Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
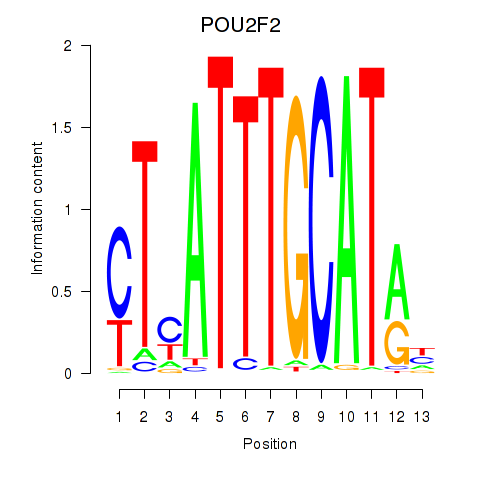
Results for POU2F2_POU3F1
Z-value: 2.83
Transcription factors associated with POU2F2_POU3F1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
POU2F2
|
ENSG00000028277.16 | POU2F2 |
|
POU3F1
|
ENSG00000185668.5 | POU3F1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| POU2F2 | hg19_v2_chr19_-_42636617_42636632 | 0.69 | 3.1e-03 | Click! |
| POU3F1 | hg19_v2_chr1_-_38512450_38512474 | 0.01 | 9.8e-01 | Click! |
Activity profile of POU2F2_POU3F1 motif
Sorted Z-values of POU2F2_POU3F1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of POU2F2_POU3F1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_+_23847339 | 27.11 |
ENST00000303531.7 |
PRKCB |
protein kinase C, beta |
| chr16_+_23847267 | 23.27 |
ENST00000321728.7 |
PRKCB |
protein kinase C, beta |
| chr16_+_32077386 | 15.46 |
ENST00000354689.6 |
IGHV3OR16-9 |
immunoglobulin heavy variable 3/OR16-9 (non-functional) |
| chr14_-_106692191 | 12.21 |
ENST00000390607.2 |
IGHV3-21 |
immunoglobulin heavy variable 3-21 |
| chr14_-_106573756 | 10.46 |
ENST00000390601.2 |
IGHV3-11 |
immunoglobulin heavy variable 3-11 (gene/pseudogene) |
| chr1_+_228645796 | 10.41 |
ENST00000369160.2 |
HIST3H2BB |
histone cluster 3, H2bb |
| chr8_+_56792355 | 10.32 |
ENST00000519728.1 |
LYN |
v-yes-1 Yamaguchi sarcoma viral related oncogene homolog |
| chr6_+_26124373 | 10.15 |
ENST00000377791.2 ENST00000602637.1 |
HIST1H2AC |
histone cluster 1, H2ac |
| chr6_-_27114577 | 9.36 |
ENST00000356950.1 ENST00000396891.4 |
HIST1H2BK |
histone cluster 1, H2bk |
| chr6_-_26124138 | 9.23 |
ENST00000314332.5 ENST00000396984.1 |
HIST1H2BC |
histone cluster 1, H2bc |
| chr8_+_56792377 | 9.10 |
ENST00000520220.2 |
LYN |
v-yes-1 Yamaguchi sarcoma viral related oncogene homolog |
| chr16_+_33605231 | 9.09 |
ENST00000570121.2 |
IGHV3OR16-12 |
immunoglobulin heavy variable 3/OR16-12 (non-functional) |
| chrX_-_48776292 | 8.98 |
ENST00000376509.4 |
PIM2 |
pim-2 oncogene |
| chr14_-_106994333 | 8.85 |
ENST00000390624.2 |
IGHV3-48 |
immunoglobulin heavy variable 3-48 |
| chr6_+_26183958 | 8.76 |
ENST00000356530.3 |
HIST1H2BE |
histone cluster 1, H2be |
| chr14_-_106926724 | 8.60 |
ENST00000434710.1 |
IGHV3-43 |
immunoglobulin heavy variable 3-43 |
| chr1_-_149783914 | 8.59 |
ENST00000369167.1 ENST00000427880.2 ENST00000545683.1 |
HIST2H2BF |
histone cluster 2, H2bf |
| chr14_-_106552755 | 7.75 |
ENST00000390600.2 |
IGHV3-9 |
immunoglobulin heavy variable 3-9 |
| chr6_-_26216872 | 7.11 |
ENST00000244601.3 |
HIST1H2BG |
histone cluster 1, H2bg |
| chr19_-_17516449 | 7.03 |
ENST00000252593.6 |
BST2 |
bone marrow stromal cell antigen 2 |
| chr15_-_20193370 | 7.01 |
ENST00000558565.2 |
IGHV3OR15-7 |
immunoglobulin heavy variable 3/OR15-7 (pseudogene) |
| chr1_-_149814478 | 6.29 |
ENST00000369161.3 |
HIST2H2AA3 |
histone cluster 2, H2aa3 |
| chr6_+_26273144 | 5.97 |
ENST00000377733.2 |
HIST1H2BI |
histone cluster 1, H2bi |
| chr14_-_106845789 | 5.86 |
ENST00000390617.2 |
IGHV3-35 |
immunoglobulin heavy variable 3-35 (non-functional) |
| chr22_+_23077065 | 5.67 |
ENST00000390310.2 |
IGLV2-18 |
immunoglobulin lambda variable 2-18 |
| chr6_+_26251835 | 5.42 |
ENST00000356350.2 |
HIST1H2BH |
histone cluster 1, H2bh |
| chr14_-_106518922 | 5.32 |
ENST00000390598.2 |
IGHV3-7 |
immunoglobulin heavy variable 3-7 |
| chr1_+_149822620 | 5.29 |
ENST00000369159.2 |
HIST2H2AA4 |
histone cluster 2, H2aa4 |
| chr22_+_22930626 | 4.98 |
ENST00000390302.2 |
IGLV2-33 |
immunoglobulin lambda variable 2-33 (non-functional) |
| chr22_+_23040274 | 4.98 |
ENST00000390306.2 |
IGLV2-23 |
immunoglobulin lambda variable 2-23 |
| chr3_+_98250743 | 4.75 |
ENST00000284311.3 |
GPR15 |
G protein-coupled receptor 15 |
| chr18_-_53089723 | 4.68 |
ENST00000561992.1 ENST00000562512.2 |
TCF4 |
transcription factor 4 |
| chr6_+_26217159 | 4.48 |
ENST00000303910.2 |
HIST1H2AE |
histone cluster 1, H2ae |
| chr16_+_33020496 | 4.32 |
ENST00000565407.2 |
IGHV3OR16-8 |
immunoglobulin heavy variable 3/OR16-8 (non-functional) |
| chr6_+_26158343 | 4.31 |
ENST00000377777.4 ENST00000289316.2 |
HIST1H2BD |
histone cluster 1, H2bd |
| chr1_+_149858461 | 4.24 |
ENST00000331380.2 |
HIST2H2AC |
histone cluster 2, H2ac |
| chr8_+_19796381 | 4.10 |
ENST00000524029.1 ENST00000522701.1 ENST00000311322.8 |
LPL |
lipoprotein lipase |
| chr17_-_47841485 | 4.06 |
ENST00000506156.1 ENST00000240364.2 |
FAM117A |
family with sequence similarity 117, member A |
| chrX_+_12993202 | 4.01 |
ENST00000451311.2 ENST00000380636.1 |
TMSB4X |
thymosin beta 4, X-linked |
| chr10_-_98031310 | 3.98 |
ENST00000427367.2 ENST00000413476.2 |
BLNK |
B-cell linker |
| chr10_-_98031265 | 3.83 |
ENST00000224337.5 ENST00000371176.2 |
BLNK |
B-cell linker |
| chr16_-_33647696 | 3.80 |
ENST00000558425.1 ENST00000569103.2 |
RP11-812E19.9 |
Uncharacterized protein |
| chr22_+_23165153 | 3.72 |
ENST00000390317.2 |
IGLV2-8 |
immunoglobulin lambda variable 2-8 |
| chr4_-_71532339 | 3.65 |
ENST00000254801.4 |
IGJ |
immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides |
| chr6_-_27100529 | 3.54 |
ENST00000607124.1 ENST00000339812.2 ENST00000541790.1 |
HIST1H2BJ |
histone cluster 1, H2bj |
| chr4_-_40517984 | 3.53 |
ENST00000381795.6 |
RBM47 |
RNA binding motif protein 47 |
| chr16_+_33629600 | 3.48 |
ENST00000562905.2 |
IGHV3OR16-13 |
immunoglobulin heavy variable 3/OR16-13 (non-functional) |
| chr1_+_167298281 | 3.19 |
ENST00000367862.5 |
POU2F1 |
POU class 2 homeobox 1 |
| chr1_-_154832316 | 3.15 |
ENST00000361147.4 |
KCNN3 |
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 |
| chr6_+_37137939 | 3.07 |
ENST00000373509.5 |
PIM1 |
pim-1 oncogene |
| chr6_+_6588902 | 3.07 |
ENST00000230568.4 |
LY86 |
lymphocyte antigen 86 |
| chrX_+_12993336 | 3.01 |
ENST00000380635.1 |
TMSB4X |
thymosin beta 4, X-linked |
| chr1_+_156119798 | 2.94 |
ENST00000355014.2 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr6_-_27860956 | 2.86 |
ENST00000359611.2 |
HIST1H2AM |
histone cluster 1, H2am |
| chr2_-_89399845 | 2.83 |
ENST00000479981.1 |
IGKV1-16 |
immunoglobulin kappa variable 1-16 |
| chr6_-_99797522 | 2.77 |
ENST00000389677.5 |
FAXC |
failed axon connections homolog (Drosophila) |
| chr6_+_27100811 | 2.72 |
ENST00000359193.2 |
HIST1H2AG |
histone cluster 1, H2ag |
| chr15_-_58357932 | 2.68 |
ENST00000347587.3 |
ALDH1A2 |
aldehyde dehydrogenase 1 family, member A2 |
| chr6_+_27114861 | 2.55 |
ENST00000377459.1 |
HIST1H2AH |
histone cluster 1, H2ah |
| chrX_+_108779004 | 2.52 |
ENST00000218004.1 |
NXT2 |
nuclear transport factor 2-like export factor 2 |
| chr14_-_107078851 | 2.49 |
ENST00000390628.2 |
IGHV1-58 |
immunoglobulin heavy variable 1-58 |
| chr8_-_33370607 | 2.49 |
ENST00000360742.5 ENST00000523305.1 |
TTI2 |
TELO2 interacting protein 2 |
| chr14_-_106586656 | 2.44 |
ENST00000390602.2 |
IGHV3-13 |
immunoglobulin heavy variable 3-13 |
| chr12_+_113344582 | 2.39 |
ENST00000202917.5 ENST00000445409.2 ENST00000452357.2 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr12_+_113344755 | 2.34 |
ENST00000550883.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr14_-_106539557 | 2.34 |
ENST00000390599.2 |
IGHV1-8 |
immunoglobulin heavy variable 1-8 |
| chr16_+_33006369 | 2.28 |
ENST00000425181.3 |
IGHV3OR16-10 |
immunoglobulin heavy variable 3/OR16-10 (non-functional) |
| chr16_+_32063311 | 2.26 |
ENST00000426099.1 |
AC142381.1 |
AC142381.1 |
| chr14_-_106816253 | 2.21 |
ENST00000390615.2 |
IGHV3-33 |
immunoglobulin heavy variable 3-33 |
| chr21_+_10862622 | 2.15 |
ENST00000302092.5 ENST00000559480.1 |
IGHV1OR21-1 |
immunoglobulin heavy variable 1/OR21-1 (non-functional) |
| chr22_+_22550113 | 2.13 |
ENST00000390285.3 |
IGLV6-57 |
immunoglobulin lambda variable 6-57 |
| chr14_-_107049312 | 2.12 |
ENST00000390627.2 |
IGHV3-53 |
immunoglobulin heavy variable 3-53 |
| chr12_-_31479045 | 2.08 |
ENST00000539409.1 ENST00000395766.1 |
FAM60A |
family with sequence similarity 60, member A |
| chr17_+_20059302 | 2.02 |
ENST00000395530.2 |
SPECC1 |
sperm antigen with calponin homology and coiled-coil domains 1 |
| chr15_+_84115868 | 2.01 |
ENST00000427482.2 |
SH3GL3 |
SH3-domain GRB2-like 3 |
| chrX_-_80457385 | 2.00 |
ENST00000451455.1 ENST00000436386.1 ENST00000358130.2 |
HMGN5 |
high mobility group nucleosome binding domain 5 |
| chr14_-_107283278 | 1.97 |
ENST00000390639.2 |
IGHV7-81 |
immunoglobulin heavy variable 7-81 (non-functional) |
| chr15_-_58357866 | 1.92 |
ENST00000537372.1 |
ALDH1A2 |
aldehyde dehydrogenase 1 family, member A2 |
| chr4_-_74088800 | 1.85 |
ENST00000509867.2 |
ANKRD17 |
ankyrin repeat domain 17 |
| chr15_-_20170354 | 1.79 |
ENST00000338912.5 |
IGHV1OR15-9 |
immunoglobulin heavy variable 1/OR15-9 (non-functional) |
| chr9_-_124991124 | 1.78 |
ENST00000394319.4 ENST00000340587.3 |
LHX6 |
LIM homeobox 6 |
| chr13_-_52027134 | 1.77 |
ENST00000311234.4 ENST00000425000.1 ENST00000463928.1 ENST00000442263.3 ENST00000398119.2 |
INTS6 |
integrator complex subunit 6 |
| chr12_+_62654119 | 1.76 |
ENST00000353364.3 ENST00000549523.1 ENST00000280377.5 |
USP15 |
ubiquitin specific peptidase 15 |
| chr2_-_172750733 | 1.76 |
ENST00000392592.4 ENST00000422440.2 |
SLC25A12 |
solute carrier family 25 (aspartate/glutamate carrier), member 12 |
| chr12_-_68696652 | 1.75 |
ENST00000539972.1 |
MDM1 |
Mdm1 nuclear protein homolog (mouse) |
| chr2_-_136678123 | 1.70 |
ENST00000422708.1 |
DARS |
aspartyl-tRNA synthetase |
| chr2_-_89340242 | 1.69 |
ENST00000480492.1 |
IGKV1-12 |
immunoglobulin kappa variable 1-12 |
| chr14_-_107114267 | 1.67 |
ENST00000454421.2 |
IGHV3-64 |
immunoglobulin heavy variable 3-64 |
| chr14_-_107219365 | 1.66 |
ENST00000424969.2 |
IGHV3-74 |
immunoglobulin heavy variable 3-74 |
| chr3_-_119276575 | 1.64 |
ENST00000383669.3 ENST00000383668.3 |
CD80 |
CD80 molecule |
| chr6_+_34204642 | 1.63 |
ENST00000347617.6 ENST00000401473.3 ENST00000311487.5 ENST00000447654.1 ENST00000395004.3 |
HMGA1 |
high mobility group AT-hook 1 |
| chr14_-_106866934 | 1.62 |
ENST00000390618.2 |
IGHV3-38 |
immunoglobulin heavy variable 3-38 (non-functional) |
| chr12_+_62654155 | 1.61 |
ENST00000312635.6 ENST00000393654.3 ENST00000549237.1 |
USP15 |
ubiquitin specific peptidase 15 |
| chr3_-_24207039 | 1.57 |
ENST00000280696.5 |
THRB |
thyroid hormone receptor, beta |
| chr15_+_84116106 | 1.56 |
ENST00000535412.1 ENST00000324537.5 |
SH3GL3 |
SH3-domain GRB2-like 3 |
| chr16_+_85645007 | 1.55 |
ENST00000405402.2 |
GSE1 |
Gse1 coiled-coil protein |
| chr22_+_22764088 | 1.54 |
ENST00000390299.2 |
IGLV1-40 |
immunoglobulin lambda variable 1-40 |
| chr14_-_106725723 | 1.54 |
ENST00000390609.2 |
IGHV3-23 |
immunoglobulin heavy variable 3-23 |
| chr6_-_27782548 | 1.54 |
ENST00000333151.3 |
HIST1H2AJ |
histone cluster 1, H2aj |
| chr7_+_26438187 | 1.49 |
ENST00000439120.1 ENST00000430548.1 ENST00000421862.1 ENST00000449537.1 ENST00000420774.1 ENST00000418758.2 |
AC004540.5 |
AC004540.5 |
| chr5_-_179045199 | 1.48 |
ENST00000523921.1 |
HNRNPH1 |
heterogeneous nuclear ribonucleoprotein H1 (H) |
| chr22_+_22712087 | 1.47 |
ENST00000390294.2 |
IGLV1-47 |
immunoglobulin lambda variable 1-47 |
| chr2_+_58134756 | 1.45 |
ENST00000435505.2 ENST00000417641.2 |
VRK2 |
vaccinia related kinase 2 |
| chr12_+_113344811 | 1.40 |
ENST00000551241.1 ENST00000553185.1 ENST00000550689.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr18_-_52989525 | 1.32 |
ENST00000457482.3 |
TCF4 |
transcription factor 4 |
| chr8_-_57123815 | 1.32 |
ENST00000316981.3 ENST00000423799.2 ENST00000429357.2 |
PLAG1 |
pleiomorphic adenoma gene 1 |
| chr22_-_29107919 | 1.29 |
ENST00000434810.1 ENST00000456369.1 |
CHEK2 |
checkpoint kinase 2 |
| chr2_+_90139056 | 1.29 |
ENST00000492446.1 |
IGKV1D-16 |
immunoglobulin kappa variable 1D-16 |
| chr4_-_144940477 | 1.28 |
ENST00000513128.1 ENST00000429670.2 ENST00000502664.1 |
GYPB |
glycophorin B (MNS blood group) |
| chr22_+_22735135 | 1.28 |
ENST00000390297.2 |
IGLV1-44 |
immunoglobulin lambda variable 1-44 |
| chr6_+_27215494 | 1.27 |
ENST00000230582.3 |
PRSS16 |
protease, serine, 16 (thymus) |
| chr22_-_24096562 | 1.27 |
ENST00000398465.3 |
VPREB3 |
pre-B lymphocyte 3 |
| chr19_-_41903161 | 1.26 |
ENST00000602129.1 ENST00000593771.1 ENST00000596905.1 ENST00000221233.4 |
EXOSC5 |
exosome component 5 |
| chr6_-_27775694 | 1.24 |
ENST00000377401.2 |
HIST1H2BL |
histone cluster 1, H2bl |
| chr8_-_16859690 | 1.24 |
ENST00000180166.5 |
FGF20 |
fibroblast growth factor 20 |
| chr2_+_90198535 | 1.22 |
ENST00000390276.2 |
IGKV1D-12 |
immunoglobulin kappa variable 1D-12 |
| chr4_-_25865159 | 1.22 |
ENST00000502949.1 ENST00000264868.5 ENST00000513691.1 ENST00000514872.1 |
SEL1L3 |
sel-1 suppressor of lin-12-like 3 (C. elegans) |
| chr15_-_66649010 | 1.20 |
ENST00000367709.4 ENST00000261881.4 |
TIPIN |
TIMELESS interacting protein |
| chr2_+_90192768 | 1.20 |
ENST00000390275.2 |
IGKV1D-13 |
immunoglobulin kappa variable 1D-13 |
| chr6_+_27215471 | 1.19 |
ENST00000421826.2 |
PRSS16 |
protease, serine, 16 (thymus) |
| chr1_+_32666188 | 1.19 |
ENST00000421922.2 |
CCDC28B |
coiled-coil domain containing 28B |
| chr11_-_11374904 | 1.19 |
ENST00000528848.2 |
CSNK2A3 |
casein kinase 2, alpha 3 polypeptide |
| chr11_-_5248294 | 1.18 |
ENST00000335295.4 |
HBB |
hemoglobin, beta |
| chr19_+_35820064 | 1.15 |
ENST00000341773.6 ENST00000600131.1 ENST00000270311.6 ENST00000595780.1 ENST00000597916.1 ENST00000593867.1 ENST00000600424.1 ENST00000599811.1 ENST00000536635.2 ENST00000085219.5 ENST00000544992.2 ENST00000419549.2 |
CD22 |
CD22 molecule |
| chr6_+_27861190 | 1.14 |
ENST00000303806.4 |
HIST1H2BO |
histone cluster 1, H2bo |
| chr3_+_157827841 | 1.13 |
ENST00000295930.3 ENST00000471994.1 ENST00000464171.1 ENST00000312179.6 ENST00000475278.2 |
RSRC1 |
arginine/serine-rich coiled-coil 1 |
| chr15_+_67841330 | 1.09 |
ENST00000354498.5 |
MAP2K5 |
mitogen-activated protein kinase kinase 5 |
| chr14_-_107131560 | 1.07 |
ENST00000390632.2 |
IGHV3-66 |
immunoglobulin heavy variable 3-66 |
| chr10_-_103880209 | 1.06 |
ENST00000425280.1 |
LDB1 |
LIM domain binding 1 |
| chr6_-_89927151 | 1.05 |
ENST00000454853.2 |
GABRR1 |
gamma-aminobutyric acid (GABA) A receptor, rho 1 |
| chr17_-_61850894 | 1.02 |
ENST00000403162.3 ENST00000582252.1 ENST00000225726.5 |
CCDC47 |
coiled-coil domain containing 47 |
| chr6_+_27775899 | 1.01 |
ENST00000358739.3 |
HIST1H2AI |
histone cluster 1, H2ai |
| chr3_-_33686743 | 1.00 |
ENST00000333778.6 ENST00000539981.1 |
CLASP2 |
cytoplasmic linker associated protein 2 |
| chr16_+_53164833 | 0.99 |
ENST00000564845.1 |
CHD9 |
chromodomain helicase DNA binding protein 9 |
| chr19_+_17858509 | 0.98 |
ENST00000594202.1 ENST00000252771.7 ENST00000389133.4 |
FCHO1 |
FCH domain only 1 |
| chr12_-_56359802 | 0.97 |
ENST00000548803.1 ENST00000449260.2 ENST00000550447.1 ENST00000547137.1 ENST00000546543.1 ENST00000550464.1 |
PMEL |
premelanosome protein |
| chr19_+_35861831 | 0.95 |
ENST00000454971.1 |
GPR42 |
G protein-coupled receptor 42 (gene/pseudogene) |
| chr6_-_43655511 | 0.95 |
ENST00000372133.3 ENST00000372116.1 ENST00000427312.1 |
MRPS18A |
mitochondrial ribosomal protein S18A |
| chr7_+_50348268 | 0.94 |
ENST00000438033.1 ENST00000439701.1 |
IKZF1 |
IKAROS family zinc finger 1 (Ikaros) |
| chr19_-_50168962 | 0.93 |
ENST00000599223.1 ENST00000593922.1 ENST00000600022.1 ENST00000596765.1 ENST00000599144.1 ENST00000596822.1 ENST00000598108.1 ENST00000601373.1 ENST00000595034.1 ENST00000601291.1 |
IRF3 |
interferon regulatory factor 3 |
| chr6_-_168479839 | 0.93 |
ENST00000283309.6 |
FRMD1 |
FERM domain containing 1 |
| chr1_+_32538492 | 0.92 |
ENST00000336294.5 |
TMEM39B |
transmembrane protein 39B |
| chr4_-_152682129 | 0.92 |
ENST00000512306.1 ENST00000508611.1 ENST00000515812.1 ENST00000263985.6 |
PET112 |
PET112 homolog (yeast) |
| chr18_-_53253112 | 0.91 |
ENST00000568673.1 ENST00000562847.1 ENST00000568147.1 |
TCF4 |
transcription factor 4 |
| chr2_+_89901292 | 0.91 |
ENST00000448155.2 |
IGKV1D-39 |
immunoglobulin kappa variable 1D-39 |
| chr2_+_114163945 | 0.91 |
ENST00000453673.3 |
IGKV1OR2-108 |
immunoglobulin kappa variable 1/OR2-108 (non-functional) |
| chr19_-_50169064 | 0.91 |
ENST00000593337.1 ENST00000598808.1 ENST00000600453.1 ENST00000593818.1 ENST00000597198.1 ENST00000601809.1 ENST00000377139.3 |
IRF3 |
interferon regulatory factor 3 |
| chr12_-_48963829 | 0.90 |
ENST00000301046.2 ENST00000549817.1 |
LALBA |
lactalbumin, alpha- |
| chr6_+_122720681 | 0.90 |
ENST00000368455.4 ENST00000452194.1 |
HSF2 |
heat shock transcription factor 2 |
| chr9_-_124990680 | 0.87 |
ENST00000541397.2 ENST00000560485.1 |
LHX6 |
LIM homeobox 6 |
| chr6_+_43139037 | 0.87 |
ENST00000265354.4 |
SRF |
serum response factor (c-fos serum response element-binding transcription factor) |
| chr1_+_166808692 | 0.85 |
ENST00000367876.4 |
POGK |
pogo transposable element with KRAB domain |
| chr22_+_22385332 | 0.84 |
ENST00000390282.2 |
IGLV4-69 |
immunoglobulin lambda variable 4-69 |
| chr22_-_17640110 | 0.84 |
ENST00000399852.3 ENST00000336737.4 |
CECR5 |
cat eye syndrome chromosome region, candidate 5 |
| chr1_-_190446759 | 0.83 |
ENST00000367462.3 |
BRINP3 |
bone morphogenetic protein/retinoic acid inducible neural-specific 3 |
| chr2_+_90248739 | 0.83 |
ENST00000468879.1 |
IGKV1D-43 |
immunoglobulin kappa variable 1D-43 |
| chr15_-_83240507 | 0.83 |
ENST00000564522.1 ENST00000398592.2 |
CPEB1 |
cytoplasmic polyadenylation element binding protein 1 |
| chr15_-_83240553 | 0.83 |
ENST00000423133.2 ENST00000398591.2 |
CPEB1 |
cytoplasmic polyadenylation element binding protein 1 |
| chr2_+_97426631 | 0.82 |
ENST00000377075.2 |
CNNM4 |
cyclin M4 |
| chr14_-_106622419 | 0.82 |
ENST00000390604.2 |
IGHV3-16 |
immunoglobulin heavy variable 3-16 (non-functional) |
| chr3_+_119316689 | 0.82 |
ENST00000273371.4 |
PLA1A |
phospholipase A1 member A |
| chr10_-_51130715 | 0.81 |
ENST00000402038.3 |
PARG |
poly (ADP-ribose) glycohydrolase |
| chr19_-_18508396 | 0.81 |
ENST00000595840.1 ENST00000339007.3 |
LRRC25 |
leucine rich repeat containing 25 |
| chr22_-_24096630 | 0.79 |
ENST00000248948.3 |
VPREB3 |
pre-B lymphocyte 3 |
| chr16_-_21314360 | 0.78 |
ENST00000219599.3 ENST00000576703.1 |
CRYM |
crystallin, mu |
| chr17_+_61851504 | 0.77 |
ENST00000359353.5 ENST00000389924.2 |
DDX42 |
DEAD (Asp-Glu-Ala-Asp) box helicase 42 |
| chr19_+_41305085 | 0.77 |
ENST00000303961.4 |
EGLN2 |
egl-9 family hypoxia-inducible factor 2 |
| chr1_-_149858227 | 0.75 |
ENST00000369155.2 |
HIST2H2BE |
histone cluster 2, H2be |
| chr18_-_52989217 | 0.75 |
ENST00000570287.2 |
TCF4 |
transcription factor 4 |
| chr6_-_55739542 | 0.75 |
ENST00000446683.2 |
BMP5 |
bone morphogenetic protein 5 |
| chrX_-_3761898 | 0.73 |
ENST00000425492.2 |
RP11-706O15.1 |
HCG1981372, isoform CRA_c; Uncharacterized protein |
| chr17_-_19651668 | 0.73 |
ENST00000494157.2 ENST00000225740.6 |
ALDH3A1 |
aldehyde dehydrogenase 3 family, member A1 |
| chr2_+_89998789 | 0.72 |
ENST00000453166.2 |
IGKV2D-28 |
immunoglobulin kappa variable 2D-28 |
| chr6_-_35656712 | 0.72 |
ENST00000357266.4 ENST00000542713.1 |
FKBP5 |
FK506 binding protein 5 |
| chr6_+_97010424 | 0.72 |
ENST00000541107.1 ENST00000450218.1 ENST00000326771.2 |
FHL5 |
four and a half LIM domains 5 |
| chr1_+_32538520 | 0.72 |
ENST00000438825.1 ENST00000456834.2 ENST00000373634.4 ENST00000427288.1 |
TMEM39B |
transmembrane protein 39B |
| chr14_-_107170409 | 0.71 |
ENST00000390633.2 |
IGHV1-69 |
immunoglobulin heavy variable 1-69 |
| chr2_-_89247338 | 0.70 |
ENST00000496168.1 |
IGKV1-5 |
immunoglobulin kappa variable 1-5 |
| chr2_-_2334888 | 0.70 |
ENST00000428368.2 ENST00000399161.2 |
MYT1L |
myelin transcription factor 1-like |
| chrX_+_129535937 | 0.68 |
ENST00000305536.6 ENST00000370947.1 |
RBMX2 |
RNA binding motif protein, X-linked 2 |
| chr9_+_108463234 | 0.68 |
ENST00000374688.1 |
TMEM38B |
transmembrane protein 38B |
| chr22_-_22221900 | 0.68 |
ENST00000215832.6 ENST00000398822.3 |
MAPK1 |
mitogen-activated protein kinase 1 |
| chr2_-_89417335 | 0.68 |
ENST00000490686.1 |
IGKV1-17 |
immunoglobulin kappa variable 1-17 |
| chr19_-_50168861 | 0.68 |
ENST00000596756.1 |
IRF3 |
interferon regulatory factor 3 |
| chr6_-_35656685 | 0.68 |
ENST00000539068.1 ENST00000540787.1 |
FKBP5 |
FK506 binding protein 5 |
| chr1_-_68962805 | 0.67 |
ENST00000370966.5 |
DEPDC1 |
DEP domain containing 1 |
| chr7_+_18535346 | 0.66 |
ENST00000405010.3 ENST00000406451.4 ENST00000428307.2 |
HDAC9 |
histone deacetylase 9 |
| chr3_+_119316721 | 0.65 |
ENST00000488919.1 ENST00000495992.1 |
PLA1A |
phospholipase A1 member A |
| chr12_+_14518598 | 0.65 |
ENST00000261168.4 ENST00000538511.1 ENST00000545723.1 ENST00000543189.1 ENST00000536444.1 |
ATF7IP |
activating transcription factor 7 interacting protein |
| chr19_-_18392422 | 0.65 |
ENST00000252818.3 |
JUND |
jun D proto-oncogene |
| chr6_-_53013620 | 0.65 |
ENST00000259803.7 |
GCM1 |
glial cells missing homolog 1 (Drosophila) |
| chrX_+_99839799 | 0.64 |
ENST00000373031.4 |
TNMD |
tenomodulin |
| chr2_-_70529180 | 0.64 |
ENST00000450256.1 ENST00000037869.3 |
FAM136A |
family with sequence similarity 136, member A |
| chr2_+_89923550 | 0.63 |
ENST00000509129.1 |
IGKV1D-37 |
immunoglobulin kappa variable 1D-37 (non-functional) |
| chr3_-_107777208 | 0.63 |
ENST00000398258.3 |
CD47 |
CD47 molecule |
| chr2_-_225811747 | 0.62 |
ENST00000409592.3 |
DOCK10 |
dedicator of cytokinesis 10 |
| chr2_-_89619904 | 0.62 |
ENST00000498574.1 |
IGKV1-39 |
immunoglobulin kappa variable 1-39 (gene/pseudogene) |
| chr17_-_19651654 | 0.61 |
ENST00000395555.3 |
ALDH3A1 |
aldehyde dehydrogenase 3 family, member A1 |
| chr20_+_62795827 | 0.61 |
ENST00000328439.1 ENST00000536311.1 |
MYT1 |
myelin transcription factor 1 |
| chr19_+_35862192 | 0.61 |
ENST00000597214.1 |
GPR42 |
G protein-coupled receptor 42 (gene/pseudogene) |
Gene Ontology Analysis
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 16.8 | 50.4 | GO:0035403 | histone kinase activity (H3-T6 specific)(GO:0035403) |
| 2.8 | 19.5 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 1.4 | 79.7 | GO:0034987 | immunoglobulin receptor binding(GO:0034987) |
| 1.4 | 4.1 | GO:0017129 | triglyceride binding(GO:0017129) |
| 0.6 | 3.1 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.6 | 6.1 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.6 | 7.7 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.6 | 3.4 | GO:0061649 | ubiquitinated histone binding(GO:0061649) |
| 0.4 | 5.9 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.4 | 7.8 | GO:0005068 | transmembrane receptor protein tyrosine kinase adaptor activity(GO:0005068) |
| 0.4 | 74.0 | GO:0003823 | antigen binding(GO:0003823) |
| 0.4 | 1.8 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.3 | 1.7 | GO:0004815 | aspartate-tRNA ligase activity(GO:0004815) |
| 0.3 | 2.5 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.3 | 7.0 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.3 | 4.1 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.2 | 0.9 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.2 | 3.1 | GO:0043024 | ribosomal small subunit binding(GO:0043024) |
| 0.2 | 1.5 | GO:0052739 | phosphatidylserine 1-acylhydrolase activity(GO:0052739) |
| 0.2 | 1.2 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.2 | 0.6 | GO:0030107 | HLA-A specific inhibitory MHC class I receptor activity(GO:0030107) |
| 0.2 | 0.5 | GO:0033858 | N-acetylgalactosamine kinase activity(GO:0033858) |
| 0.2 | 0.5 | GO:0015235 | cobalamin transporter activity(GO:0015235) |
| 0.2 | 0.5 | GO:0016899 | oxidoreductase activity, acting on the CH-OH group of donors, oxygen as acceptor(GO:0016899) |
| 0.2 | 0.5 | GO:0004877 | complement component C4b receptor activity(GO:0001861) complement component C3b receptor activity(GO:0004877) |
| 0.1 | 1.6 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.1 | 5.5 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 1.7 | GO:0035925 | mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.1 | 3.5 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 2.5 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 0.9 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
| 0.1 | 68.0 | GO:0046982 | protein heterodimerization activity(GO:0046982) |
| 0.1 | 0.9 | GO:0019103 | pyrimidine nucleotide binding(GO:0019103) |
| 0.1 | 1.1 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.1 | 1.1 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.1 | 0.8 | GO:0019826 | oxygen sensor activity(GO:0019826) |
| 0.1 | 0.4 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.1 | 0.6 | GO:0070053 | thrombospondin receptor activity(GO:0070053) |
| 0.1 | 1.0 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.1 | 1.5 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.1 | 0.4 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.1 | 1.8 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 0.8 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 0.3 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 0.0 | 1.2 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.0 | 0.5 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.0 | 1.4 | GO:0005528 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.0 | 0.7 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 1.0 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.0 | 0.3 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.0 | 0.8 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 1.3 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.0 | 0.4 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.0 | 0.1 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.0 | 1.6 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.0 | 0.6 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 1.1 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.1 | GO:0051731 | polynucleotide 5'-hydroxyl-kinase activity(GO:0051731) |
| 0.0 | 0.2 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.0 | 0.3 | GO:1990948 | ligase inhibitor activity(GO:0055104) ubiquitin ligase inhibitor activity(GO:1990948) |
| 0.0 | 0.6 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.0 | 0.4 | GO:0008656 | cysteine-type endopeptidase activator activity involved in apoptotic process(GO:0008656) |
| 0.0 | 0.1 | GO:0004704 | NF-kappaB-inducing kinase activity(GO:0004704) |
| 0.0 | 0.3 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 3.1 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 0.6 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 0.3 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.2 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.0 | 0.4 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 0.3 | GO:0070888 | E-box binding(GO:0070888) |
| 0.0 | 0.1 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.0 | 0.4 | GO:0005123 | death receptor binding(GO:0005123) |
| 0.0 | 0.3 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.0 | 0.4 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 0.1 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.9 | 19.4 | GO:0034666 | integrin alpha2-beta1 complex(GO:0034666) |
| 1.6 | 79.7 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 1.4 | 80.7 | GO:0000786 | nucleosome(GO:0000786) |
| 0.3 | 1.2 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.2 | 1.8 | GO:0032039 | integrator complex(GO:0032039) |
| 0.2 | 4.1 | GO:0042627 | chylomicron(GO:0042627) |
| 0.2 | 1.5 | GO:0031298 | replication fork protection complex(GO:0031298) |
| 0.1 | 1.6 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.1 | 8.0 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.1 | 1.3 | GO:0000176 | nuclear exosome (RNase complex)(GO:0000176) |
| 0.1 | 0.5 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.1 | 1.0 | GO:0045180 | kinetochore microtubule(GO:0005828) basal cortex(GO:0045180) |
| 0.1 | 2.1 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.1 | 0.4 | GO:0043293 | apoptosome(GO:0043293) |
| 0.1 | 1.7 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.1 | 0.4 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) |
| 0.1 | 0.6 | GO:0035976 | AP1 complex(GO:0035976) |
| 0.1 | 1.7 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.1 | 0.4 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.1 | 0.9 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.1 | 0.2 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.1 | 1.5 | GO:0002080 | acrosomal membrane(GO:0002080) |
| 0.1 | 7.0 | GO:0031093 | platelet alpha granule lumen(GO:0031093) |
| 0.1 | 0.2 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.1 | 7.6 | GO:0032993 | protein-DNA complex(GO:0032993) |
| 0.0 | 0.9 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.2 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.0 | 1.6 | GO:0098636 | protein complex involved in cell adhesion(GO:0098636) |
| 0.0 | 1.1 | GO:1902711 | GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.7 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 3.9 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 1.8 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 0.1 | GO:0032444 | activin responsive factor complex(GO:0032444) |
| 0.0 | 1.9 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 2.9 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.2 | GO:0097197 | tetraspanin-enriched microdomain(GO:0097197) |
| 0.0 | 0.3 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.0 | 1.5 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.4 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 0.7 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.8 | GO:0005913 | cell-cell adherens junction(GO:0005913) |
| 0.0 | 0.6 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.6 | GO:0005871 | kinesin complex(GO:0005871) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 99.1 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 1.8 | 50.8 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 1.1 | 19.4 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.3 | 7.8 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.2 | 4.1 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.2 | 1.6 | REACTOME APOBEC3G MEDIATED RESISTANCE TO HIV1 INFECTION | Genes involved in APOBEC3G mediated resistance to HIV-1 infection |
| 0.1 | 1.3 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.1 | 2.9 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 2.5 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.1 | 1.6 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.1 | 5.5 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.1 | 1.2 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.1 | 3.2 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 3 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 3 Promoter |
| 0.1 | 0.9 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.1 | 6.1 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.1 | 1.3 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.1 | 0.7 | REACTOME RAF MAP KINASE CASCADE | Genes involved in RAF/MAP kinase cascade |
| 0.0 | 7.0 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 1.7 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 0.5 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.0 | 0.5 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 1.6 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 3.4 | REACTOME TOLL RECEPTOR CASCADES | Genes involved in Toll Receptor Cascades |
| 0.0 | 1.1 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.0 | 0.9 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 0.6 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.0 | 0.7 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.6 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.0 | 1.9 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.5 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.0 | 0.7 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 1.4 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.4 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 1.7 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.2 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.3 | REACTOME OXYGEN DEPENDENT PROLINE HYDROXYLATION OF HYPOXIA INDUCIBLE FACTOR ALPHA | Genes involved in Oxygen-dependent Proline Hydroxylation of Hypoxia-inducible Factor Alpha |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.3 | 51.1 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
| 0.9 | 22.4 | PID IL5 PATHWAY | IL5-mediated signaling events |
| 0.3 | 7.8 | SA B CELL RECEPTOR COMPLEXES | Antigen binding to B cell receptors activates protein tyrosine kinases, such as the Src family, which ultimate activate MAP kinases. |
| 0.1 | 3.3 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.0 | 0.9 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.0 | 1.6 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 1.9 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.9 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 1.1 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 1.2 | PID ATR PATHWAY | ATR signaling pathway |
| 0.0 | 1.2 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 1.3 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 1.1 | PID BCR 5PATHWAY | BCR signaling pathway |
| 0.0 | 1.6 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.0 | 0.9 | PID NFAT TFPATHWAY | Calcineurin-regulated NFAT-dependent transcription in lymphocytes |
| 0.0 | 1.1 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 0.4 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.0 | 1.3 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.0 | 1.4 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 0.6 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 1.7 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 0.6 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.0 | 0.4 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.6 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 0.9 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.0 | 0.3 | PID HIF1A PATHWAY | Hypoxic and oxygen homeostasis regulation of HIF-1-alpha |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 16.8 | 50.4 | GO:0035408 | histone H3-T6 phosphorylation(GO:0035408) |
| 6.5 | 19.4 | GO:0070668 | positive regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060369) regulation of mast cell proliferation(GO:0070666) positive regulation of mast cell proliferation(GO:0070668) |
| 4.1 | 4.1 | GO:0055096 | lipoprotein particle mediated signaling(GO:0055095) low-density lipoprotein particle mediated signaling(GO:0055096) |
| 2.3 | 7.0 | GO:1901253 | negative regulation of dendritic cell cytokine production(GO:0002731) negative regulation of intracellular transport of viral material(GO:1901253) |
| 1.2 | 22.9 | GO:0002227 | innate immune response in mucosa(GO:0002227) |
| 1.1 | 77.6 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 1.0 | 7.0 | GO:2000483 | negative regulation of interleukin-8 secretion(GO:2000483) |
| 0.8 | 3.4 | GO:0035616 | histone H2B conserved C-terminal lysine deubiquitination(GO:0035616) |
| 0.6 | 0.6 | GO:0035992 | tendon cell differentiation(GO:0035990) tendon formation(GO:0035992) |
| 0.6 | 63.4 | GO:0006958 | complement activation, classical pathway(GO:0006958) |
| 0.6 | 3.5 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.6 | 1.8 | GO:0043490 | malate-aspartate shuttle(GO:0043490) |
| 0.6 | 1.7 | GO:0006422 | aspartyl-tRNA aminoacylation(GO:0006422) |
| 0.6 | 1.7 | GO:0008050 | female courtship behavior(GO:0008050) |
| 0.5 | 4.6 | GO:0035799 | ureter maturation(GO:0035799) |
| 0.5 | 1.9 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.4 | 1.3 | GO:1903925 | signal transduction involved in intra-S DNA damage checkpoint(GO:0072428) response to bisphenol A(GO:1903925) cellular response to bisphenol A(GO:1903926) |
| 0.4 | 1.3 | GO:0071051 | polyadenylation-dependent snoRNA 3'-end processing(GO:0071051) |
| 0.4 | 1.6 | GO:0090402 | oncogene-induced cell senescence(GO:0090402) |
| 0.4 | 1.1 | GO:0071676 | negative regulation of mononuclear cell migration(GO:0071676) |
| 0.3 | 3.6 | GO:2000138 | positive regulation of cell proliferation involved in heart morphogenesis(GO:2000138) |
| 0.3 | 0.6 | GO:0060018 | astrocyte fate commitment(GO:0060018) |
| 0.3 | 2.5 | GO:0039530 | MDA-5 signaling pathway(GO:0039530) |
| 0.3 | 0.9 | GO:0005989 | lactose metabolic process(GO:0005988) lactose biosynthetic process(GO:0005989) |
| 0.3 | 1.2 | GO:0030185 | nitric oxide transport(GO:0030185) |
| 0.3 | 26.6 | GO:0006342 | chromatin silencing(GO:0006342) |
| 0.3 | 1.1 | GO:2000342 | negative regulation of chemokine (C-X-C motif) ligand 2 production(GO:2000342) |
| 0.3 | 31.2 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.2 | 2.5 | GO:0021853 | cerebral cortex GABAergic interneuron migration(GO:0021853) interneuron migration(GO:1904936) |
| 0.2 | 1.2 | GO:0048478 | replication fork protection(GO:0048478) |
| 0.2 | 1.6 | GO:0045425 | regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045423) positive regulation of granulocyte macrophage colony-stimulating factor biosynthetic process(GO:0045425) |
| 0.2 | 0.7 | GO:0019858 | cytosine metabolic process(GO:0019858) |
| 0.2 | 0.9 | GO:0003257 | positive regulation of transcription from RNA polymerase II promoter involved in myocardial precursor cell differentiation(GO:0003257) positive regulation of transcription from RNA polymerase II promoter involved in heart development(GO:1901228) |
| 0.2 | 1.7 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.2 | 1.1 | GO:0046985 | positive regulation of hemoglobin biosynthetic process(GO:0046985) |
| 0.2 | 1.7 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.2 | 1.2 | GO:1904338 | regulation of dopaminergic neuron differentiation(GO:1904338) |
| 0.2 | 0.2 | GO:0060028 | convergent extension involved in axis elongation(GO:0060028) mediolateral intercalation(GO:0060031) planar cell polarity pathway involved in gastrula mediolateral intercalation(GO:0060775) |
| 0.2 | 0.9 | GO:0051941 | regulation of amino acid uptake involved in synaptic transmission(GO:0051941) regulation of glutamate uptake involved in transmission of nerve impulse(GO:0051946) regulation of L-glutamate import(GO:1900920) |
| 0.2 | 0.5 | GO:0048733 | sebaceous gland development(GO:0048733) |
| 0.2 | 0.5 | GO:0060370 | susceptibility to T cell mediated cytotoxicity(GO:0060370) |
| 0.1 | 0.6 | GO:2001193 | gamma-delta T cell activation involved in immune response(GO:0002290) negative regulation of interferon-beta secretion(GO:0035548) regulation of gamma-delta T cell activation involved in immune response(GO:2001191) positive regulation of gamma-delta T cell activation involved in immune response(GO:2001193) |
| 0.1 | 1.7 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.1 | 0.4 | GO:0021913 | regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification(GO:0021913) |
| 0.1 | 1.0 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.1 | 2.9 | GO:0045063 | T-helper 1 cell differentiation(GO:0045063) |
| 0.1 | 1.5 | GO:2000659 | regulation of interleukin-1-mediated signaling pathway(GO:2000659) |
| 0.1 | 9.0 | GO:0016239 | positive regulation of macroautophagy(GO:0016239) |
| 0.1 | 0.8 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.1 | 1.0 | GO:0006983 | ER overload response(GO:0006983) |
| 0.1 | 4.7 | GO:0072678 | T cell migration(GO:0072678) |
| 0.1 | 0.4 | GO:0072369 | regulation of lipid transport by positive regulation of transcription from RNA polymerase II promoter(GO:0072369) |
| 0.1 | 0.4 | GO:1902748 | positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.1 | 0.4 | GO:0072396 | response to cell cycle checkpoint signaling(GO:0072396) response to DNA integrity checkpoint signaling(GO:0072402) response to DNA damage checkpoint signaling(GO:0072423) |
| 0.1 | 0.4 | GO:0060743 | epithelial cell maturation involved in prostate gland development(GO:0060743) |
| 0.1 | 0.5 | GO:0009441 | glycolate metabolic process(GO:0009441) |
| 0.1 | 1.3 | GO:0060252 | positive regulation of glial cell proliferation(GO:0060252) |
| 0.1 | 1.3 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 0.3 | GO:0045872 | positive regulation of rhodopsin gene expression(GO:0045872) |
| 0.1 | 6.1 | GO:0045071 | negative regulation of viral genome replication(GO:0045071) |
| 0.1 | 0.4 | GO:0030885 | regulation of myeloid dendritic cell activation(GO:0030885) positive regulation of interferon-gamma secretion(GO:1902715) |
| 0.1 | 11.3 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.1 | 1.5 | GO:0036150 | phosphatidylserine acyl-chain remodeling(GO:0036150) |
| 0.1 | 0.3 | GO:0060010 | Sertoli cell fate commitment(GO:0060010) |
| 0.1 | 0.7 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.1 | 7.9 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.1 | 0.3 | GO:2000434 | regulation of protein neddylation(GO:2000434) negative regulation of protein neddylation(GO:2000435) |
| 0.1 | 0.6 | GO:0008228 | opsonization(GO:0008228) |
| 0.1 | 0.9 | GO:0046135 | pyrimidine nucleoside catabolic process(GO:0046135) |
| 0.1 | 0.8 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 0.1 | GO:1904294 | positive regulation of ERAD pathway(GO:1904294) |
| 0.1 | 0.3 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.1 | 0.3 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.1 | 7.4 | GO:0065004 | protein-DNA complex assembly(GO:0065004) |
| 0.0 | 3.2 | GO:0042795 | snRNA transcription(GO:0009301) snRNA transcription from RNA polymerase II promoter(GO:0042795) |
| 0.0 | 0.1 | GO:0048867 | stem cell fate determination(GO:0048867) |
| 0.0 | 0.2 | GO:1903803 | glutamine secretion(GO:0010585) L-glutamine import(GO:0036229) L-glutamine import into cell(GO:1903803) |
| 0.0 | 0.3 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.0 | 0.2 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.0 | 0.0 | GO:0010165 | response to X-ray(GO:0010165) response to ionizing radiation(GO:0010212) |
| 0.0 | 0.1 | GO:1900224 | positive regulation of nodal signaling pathway involved in determination of lateral mesoderm left/right asymmetry(GO:1900224) |
| 0.0 | 0.2 | GO:0038031 | non-canonical Wnt signaling pathway via JNK cascade(GO:0038031) |
| 0.0 | 1.0 | GO:0032438 | melanosome organization(GO:0032438) pigment granule organization(GO:0048753) |
| 0.0 | 0.4 | GO:0042756 | drinking behavior(GO:0042756) |
| 0.0 | 0.3 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.0 | 0.4 | GO:0071372 | cellular response to follicle-stimulating hormone stimulus(GO:0071372) |
| 0.0 | 0.3 | GO:0061077 | chaperone-mediated protein folding(GO:0061077) |
| 0.0 | 1.0 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.0 | 3.1 | GO:1903955 | positive regulation of protein targeting to mitochondrion(GO:1903955) |
| 0.0 | 0.5 | GO:0006012 | galactose metabolic process(GO:0006012) |
| 0.0 | 1.4 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.1 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 1.1 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 0.2 | GO:2000210 | positive regulation of anoikis(GO:2000210) |
| 0.0 | 0.1 | GO:0035871 | protein K11-linked deubiquitination(GO:0035871) |
| 0.0 | 1.3 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.0 | 0.1 | GO:0098908 | regulation of neuronal action potential(GO:0098908) |
| 0.0 | 0.2 | GO:0019321 | pentose metabolic process(GO:0019321) neuron cellular homeostasis(GO:0070050) |
| 0.0 | 2.9 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.0 | 1.1 | GO:0043484 | regulation of RNA splicing(GO:0043484) |
| 0.0 | 0.5 | GO:0010501 | RNA secondary structure unwinding(GO:0010501) |
| 0.0 | 0.7 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.0 | 0.8 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.9 | GO:0070125 | mitochondrial translational elongation(GO:0070125) mitochondrial translational termination(GO:0070126) |
| 0.0 | 1.8 | GO:0051591 | response to cAMP(GO:0051591) |
| 0.0 | 0.1 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 0.3 | GO:0048665 | neuron fate specification(GO:0048665) |
| 0.0 | 0.4 | GO:0009948 | anterior/posterior axis specification(GO:0009948) |
| 0.0 | 0.5 | GO:0048488 | synaptic vesicle endocytosis(GO:0048488) |
| 0.0 | 0.2 | GO:2000144 | positive regulation of DNA-templated transcription, initiation(GO:2000144) |
| 0.0 | 0.5 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 0.4 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.0 | 0.3 | GO:0032878 | regulation of establishment or maintenance of cell polarity(GO:0032878) |
| 0.0 | 0.0 | GO:2001183 | negative regulation of interleukin-10 secretion(GO:2001180) negative regulation of interleukin-12 secretion(GO:2001183) |
| 0.0 | 0.1 | GO:1903800 | positive regulation of production of miRNAs involved in gene silencing by miRNA(GO:1903800) |