEAVAGE OF CYTOSKELETAL PROTEINS | Genes involved in Caspase-mediated cleavage of cytoskeletal proteins |
| 0.0 | 0.5 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.0 | 0.4 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.4 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.0 | 0.2 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.0 | 1.7 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.5 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.0 | 0.7 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 4.2 | GO:0003330 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.5 | 1.9 | GO:0002215 | defense response to nematode(GO:0002215) |
| 0.4 | 1.3 | GO:0072428 | signal transduction involved in intra-S DNA damage checkpoint(GO:0072428) response to bisphenol A(GO:1903925) cellular response to bisphenol A(GO:1903926) |
| 0.4 | 1.9 | GO:0071279 | cellular response to cobalt ion(GO:0071279) |
| 0.3 | 3.2 | GO:0021902 | commitment of neuronal cell to specific neuron type in forebrain(GO:0021902) |
| 0.3 | 2.5 | GO:0038028 | insulin receptor signaling pathway via phosphatidylinositol 3-kinase(GO:0038028) |
| 0.2 | 1.2 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.2 | 1.6 | GO:0070269 | pyroptosis(GO:0070269) |
| 0.2 | 0.7 | GO:0000354 | cis assembly of pre-catalytic spliceosome(GO:0000354) |
| 0.2 | 1.5 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.2 | 0.6 | GO:0006478 | peptidyl-tyrosine sulfation(GO:0006478) |
| 0.2 | 0.6 | GO:0006106 | fumarate metabolic process(GO:0006106) aspartate catabolic process(GO:0006533) |
| 0.2 | 0.6 | GO:1903939 | negative regulation of histone H3-K9 dimethylation(GO:1900110) regulation of TORC2 signaling(GO:1903939) |
| 0.2 | 1.7 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.2 | 1.8 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.2 | 1.0 | GO:0060770 | negative regulation of epithelial cell proliferation involved in prostate gland development(GO:0060770) |
| 0.2 | 1.6 | GO:0072025 | distal convoluted tubule development(GO:0072025) metanephric distal convoluted tubule development(GO:0072221) |
| 0.2 | 1.5 | GO:0032000 | positive regulation of fatty acid beta-oxidation(GO:0032000) |
| 0.1 | 1.8 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.1 | 1.4 | GO:1903943 | regulation of hepatocyte apoptotic process(GO:1903943) negative regulation of hepatocyte apoptotic process(GO:1903944) |
| 0.1 | 1.8 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 0.8 | GO:1900748 | positive regulation of vascular endothelial growth factor signaling pathway(GO:1900748) |
| 0.1 | 0.4 | GO:0060974 | neural crest cell migration involved in heart formation(GO:0003147) cell migration involved in heart formation(GO:0060974) anterior neural tube closure(GO:0061713) |
| 0.1 | 0.7 | GO:0006499 | N-terminal protein myristoylation(GO:0006499) |
| 0.1 | 0.5 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 0.1 | 0.3 | GO:0006419 | alanyl-tRNA aminoacylation(GO:0006419) |
| 0.1 | 0.3 | GO:0007231 | osmosensory signaling pathway(GO:0007231) |
| 0.1 | 0.3 | GO:0007227 | signal transduction downstream of smoothened(GO:0007227) |
| 0.1 | 4.1 | GO:0008209 | androgen metabolic process(GO:0008209) |
| 0.1 | 1.4 | GO:0006744 | ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.1 | 0.8 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.1 | 0.2 | GO:0003095 | pressure natriuresis(GO:0003095) negative regulation of icosanoid secretion(GO:0032304) |
| 0.1 | 0.1 | GO:0060164 | regulation of timing of neuron differentiation(GO:0060164) |
| 0.1 | 0.5 | GO:0051533 | positive regulation of NFAT protein import into nucleus(GO:0051533) |
| 0.1 | 0.2 | GO:2000439 | positive regulation of monocyte extravasation(GO:2000439) |
| 0.1 | 1.0 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 0.1 | 0.3 | GO:0006531 | aspartate metabolic process(GO:0006531) |
| 0.1 | 0.5 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 0.0 | 0.3 | GO:0006537 | glutamate biosynthetic process(GO:0006537) |
| 0.0 | 0.4 | GO:1901409 | positive regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901409) |
| 0.0 | 1.2 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 0.0 | 1.4 | GO:0050855 | regulation of B cell receptor signaling pathway(GO:0050855) |
| 0.0 | 0.2 | GO:0015811 | L-cystine transport(GO:0015811) |
| 0.0 | 0.6 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.0 | 0.2 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.0 | 0.2 | GO:0042091 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.0 | 0.8 | GO:0051457 | maintenance of protein location in nucleus(GO:0051457) |
| 0.0 | 0.1 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.0 | 0.6 | GO:0061154 | endothelial tube morphogenesis(GO:0061154) |
| 0.0 | 0.2 | GO:0010908 | regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010908) positive regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010909) canonical Wnt signaling pathway involved in positive regulation of epithelial to mesenchymal transition(GO:0044334) positive regulation of proteoglycan biosynthetic process(GO:1902730) |
| 0.0 | 0.7 | GO:0034453 | microtubule anchoring(GO:0034453) |
| 0.0 | 0.1 | GO:0008065 | establishment of blood-nerve barrier(GO:0008065) |
| 0.0 | 0.2 | GO:1901525 | negative regulation of macromitophagy(GO:1901525) |
| 0.0 | 0.2 | GO:0032071 | regulation of endodeoxyribonuclease activity(GO:0032071) |
| 0.0 | 0.4 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.0 | 0.2 | GO:0097167 | circadian regulation of translation(GO:0097167) |
| 0.0 | 0.6 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 1.5 | GO:0006342 | chromatin silencing(GO:0006342) |
| 0.0 | 1.2 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.0 | 0.1 | GO:0008628 | hormone-mediated apoptotic signaling pathway(GO:0008628) |
| 0.0 | 0.1 | GO:0034201 | response to oleic acid(GO:0034201) |
| 0.0 | 0.2 | GO:0030321 | transepithelial chloride transport(GO:0030321) |
| 0.0 | 0.1 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.0 | 0.3 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 0.6 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.0 | 0.2 | GO:0071688 | striated muscle myosin thick filament assembly(GO:0071688) |
| 0.0 | 0.9 | GO:0008542 | visual learning(GO:0008542) |
| 0.0 | 0.2 | GO:1902260 | diaphragm development(GO:0060539) negative regulation of delayed rectifier potassium channel activity(GO:1902260) |
| 0.0 | 0.4 | GO:0030517 | negative regulation of axon extension(GO:0030517) |
| 0.0 | 0.7 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.8 | GO:0070126 | mitochondrial translational elongation(GO:0070125) mitochondrial translational termination(GO:0070126) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.7 | GO:0071062 | rough endoplasmic reticulum lumen(GO:0048237) alphav-beta3 integrin-vitronectin complex(GO:0071062) |
| 0.2 | 1.9 | GO:0043203 | axon hillock(GO:0043203) |
| 0.2 | 1.5 | GO:0045179 | apical cortex(GO:0045179) |
| 0.1 | 0.4 | GO:0038039 | G-protein coupled receptor heterodimeric complex(GO:0038039) |
| 0.1 | 0.4 | GO:0070985 | TFIIK complex(GO:0070985) |
| 0.1 | 1.8 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.1 | 1.0 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.1 | 1.4 | GO:0097342 | ripoptosome(GO:0097342) |
| 0.1 | 0.5 | GO:0001740 | Barr body(GO:0001740) |
| 0.1 | 0.9 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.1 | 0.6 | GO:0070761 | pre-snoRNP complex(GO:0070761) |
| 0.1 | 0.5 | GO:0098554 | cytoplasmic side of endoplasmic reticulum membrane(GO:0098554) |
| 0.1 | 0.3 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.1 | 2.6 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.5 | GO:0045254 | pyruvate dehydrogenase complex(GO:0045254) |
| 0.0 | 3.3 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 0.8 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.2 | GO:0030667 | secretory granule membrane(GO:0030667) |
| 0.0 | 0.4 | GO:0002199 | zona pellucida receptor complex(GO:0002199) |
| 0.0 | 0.3 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.0 | 0.2 | GO:0070369 | beta-catenin-TCF7L2 complex(GO:0070369) |
| 0.0 | 0.2 | GO:1990635 | proximal dendrite(GO:1990635) |
| 0.0 | 0.4 | GO:0031362 | anchored component of external side of plasma membrane(GO:0031362) |
| 0.0 | 0.7 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 0.5 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.2 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.0 | 2.5 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.0 | 0.3 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.0 | 0.6 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.0 | 1.9 | GO:0005746 | mitochondrial respiratory chain(GO:0005746) |
| 0.0 | 1.5 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.2 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 1.2 | GO:0035578 | azurophil granule lumen(GO:0035578) |
| 0.0 | 0.1 | GO:0042788 | polysomal ribosome(GO:0042788) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.7 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 1.4 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 1.0 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.0 | 2.2 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 2.5 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 2.1 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 0.9 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
| 0.0 | 0.9 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.3 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 1.0 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.8 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 0.4 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.0 | 2.7 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |