Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
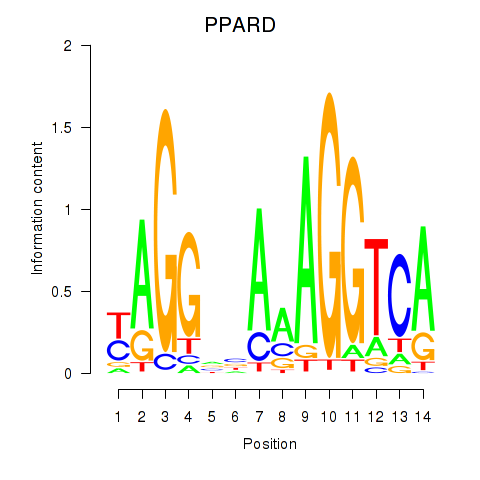
Results for PPARD
Z-value: 0.62
Transcription factors associated with PPARD
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PPARD
|
ENSG00000112033.9 | PPARD |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| PPARD | hg19_v2_chr6_+_35310312_35310359 | -0.31 | 2.4e-01 | Click! |
Activity profile of PPARD motif
Sorted Z-values of PPARD motif
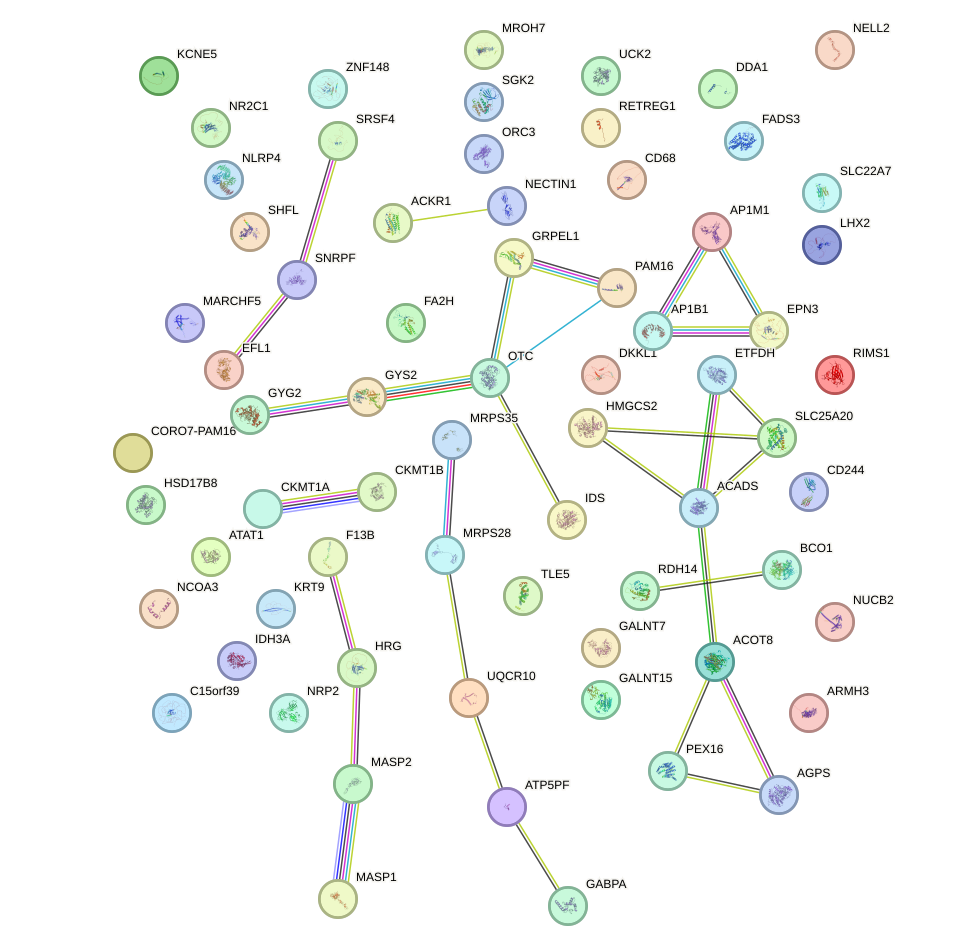
Network of associatons between targets according to the STRING database.
First level regulatory network of PPARD
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr15_+_43985084 | 1.39 |
ENST00000434505.1 ENST00000411750.1 |
CKMT1A |
creatine kinase, mitochondrial 1A |
| chr15_+_43885252 | 1.29 |
ENST00000453782.1 ENST00000300283.6 ENST00000437924.1 ENST00000450086.2 |
CKMT1B |
creatine kinase, mitochondrial 1B |
| chr6_+_33172407 | 0.67 |
ENST00000374662.3 |
HSD17B8 |
hydroxysteroid (17-beta) dehydrogenase 8 |
| chr4_+_174089904 | 0.64 |
ENST00000265000.4 |
GALNT7 |
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 7 (GalNAc-T7) |
| chr1_+_156123318 | 0.61 |
ENST00000368285.3 |
SEMA4A |
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A |
| chr19_+_10197463 | 0.54 |
ENST00000590378.1 ENST00000397881.3 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr15_-_82555000 | 0.54 |
ENST00000557844.1 ENST00000359445.3 ENST00000268206.7 |
EFTUD1 |
elongation factor Tu GTP binding domain containing 1 |
| chr19_-_10687983 | 0.52 |
ENST00000587069.1 |
AP1M2 |
adaptor-related protein complex 1, mu 2 subunit |
| chr19_-_10687907 | 0.52 |
ENST00000589348.1 |
AP1M2 |
adaptor-related protein complex 1, mu 2 subunit |
| chr19_+_10196981 | 0.49 |
ENST00000591813.1 |
C19orf66 |
chromosome 19 open reading frame 66 |
| chr5_-_16509101 | 0.49 |
ENST00000399793.2 |
FAM134B |
family with sequence similarity 134, member B |
| chr15_+_78441663 | 0.46 |
ENST00000299518.2 ENST00000558554.1 ENST00000557826.1 ENST00000561279.1 ENST00000559186.1 ENST00000560770.1 ENST00000559881.1 ENST00000559205.1 ENST00000441490.2 |
IDH3A |
isocitrate dehydrogenase 3 (NAD+) alpha |
| chrX_-_148571884 | 0.46 |
ENST00000537071.1 |
IDS |
iduronate 2-sulfatase |
| chr1_-_197036364 | 0.39 |
ENST00000367412.1 |
F13B |
coagulation factor XIII, B polypeptide |
| chr3_-_48936272 | 0.38 |
ENST00000544097.1 ENST00000430379.1 ENST00000319017.4 |
SLC25A20 |
solute carrier family 25 (carnitine/acylcarnitine translocase), member 20 |
| chr17_-_39728303 | 0.36 |
ENST00000588431.1 ENST00000246662.4 |
KRT9 |
keratin 9 |
| chr8_-_80942467 | 0.34 |
ENST00000518271.1 ENST00000276585.4 ENST00000521605.1 |
MRPS28 |
mitochondrial ribosomal protein S28 |
| chr9_+_126773880 | 0.34 |
ENST00000373615.4 |
LHX2 |
LIM homeobox 2 |
| chr8_-_80942061 | 0.33 |
ENST00000519386.1 |
MRPS28 |
mitochondrial ribosomal protein S28 |
| chr19_-_10687948 | 0.33 |
ENST00000592285.1 |
AP1M2 |
adaptor-related protein complex 1, mu 2 subunit |
| chr12_-_95467267 | 0.32 |
ENST00000330677.7 |
NR2C1 |
nuclear receptor subfamily 2, group C, member 1 |
| chr8_-_80942139 | 0.30 |
ENST00000521434.1 ENST00000519120.1 ENST00000520946.1 |
MRPS28 |
mitochondrial ribosomal protein S28 |
| chr16_+_2255710 | 0.29 |
ENST00000397124.1 ENST00000565250.1 |
MLST8 |
MTOR associated protein, LST8 homolog (S. cerevisiae) |
| chr22_-_51017084 | 0.27 |
ENST00000360719.2 ENST00000457250.1 ENST00000440709.1 |
CPT1B |
carnitine palmitoyltransferase 1B (muscle) |
| chr22_-_51016846 | 0.27 |
ENST00000312108.7 ENST00000395650.2 |
CPT1B |
carnitine palmitoyltransferase 1B (muscle) |
| chr11_-_119599794 | 0.27 |
ENST00000264025.3 |
PVRL1 |
poliovirus receptor-related 1 (herpesvirus entry mediator C) |
| chr11_-_116968987 | 0.27 |
ENST00000434315.2 ENST00000292055.4 ENST00000375288.1 ENST00000542607.1 ENST00000445177.1 ENST00000375300.1 ENST00000446921.2 |
SIK3 |
SIK family kinase 3 |
| chr16_+_2255841 | 0.27 |
ENST00000301725.7 |
MLST8 |
MTOR associated protein, LST8 homolog (S. cerevisiae) |
| chr6_-_33547975 | 0.26 |
ENST00000442998.2 ENST00000360661.5 |
BAK1 |
BCL2-antagonist/killer 1 |
| chr22_-_29784519 | 0.26 |
ENST00000357586.2 ENST00000356015.2 ENST00000432560.2 ENST00000317368.7 |
AP1B1 |
adaptor-related protein complex 1, beta 1 subunit |
| chr4_-_7069760 | 0.25 |
ENST00000264954.4 |
GRPEL1 |
GrpE-like 1, mitochondrial (E. coli) |
| chr6_-_33548006 | 0.24 |
ENST00000374467.3 |
BAK1 |
BCL2-antagonist/killer 1 |
| chr22_+_30163340 | 0.24 |
ENST00000330029.6 ENST00000401406.3 |
UQCR10 |
ubiquinol-cytochrome c reductase, complex III subunit X |
| chr17_+_48610074 | 0.24 |
ENST00000503690.1 ENST00000514874.1 ENST00000537145.1 ENST00000541226.1 |
EPN3 |
epsin 3 |
| chr11_+_64073699 | 0.23 |
ENST00000405666.1 ENST00000468670.1 |
ESRRA |
estrogen-related receptor alpha |
| chr2_+_219433281 | 0.22 |
ENST00000273064.6 ENST00000509807.2 ENST00000542068.1 |
RQCD1 |
RCD1 required for cell differentiation1 homolog (S. pombe) |
| chr19_-_3062465 | 0.22 |
ENST00000327141.4 |
AES |
amino-terminal enhancer of split |
| chr19_+_17420340 | 0.21 |
ENST00000359866.4 |
DDA1 |
DET1 and DDB1 associated 1 |
| chr1_-_29508321 | 0.21 |
ENST00000546138.1 |
SRSF4 |
serine/arginine-rich splicing factor 4 |
| chr16_+_81272287 | 0.21 |
ENST00000425577.2 ENST00000564552.1 |
BCMO1 |
beta-carotene 15,15'-monooxygenase 1 |
| chr16_-_4401258 | 0.19 |
ENST00000577031.1 |
PAM16 |
presequence translocase-associated motor 16 homolog (S. cerevisiae) |
| chr11_-_45939565 | 0.19 |
ENST00000525192.1 ENST00000378750.5 |
PEX16 |
peroxisomal biogenesis factor 16 |
| chr11_-_45939374 | 0.18 |
ENST00000533151.1 ENST00000241041.3 |
PEX16 |
peroxisomal biogenesis factor 16 |
| chrX_-_108868390 | 0.18 |
ENST00000372101.2 |
KCNE1L |
KCNE1-like |
| chr15_+_75494214 | 0.17 |
ENST00000394987.4 |
C15orf39 |
chromosome 15 open reading frame 39 |
| chr1_-_161102421 | 0.16 |
ENST00000490843.2 ENST00000368006.3 ENST00000392188.1 ENST00000545495.1 |
DEDD |
death effector domain containing |
| chr19_+_39616410 | 0.16 |
ENST00000602004.1 ENST00000599470.1 ENST00000321944.4 ENST00000593480.1 ENST00000358301.3 ENST00000593690.1 ENST00000599386.1 |
PAK4 |
p21 protein (Cdc42/Rac)-activated kinase 4 |
| chr1_-_161102367 | 0.15 |
ENST00000464113.1 |
DEDD |
death effector domain containing |
| chr1_-_11107280 | 0.14 |
ENST00000400897.3 ENST00000400898.3 |
MASP2 |
mannan-binding lectin serine peptidase 2 |
| chr20_-_44485835 | 0.14 |
ENST00000457981.1 ENST00000426915.1 ENST00000217455.4 |
ACOT8 |
acyl-CoA thioesterase 8 |
| chr20_+_46130601 | 0.14 |
ENST00000341724.6 |
NCOA3 |
nuclear receptor coactivator 3 |
| chr11_+_17281900 | 0.14 |
ENST00000530527.1 |
NUCB2 |
nucleobindin 2 |
| chr12_+_60058458 | 0.13 |
ENST00000548610.1 |
SLC16A7 |
solute carrier family 16 (monocarboxylate transporter), member 7 |
| chr1_+_165864800 | 0.13 |
ENST00000469256.2 |
UCK2 |
uridine-cytidine kinase 2 |
| chr10_-_103815874 | 0.12 |
ENST00000370033.4 ENST00000311122.5 |
C10orf76 |
chromosome 10 open reading frame 76 |
| chr12_+_121163538 | 0.12 |
ENST00000242592.4 |
ACADS |
acyl-CoA dehydrogenase, C-2 to C-3 short chain |
| chr3_+_48507210 | 0.12 |
ENST00000433541.1 ENST00000422277.2 ENST00000436480.2 ENST00000444177.1 |
TREX1 |
three prime repair exonuclease 1 |
| chr10_+_94050913 | 0.12 |
ENST00000358935.2 |
MARCH5 |
membrane-associated ring finger (C3HC4) 5 |
| chr2_+_206547215 | 0.11 |
ENST00000360409.3 ENST00000540178.1 ENST00000540841.1 ENST00000355117.4 ENST00000450507.1 ENST00000417189.1 |
NRP2 |
neuropilin 2 |
| chr6_+_30594619 | 0.11 |
ENST00000318999.7 ENST00000376485.4 ENST00000376478.2 ENST00000319027.5 ENST00000376483.4 ENST00000329992.8 ENST00000330083.5 |
ATAT1 |
alpha tubulin acetyltransferase 1 |
| chr17_+_7483761 | 0.11 |
ENST00000584180.1 |
CD68 |
CD68 molecule |
| chr12_+_96252706 | 0.11 |
ENST00000266735.5 ENST00000553192.1 ENST00000552085.1 |
SNRPF |
small nuclear ribonucleoprotein polypeptide F |
| chr3_-_187009798 | 0.10 |
ENST00000337774.5 |
MASP1 |
mannan-binding lectin serine peptidase 1 (C4/C2 activating component of Ra-reactive factor) |
| chr21_-_27107198 | 0.10 |
ENST00000400094.1 |
ATP5J |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr4_+_140586922 | 0.10 |
ENST00000265498.1 ENST00000506797.1 |
MGST2 |
microsomal glutathione S-transferase 2 |
| chr3_+_186383741 | 0.10 |
ENST00000232003.4 |
HRG |
histidine-rich glycoprotein |
| chrX_+_2746818 | 0.10 |
ENST00000398806.3 |
GYG2 |
glycogenin 2 |
| chr20_+_42187682 | 0.09 |
ENST00000373092.3 ENST00000373077.1 |
SGK2 |
serum/glucocorticoid regulated kinase 2 |
| chr1_+_159175201 | 0.09 |
ENST00000368121.2 |
DARC |
Duffy blood group, atypical chemokine receptor |
| chr16_-_4401284 | 0.09 |
ENST00000318059.3 |
PAM16 |
presequence translocase-associated motor 16 homolog (S. cerevisiae) |
| chr1_+_165864821 | 0.08 |
ENST00000470820.1 |
UCK2 |
uridine-cytidine kinase 2 |
| chr20_+_42187608 | 0.08 |
ENST00000373100.1 |
SGK2 |
serum/glucocorticoid regulated kinase 2 |
| chr21_-_27107283 | 0.08 |
ENST00000284971.3 ENST00000400099.1 |
ATP5J |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr1_+_55107449 | 0.07 |
ENST00000421030.2 ENST00000545244.1 ENST00000339553.5 ENST00000409996.1 ENST00000454855.2 |
MROH7 |
maestro heat-like repeat family member 7 |
| chr12_-_21757774 | 0.07 |
ENST00000261195.2 |
GYS2 |
glycogen synthase 2 (liver) |
| chr6_+_43265992 | 0.07 |
ENST00000449231.1 ENST00000372589.3 ENST00000372585.5 |
SLC22A7 |
solute carrier family 22 (organic anion transporter), member 7 |
| chr21_-_27107344 | 0.07 |
ENST00000457143.2 |
ATP5J |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr21_+_27107672 | 0.06 |
ENST00000400075.3 |
GABPA |
GA binding protein transcription factor, alpha subunit 60kDa |
| chr2_+_178257372 | 0.06 |
ENST00000264167.4 ENST00000409888.1 |
AGPS |
alkylglycerone phosphate synthase |
| chr12_-_56727487 | 0.06 |
ENST00000548043.1 ENST00000425394.2 |
PAN2 |
PAN2 poly(A) specific ribonuclease subunit homolog (S. cerevisiae) |
| chr1_-_160832642 | 0.05 |
ENST00000368034.4 |
CD244 |
CD244 molecule, natural killer cell receptor 2B4 |
| chr21_-_27107881 | 0.05 |
ENST00000400090.3 ENST00000400087.3 ENST00000400093.3 |
ATP5J |
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit F6 |
| chr12_-_53320245 | 0.05 |
ENST00000552150.1 |
KRT8 |
keratin 8 |
| chr1_-_120311517 | 0.04 |
ENST00000369406.3 ENST00000544913.2 |
HMGCS2 |
3-hydroxy-3-methylglutaryl-CoA synthase 2 (mitochondrial) |
| chr6_+_88299833 | 0.04 |
ENST00000392844.3 ENST00000257789.4 ENST00000546266.1 ENST00000417380.2 |
ORC3 |
origin recognition complex, subunit 3 |
| chr4_+_159593271 | 0.04 |
ENST00000512251.1 ENST00000511912.1 |
ETFDH |
electron-transferring-flavoprotein dehydrogenase |
| chr16_-_74808710 | 0.03 |
ENST00000219368.3 ENST00000544337.1 |
FA2H |
fatty acid 2-hydroxylase |
| chrX_+_38211777 | 0.03 |
ENST00000039007.4 |
OTC |
ornithine carbamoyltransferase |
| chr1_+_9005917 | 0.03 |
ENST00000549778.1 ENST00000480186.3 ENST00000377443.2 ENST00000377436.3 ENST00000377442.2 |
CA6 |
carbonic anhydrase VI |
| chr1_-_11863171 | 0.03 |
ENST00000376592.1 |
MTHFR |
methylenetetrahydrofolate reductase (NAD(P)H) |
| chr11_-_61658853 | 0.02 |
ENST00000525588.1 ENST00000540820.1 |
FADS3 |
fatty acid desaturase 3 |
| chr19_+_49867181 | 0.02 |
ENST00000597546.1 |
DKKL1 |
dickkopf-like 1 |
| chr1_+_62417957 | 0.01 |
ENST00000307297.7 ENST00000543708.1 |
INADL |
InaD-like (Drosophila) |
| chr6_+_43266063 | 0.00 |
ENST00000372574.3 |
SLC22A7 |
solute carrier family 22 (organic anion transporter), member 7 |
| chr3_-_125094093 | 0.00 |
ENST00000484491.1 ENST00000492394.1 ENST00000471196.1 ENST00000468369.1 ENST00000544464.1 ENST00000485866.1 ENST00000360647.4 |
ZNF148 |
zinc finger protein 148 |
| chr6_+_72922505 | 0.00 |
ENST00000401910.3 |
RIMS1 |
regulating synaptic membrane exocytosis 1 |
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0002352 | B cell negative selection(GO:0002352) post-embryonic camera-type eye morphogenesis(GO:0048597) |
| 0.1 | 1.4 | GO:0006600 | creatine metabolic process(GO:0006600) |
| 0.1 | 0.4 | GO:0032581 | ER-dependent peroxisome organization(GO:0032581) |
| 0.1 | 0.2 | GO:0009224 | CMP salvage(GO:0006238) CMP biosynthetic process(GO:0009224) CMP metabolic process(GO:0046035) |
| 0.0 | 0.9 | GO:0006853 | carnitine shuttle(GO:0006853) |
| 0.0 | 0.6 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.0 | 0.3 | GO:0048386 | positive regulation of retinoic acid receptor signaling pathway(GO:0048386) |
| 0.0 | 0.5 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 1.6 | GO:0050690 | regulation of defense response to virus by virus(GO:0050690) |
| 0.0 | 0.2 | GO:0017062 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.0 | 0.3 | GO:0021978 | telencephalon regionalization(GO:0021978) |
| 0.0 | 0.1 | GO:2001027 | negative regulation of endothelial cell chemotaxis(GO:2001027) |
| 0.0 | 0.7 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.0 | 0.2 | GO:2000210 | positive regulation of anoikis(GO:2000210) |
| 0.0 | 0.1 | GO:0046504 | ether lipid biosynthetic process(GO:0008611) glycerol ether biosynthetic process(GO:0046504) ether biosynthetic process(GO:1901503) |
| 0.0 | 0.1 | GO:0035624 | receptor transactivation(GO:0035624) |
| 0.0 | 0.1 | GO:2000566 | positive regulation of CD8-positive, alpha-beta T cell proliferation(GO:2000566) |
| 0.0 | 0.1 | GO:1901475 | pyruvate transport(GO:0006848) pyruvate transmembrane transport(GO:1901475) |
| 0.0 | 0.4 | GO:0045109 | intermediate filament organization(GO:0045109) |
| 0.0 | 0.5 | GO:0038202 | TORC1 signaling(GO:0038202) |
| 0.0 | 0.5 | GO:0030207 | chondroitin sulfate catabolic process(GO:0030207) |
| 0.0 | 0.0 | GO:0006267 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 0.0 | 1.0 | GO:0070126 | mitochondrial translational elongation(GO:0070125) mitochondrial translational termination(GO:0070126) |
| 0.0 | 0.5 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 0.1 | GO:0090141 | positive regulation of mitochondrial fission(GO:0090141) |
| 0.0 | 0.1 | GO:1900227 | positive regulation of NLRP3 inflammasome complex assembly(GO:1900227) |
| 0.0 | 0.0 | GO:0070781 | arginine biosynthetic process via ornithine(GO:0042450) response to biotin(GO:0070781) |
| 0.0 | 0.1 | GO:0016559 | peroxisome fission(GO:0016559) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.4 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.1 | 0.7 | GO:0070404 | NADH binding(GO:0070404) |
| 0.1 | 0.4 | GO:0015199 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 0.1 | 0.5 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.1 | 0.5 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) |
| 0.0 | 0.6 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.6 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.0 | 0.1 | GO:0001855 | complement component C4b binding(GO:0001855) |
| 0.0 | 0.2 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.0 | 0.1 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 0.0 | 0.1 | GO:0005477 | pyruvate secondary active transmembrane transporter activity(GO:0005477) |
| 0.0 | 0.2 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.0 | 0.5 | GO:0051400 | BH domain binding(GO:0051400) |
| 0.0 | 0.1 | GO:0004468 | lysine N-acetyltransferase activity, acting on acetyl phosphate as donor(GO:0004468) |
| 0.0 | 0.5 | GO:0008484 | sulfuric ester hydrolase activity(GO:0008484) |
| 0.0 | 0.2 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.0 | 0.1 | GO:0008853 | exodeoxyribonuclease III activity(GO:0008853) |
| 0.0 | 0.2 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.0 | 0.6 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.1 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.0 | 0.9 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.0 | 0.6 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.6 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.4 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.5 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.5 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |
| 0.0 | 0.3 | REACTOME FORMATION OF ATP BY CHEMIOSMOTIC COUPLING | Genes involved in Formation of ATP by chemiosmotic coupling |
| 0.0 | 0.5 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.5 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 0.1 | 0.6 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 0.5 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.0 | 1.0 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 1.6 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 0.2 | GO:0045275 | mitochondrial respiratory chain complex III(GO:0005750) respiratory chain complex III(GO:0045275) |
| 0.0 | 0.3 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 0.4 | GO:0031231 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.0 | 0.1 | GO:0005683 | U7 snRNP(GO:0005683) pICln-Sm protein complex(GO:0034715) |
| 0.0 | 0.0 | GO:0005656 | nuclear pre-replicative complex(GO:0005656) pre-replicative complex(GO:0036387) |
| 0.0 | 0.0 | GO:0031251 | PAN complex(GO:0031251) |
| 0.0 | 0.1 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.0 | 0.1 | GO:0002116 | semaphorin receptor complex(GO:0002116) |