Project
ENCODE cell lines, expression (Ernst 2011)
Navigation
Downloads
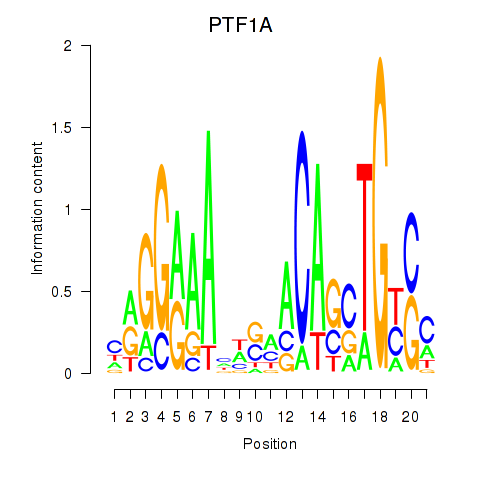
Results for PTF1A
Z-value: 0.75
Transcription factors associated with PTF1A
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
PTF1A
|
ENSG00000168267.5 | PTF1A |
Activity profile of PTF1A motif
Sorted Z-values of PTF1A motif
Network of associatons between targets according to the STRING database.
First level regulatory network of PTF1A
| Promoter | Score | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_+_40198527 | 1.91 |
ENST00000381799.5 |
RHOH |
ras homolog family member H |
| chr22_+_23101182 | 1.74 |
ENST00000390312.2 |
IGLV2-14 |
immunoglobulin lambda variable 2-14 |
| chr1_-_9129735 | 1.63 |
ENST00000377424.4 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr8_+_11351494 | 1.53 |
ENST00000259089.4 |
BLK |
B lymphoid tyrosine kinase |
| chr1_-_9129598 | 1.40 |
ENST00000535586.1 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr2_-_160761179 | 1.24 |
ENST00000554112.1 ENST00000553424.1 ENST00000263636.4 ENST00000504764.1 ENST00000505052.1 |
LY75 LY75-CD302 |
lymphocyte antigen 75 LY75-CD302 readthrough |
| chr8_+_11351876 | 1.21 |
ENST00000529894.1 |
BLK |
B lymphoid tyrosine kinase |
| chr1_-_9129895 | 1.13 |
ENST00000473209.1 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr11_+_7597639 | 1.13 |
ENST00000533792.1 |
PPFIBP2 |
PTPRF interacting protein, binding protein 2 (liprin beta 2) |
| chr1_-_9129631 | 1.07 |
ENST00000377414.3 |
SLC2A5 |
solute carrier family 2 (facilitated glucose/fructose transporter), member 5 |
| chr6_-_24877490 | 1.03 |
ENST00000540914.1 ENST00000378023.4 |
FAM65B |
family with sequence similarity 65, member B |
| chr17_+_67498538 | 0.98 |
ENST00000589647.1 |
MAP2K6 |
mitogen-activated protein kinase kinase 6 |
| chr10_-_74856608 | 0.93 |
ENST00000307116.2 ENST00000373008.2 ENST00000412021.2 ENST00000394890.2 ENST00000263556.3 ENST00000440381.1 |
P4HA1 |
prolyl 4-hydroxylase, alpha polypeptide I |
| chr7_+_7606497 | 0.54 |
ENST00000340080.4 ENST00000405785.1 ENST00000433635.1 |
MIOS |
missing oocyte, meiosis regulator, homolog (Drosophila) |
| chr1_+_12185949 | 0.52 |
ENST00000413146.2 |
TNFRSF8 |
tumor necrosis factor receptor superfamily, member 8 |
| chr3_+_49027308 | 0.48 |
ENST00000383729.4 ENST00000343546.4 |
P4HTM |
prolyl 4-hydroxylase, transmembrane (endoplasmic reticulum) |
| chr3_+_35721106 | 0.43 |
ENST00000474696.1 ENST00000412048.1 ENST00000396482.2 ENST00000432682.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr3_+_167453493 | 0.42 |
ENST00000295777.5 ENST00000472747.2 |
SERPINI1 |
serpin peptidase inhibitor, clade I (neuroserpin), member 1 |
| chr12_-_90102594 | 0.41 |
ENST00000428670.3 |
ATP2B1 |
ATPase, Ca++ transporting, plasma membrane 1 |
| chr1_-_231114542 | 0.40 |
ENST00000522821.1 ENST00000366661.4 ENST00000366662.4 ENST00000414259.1 ENST00000522399.1 |
TTC13 |
tetratricopeptide repeat domain 13 |
| chr12_+_113344755 | 0.40 |
ENST00000550883.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr3_+_35681081 | 0.38 |
ENST00000428373.1 |
ARPP21 |
cAMP-regulated phosphoprotein, 21kDa |
| chr17_+_12692774 | 0.38 |
ENST00000379672.5 ENST00000340825.3 |
ARHGAP44 |
Rho GTPase activating protein 44 |
| chr11_+_96123158 | 0.38 |
ENST00000332349.4 ENST00000458427.1 |
JRKL |
jerky homolog-like (mouse) |
| chr18_-_53177984 | 0.37 |
ENST00000543082.1 |
TCF4 |
transcription factor 4 |
| chr20_+_44441215 | 0.37 |
ENST00000356455.4 ENST00000405520.1 |
UBE2C |
ubiquitin-conjugating enzyme E2C |
| chr2_-_89513402 | 0.36 |
ENST00000498435.1 |
IGKV1-27 |
immunoglobulin kappa variable 1-27 |
| chr19_-_48389651 | 0.35 |
ENST00000222002.3 |
SULT2A1 |
sulfotransferase family, cytosolic, 2A, dehydroepiandrosterone (DHEA)-preferring, member 1 |
| chr17_+_53342311 | 0.34 |
ENST00000226067.5 |
HLF |
hepatic leukemia factor |
| chr20_+_44441304 | 0.33 |
ENST00000352551.5 |
UBE2C |
ubiquitin-conjugating enzyme E2C |
| chrX_-_107019181 | 0.31 |
ENST00000315660.4 ENST00000372384.2 ENST00000502650.1 ENST00000506724.1 |
TSC22D3 |
TSC22 domain family, member 3 |
| chr7_-_99097863 | 0.31 |
ENST00000426306.2 ENST00000337673.6 |
ZNF394 |
zinc finger protein 394 |
| chr12_+_113344582 | 0.31 |
ENST00000202917.5 ENST00000445409.2 ENST00000452357.2 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr13_+_113633620 | 0.29 |
ENST00000421756.1 ENST00000375601.3 |
MCF2L |
MCF.2 cell line derived transforming sequence-like |
| chr7_-_29186008 | 0.28 |
ENST00000396276.3 ENST00000265394.5 |
CPVL |
carboxypeptidase, vitellogenic-like |
| chr5_-_118324200 | 0.27 |
ENST00000515439.3 ENST00000510708.1 |
DTWD2 |
DTW domain containing 2 |
| chr12_-_48398104 | 0.27 |
ENST00000337299.6 ENST00000380518.3 |
COL2A1 |
collagen, type II, alpha 1 |
| chr7_-_152373216 | 0.25 |
ENST00000359321.1 |
XRCC2 |
X-ray repair complementing defective repair in Chinese hamster cells 2 |
| chr6_+_57182400 | 0.25 |
ENST00000607273.1 |
PRIM2 |
primase, DNA, polypeptide 2 (58kDa) |
| chr22_+_24999114 | 0.24 |
ENST00000412658.1 ENST00000445029.1 ENST00000419133.1 ENST00000400382.1 ENST00000438643.2 ENST00000452551.1 ENST00000400383.1 ENST00000412898.1 ENST00000400380.1 ENST00000455483.1 ENST00000430289.1 |
GGT1 |
gamma-glutamyltransferase 1 |
| chr10_-_101841588 | 0.24 |
ENST00000370418.3 |
CPN1 |
carboxypeptidase N, polypeptide 1 |
| chr10_-_1779663 | 0.22 |
ENST00000381312.1 |
ADARB2 |
adenosine deaminase, RNA-specific, B2 (non-functional) |
| chr5_+_169532896 | 0.22 |
ENST00000306268.6 ENST00000449804.2 |
FOXI1 |
forkhead box I1 |
| chr6_-_41715128 | 0.22 |
ENST00000356667.4 ENST00000373025.3 ENST00000425343.2 |
PGC |
progastricsin (pepsinogen C) |
| chr2_-_27435125 | 0.21 |
ENST00000414408.1 ENST00000310574.3 |
SLC5A6 |
solute carrier family 5 (sodium/multivitamin and iodide cotransporter), member 6 |
| chr2_+_97426631 | 0.21 |
ENST00000377075.2 |
CNNM4 |
cyclin M4 |
| chr17_-_73892992 | 0.21 |
ENST00000540128.1 ENST00000269383.3 |
TRIM65 |
tripartite motif containing 65 |
| chr1_-_45956822 | 0.20 |
ENST00000372086.3 ENST00000341771.6 |
TESK2 |
testis-specific kinase 2 |
| chr7_-_100844193 | 0.20 |
ENST00000440203.2 ENST00000379423.3 ENST00000223114.4 |
MOGAT3 |
monoacylglycerol O-acyltransferase 3 |
| chr11_-_111794446 | 0.20 |
ENST00000527950.1 |
CRYAB |
crystallin, alpha B |
| chr15_-_89764929 | 0.20 |
ENST00000268125.5 |
RLBP1 |
retinaldehyde binding protein 1 |
| chr12_+_53689309 | 0.19 |
ENST00000351500.3 ENST00000550846.1 ENST00000334478.4 ENST00000549759.1 |
PFDN5 |
prefoldin subunit 5 |
| chr9_+_4985228 | 0.17 |
ENST00000381652.3 |
JAK2 |
Janus kinase 2 |
| chr12_+_113344811 | 0.17 |
ENST00000551241.1 ENST00000553185.1 ENST00000550689.1 |
OAS1 |
2'-5'-oligoadenylate synthetase 1, 40/46kDa |
| chr13_-_79979919 | 0.15 |
ENST00000267229.7 |
RBM26 |
RNA binding motif protein 26 |
| chr2_-_89417335 | 0.15 |
ENST00000490686.1 |
IGKV1-17 |
immunoglobulin kappa variable 1-17 |
| chr1_-_76076759 | 0.15 |
ENST00000370855.5 |
SLC44A5 |
solute carrier family 44, member 5 |
| chr1_-_78149041 | 0.14 |
ENST00000414381.1 ENST00000370798.1 |
ZZZ3 |
zinc finger, ZZ-type containing 3 |
| chrX_+_154299690 | 0.13 |
ENST00000340647.4 ENST00000330045.7 |
BRCC3 |
BRCA1/BRCA2-containing complex, subunit 3 |
| chr19_-_10491234 | 0.13 |
ENST00000524462.1 ENST00000531836.1 ENST00000525621.1 |
TYK2 |
tyrosine kinase 2 |
| chr17_-_33416231 | 0.13 |
ENST00000584655.1 ENST00000447669.2 ENST00000315249.7 |
RFFL |
ring finger and FYVE-like domain containing E3 ubiquitin protein ligase |
| chr16_+_89696692 | 0.13 |
ENST00000261615.4 |
DPEP1 |
dipeptidase 1 (renal) |
| chr13_-_79979952 | 0.13 |
ENST00000438724.1 |
RBM26 |
RNA binding motif protein 26 |
| chr17_-_3461092 | 0.13 |
ENST00000301365.4 ENST00000572519.1 |
TRPV3 |
transient receptor potential cation channel, subfamily V, member 3 |
| chr16_-_75569068 | 0.12 |
ENST00000336257.3 ENST00000565039.1 |
CHST5 |
carbohydrate (N-acetylglucosamine 6-O) sulfotransferase 5 |
| chr1_-_45956800 | 0.12 |
ENST00000538496.1 |
TESK2 |
testis-specific kinase 2 |
| chr10_-_5855350 | 0.12 |
ENST00000456041.1 ENST00000380181.3 ENST00000418688.1 ENST00000380132.4 ENST00000609712.1 ENST00000380191.4 |
GDI2 |
GDP dissociation inhibitor 2 |
| chr11_-_207221 | 0.12 |
ENST00000486280.1 ENST00000332865.6 ENST00000529614.2 ENST00000325147.9 ENST00000410108.1 ENST00000382762.3 |
BET1L |
Bet1 golgi vesicular membrane trafficking protein-like |
| chr10_+_76871454 | 0.12 |
ENST00000372687.4 |
SAMD8 |
sterile alpha motif domain containing 8 |
| chr2_-_89278535 | 0.11 |
ENST00000390247.2 |
IGKV3-7 |
immunoglobulin kappa variable 3-7 (non-functional) |
| chr6_-_62996066 | 0.10 |
ENST00000281156.4 |
KHDRBS2 |
KH domain containing, RNA binding, signal transduction associated 2 |
| chr17_-_42143963 | 0.10 |
ENST00000585388.1 ENST00000293406.3 |
LSM12 |
LSM12 homolog (S. cerevisiae) |
| chr2_-_136288113 | 0.10 |
ENST00000401392.1 |
ZRANB3 |
zinc finger, RAN-binding domain containing 3 |
| chrX_+_154299753 | 0.10 |
ENST00000369459.2 ENST00000369462.1 ENST00000411985.1 ENST00000399042.1 |
BRCC3 |
BRCA1/BRCA2-containing complex, subunit 3 |
| chr20_+_30697298 | 0.09 |
ENST00000398022.2 |
TM9SF4 |
transmembrane 9 superfamily protein member 4 |
| chr5_+_140557371 | 0.08 |
ENST00000239444.2 |
PCDHB8 |
protocadherin beta 8 |
| chr12_+_50451331 | 0.08 |
ENST00000228468.4 |
ASIC1 |
acid-sensing (proton-gated) ion channel 1 |
| chr10_+_99332198 | 0.08 |
ENST00000307518.5 ENST00000298808.5 ENST00000370655.1 |
ANKRD2 |
ankyrin repeat domain 2 (stretch responsive muscle) |
| chr6_-_56258892 | 0.08 |
ENST00000370819.1 |
COL21A1 |
collagen, type XXI, alpha 1 |
| chr18_+_5238549 | 0.07 |
ENST00000580684.1 |
LINC00667 |
long intergenic non-protein coding RNA 667 |
| chr14_-_77737543 | 0.07 |
ENST00000298352.4 |
NGB |
neuroglobin |
| chr12_+_112451120 | 0.07 |
ENST00000261735.3 ENST00000455836.1 |
ERP29 |
endoplasmic reticulum protein 29 |
| chr17_+_40996590 | 0.07 |
ENST00000253799.3 ENST00000452774.2 |
AOC2 |
amine oxidase, copper containing 2 (retina-specific) |
| chr13_+_113812956 | 0.06 |
ENST00000375547.2 ENST00000342783.4 |
PROZ |
protein Z, vitamin K-dependent plasma glycoprotein |
| chr13_+_49794474 | 0.06 |
ENST00000218721.1 ENST00000398307.1 |
MLNR |
motilin receptor |
| chrX_+_155110956 | 0.05 |
ENST00000286448.6 ENST00000262640.6 ENST00000460621.1 |
VAMP7 |
vesicle-associated membrane protein 7 |
| chr3_+_51663407 | 0.05 |
ENST00000432863.1 ENST00000296477.3 |
RAD54L2 |
RAD54-like 2 (S. cerevisiae) |
| chr10_+_43572475 | 0.05 |
ENST00000355710.3 ENST00000498820.1 ENST00000340058.5 |
RET |
ret proto-oncogene |
| chr16_-_15950868 | 0.05 |
ENST00000396324.3 ENST00000452625.2 ENST00000576790.2 ENST00000300036.5 |
MYH11 |
myosin, heavy chain 11, smooth muscle |
| chr16_+_2802316 | 0.05 |
ENST00000301740.8 |
SRRM2 |
serine/arginine repetitive matrix 2 |
| chr3_+_40518599 | 0.05 |
ENST00000314686.5 ENST00000447116.2 ENST00000429348.2 ENST00000456778.1 |
ZNF619 |
zinc finger protein 619 |
| chrX_+_15808569 | 0.05 |
ENST00000380308.3 ENST00000307771.7 |
ZRSR2 |
zinc finger (CCCH type), RNA-binding motif and serine/arginine rich 2 |
| chr20_-_48732472 | 0.04 |
ENST00000340309.3 ENST00000415862.2 ENST00000371677.3 ENST00000420027.2 |
UBE2V1 |
ubiquitin-conjugating enzyme E2 variant 1 |
| chr12_+_75784850 | 0.04 |
ENST00000550916.1 ENST00000435775.1 ENST00000378689.2 ENST00000378692.3 ENST00000320460.4 ENST00000547164.1 |
GLIPR1L2 |
GLI pathogenesis-related 1 like 2 |
| chr6_+_27107053 | 0.04 |
ENST00000354348.2 |
HIST1H4I |
histone cluster 1, H4i |
| chr18_+_33877654 | 0.04 |
ENST00000257209.4 ENST00000445677.1 ENST00000590592.1 ENST00000359247.4 |
FHOD3 |
formin homology 2 domain containing 3 |
| chr8_-_22089533 | 0.04 |
ENST00000321613.3 |
PHYHIP |
phytanoyl-CoA 2-hydroxylase interacting protein |
| chr15_+_68924327 | 0.04 |
ENST00000543950.1 |
CORO2B |
coronin, actin binding protein, 2B |
| chr22_-_17640110 | 0.04 |
ENST00000399852.3 ENST00000336737.4 |
CECR5 |
cat eye syndrome chromosome region, candidate 5 |
| chr4_+_156588115 | 0.04 |
ENST00000455639.2 |
GUCY1A3 |
guanylate cyclase 1, soluble, alpha 3 |
| chr22_-_50523760 | 0.04 |
ENST00000395876.2 |
MLC1 |
megalencephalic leukoencephalopathy with subcortical cysts 1 |
| chr12_+_12966250 | 0.03 |
ENST00000352940.4 ENST00000358007.3 ENST00000544400.1 |
DDX47 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 |
| chr4_-_83719884 | 0.03 |
ENST00000282709.4 ENST00000273908.4 |
SCD5 |
stearoyl-CoA desaturase 5 |
| chr4_-_89744314 | 0.03 |
ENST00000508369.1 |
FAM13A |
family with sequence similarity 13, member A |
| chr19_-_40331345 | 0.03 |
ENST00000597224.1 |
FBL |
fibrillarin |
| chr4_-_89744365 | 0.03 |
ENST00000513837.1 ENST00000503556.1 |
FAM13A |
family with sequence similarity 13, member A |
| chr12_+_120884222 | 0.03 |
ENST00000551765.1 ENST00000229384.5 |
GATC |
glutamyl-tRNA(Gln) amidotransferase, subunit C |
| chr21_+_44073916 | 0.03 |
ENST00000349112.3 ENST00000398224.3 |
PDE9A |
phosphodiesterase 9A |
| chr21_+_44073860 | 0.03 |
ENST00000335512.4 ENST00000539837.1 ENST00000291539.6 ENST00000380328.2 ENST00000398232.3 ENST00000398234.3 ENST00000398236.3 ENST00000328862.6 ENST00000335440.6 ENST00000398225.3 ENST00000398229.3 ENST00000398227.3 |
PDE9A |
phosphodiesterase 9A |
| chr16_-_46723066 | 0.02 |
ENST00000299138.7 |
VPS35 |
vacuolar protein sorting 35 homolog (S. cerevisiae) |
| chr4_-_175750364 | 0.02 |
ENST00000340217.5 ENST00000274093.3 |
GLRA3 |
glycine receptor, alpha 3 |
| chr11_+_71238313 | 0.02 |
ENST00000398536.4 |
KRTAP5-7 |
keratin associated protein 5-7 |
| chr13_+_49280951 | 0.01 |
ENST00000282018.3 |
CYSLTR2 |
cysteinyl leukotriene receptor 2 |
| chr19_+_45254529 | 0.00 |
ENST00000444487.1 |
BCL3 |
B-cell CLL/lymphoma 3 |
| chr4_-_123843597 | 0.00 |
ENST00000510735.1 ENST00000304430.5 |
NUDT6 |
nudix (nucleoside diphosphate linked moiety X)-type motif 6 |
Gene Ontology Analysis
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.7 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 0.3 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.0 | 0.5 | SA MMP CYTOKINE CONNECTION | Cytokines can induce activation of matrix metalloproteinases, which degrade extracellular matrix. |
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 5.2 | GO:0032445 | fructose transport(GO:0015755) fructose import(GO:0032445) carbohydrate import into cell(GO:0097319) carbohydrate import across plasma membrane(GO:0098704) fructose import across plasma membrane(GO:1990539) |
| 0.3 | 1.0 | GO:0072709 | cellular response to sorbitol(GO:0072709) |
| 0.2 | 0.5 | GO:0032639 | TRAIL production(GO:0032639) regulation of TRAIL production(GO:0032679) positive regulation of TRAIL production(GO:0032759) |
| 0.1 | 0.9 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.1 | 0.7 | GO:0010993 | regulation of ubiquitin homeostasis(GO:0010993) free ubiquitin chain polymerization(GO:0010994) positive regulation of exit from mitosis(GO:0031536) |
| 0.1 | 0.2 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.1 | 0.4 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.1 | 2.7 | GO:0050855 | regulation of B cell receptor signaling pathway(GO:0050855) |
| 0.1 | 0.2 | GO:0060398 | regulation of growth hormone receptor signaling pathway(GO:0060398) |
| 0.1 | 0.4 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) |
| 0.0 | 0.2 | GO:0030070 | insulin processing(GO:0030070) |
| 0.0 | 0.2 | GO:0051182 | coenzyme transport(GO:0051182) |
| 0.0 | 0.1 | GO:0016999 | antibiotic metabolic process(GO:0016999) cellular amide catabolic process(GO:0043605) |
| 0.0 | 0.3 | GO:0000707 | meiotic DNA recombinase assembly(GO:0000707) |
| 0.0 | 0.3 | GO:0070236 | regulation of activation-induced cell death of T cells(GO:0070235) negative regulation of activation-induced cell death of T cells(GO:0070236) |
| 0.0 | 0.2 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.0 | 0.2 | GO:0002803 | positive regulation of antimicrobial peptide production(GO:0002225) positive regulation of antimicrobial humoral response(GO:0002760) positive regulation of antibacterial peptide production(GO:0002803) |
| 0.0 | 0.2 | GO:0019344 | cysteine biosynthetic process(GO:0019344) |
| 0.0 | 1.9 | GO:0034260 | negative regulation of GTPase activity(GO:0034260) |
| 0.0 | 0.1 | GO:2001271 | negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.0 | 0.2 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.0 | 0.3 | GO:0060174 | limb bud formation(GO:0060174) |
| 0.0 | 0.1 | GO:0048478 | replication fork protection(GO:0048478) |
| 0.0 | 0.1 | GO:1903595 | positive regulation of histamine secretion by mast cell(GO:1903595) |
| 0.0 | 0.1 | GO:0042636 | negative regulation of hair cycle(GO:0042636) |
| 0.0 | 0.1 | GO:0007497 | posterior midgut development(GO:0007497) |
| 0.0 | 0.1 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) |
| 0.0 | 0.5 | GO:0051923 | sulfation(GO:0051923) |
| 0.0 | 0.1 | GO:0043335 | protein unfolding(GO:0043335) |
| 0.0 | 0.2 | GO:0006776 | vitamin A metabolic process(GO:0006776) |
| 0.0 | 0.5 | GO:0034629 | cellular protein complex localization(GO:0034629) |
| 0.0 | 0.1 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.0 | 1.9 | GO:0030449 | regulation of complement activation(GO:0030449) |
| 0.0 | 0.2 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.0 | 0.1 | GO:0015871 | choline transport(GO:0015871) |
| 0.0 | 0.2 | GO:0006071 | glycerol metabolic process(GO:0006071) |
| 0.0 | 0.1 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 5.2 | GO:0005353 | fructose transmembrane transporter activity(GO:0005353) |
| 0.2 | 0.9 | GO:0019798 | procollagen-proline 4-dioxygenase activity(GO:0004656) procollagen-proline dioxygenase activity(GO:0019798) |
| 0.1 | 0.9 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.1 | 1.9 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.1 | 0.2 | GO:0008523 | sodium-dependent multivitamin transmembrane transporter activity(GO:0008523) |
| 0.1 | 0.2 | GO:0003846 | 2-acylglycerol O-acyltransferase activity(GO:0003846) |
| 0.1 | 0.3 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.1 | 0.5 | GO:0048406 | nerve growth factor binding(GO:0048406) |
| 0.0 | 0.3 | GO:0050294 | steroid sulfotransferase activity(GO:0050294) |
| 0.0 | 0.2 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.0 | 0.1 | GO:0033188 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.0 | 0.1 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.0 | 0.3 | GO:0043426 | MRF binding(GO:0043426) |
| 0.0 | 1.0 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 3.0 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.5 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.0 | 0.4 | GO:0001087 | transcription factor activity, sequence-specific DNA binding, RNA polymerase recruiting(GO:0001011) transcription factor activity, TFIIB-class binding(GO:0001087) |
| 0.0 | 0.2 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 0.0 | 0.1 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 0.0 | 0.5 | GO:0031418 | L-ascorbic acid binding(GO:0031418) |
| 0.0 | 0.2 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.3 | GO:0000150 | recombinase activity(GO:0000150) |
| 0.0 | 0.1 | GO:0052595 | tryptamine:oxygen oxidoreductase (deaminating) activity(GO:0052593) aminoacetone:oxygen oxidoreductase(deaminating) activity(GO:0052594) aliphatic-amine oxidase activity(GO:0052595) phenethylamine:oxygen oxidoreductase (deaminating) activity(GO:0052596) |
| 0.0 | 0.1 | GO:0070573 | metallodipeptidase activity(GO:0070573) |
| 0.0 | 0.2 | GO:0036374 | glutathione hydrolase activity(GO:0036374) |
| 0.0 | 0.1 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.0 | 0.7 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.1 | GO:0016520 | growth hormone-releasing hormone receptor activity(GO:0016520) |
| 0.0 | 0.4 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.0 | GO:0004768 | stearoyl-CoA 9-desaturase activity(GO:0004768) acyl-CoA desaturase activity(GO:0016215) |
| 0.0 | 0.1 | GO:0036310 | annealing helicase activity(GO:0036310) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.9 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.1 | 0.5 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.0 | 0.3 | GO:0033063 | DNA recombinase mediator complex(GO:0033061) Rad51B-Rad51C-Rad51D-XRCC2 complex(GO:0033063) |
| 0.0 | 0.2 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.0 | 5.1 | GO:0035579 | specific granule membrane(GO:0035579) |
| 0.0 | 1.9 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 0.2 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 0.2 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 0.7 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 0.3 | GO:0005583 | fibrillar collagen trimer(GO:0005583) banded collagen fibril(GO:0098643) |
| 0.0 | 0.4 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 3.0 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 0.2 | GO:0097512 | cardiac myofibril(GO:0097512) |
| 0.0 | 0.1 | GO:0043596 | nuclear replication fork(GO:0043596) |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 5.2 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.1 | 2.7 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 1.0 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.0 | 0.7 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.0 | 0.3 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 0.2 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.0 | 0.2 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.0 | 2.4 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.3 | REACTOME CYTOSOLIC SULFONATION OF SMALL MOLECULES | Genes involved in Cytosolic sulfonation of small molecules |
| 0.0 | 0.9 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |