Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
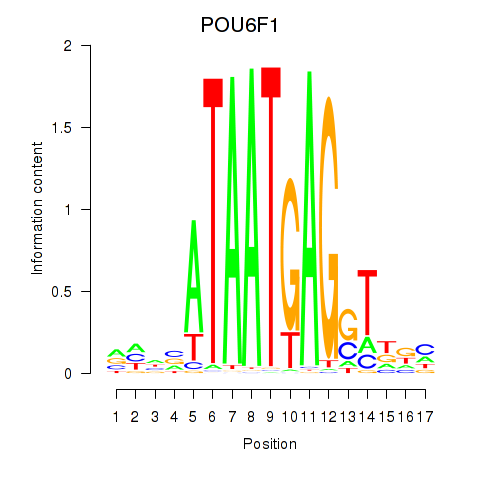
Results for POU6F1
Z-value: 0.55
Transcription factors associated with POU6F1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
POU6F1
|
ENSG00000184271.11 | POU6F1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| POU6F1 | hg19_v2_chr12_-_51611477_51611612 | 0.43 | 9.3e-02 | Click! |
Activity profile of POU6F1 motif
Sorted Z-values of POU6F1 motif
Network of associatons between targets according to the STRING database.
First level regulatory network of POU6F1
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_-_91546926 | 1.80 |
ENST00000550758.1 |
DCN |
decorin |
| chr6_+_151646800 | 1.61 |
ENST00000354675.6 |
AKAP12 |
A kinase (PRKA) anchor protein 12 |
| chr3_-_127541194 | 1.45 |
ENST00000453507.2 |
MGLL |
monoglyceride lipase |
| chr2_-_217560248 | 0.99 |
ENST00000233813.4 |
IGFBP5 |
insulin-like growth factor binding protein 5 |
| chr2_+_102721023 | 0.95 |
ENST00000409589.1 ENST00000409329.1 |
IL1R1 |
interleukin 1 receptor, type I |
| chr18_+_61554932 | 0.81 |
ENST00000299502.4 ENST00000457692.1 ENST00000413956.1 |
SERPINB2 |
serpin peptidase inhibitor, clade B (ovalbumin), member 2 |
| chr7_-_83824169 | 0.76 |
ENST00000265362.4 |
SEMA3A |
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A |
| chr7_+_80267973 | 0.48 |
ENST00000394788.3 ENST00000447544.2 |
CD36 |
CD36 molecule (thrombospondin receptor) |
| chr9_-_13165457 | 0.36 |
ENST00000542239.1 ENST00000538841.1 ENST00000433359.2 |
MPDZ |
multiple PDZ domain protein |
| chr14_-_70883708 | 0.36 |
ENST00000256366.4 |
SYNJ2BP |
synaptojanin 2 binding protein |
| chr3_-_114343768 | 0.29 |
ENST00000393785.2 |
ZBTB20 |
zinc finger and BTB domain containing 20 |
| chr9_+_128509624 | 0.29 |
ENST00000342287.5 ENST00000373487.4 |
PBX3 |
pre-B-cell leukemia homeobox 3 |
| chr11_-_89224139 | 0.25 |
ENST00000413594.2 |
NOX4 |
NADPH oxidase 4 |
| chr13_+_58206655 | 0.24 |
ENST00000377918.3 |
PCDH17 |
protocadherin 17 |
| chr19_+_52076425 | 0.24 |
ENST00000436511.2 |
ZNF175 |
zinc finger protein 175 |
| chr6_-_111804905 | 0.22 |
ENST00000358835.3 ENST00000435970.1 |
REV3L |
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr5_+_115177178 | 0.22 |
ENST00000316788.7 |
AP3S1 |
adaptor-related protein complex 3, sigma 1 subunit |
| chr19_-_58662139 | 0.21 |
ENST00000598312.1 |
ZNF329 |
zinc finger protein 329 |
| chr11_-_89223883 | 0.20 |
ENST00000528341.1 |
NOX4 |
NADPH oxidase 4 |
| chr20_-_58515344 | 0.19 |
ENST00000370996.3 |
PPP1R3D |
protein phosphatase 1, regulatory subunit 3D |
| chr15_-_85259330 | 0.19 |
ENST00000560266.1 |
SEC11A |
SEC11 homolog A (S. cerevisiae) |
| chr15_-_85259294 | 0.18 |
ENST00000558217.1 ENST00000558196.1 ENST00000558134.1 |
SEC11A |
SEC11 homolog A (S. cerevisiae) |
| chr15_-_85259384 | 0.15 |
ENST00000455959.3 |
SEC11A |
SEC11 homolog A (S. cerevisiae) |
| chr15_+_59279851 | 0.13 |
ENST00000348370.4 ENST00000434298.1 ENST00000559160.1 |
RNF111 |
ring finger protein 111 |
| chr6_-_110501200 | 0.12 |
ENST00000392586.1 ENST00000419252.1 ENST00000392589.1 ENST00000392588.1 ENST00000359451.2 |
WASF1 |
WAS protein family, member 1 |
| chr15_-_85259360 | 0.10 |
ENST00000559729.1 |
SEC11A |
SEC11 homolog A (S. cerevisiae) |
| chr18_+_9475001 | 0.10 |
ENST00000019317.4 |
RALBP1 |
ralA binding protein 1 |
| chr2_+_242289502 | 0.10 |
ENST00000451310.1 |
SEPT2 |
septin 2 |
| chr20_+_34129770 | 0.10 |
ENST00000348547.2 ENST00000357394.4 ENST00000447986.1 ENST00000279052.6 ENST00000416206.1 ENST00000411577.1 ENST00000413587.1 |
ERGIC3 |
ERGIC and golgi 3 |
| chr15_+_80351977 | 0.10 |
ENST00000559157.1 ENST00000561012.1 ENST00000564367.1 ENST00000558494.1 |
ZFAND6 |
zinc finger, AN1-type domain 6 |
| chr5_+_162864575 | 0.09 |
ENST00000512163.1 ENST00000393929.1 ENST00000340828.2 ENST00000511683.2 ENST00000510097.1 ENST00000511490.2 ENST00000510664.1 |
CCNG1 |
cyclin G1 |
| chr2_+_128403720 | 0.08 |
ENST00000272644.3 |
GPR17 |
G protein-coupled receptor 17 |
| chr4_+_113970772 | 0.07 |
ENST00000504454.1 ENST00000394537.3 ENST00000357077.4 ENST00000264366.6 |
ANK2 |
ankyrin 2, neuronal |
| chrX_+_119737806 | 0.07 |
ENST00000371317.5 |
MCTS1 |
malignant T cell amplified sequence 1 |
| chr15_+_80351910 | 0.07 |
ENST00000261749.6 ENST00000561060.1 |
ZFAND6 |
zinc finger, AN1-type domain 6 |
| chr2_+_128403439 | 0.05 |
ENST00000544369.1 |
GPR17 |
G protein-coupled receptor 17 |
| chr6_+_25754927 | 0.05 |
ENST00000377905.4 ENST00000439485.2 |
SLC17A4 |
solute carrier family 17, member 4 |
| chr6_+_42123141 | 0.05 |
ENST00000418175.1 ENST00000541991.1 ENST00000053469.4 ENST00000394237.1 ENST00000372963.1 |
GUCA1A RP1-139D8.6 |
guanylate cyclase activator 1A (retina) RP1-139D8.6 |
| chr6_+_110501621 | 0.05 |
ENST00000368930.1 ENST00000307731.1 |
CDC40 |
cell division cycle 40 |
| chr22_-_32767017 | 0.04 |
ENST00000400234.1 |
RFPL3S |
RFPL3 antisense |
| chr11_+_85339623 | 0.04 |
ENST00000358867.6 ENST00000534341.1 ENST00000393375.1 ENST00000531274.1 |
TMEM126B |
transmembrane protein 126B |
| chr9_-_86955598 | 0.03 |
ENST00000376238.4 |
SLC28A3 |
solute carrier family 28 (concentrative nucleoside transporter), member 3 |
| chr11_+_2323349 | 0.03 |
ENST00000381121.3 |
TSPAN32 |
tetraspanin 32 |
| chr7_-_38407770 | 0.03 |
ENST00000390348.2 |
TRGV1 |
T cell receptor gamma variable 1 (non-functional) |
| chr1_-_21377383 | 0.03 |
ENST00000374935.3 |
EIF4G3 |
eukaryotic translation initiation factor 4 gamma, 3 |
| chr3_-_108248169 | 0.02 |
ENST00000273353.3 |
MYH15 |
myosin, heavy chain 15 |
| chr6_+_110501344 | 0.02 |
ENST00000368932.1 |
CDC40 |
cell division cycle 40 |
| chr22_+_31518938 | 0.02 |
ENST00000412985.1 ENST00000331075.5 ENST00000412277.2 ENST00000420017.1 ENST00000400294.2 ENST00000405300.1 ENST00000404390.3 |
INPP5J |
inositol polyphosphate-5-phosphatase J |
| chr10_-_94257512 | 0.02 |
ENST00000371581.5 |
IDE |
insulin-degrading enzyme |
| chr16_-_87350970 | 0.02 |
ENST00000567970.1 |
C16orf95 |
chromosome 16 open reading frame 95 |
| chr4_+_110749143 | 0.01 |
ENST00000317735.4 |
RRH |
retinal pigment epithelium-derived rhodopsin homolog |
| chr11_+_2323236 | 0.01 |
ENST00000182290.4 |
TSPAN32 |
tetraspanin 32 |
| chr1_+_176432298 | 0.01 |
ENST00000367661.3 ENST00000367662.3 |
PAPPA2 |
pappalysin 2 |
| chr9_-_118687405 | 0.00 |
ENST00000374014.3 |
LINC00474 |
long intergenic non-protein coding RNA 474 |
| chr2_-_166060571 | 0.00 |
ENST00000360093.3 |
SCN3A |
sodium channel, voltage-gated, type III, alpha subunit |
| chr8_-_146176199 | 0.00 |
ENST00000532351.1 ENST00000276816.4 ENST00000394909.2 |
ZNF16 |
zinc finger protein 16 |
| chr17_-_40337470 | 0.00 |
ENST00000293330.1 |
HCRT |
hypocretin (orexin) neuropeptide precursor |
| chr1_-_47655686 | 0.00 |
ENST00000294338.2 |
PDZK1IP1 |
PDZK1 interacting protein 1 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.8 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.9 | PID IL1 PATHWAY | IL1-mediated signaling events |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 1.4 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 1.0 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 0.8 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.6 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.0 | 0.9 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | GO:0005589 | collagen type VI trimer(GO:0005589) collagen beaded filament(GO:0098647) |
| 0.1 | 1.0 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.1 | 0.6 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.0 | 0.4 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.0 | 0.2 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.0 | 0.2 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.0 | 0.4 | GO:0043220 | Schmidt-Lanterman incisure(GO:0043220) |
| 0.0 | 0.2 | GO:0042575 | DNA polymerase complex(GO:0042575) |
| 0.0 | 0.5 | GO:0031092 | platelet alpha granule membrane(GO:0031092) |
| 0.0 | 0.4 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:1904204 | regulation of skeletal muscle hypertrophy(GO:1904204) |
| 0.3 | 0.9 | GO:2000661 | positive regulation of interleukin-1-mediated signaling pathway(GO:2000661) |
| 0.2 | 1.4 | GO:2000124 | regulation of endocannabinoid signaling pathway(GO:2000124) |
| 0.2 | 0.8 | GO:0048880 | sensory system development(GO:0048880) |
| 0.1 | 1.8 | GO:1900747 | negative regulation of vascular endothelial growth factor signaling pathway(GO:1900747) |
| 0.1 | 1.6 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.1 | 0.5 | GO:0070543 | response to linoleic acid(GO:0070543) |
| 0.0 | 0.3 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.0 | 0.4 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.0 | 0.2 | GO:2000807 | regulation of synaptic vesicle clustering(GO:2000807) |
| 0.0 | 0.6 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.0 | 0.8 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.0 | 0.1 | GO:0036309 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.0 | 0.1 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.0 | 0.1 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.0 | 0.1 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.0 | 0.2 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 0.0 | 0.2 | GO:0099514 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.0 | 0.0 | GO:0030886 | negative regulation of myeloid dendritic cell activation(GO:0030886) |
| 0.0 | 0.4 | GO:0050667 | homocysteine metabolic process(GO:0050667) |
| 0.0 | 0.1 | GO:0002862 | negative regulation of inflammatory response to antigenic stimulus(GO:0002862) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | GO:0004909 | interleukin-1, Type I, activating receptor activity(GO:0004909) |
| 0.1 | 1.4 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.1 | 1.6 | GO:0008179 | adenylate cyclase binding(GO:0008179) |
| 0.1 | 1.0 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.1 | 0.5 | GO:0070892 | lipoteichoic acid receptor activity(GO:0070892) |
| 0.1 | 0.8 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.4 | GO:0019826 | oxygen sensor activity(GO:0019826) |
| 0.0 | 0.1 | GO:0032184 | SUMO polymer binding(GO:0032184) |
| 0.0 | 1.8 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.0 | GO:0015389 | pyrimidine- and adenine-specific:sodium symporter activity(GO:0015389) |