Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
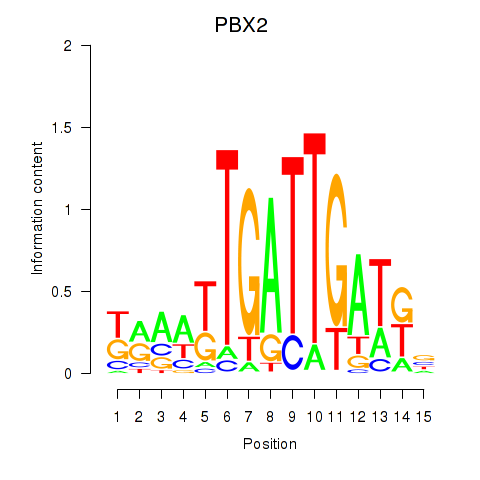
Results for PBX2
Z-value: 0.61
Transcription factors associated with PBX2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PBX2
|
ENSG00000204304.7 | PBX2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PBX2 | hg19_v2_chr6_-_32157947_32157992 | -0.63 | 9.2e-03 | Click! |
Activity profile of PBX2 motif
Sorted Z-values of PBX2 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PBX2
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_-_6420930 | 1.49 |
ENST00000325203.5 |
ANGPT2 |
angiopoietin 2 |
| chr2_-_190044480 | 1.48 |
ENST00000374866.3 |
COL5A2 |
collagen, type V, alpha 2 |
| chr12_-_28124903 | 1.32 |
ENST00000395872.1 ENST00000354417.3 ENST00000201015.4 |
PTHLH |
parathyroid hormone-like hormone |
| chr2_+_234601512 | 0.88 |
ENST00000305139.6 |
UGT1A6 |
UDP glucuronosyltransferase 1 family, polypeptide A6 |
| chr1_+_101702417 | 0.87 |
ENST00000305352.6 |
S1PR1 |
sphingosine-1-phosphate receptor 1 |
| chr2_+_234602305 | 0.72 |
ENST00000406651.1 |
UGT1A6 |
UDP glucuronosyltransferase 1 family, polypeptide A6 |
| chr7_+_134528635 | 0.69 |
ENST00000445569.2 |
CALD1 |
caldesmon 1 |
| chrX_+_119737806 | 0.65 |
ENST00000371317.5 |
MCTS1 |
malignant T cell amplified sequence 1 |
| chr12_-_28125638 | 0.53 |
ENST00000545234.1 |
PTHLH |
parathyroid hormone-like hormone |
| chr2_+_20646824 | 0.48 |
ENST00000272233.4 |
RHOB |
ras homolog family member B |
| chr13_-_107220455 | 0.44 |
ENST00000400198.3 |
ARGLU1 |
arginine and glutamate rich 1 |
| chr2_+_33661382 | 0.36 |
ENST00000402538.3 |
RASGRP3 |
RAS guanyl releasing protein 3 (calcium and DAG-regulated) |
| chr3_+_129158926 | 0.36 |
ENST00000347300.2 ENST00000296266.3 |
IFT122 |
intraflagellar transport 122 homolog (Chlamydomonas) |
| chr3_+_129159039 | 0.34 |
ENST00000507564.1 ENST00000431818.2 ENST00000504021.1 ENST00000349441.2 ENST00000348417.2 ENST00000440957.2 |
IFT122 |
intraflagellar transport 122 homolog (Chlamydomonas) |
| chr14_-_55658323 | 0.32 |
ENST00000554067.1 ENST00000247191.2 |
DLGAP5 |
discs, large (Drosophila) homolog-associated protein 5 |
| chr3_-_129158676 | 0.31 |
ENST00000393278.2 |
MBD4 |
methyl-CpG binding domain protein 4 |
| chr2_+_113033164 | 0.31 |
ENST00000409871.1 ENST00000343936.4 |
ZC3H6 |
zinc finger CCCH-type containing 6 |
| chr15_+_67418047 | 0.28 |
ENST00000540846.2 |
SMAD3 |
SMAD family member 3 |
| chrX_+_46696372 | 0.27 |
ENST00000218340.3 |
RP2 |
retinitis pigmentosa 2 (X-linked recessive) |
| chr2_+_12857043 | 0.24 |
ENST00000381465.2 |
TRIB2 |
tribbles pseudokinase 2 |
| chrX_+_56590002 | 0.24 |
ENST00000338222.5 |
UBQLN2 |
ubiquilin 2 |
| chr3_-_129158850 | 0.23 |
ENST00000503197.1 ENST00000249910.1 ENST00000429544.2 ENST00000507208.1 |
MBD4 |
methyl-CpG binding domain protein 4 |
| chr14_-_55658252 | 0.21 |
ENST00000395425.2 |
DLGAP5 |
discs, large (Drosophila) homolog-associated protein 5 |
| chr12_+_53835383 | 0.21 |
ENST00000429243.2 |
PRR13 |
proline rich 13 |
| chr10_+_124320156 | 0.21 |
ENST00000338354.3 ENST00000344338.3 ENST00000330163.4 ENST00000368909.3 ENST00000368955.3 ENST00000368956.2 |
DMBT1 |
deleted in malignant brain tumors 1 |
| chr11_+_69455855 | 0.20 |
ENST00000227507.2 ENST00000536559.1 |
CCND1 |
cyclin D1 |
| chr10_+_124320195 | 0.20 |
ENST00000359586.6 |
DMBT1 |
deleted in malignant brain tumors 1 |
| chr11_-_111794446 | 0.19 |
ENST00000527950.1 |
CRYAB |
crystallin, alpha B |
| chr17_-_29624343 | 0.18 |
ENST00000247271.4 |
OMG |
oligodendrocyte myelin glycoprotein |
| chr10_+_47894572 | 0.15 |
ENST00000355876.5 |
FAM21B |
family with sequence similarity 21, member B |
| chr2_+_12857015 | 0.15 |
ENST00000155926.4 |
TRIB2 |
tribbles pseudokinase 2 |
| chr9_-_128246769 | 0.15 |
ENST00000444226.1 |
MAPKAP1 |
mitogen-activated protein kinase associated protein 1 |
| chr12_+_53835539 | 0.14 |
ENST00000547368.1 ENST00000379786.4 ENST00000551945.1 |
PRR13 |
proline rich 13 |
| chr17_+_8191815 | 0.13 |
ENST00000226105.6 ENST00000407006.4 ENST00000580434.1 ENST00000439238.3 |
RANGRF |
RAN guanine nucleotide release factor |
| chr17_-_29151794 | 0.07 |
ENST00000324238.6 |
CRLF3 |
cytokine receptor-like factor 3 |
| chr8_+_110098850 | 0.07 |
ENST00000518632.1 |
TRHR |
thyrotropin-releasing hormone receptor |
| chr1_-_39339777 | 0.05 |
ENST00000397572.2 |
MYCBP |
MYC binding protein |
| chr6_+_26045603 | 0.02 |
ENST00000540144.1 |
HIST1H3C |
histone cluster 1, H3c |
| chr11_+_68228186 | 0.02 |
ENST00000393799.2 ENST00000393800.2 ENST00000528635.1 ENST00000533127.1 ENST00000529907.1 ENST00000529344.1 ENST00000534534.1 ENST00000524845.1 ENST00000265637.4 ENST00000524904.1 ENST00000393801.3 ENST00000265636.5 ENST00000529710.1 |
PPP6R3 |
protein phosphatase 6, regulatory subunit 3 |
| chr22_+_42017987 | 0.02 |
ENST00000405506.1 |
XRCC6 |
X-ray repair complementing defective repair in Chinese hamster cells 6 |
| chr16_+_1128781 | 0.02 |
ENST00000293897.4 ENST00000562758.1 |
SSTR5 |
somatostatin receptor 5 |
| chr10_-_45474237 | 0.02 |
ENST00000448778.1 ENST00000298295.3 |
C10orf10 |
chromosome 10 open reading frame 10 |
| chr12_+_53835508 | 0.01 |
ENST00000551003.1 ENST00000549068.1 ENST00000549740.1 ENST00000546581.1 ENST00000549581.1 ENST00000541275.1 |
PRR13 PCBP2 |
proline rich 13 poly(rC) binding protein 2 |
| chr3_-_52864680 | 0.01 |
ENST00000406595.1 ENST00000485816.1 ENST00000434759.3 ENST00000346281.5 ENST00000266041.4 |
ITIH4 |
inter-alpha-trypsin inhibitor heavy chain family, member 4 |
| chr14_-_23624511 | 0.01 |
ENST00000529705.2 |
SLC7A8 |
solute carrier family 7 (amino acid transporter light chain, L system), member 8 |
| chr16_+_1832902 | 0.00 |
ENST00000262302.9 ENST00000563136.1 ENST00000565987.1 ENST00000543305.1 ENST00000568287.1 ENST00000565134.1 |
NUBP2 |
nucleotide binding protein 2 |
| chr4_+_37962018 | 0.00 |
ENST00000504686.1 |
PTTG2 |
pituitary tumor-transforming 2 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.5 | GO:0050928 | negative regulation of positive chemotaxis(GO:0050928) |
| 0.4 | 1.5 | GO:1903224 | regulation of endodermal cell differentiation(GO:1903224) |
| 0.3 | 0.9 | GO:0003241 | growth involved in heart morphogenesis(GO:0003241) |
| 0.2 | 0.6 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.2 | 0.7 | GO:0035720 | signal transduction downstream of smoothened(GO:0007227) intraciliary anterograde transport(GO:0035720) ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.1 | 1.6 | GO:0052697 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.1 | 0.5 | GO:0007079 | mitotic chromosome movement towards spindle pole(GO:0007079) |
| 0.1 | 0.4 | GO:0042091 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.1 | 2.1 | GO:0002076 | osteoblast development(GO:0002076) |
| 0.1 | 0.2 | GO:1900186 | negative regulation of clathrin-mediated endocytosis(GO:1900186) |
| 0.0 | 0.3 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 0.0 | 0.1 | GO:0098905 | regulation of bundle of His cell action potential(GO:0098905) |
| 0.0 | 0.5 | GO:0045008 | depyrimidination(GO:0045008) |
| 0.0 | 0.4 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.0 | 0.1 | GO:0030950 | establishment or maintenance of actin cytoskeleton polarity(GO:0030950) |
| 0.0 | 0.2 | GO:0070141 | response to UV-A(GO:0070141) |
| 0.0 | 0.5 | GO:0061154 | endothelial tube morphogenesis(GO:0061154) |
| 0.0 | 0.2 | GO:0007021 | tubulin complex assembly(GO:0007021) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.1 | 0.7 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.1 | 0.3 | GO:1990075 | periciliary membrane compartment(GO:1990075) |
| 0.0 | 0.7 | GO:0030478 | actin cap(GO:0030478) |
| 0.0 | 0.3 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.5 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 0.4 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.0 | 0.4 | GO:0042589 | zymogen granule membrane(GO:0042589) |
| 0.0 | 0.2 | GO:0097512 | cardiac myofibril(GO:0097512) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | REACTOME GLUCURONIDATION | Genes involved in Glucuronidation |
| 0.0 | 1.5 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.0 | 0.5 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.0 | 1.5 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 1.9 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.5 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 0.7 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.9 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 1.5 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.0 | 0.9 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.0 | 1.5 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.5 | PID AURORA A PATHWAY | Aurora A signaling |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0008263 | pyrimidine-specific mismatch base pair DNA N-glycosylase activity(GO:0008263) |
| 0.1 | 0.9 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) sphingolipid binding(GO:0046625) |
| 0.1 | 1.9 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.1 | 0.3 | GO:0031962 | mineralocorticoid receptor binding(GO:0031962) |
| 0.0 | 1.6 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.7 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.0 | 0.4 | GO:0055106 | ubiquitin-protein transferase regulator activity(GO:0055106) |
| 0.0 | 0.4 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.0 | 0.1 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.0 | 1.5 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.4 | GO:0038187 | signaling pattern recognition receptor activity(GO:0008329) pattern recognition receptor activity(GO:0038187) |
| 0.0 | 1.5 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |